Abstract
During the course of evolution, amino acid shifts might have resulted in mitochondrial proteomes better endowed to resist oxidative stress. However, owing to the problem of distinguishing between functional constraints/adaptations in protein sequences and mutation-driven biases in the composition of these sequences, the adaptive value of such amino acid shifts remains under discussion. We have analyzed the coding sequences of mtDNA from 173 mammalian species, dissecting the effect of nucleotide composition on amino acid usages. We found remarkable cysteine avoidance in mtDNA-encoded proteins. However, no effect of longevity on cysteine content could be detected. On the other hand, nucleotide compositional shifts fully accounted for threonine usages. In spite of a strong effect of mutational bias on methionine abundances, our results suggest a role of selection in determining the composition of methionine. Whether this selective effect is linked or not to protection against oxidative stress is still a subject of debate.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The free radical theory of aging asserts that buildup of macromolecular damage caused by reactive oxygen species (ROS) leads to the functional decline associated with aging in animals (Harman 2006). According to the current theory, mitochondria play a key role in aging because they are both a source of ROS (through leakage of the electron transport chain) and a major target for damage that could lead to a reduction in metabolic function (Balaban et al. 2005). Consistent with this theory, a number of mitochondrial adaptations to oxidative stress related to longevity have been described in the literature (for recent reviews, see Pamplona and Barja 2011; Pamplona 2011; Pamplona and Costantini 2011).
A first line of defense against ROS-related damage consists of ameliorating the rate of ROS production (Barja 1998). In this sense, comparative studies across-species have pointed to complex I of the electron transport chain (Pamplona et al. 2005; Lambert et al. 2007, 2010) and uncoupling proteins (Brand 2000; Speakman et al. 2004), as two potential targets of natural selection. Another strategy that animals seem to have adopted during the course of evolution consists of favoring macromolecular constituents less susceptible to oxidative modification. This point is well illustrated by the finding that the mitochondrial membrane peroxidizability index is inversely related to maximum life span in mammals (Pamplona et al. 1998). Along the same lines, different authors have suggested that concerted changes in mtDNA-encoded proteins leading to mitochondrial proteomes better endowed to resist oxidative stress may have evolved in response to selective pressures (Moosmann and Behl 2008; Kitazoe et al. 2008; Aledo et al. 2011).
Although all amino acid residues are potential targets of oxidative damage, the sulfur-containing residues cysteine and methionine are particularly sensitive to oxidation (Berlett and Stadtman 1997). Interestingly, mitochondrially encoded cysteine and methionine have both been correlated negatively with longevity (Moosmann and Behl 2008; Aledo et al. 2011). Even though both cysteine and methionine are negatively related to longevity, it may be that cysteine is a pro-oxidant while methionine is an anti-oxidant, as we have previously argued (Aledo et al. 2011). In brief, it is the detrimental capacity of cysteine thiyl radicals and their potential to initiate irreversible protein cross-linking that might have caused a selection against cysteine in mtDNA-encoded proteins. In contrast, the major oxidation product of protein-bond methionine is methionine sulfoxide, which can be reduced back to methionine by methionine sulfoxide reductases at the low metabolic cost of one molecule of NADPH (Stadtman et al. 2005). In this way, one equivalent of ROS is destroyed for every equivalent of methionine residue repaired (Levine et al. 1996). As such, a methionine enrichment of mitochondrial proteins may represent an adaptive response to oxidative stress (Bender et al. 2008; Aledo et al. 2011). Therefore, those animals that exhibit higher rates of ROS generation (short-lived animals) might have been subjected to higher selective pressures to increase the methionine content of their mitochondrial proteins. In other words, if mitochondrial methionine residues serve as a ROS sink, then the proteins from animals subjected to high oxidative stress should accumulate methionine more effectively than their orthologous proteins from species exposed to lower oxidative stress. That is, the relationship between methionine usage and longevity is somehow reminiscent of the well documented negative relationship between endogenous antioxidant levels and longevity (reviewed in Pamplona and Costantini 2011).
On the other hand, a positive correlation between threonine abundance in transmembrane regions of mitochondrial proteins and longevity has been reported (Kitazoe et al. 2008; Aledo et al. 2011). Since threonine is thought to provide extra intrahelical hydrogen bonding, thereby reinforcing protein stability, these authors argue that increased threonine content could be beneficial to achieve longer lifespan (Kitazoe et al. 2011).
Although these correlations of longevity with cysteine and methionine (negative) and threonine (positive) were originally interpreted in terms of Darwinian selection, Jobson et al. (2010) have recently raised doubts about the adaptive character of the above-mentioned amino acid compositional shifts. These authors point out that the reported correlations between longevity and amino acid usages might be explained by neutral mutational shifts of nucleotide composition and codon biases, which tend to be prominent in mitochondrial genomes (Albu et al. 2008). Indeed, a plethora of studies have established that protein evolution is affected by the nucleotide composition of the encoding genes (Sueoka 1961; D’Onofrio et al. 1991; Collins and Jukes 1993; Singer and Hickey 2000; Wang et al. 2004). Mitochondrial genomes are not an exception to this general rule (Foster et al. 1997; Nikolaou and Almirantis 2006; Min and Hickey 2007; Jia and Higgs 2008; Jobson et al. 2010). In fact, it has been documented that uncorrected nucleotide bias in mtDNA can mimic the effects of positive selection (Albu et al. 2008). These precedents emphasize the difficulties of distinguishing between functional constraints in protein sequences and mutation-driven biases in the composition of these same sequences. This means that extreme caution should be used when comparing results between taxa that differ in their nucleotide contents, at the same time that it underscores the need for methodologies allowing to discriminate between neutral and selective forces.
In the current study, we have addressed the potential contribution of natural selection to the observed link between amino acid compositional shifts in mitochondrial proteomes and longevity. To this end, we have developed a framework that accounts for the effects of nucleotide compositional biases.
Materials and Methods
Data Sampling
Sequences for mitochondrial genomes and proteome analysis were obtained from the National Center for Biotechnology Information (NCBI) genome database (www.ncbi.nlm.nih.gov). A collection of 173 mammalian species was assembled based on neutral selection criteria, completely unrelated to longevity or nucleotide/amino acid frequencies. Annotated complete mitochondrial sequences had to be available from NCBI genome database, and longevity information had to be given in a reliable source such as the AnAge database at http://genomics.senescence.info/species/ (de Magalhães and Costa 2009).
Computations
The sense sequences (L-strand) corresponding to the 12 protein-coding mitochondrial genes, without stop codons, were concatenated into a single nucleotide sequence. This sequence was translated into amino acid sequence using the vertebrate mitochondrial code. Six different nucleotide random sequence models were generated using a customized Perl script. In the so-called homogeneous model, only two constrains were imposed: (i) the nucleotide frequencies in the random sequence should be the same as in the actual sequence, and (ii) no stop codons were allowed. When a stop codon appeared during the randomization procedure, the nucleotides from that triplet were, in turn, randomly shuffled until they coded for an amino acid. Beside this homogeneous model, we also considered modeling approaches where codon positions were treated as separate categories. For instance, for the model 1-2-3 the nucleotides at the three positions were shuffled in such a way that the frequencies at the first, second, and third position in the random sequence should be the same as the frequencies at the first, second, and third codon position of the actual sequence, respectively. In the models 1, 2, and 3, only the first, the second or the third position was shuffled, respectively. Finally, in the model 1-3, both the first and third positions were shuffled in such a way that the nucleotide frequencies at each position in the random sequence were the same as in the actual sequence. The calculations of the nucleotide and amino acid abundances were carried out with simple Perl code. All the scripts are available from the authors on request.
Statistical Treatment
For each species (i), the occurrences of the amino acid under investigation (methionine, cysteine or threonine) in the actual (x i ) and random (y i ) sequences were computed, yielding a pair of measurement (x i , y i ) for each amino acid. In the absence of functional constraints in the protein sequences, the null hypothesis states that x i and y i are equally likely to be larger than the other (P[X < Y] = P[X > Y] = 0.5). Pairs were omitted for which there was no difference. In this way, the number of valid pairs was n = 171 for the methionine analysis and n = 167 for the case of threonine. Then, the number of pairs W for which x i − y i > 0 was calculated. Under the null hypothesis conditions, W follows a binomial distribution, \( W\,\sim \,B(n,\,0.5). \) However, since n is large enough, the normal approximation to the binomial distribution can be used. For this purpose, the variable W was typified taking into consideration that the mean is given by n/2 and the variance is n/4 (E[W] = np = n/2; Var[W] = np(1 − p) = n/4). The typified variable, let us call it Z, follows now a normal distribution, \( Z\,\sim \,N(0,\,1). \)
Probability calculations were assisted by Wolfram Mathematica 8.0. All other statistic analyses were done with SPSS 15.0.
Results and Discussion
Since it has been suggested that the link between longevity and amino acid frequencies in mtDNA-encoded proteins may be a consequence of nucleotides biases rather than reflect an adaptive process, we started by addressing the relationship of nucleotide abundances to maximum life span in a set of 173 mammalian species (Fig. 1). While adenine and guanine abundances showed weak, if any, relationships with longevity (p values: 0.001 and 0.070, respectively), thymine and cytosine abundances exhibited a strong relationship with lifespan (p values: 7 × 10−15 and 10−12, respectively). The clear pattern in these data is that increases in longevity are accompanied by increases in the frequencies of C, which are paralleled by decreases in the frequencies of T. A similar trend has previously been reported (Samuels 2005). In this study 76 mammalian species were analyzed in the context of a differential susceptibility of mtDNA to damage, which according to this author may be brought about by nucleotide composition and be related to lifespan. While we confirm and extent this initial observation using a much wider set of species, that was not the main focus of the current paper.
Relationships between nucleotide abundance on the mtDNA L-strand and longevity. The 12 protein-coding genes on the L-strand from mtDNA were used to compute the base content. Then, the correlations between these absolute frequencies and log MLSP were addressed, using the data set formed by the 173 species analyzed in the current study. Thymine and cytosine abundances exhibited a strong relationship with lifespan (p values: 7 × 10−15 and 10−12, respectively), while adenine and guanine exhibited little, if any, relationship with longevity (p values: 0.001 and 0.070, respectively)
It seems obvious that these nucleotide biases, which may be due to differences in generation times since long-lived species also have longer generation times and thus mutation biases will be skewed according to generation times, may bring about biases in the amino acid composition of the encoded proteins. Therefore, we next asked if the observed correlation between nucleotide composition and longevity may account for the previously reported links between longevity and cysteine, methionine and threonine usages in mtDNA-encoded proteins. To this end, for each species, the sense sequences encompassing the 12 proteins encoded by the H-strand were used to generate random sequences with the same nucleotide frequencies as the actual sequence. After translating these random sequences, the number of times a given residue appeared was plotted against the longevity of the species being analyzed. The results of such analyses showed that the nucleotide bias might be a significant driving force in methionine and threonine composition of the encoded proteins (Fig. 2). At this point, it may be argued that using a homogeneous model where the overall mitochondrial sequence is randomized, any constraints depending on codon position may be overlooked. Therefore, we next considered modeling approaches where codon positions were treated as separate categories of sites (see “Materials and Methods” section). The results derived from such models were qualitatively the same as those obtained with the homogenous model (see Supplementary Fig. 1). This congruency between models was not unexpected since Jobson et al. (2010) noted that the covariance between nucleotide usages and longevity showed a remarkable similar pattern regardless of the codon position, which was interpreted by these authors as evidence that the overall mitochondrial compositional pattern is driven by mutation biases.
Potential influence of the nucleotide composition on the cysteine, methionine, and threonine usages in mtDNA-encoded proteins. For each species, the sense sequence encompassing the 12 proteins encoded by the H-strand was used to generate random sequences with the same nucleotide frequencies than the actual sequence. Afterward, these random DNA sequences were translated using the vertebrate mitochondrial code and the number of times a cysteine (a), methionine (b) or threonine (c) residue appeared was plotted against the longevity of the species under analysis
Whatever the underlying causes, nucleotide bias seems to play an important role in determining the methionine and threonine content. However, the question we wanted to answer was: Does nucleotide composition fully account for the observed amino acid usages? In other words, can we rule out selective forces contributing to shape the methionine and threonine composition of mtDNA-encoded proteins?
As a first approach to the above questions, we computed the number of times a given residue appeared in the 12 proteins encoded by the H-strand, plotted it against the number of times this residue appeared in a random sequence with abundances for the four bases equal to those observed for the species being considered (Fig. 3). As expected, there was a positive covariance between actual and random occurrences for the three amino acids. However, each amino acid behaved in a different way. As such, to evaluate the influence of nucleotide composition on each amino acid usage, two aspects need to be considered. On one hand, the slopes in these plots inform us about the inertia of amino acid usages with respect to mutational bias. On the other hand, the distribution of the points with respect to the bisectrix can also be informative.
Departures of the actual amino acid usages from random expectations. For each of the 173 mammalian species analysed in the current work, the number of cysteine (a), methionine (b) and threonine (c) residues present in the 12 concatenated proteins encoded in the same mtDNA-strand, were computed and plotted against the number of times that the corresponding amino acid appeared in a random sequence with abundances for the four bases equal to those observed for the species under consideration. We divided the whole sample into two groups, with short-lived mammals (triangles) having log MLSP <2.46 and long-lived mammals (circles) having log MLSP >2.46, where 2.46 (corresponding to 228.4 months) is the median of log MLSP
The nearly null slope exhibited by cysteine suggests the existence of strong constraints that buffer changes in the abundance of cysteinyl residues. Furthermore, the low variability of the actual cysteine usage variable, particularly true within long-lived animals (Fig. 3a), also suggests the action of a strong purifying selection. Even more, the fact that all the points lie below the bisectrix, points to a strong selection against cysteine. However, no effect of longevity on the departure from neutral mutation could be detected for this amino acid (Fig. 4a). In other words, we found that although cysteinyl residues were actively avoided, there were no differences between short- or long-lived species. This observation contrasts with previous results reporting a negative relationship between cysteine usage and longevity, across a wide range of animals covering mammals, birds, reptiles, amphibians, fishes, insects, crustaceans, and arachnids (Moosmann and Behl 2008). Nonetheless, when the correlation analyses were focused on the class Mammalia after correction for the effects of phylogeny, no significant correlation between longevity and cysteine usage could be found (Aledo et al. 2011), which is line with the current observations (Fig. 4a). Thus, a note of caution should be sounded concerning a possible link between longevity and cysteine abundance in mtDNA-encoded proteins.
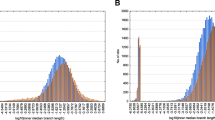
mtDNA-encoded proteins incorporate more methionyl residues than expected from the influence of nucleotide bias. The differences between the numbers of occurrences of given residue in the actual and random sequences were computed for each species. The distribution of such a variable, ΔAa, is shown in a. While ΔThr is distributed around zero indicating the absence of selective constraints, ΔCys and ΔMet departure from random in opposite directions. Since most mtDNA-encoded proteins use the AUG start codon, which encodes for methionine, we carried out the analyses either including (Met + start) or excluding (Met − start) start codons. b The number of species for which methionine frequency is higher in the actual sequence with respect to its random sequence, can be considered as a random variable, which after typification yield the so-called Z statistic. In the absence of constraints, the Z statistic follows a normal distribution (see technical details in the “Material and Methods” section). The calculated value of the Z statistic for methionine was 9.2 when start codons were included in the analysis, or 6.6 when these start codons were excluded. Thus, the probability of the observed departure of ΔMet from random happening by chance is less than 2 × 10−11. For comparative purposes, the Z statistic and p values for threonine were also calculated and indicated in the figure
On the other hand, threonine frequency showed a great variability that seemed to be well accounted by the nucleotide composition, as indicated by a high slope (Fig. 3c). In addition, the points are equidistantly distributed around the bisectrix. These results, taken all together, suggest that threonine usages in mtDNA-encoded proteins are shaped by mutational bias and the corresponding nucleotide composition of the coding genes. These observations, while not ruling out the adaptive character of increased threonine usages in long-lived mammals (Kitazoe et al. 2008), throw doubts on it.
With regard to methionine usages, we observed a behavior that was intermediate between that described for cysteinyl residues and that noted for threonyl residues. That is, although the response to nucleotide composition seemed to be less constrained than that observed for cysteine, a certain degree of inertia became evident when compared to the response of threonine frequencies. Nevertheless, the fact to be emphasized is that most species carried a higher number of methionyl residues into their proteins with respect to that expected from the influence of nucleotide bias (Fig. 3b). This conclusion was quantitatively confirmed by performing further analyses.
For a given amino acid, the differences between the numbers of occurrences in the actual and random sequences were computed for each species. These differences (ΔAa) can be interpreted as a measurement of the departure from neutral mutational effects. For instance, the distribution of ΔThr was centered around zero, indicating that threonine frequencies are mainly shaped by random forces with little constraints. In contrast, ΔCys took negative values for all the analyzed species without exception, which suggests a selection against this amino acid. On the other hand, ΔMet showed a distribution centered around 30, with only a few species exhibiting negative values, indicating a certain degree of positive selection favoring methionine incorporation into mitochondrial proteins (Fig. 4a).
To substantiate this claim, we next carried out a sign test. For this purpose, we defined a random variable as the number of ΔAa >0 occurrences. In this way, the null hypothesis states that in the absence of constraints, the typified random variable should follow a normal distribution, \( Z\,\sim \,N(0,\,1). \) While for threonine the null hypothesis was accepted (Z statistic = 0.0, p value = 0.5), it was clearly rejected for methionine (Z statistic = 9.2, p value = 2 × 10−20). Since most mtDNA-encoded proteins use the AUG start codon, which encodes for methionine, ΔMet may well be inflated because of this translational constraint. Therefore, we repeated the analyses, this time excluding start codons. As it can be seen in Fig. 4a, a non-negligible fraction of the described methionine excess could be accounted for the translational effect. Nevertheless, once this translational effect was discounted, the null hypothesis that the typified random variable Z follows a normal distribution centered at zero, was still rejected (Z = 6.6, p value = 2 × 10−11). These observations suggest that mtDNA-encoded proteins may incorporate more methionyl residues than expected from the neutral influence of nucleotide bias (Fig. 4b), which would be in line with the anti-oxidant role proposed for this amino acid (Levine et al. 1996; Bender et al. 2008; Aledo et al. 2011). Nevertheless, we acknowledge that further work and corroborating evidences will be required to fully support such a hypothesis.
A remarkable observation was that among the few species (21 species out of 173) that did not carry more methionine into their proteins than expected from random (those species that are distributed below the bisectrix from Fig. 3b), all, but one, belonged to the group of short-lived mammals (species with longevities below the median, log MLSP <2.46). Although we do not know the reasons behind this intriguing result, one is tempted to speculate that because the sense mtDNA-strand from long-lived mammals exhibit lower frequencies of T and higher frequencies of C (Fig. 1), which seems to favor the presence of the codons ACA and ACG (coding for threonine) at expenses of AUA and AUG (coding for methionine), long-lived animals are more in need of mechanisms, other than mutational bias, leading to increased methionine usages. To investigate this working hypothesis, we next addressed the correlation between ΔAa and longevity. As it can be observed in Fig. 5, there was a highly significant correlation between ΔMet and log MLSP (r = 0.456, p value = 3 × 10−10, n = 173). In contrast, ΔCys and ΔThr did not show any relationship with longevity (p values = 0.3 and 0.5, respectively).
The lack of an association of ΔThr with longevity is congruent with the conclusion that threonine usage is mainly determined by the nucleotide composition of the considered mitochondrial genome. On the other hand, the lack of correlation between ΔCys and longevity could simply be due to an inverse ceiling effect. To this respect, it should be noted that mtDNA-encoded proteins exhibit a remarkable cysteine avoidance when compared to their nDNA-encoded counterparts (relative frequencies around 0.6 and 1.7 %, respectively). Furthermore, evolutionary pressure towards reduced global cysteine usage cannot essentially affect functionally indispensable residues, such as those that bind the numerous iron–sulfur clusters of the respiratory chain. Therefore, it may be that current mtDNA-encoded proteomes have reached a limit beyond which is not possible any further lifespan-dependent cysteine depletion.
With respect to the highly significant correlation observed between ΔMet and longevity, this result is in agreement with the following hypothesis. If methionyl residues fulfill an anti-oxidant role, then increased methionine usages may confer an adaptive advantage for both short- and long-lived species. In this context, short-lived mammals that achieve high methionine frequencies through biased mutational processes do not experience the need to depart from randomness in the same extent as long-lived animals do.
In any event, the current results challenge the view of an active selection against methionine in long-lived animals. Previous authors have pointed out a pro-oxidant role for methionyl residues (Ruiz et al. 2005). They argue that the sensitivity of proteins to oxidative stress may increase as a function of the number of methionine residues and, consequently, a lower abundance of this amino acid in proteins from long-lived animals likely contributes to the superior longevity of these species. However, our results suggest that methionine residues are not selected against in the mitochondrial proteins of long-lived mammals; on the contrary, long-lived animals seem to actively add methionine residues into their mtDNA-encoded proteins. Furthermore, we have observed a positive relationship among longevity and the departure from random methionine content (Fig. 5b). It is important to realize that this gain in methionine residues, preferentially observed in long-lived animals, is relative to random sequences with the same nucleotide composition as the actual sequences. In fact, when methionine-adding events are analyzed regardless of the nucleotide composition of the coding genes, short-lived animals exhibit higher numbers of additions than their long-lived counterparts (Aledo et al. 2011). Thus, an integrated, though somewhat speculative, interpretation of all these data may be as follows.
Because of the anti-oxidant role of methionine, both short- and long-lived mammals may benefit from incorporating this amino acid into their proteins. However, the strategies followed to achieve increased methionine usages can differ from one species to other. In this respect, short-lived mammals exhibit biases in nucleotide composition of their genes favoring the incorporation of methionine into their mitochondrial proteins (Fig. 2b). Whether these favorable biases are driven by shorter generation times (de Magalhães et al. 2007) or higher mutation rates (Nabholz et al. 2008) with respect to their long-lived counterparts are, however, issues that remain unresolved. On the other hand, long-lived mammals possess mitochondrial genes with nucleotide abundances less prone to form methionine codons when randomly reorganized into triplets (Fig. 5). Thus, if incorporating methionine into mtDNA-encoded proteins has any beneficial effect on fighting oxidative stress, this group of long-lived animals may rely more heavily on post mutational mechanisms to increase methionine usages, and therefore departure from random expectation (Fig. 5b).
In a previous work, Jobson et al. (2010) failed to find a significant correlation between cysteine, methionine, and threonine composition and lifespan in 40 nDNA-encoded OXPHOS proteins. Herein, we have carried out the randomization analyses described above, using 61 nDNA-encoded OXPHOS genes from a dozen of mammalian species (Supplementary Table 1). As it can be deduced from Supplementary Fig. 1, while ΔMet >0 for all the analyzed species, ΔCys and ΔThr were negative in all the cases. These results may suggest that also in nDNA-encoded OXPHOS proteins methionyl residues are favored, while cysteine is avoided. However, we found no significant correlation between ΔAa and lifespan, regardless of the residue being considered. This lack of relationship between lifespan and ΔMet in nDNA-encoded proteins may be seen as a circumstantial evidence against a role of adaptation in shaping methionine content in mtDNA-encoded proteins. However, a note of caution should be made to this respect: because mtDNA-encoded proteins often evolve under different selective constraints to those of nDNA-encoded proteins. This point is well illustrated by the observation that, while nDNA-encoded residues in the interface of OXPHOS protein complexes are highly constrained, their mtDNA-encoded counterparts evolve even faster than other mtDNA-encoded residues (Schmidt et al. 2001).
In summary, methionine content in mtDNA-encoded proteins seems to be shaped by the contribution of two factors. On one hand, nucleotide composition bias brought about by a directional mutation bias is a significant driving force in methionine abundance. On the other hand, most species carry a higher number of methionyl residues into their proteins, with respect to that expected from the influence of nucleotide bias, suggesting the existence of selective forces favoring such outcome. Nevertheless, the circumstantiality of the evidences advices for caution.
Although there is a wide consensus that directional mutation bias may be related to the particular mode of replication of the mitochondrial chromosomes (Nikolaou and Almirantis 2006; Fonseca et al. 2008), the causes leading to the remarkable inter-special difference between mitochondrial genomes are largely unknown. Thus, the question of whether the mechanisms underlying this directional mutation bias are selectively neutral or not, remains an open issue that will provide a future challenge for molecular evolutionary biologists.
Conclusion
We have presented a comparative study of mitochondrial genomes across multiple species, aiming to address the long-standing question of whether the link between amino acid usages and longevity is due to an adaptive response and/or nucleotide mutation bias. We found that the nucleotide composition bias is the main, if not the only, driving force in threonine composition of the encoded proteins. In contrast, nucleotide composition has no effect at all on the cysteine usages, which seem to be kept at low values by purifying selection. With respect to methionine, the results suggest that both nucleotide bias and selective forces unrelated to the nucleotide composition, contribute to shape the methionine content in mtDNA-encoded proteins. Overall, our results demonstrate a role of selection in determining the composition of cysteine and methionine in the mitochondrial genome of mammals. Whether there is or not a link between the content of these sulfur-containing amino acids and the protection against oxidative stress, is an issue that remains open and deserves further attention.
References
Albu M, Min XJ, Hickey D, Golding B (2008) Uncorrected nucleotide bias in mtDNA can mimic the effects of positive Darwinian selection. Mol Biol Evol 25:2521–2524
Aledo JC, Li Y, de Magalhães JP, Ruíz-Camacho M, Pérez-Claros JA (2011) Mitochondrially encoded methionine is inversely related to longevity in mammals. Aging Cell 10:198–207
Balaban RS, Nemoto S, Finkel T (2005) Mitochondria, oxidants, and aging. Cell 120:483–495
Barja G (1998) Mitochondrial free radical production and aging in mammals and birds. Ann N Y Acad Sci 854:224–238
Bender A, Hajieva P, Moosmann B (2008) Adaptive antioxidant methionine accumulation in respiratory chain complexes explains the use of a deviant genetic code in mitochondria. Proc Natl Acad Sci USA 105:16496–16501
Berlett BS, Stadtman ER (1997) Protein oxidation in aging, disease, and oxidative stress. J Biol Chem 272:20313–20316
Brand MD (2000) Uncoupling to survive? The role of mitochondrial inefficiency in ageing. Exp Gerontol 35:811–820
Collins D, Jukes T (1993) Relationship between G + C in silent sites of codons and amino acid composition of human proteins. J Mol Evol 36:201–213
D’Onofrio G, Mouchiroud D, Aïssani B, Gautier C, Bernardi G (1991) Correlations between the compositional properties of human genes, codon usage, and amino acid composition of proteins. J Mol Evol 32:504–510
de Magalhães JP, Costa J (2009) A database of vertebrate longevity records and their relation to other life-history traits. J Evol Biol 22:1770–1774
de Magalhães JP, Costa J, Church GM (2007) An analysis of the relationship between metabolism, developmental schedules, and longevity using phylogenetic independent contrasts. J Gerontol A 62:149–160
Fonseca MM, Posada D, Harris DJ (2008) Inverted replication of vertebrate mitochondria. Mol Biol Evol 25:805–808
Foster PG, Jermiin LS, Hickey DA (1997) Nucleotide composition bias affects amino acid content in proteins coded by animal mitochondria. J Mol Evol 44:282–288
Harman D (2006) Free radical theory of aging: an update: increasing the functional life span. Ann N Y Acad Sci 1067:10–21
Jia W, Higgs PG (2008) Codon usage in mitochondrial genomes: distinguishing context-dependent mutation from translational selection. Mol Biol Evol 25:339–351
Jobson RW, Dehne-Garcia A, Galtier N (2010) Apparent longevity-related adaptation of mitochondrial amino acid content is due to nucleotide compositional shifts. Mitochondrion 10:540–547
Kitazoe Y, Kishino H, Hasegawa M, Nakajima N, Thorne JL, Tanaka M (2008) Adaptive threonine increase in transmembrane regions of mitochondrial proteins in higher primates. PLoS ONE 3:e3343
Kitazoe Y, Kishino H, Hasegawa M, Matsui A, Lane N, Tanaka M (2011) Stability of mitochondrial membrane proteins in terrestrial vertebrates predicts aerobic capacity and longevity. Genome Biol Evol 3:1233–1244
Lambert AJ, Boysen HM, Buckingham JA, Yang T, Podlutsky A, Austad SN, Kunz TH, Buffenstein R, Brand MD (2007) Low rates of hydrogen peroxide production by isolated heart mitochondria associate with long maximum lifespan in vertebrate homeotherms. Aging Cell 6:607–618
Lambert AJ, Buckingham JA, Boysen HM, Brand MD (2010) Low complex I content explains the low hydrogen peroxide production rate of heart mitochondria from the long-lived pigeon, Columba livia. Aging Cell 9:78–91
Levine RL, Mosoni L, Berlett BS, Stadtman ER (1996) Methionine residues as endogenous antioxidants in proteins. Proc Natl Acad Sci USA 93:15036–15040
Min XJ, Hickey DA (2007) DNA asymmetric strand bias affects the amino acid composition of mitochondrial proteins. DNA Res 14:201–206
Moosmann B, Behl C (2008) Mitochondrially encoded cysteine predicts animal lifespan. Aging Cell 7:32–46
Nabholz B, Glémin S, Galtier N (2008) Strong variations of mitochondrial mutation rate across mammals—the longevity hypothesis. Mol Biol Evol 25:120–130
Nikolaou C, Almirantis Y (2006) Deviations from Chargaff’s second parity rule in organellar DNA insights into the evolution of organellar genomes. Gene 381:34–41
Pamplona R (2011) Mitochondrial DNA damage and animal longevity: insights from comparative studies. J Aging Res 2011:807108
Pamplona R, Barja G (2011) An evolutionary comparative scan for longevity-related oxidative stress resistance mechanisms in homeotherms. Biogerontology 12:409–435
Pamplona R, Costantini D (2011) Molecular and structural antioxidant defenses against oxidative stress in animals. AJP Regul Integr Comp Physiol 301:R843–R863
Pamplona R, Portero-Otín M, Riba D, Ruíz C, Prat J, Bellmunt MJ, Barja G (1998) Mitochondrial membrane peroxidizability index is inversely related to maximum life span in mammals. J Lipid Res 39:1989–1994
Pamplona R, Portero-Otín M, Sanz A, Ayala V, Vasileva E, Barja G (2005) Protein and lipid oxidative damage and complex I content are lower in the brain of budgerigars and canaries than in mice. Relation to aging rate. Age 27:267–280
Ruiz MC, Ayala V, Portero-Otín M, Requena JR, Barja G, Pamplona R (2005) Protein methionine content and MDA-lysine adducts are inversely related to maximum life span in the heart of mammals. Mech Ageing Dev 126:1106–1114
Samuels DC (2005) Life span is related to the free energy of mitochondrial DNA. Mech Ageing Dev 126:1123–1129
Schmidt TR, Wu W, Goodman M, Grossman LI (2001) Evolution of nuclear- and mitochondrial-encoded subunit interaction in cytochrome c oxidase. Mol Biol Evol 18:563–569
Singer GA, Hickey DA (2000) Nucleotide bias causes a genomewide bias in the amino acid composition of proteins. Mol Biol Evol 17:1581–1588
Speakman JR, Talbot DA, Selman C, Snart S, McLaren JS, Redman P, Krol E, Jackson DM, Johnson MS, Brand MD (2004) Uncoupled and surviving: individual mice with high metabolism have greater mitochondrial uncoupling and live longer. Aging Cell 3:87–95
Stadtman ER, Van Remmen H, Richardson A, Wehr NB, Levine RL (2005) Methionine oxidation and aging. Biochim Biophys Acta 1703:135–140
Sueoka N (1961) Compositional correlation between deoxyribonucleic acid and protein. Cold Spring Harbor symposium on quantitative biology
Wang H-C, Singer GAC, Hickey DA (2004) Mutational bias affects protein evolution in flowering plants. Mol Biol Evol 21:90–96
Acknowledgments
Thanks to Miguel Ángel Medina and Alicia Esteban del Valle for critical reading of the manuscript. We would also like to thank Richard W. Jobson for assistance in collecting data related to nDNA-encoded genes. JCA gratefully acknowledges the support of Grant CGL2010-18124 from the Ministerio de Ciencia e Innovación, Spain. JPM is supported by the BBSRC, the Wellcome Trust, the Ellison Medical Foundation and a Marie Curie International Reintegration Grant within EC-FP7.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Aledo, J.C., Valverde, H. & de Magalhães, J.P. Mutational Bias Plays an Important Role in Shaping Longevity-Related Amino Acid Content in Mammalian mtDNA-Encoded Proteins. J Mol Evol 74, 332–341 (2012). https://doi.org/10.1007/s00239-012-9510-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-012-9510-7