Abstract
Little is known of intron sequences’ variation in cases where eukaryotic gene coding regions undergo strong balancing selection. Phosphoglucose isomerase, PGI, of Colias butterflies offers such a case. Its 11 introns include many point mutations, insertions, and deletions. This variation changes with intron position and length, and may leave little evidence of homology within introns except for their first and last few basepairs. Intron position is conserved between PGIs of Colias and the silkmoth, but no intron sequence homology remains. % GC content and length are functional properties of introns which can affect whole-gene transcription; we find a relationship between these properties which may indicate selection on transcription speed. Intragenic recombination is active in these introns, as in coding sequences. The small extent of linkage disequilibrium (LD) in the introns decays over a few hundred basepairs. Subsequences of Colias introns match subsequences of other introns, untranslated regions of cDNAs, and insect-related transposons and pathogens, showing that a diverse pool of sequence fragments is the source of intron contents via turnover due to deletion, recombination, and transposition. Like Colias PGI’s coding sequences, the introns evolve reticulately with little phylogenetic signal. Exceptions are coding-region allele clades defined by multiple amino acid variants in strong LD, whose introns are closely related but less so than their exons. Similarity of GC content between introns and flanking exons, lack of small introns despite mutational bias toward deletion, and findings already mentioned suggest constraining selection on introns, possibly balancing transcription performance against advantages of higher recombination rate conferred by intron length.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
“Spliceosomal” introns are non-coding intervening sequences commonly found in protein-coding genes of eukaryotes, which comprise a large part of their genomes (Doolittle 1978; Gilbert 1978). They are transcribed but later removed from precursor mRNA by spliceosome processing. At first, since they are not translated, introns were assumed to be non-functional, hence selectively neutral. In recent years, perception has changed, as some important regulatory functions for introns have been found (e.g., Shabalina and Spiridonov 2004; Belshaw and Bensasson 2006). Further, whole-genome scan studies have indicated that selection affects far more non-coding intronic and intergenic sequences than previously thought (Haddrill et al. 2008; Wright and Andolfatto 2008), so intron sequences are becoming important in studies of genome evolution.
Introns generally show higher levels of variation than coding sequences since they may have less evolutionary constraint and thus accumulate mutations at a faster rate (reviewed by Friesen 2000). For example, Feltus et al. (2006) showed that the polymorphism rate of rice introns is more than three times higher than that of the exons. As a result, intron polymorphism markers, including both length and single-nucleotide polymorphisms, have been used in genomic studies (e.g., Wang et al. 2006; Yang et al. 2007). However, intronic sequences can be quite conserved and may contain functionally constrained elements maintained by purifying selection (Casillas et al. 2007). In fact, a minimum of 20–30% of Drosophila’s non-coding sequences, including introns and intergenic regions, are highly conserved among other insects (Bergman and Kreitman 2001; Siepel et al. 2005).
Thus, diverse research has studied general patterns of intron properties and thoroughly explored the basic exon–intron structure of eukaryotic genes (Rogozin et al. 2005; Bradnam and Korf 2008; Zhu et al. 2009). However, there is little information about the evolution of intron diversity for genes whose coding regions display strongly selected natural polymorphism of amino acid (AA) sequence. Might such polymorphism exhibit strong interactions with variability of introns included in its gene?
The enzyme PGI (E.C. 5.3.1.9), which catalyzes the interconversion of glucose 6-phosphate and fructose 6-phosphate in glycolysis, is a case in point. It is polymorphic in many prokaryotes and eukaryotes, and its variants often have large genotypic functional and fitness differences (Gillespie 1991; Watt and Dean 2000). Specifically, large effects of PGI variants on enzyme function, thence organismal flight performance, and finally on all components of adult fitness, have been studied in Colias butterflies (e.g., Watt 2003). Emerging molecular perspectives on this variation (Wheat et al. 2006) offer a unique context for investigating PGI intron variation and evolution. A large database of Colias PGI coding haplotypes shows a remarkable degree of variability at both AA and nucleotide levels (Wang et al. 2009). Here, we study intron sequences from 31 PGI alleles of Colias eurytheme to investigate evolution of PGI introns and their interactions with the rest of the PGI gene system. We address these questions:
-
What is the natural genetic variation of Colias PGI introns? To what extent is there homology of sequence within introns?
-
Are there relationships between variation of PGI introns and of adjacent exon sequences?
-
What is the extent of intragenic recombination (IGR) within PGI introns, compared to its rate in coding regions? Is there linkage disequilibrium, LD, between polymorphic sites of these introns?
-
What are the sources of the variant sequences found in PGI introns?
-
What is the pattern of phyletic relationship among PGI introns? Is it consistent with that of PGI coding sequences?
-
Is there evidence of natural selection, based on either adaptation or constraint of functional properties, on the PGI introns? How does their variation interact with known selection or other evolutionary processes acting on PGI’s coding sequences?
Materials and Methods
Animals, Allozyme Genotyping, and Crossing Designs
Wild Colias eurytheme adults were sampled randomly from sites near Tracy, CA, and bred into our laboratory colony for study of their PGI introns. Colias were bred to isolate identical-by-descent (IBD) PGI alleles (see Wang et al. 2009 for details): a first set of alleles were sequenced from homozygous IBD larvae, and after that, individuals heterozygous with one IBD allele and one unknown allele each were sequenced to obtain the unknown alleles’ haplotypes. 31 allelic PGI copies were studied, many of whose coding regions were assessed by Wang et al. (2009), but their sampling remained roughly random as no selection for representation of particular alleles was done.
Genomic DNA Extraction, Amplification, and Sequencing
Larvae of interest were dissected and then kept at −80°C. Genomic DNA was extracted from 30 mg of stored larval tissue using DNeasy kits (Qiagen, Inc.). For each PGI allele, all 11 PGI introns were amplified separately from these DNA extracts using primers (Supplemental Table 1) designed using OLIGO 6 software (Molecular Biology Insights, Inc.). PCR amplifications were done in 50-μl reactions containing 1 unit of HiFi Platinum Taq polymerase (Invitrogen, Inc.), 2 mM MgSO4, 10 mM dNTPs, and 10 μM forward and reverse primers, and a cycle regime of 94°C 3′ 30″, 54°C 1′ 30″, 68°C 3′ 30″, then 35 cycles of 94°C 1′ 30″, 54°C 1′ 30″, 68°C 3′ 30″, with a final extension at 68°C for 10′. PCR products from IBD homozygote larvae were gel purified using QIAquick kits (Qiagen, Inc.), then cycle sequenced using ABI BigDye 3.1 chemistry and an ABI 377 sequencer. For heterozygotes, the two allele-specific PCR products, if different enough in size, were separated by agarose gel electrophoresis, then purified and sequenced as above. If alleles were not separable by size, we often could resolve the phases of the dual-peak sites, based on the known haplotype, when both alleles were short (<500 bp). Finally, to obtain the most “difficult” alleles, we cloned the mixed amplification products using TOPO TA kits (Invitrogen). Inserts were amplified using lysed bacteria as template along with the corresponding primers, and then gel purified and sequenced as above.
Data Analysis
Raw sequence data were processed with BioEdit 7.0 (Hall 2004). Each of the 11 PGI introns from all alleles studied was aligned separately using the ClustalW multiple alignment program (version 1.5, Thompson et al. 1994) and sometimes refined manually. Finalized sequences have been deposited in GenBank (Accession numbers HQ678106–678130 for intron 1 sequences, HQ698311–698337 for those coding sequences new for this study, and HQ717445–717717 for intron 2–11 sequences). For each intron, a representative subset of haplotypes reflecting the range of sequence variation was chosen as query sequences in BLAST searches for homologs in various sequence databases.
Evolutionary-genetic statistics of sequences were calculated with BioEdit or DnaSP 5.1 (Librado and Rozas 2009) and CLC Genomics Workbench (CLC bio A/S, Denmark). Indels were coded using the GapCoder program (Young and Healy 2003). To find relationships among sequence haplotypes, phylogenetic trees were constructed with PHYML (Guindon and Gascuel 2003) and PHYLIP software (Felsenstein 2005), with branch support inferred by bootstrapping (1,000 replications). Topologies of the trees were visualized with TREEVIEW (v. 1.6.6, Page 2001). For general statistical tests, data were first imported into Microsoft Excel for data sorting and manipulations, then processed in Statistica 8.0 (StatSoft Inc.) for testing and to make graphs. Sorting and plotting of LD data was done as by Wang et al. (2009).
Results
Description and Genetic Statistics of Colias PGI Intron Polymorphism
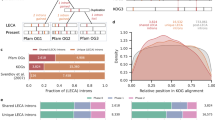
In Colias, the PGI gene, with 1,668 bp of cDNA, encodes a 556 AA, ~62 kDa enzyme monomer, but the enzyme is only functional as a dimer; the gene is autosomal and has 12 exons separated by 11 introns (Fig. 1; Wheat et al. 2006). On average, the 11 introns add 6,626 bp to the PGI gene’s size (Table 1). All C. eurytheme PGI introns begin with GT and end with AG, obeying the “GT–AG rule” which identifies the recognition sites for intron removal by spliceosomes. They also have relatively conserved short sequences at each end, as shown in Table 1; these are comparable to the consensus sequences at the intron–exon borders of vertebrates (Mount 1982).
Schematic representation of the genomic structure of the Colias PGI gene (to scale). Exons are represented as black boxes and introns as bridged gaps. Unfilled boxes are untranslated regions of the mRNA (UTRs). Lengths (in bp) of exons and introns (averaged over all alleles sequenced) are indicated above and below the diagram, respectively
Wheat et al. (2006) found initial evidence of length polymorphism in Colias PGI introns, and our data extend this finding (Table 1). Many insertions and deletions (indels), with sizes ranging from 1 bp to over 1 kb, are seen among the PGI introns; there are rarely two allelic copies of any intron with the same length. The length of the fully sequenced introns varies between 236 (intron 3 of alleles 2-38) and 2,446 bp (intron 1 of allele 4-A). There are also several introns longer than 2.5 kb (estimated by agarose gel electrophoresis, e.g., intron 1 of 3-277 and intron 10 of 4-18b), and only partial sequences were obtained from them; these were not analyzed further. (“Primer walking” would eventually yield full sequences, but this was not deemed critical to this study.) On average, intron 1 is the longest, and intron 7 is the shortest. Short introns have significantly smaller length variance than larger ones by Levene’s test for homogeneity of variance (Levene statistic = 2.785, P = 0.003, data scaled to equalize intron means). The introns’ extensive length variability contrasts to the exons’ narrow range of lengths (Fig. 1) and complete within-exon length uniformity.
These PGI introns also harbor much substitutional variation alongside their length polymorphism (see Fig. 2 for an example). With indels excluded, they display high overall nucleotide diversities π (and site diversities θ; Table 2); intron 8, with π = 0.1325, is even more variable than synonymous sites of PGI’s coding sequence (πss = 0.0993; Wang et al. 2009).
Multiple sequence alignment of intron 7 of the Colias PGI gene. DNA sequences of 31 haplotypes of the intron were aligned using the ClustalW program and manual adjustments. Only the first 100 bps are shown. Alleles were sorted by their names. Gaps introduced for optimal alignment are indicated by hyphens. Substantial amounts of both substitutional and length polymorphism can be seen from the alignment (see text for more details)
When intron 1, which has low nucleotide variability but high indel variability (Table 2), is excluded from the analysis, there is positive correlation between the nucleotide site variability and the indel variability of the introns (nucleotide θ vs. indel θ, correlation coefficient r = 0.616 and P = 0.03).
The ratio of nucleotide substitutions to indels (Chen et al. 2009) is given in Supplemental Table 2. On average, the Colias PGI introns are quite similar in this ratio to the average of introns of diverse genes of several other taxa, but both sides of the comparison show a wide range of values.
Comparison of PGI Introns Between Colias and the Silkmoth Bombyx
The PGI gene of the Bombyx genome assembly (gene BGIBMGA004221-TA; Duan et al. 2010) has the same coding region length as in Colias, 1,668 bp, but is even larger as a whole (15.2 kb) than in Colias (average 8.3 kb), owing to greater length of its introns, and its GC content (39.8%) is higher than in Colias (33.7%). No homology is seen between the one set of introns in the Bombyx genome assembly and the Colias introns studied here; their intron–exon junctions do not share any conserved sequences other than the GT–AG ends themselves. Thus, turnover of intron contents has been complete between these taxa, in contrast to 76.6% identity between coding-region nucleotide sequences, and 88.3% identity of AA sequences, of Bombyx PGI and Colias PGI (allele 4-1). But, the positions of introns in the PGI gene are exactly the same between Colias and Bombyx. This implies consistent selection to retain the 11 introns without loss, but without any preservation of sequence contents, at least as far back as the common ancestry of superfamilies Bombycoidea (bombycoid moths) and Papilionoidea (butterflies), thought to date from the Cretaceous/Tertiary boundary (Grimaldi and Engel 2005).
Pattern of Sequence Identity Among Alleles Within Introns
The extent of intron sequence homology or identity among Colias PGI alleles varies among introns. Introns 1, 2, and 7 are more conserved than others: in these cases, all sequenced alleles share a large fraction of homologous sequence (Fig. 3). Other introns (introns 3–6 and 8–11) naturally sort out during the ClustalW alignment process into two or three groups based on within-group homology, often sharing sequence only at their extreme ends. We also find singleton alleles which show more or less homology to one of the groups, but contain large, unique indels (marked in Fig. 3 as “U”).
A diagram showing patterns of sequence homology within each intron of different PGI alleles in Colias butterflies. For introns 1, 2, and 7, all alleles contain a large portion of aligned homologous sequences. Most other introns (Nos. 3, 4, 5, 6, 8, 10, and 11) can be divided into two distinct groups, each with alleles more homologous to those of the same group than those of the other group. Intron 9 has three distinct groups. Different groups are indicated by different graphic types of the boxes. Unique alleles, which may show some homology to one of the groups, are indicated by U inside boxes. Alleles that were not fully sequenced are indicated by question mark inside boxes. Note that graphic types of the boxes indicate homology among alleles of a particular intron (column), not among introns of a particular allele (row)
For introns with multiple groups, levels of sequence identity between groups vary from case to case (Table 3). For example, sequences from different groups of intron 9 or 11 show identity values significantly less than those expected from random sequences (Table 3 gives expectations based on average base frequencies for each intron). But for others such as introns 3 and 6, the sequence identities between alleles of different groups closely match random expectations. The most between-group similarity is seen in intron 5.
The grouping of alleles by sequence homology at one intron almost always breaks apart at some other intron (or introns) of the gene (cf. Fig. 3). For instance, even alleles 5-54 and 6-38, which are grouped together at introns 1, 2, and 3, belong to different groups at intron 4. Two groups of exceptions are discussed below. Setting these exceptions aside, the lack of covariation of groups among introns underscores the happenstance nature of processes—unequal recombination, transposable element dynamics, DNA replication errors, etc.—leading to intron content turnover.
Relation of Intron Variation to Coding Sequence Variation
We find no significant correlation for any comparison of introns’ substitution or indel variation to flanking exons’ values of nucleotide variation (π or θ, as summarized in Table 4), even with intron 1 excluded in the test (P > 0.05 for all correlation coefficients).
Figure 4 shows the patterns of average GC content of exons and introns at different ordinal positions along the Colias PGI gene. The overall GC content of the introns (28.8%) is much lower than that of the exons (48.2%). However, the variation patterns of GC content are similar between introns and flanking exons: the correlation coefficient between intron GC content and average GC content of the flanking exons is r = 0.582, P = 0.03.
The coding region AA polymorphs of Colias eurytheme PGI are organized into “charge macrostates” of the same relative charge (Wang et al. 2009), which correspond to “allozyme” allele categories. The most common of these, 3, displays only one “microstate,” or combination of charged AA side chains summing to the same macrostate charge, while macrostates 2, 4, and 5 have several microstates. Table 5 partitions the nucleotide diversities, π, of PGI introns according to the macrostate of their surrounding coding sequences. In 7 of 11 intron cases, the three-macrostate introns show the lowest π values among the four common macrostates—but still roughly track the intron-by-intron rise and fall of introns’ π averaged over the macrostates (Table 2). Thus, at most, the introns’ sequence contents only weakly reflect constraints on their “parent” coding sequences.
Possibly Functional Properties of Introns
Two intron properties, % GC content and length, may affect speed of transcription—% GC because there are three hydrogen bonds per GC pair compared to 2 per AT pair, slowing polymerase function (cf. Urrutia and Hurst 2003), and length due to the dependence of transcription time on length. Both these properties would alter transcription for the whole PGI gene, because on average introns comprise 80% of PGI gene length (above, Table 1). Among the 11 PGI introns, introns 2 and 9 have the highest % GC content, while introns 3 and 7 have the lowest (Figs. 4, 5). There is a positive relation between intron length and % GC content over the whole data set, and in many but not all cases intron-by-intron (Figs. 5, 6; Supplemental Table 3). But very long intron alleles (>1,480 bps) always have intermediate GC content (~30%). This agrees with the findings of Zhu et al. (2009) from study of a multi-gene, multi-genome data set. We propose a hypothesis to explain these findings:
Average intron length and GC content of introns at different ordinal positions along the Colias PGI gene. Error bars indicate standard deviation. Note that short introns (Nos. 3 and 7) have significantly smaller length variance than larger ones (F test, P < 0.001). They also have the lowest GC content
-
(a)
selection for rapid transcription favors shorter introns and lower % GC content;
-
(b)
relaxation of such selection, and/or occurrence of opposed selection favoring greater intron length (cf. below), allows persistence of longer introns with higher % GC content;
-
(c)
the longest intron insertions persist in any case only if they have moderate % GC, reducing time for their transcription.
IGR and LD in Introns
Wang et al. (2009), using the four-gamete test (Hudson and Kaplan 1985) to study PGI coding region haplotypes, found that each intron position marked a site of IGR. Using the same method, we find high rates of IGR within the major groups of each intron (Table 6): there are at least 3, and up to 13, recombination events per intron. These results more than double the total of recombination events in the whole PGI gene: 71 from the introns, and 45 from the exons, in haplotypes studied here. We compared introns’ numbers of recombinations to their lengths and nucleotide variation (π or θ), but found no significant correlation for any of the comparisons (P > 0.05 for all correlation coefficients), though the range of recombination event numbers in the three longest introns (1, 6, 9) is 6-13 as compared to the others, whose range is 3–8. Using DnaSP, we calculated LD as D (=pX 1pX 4 − pX 2pX 3, where pX i = frequency of the ith gamete) for all pairwise combinations of polymorphic sites across each intron. LD values and bootstrapped regression equations for LD versus site separation distances in sequence are reported in Table 7. For most introns, intercept values of LD are small, and there are negative regression slopes between LD and nucleotide distances, showing rapid LD decay over distance.
Sources of Intron Sequence Contents
The extensive variability of Colias’ PGI introns led us to ask what might be the sources of their sequences. We used the familiar BLASTN algorithm to compare sequences among Colias’ PGI introns, other Colias genes, and genes of Bombyx and other insects. Query-sequences included one representative from each major sequence group of each intron, plus each unique sequence intron by intron. In all cases we required a chance expectation E ≤ 10−12 to recognize a BLAST match.
The comparison set was first BLASTed against the general nucleotide data base of GenBank. Numbers of query-sequence matches to “target” sequences of other insects (mostly Lepidoptera) are listed in Table 8, and identities of the target sequences are listed in Supplemental Table 4. In a number of cases, different regions of an intron matched to different target sequences. Diverse gene targets were found; usually, in cases of identified protein coding genes, the Colias intron match was to introns and/or untranslated flanking regions rather than coding sequence. There were also a number of matches to transposons or to insect viral pathogens, e.g., nuclear polyhedrosis virus, or “bracoviruses,” viruses harbored by parasitoid wasps of the family Braconidae, which are symbiotic with these wasps in facilitating the attack of their larvae on targetted Lepidopteran larvae or other insect prey (e.g., Desjardins et al. 2007; Bézier et al. 2009).
To see if Colias intron sequences occur among presently expressed Colias mRNAs, we BLASTed our representative intron set against the sequence data base of a Colias expressed sequence tag library (prepared in collaboration with H. Vogel and C. W. Wheat, unpublished results). Numbers of matches are listed in Table 8. In a few cases of the highest match numbers, we then BLASTed the entire matching EST sequence against the Bombyx genome assembly (Duan et al. 2010), if possible to identify the gene in question and locate the intron-matching sequence in the gene’s structure. Three examples suffice to illustrate patterns found:
-
6 intron subsequences (* in Table 8) match to subsequences of EST contig 1145, and 3 of them also match to contigs 188, 1529, 1751, and 2309. A subset of contig 1145 matches the 3′-terminal coding sequence of a “hypothetical conserved B. mori protein” (BGIBMGA001660-TA), but none of the original intron subsequences are included in this match. None of the other four matched contigs show significant match to the B. mori genome (E > 0.1 in all cases).
-
4 intron subsequences (** in Table 8) match to subsequences of EST contigs 54, 3322, and 3600, and 2 of these also match to contig 2275. The matched subsequence of contig 54 matches the apparent penultimate exon of a Bombyx peptidase (peptidase C1, BGIBMGA009146-TA), and also the reverse complement of a predicted transposon sequence (BGIBMGA003628-TA). The other three contigs match the reverse complements of these Bombyx sequences in turn.
-
5 intron subsequences (*** in Table 8) match to subsequences of EST contig 2551, which includes a subsequence homologous to an entire B. mori “hypothetical conserved protein” coding region (BGIBMGA003243-TA, annotated as homologous to several unidentified protein sequences of other insects). None of the Colias intron subsequences are included in that B. mori coding region. However, 3 of the 5 subsequences do form part of the apparent 3′-untranslated region of the contig 2551 protein, two (6_3-2121 and 7_4-2521) as direct matches and one (9_4-4510) as a reverse-complement match. Remarkably, part of the matching 7_4-2521 sequence also matches overlapping subsequences identified in the original GenBank searches as related to Bombyx transposons (AB032718.1, AB126052.1) and bracovirus sequences (EF710633.1, EF710634.1).
Finally, we arbitrarily chose a subset of the PGI intron sequences (including at least one for each of the 11 introns) with matches in GenBank, whether first matched to viruses, transposons, or to other kinds of genes, to BLAST search for repetition of sequences among other Colias PGI introns not of the query sequence’s own group; e.g., for an allele sequence of intron 6, we made a data base for search of all our intron sequences except those of intron 6. Numbers of matches for the 21 sequences thus tested are listed in Table 8; they range from 0 to 8 with a mean of 2.81, which would be much higher if shorter sequences, thus with less stringent E values, were listed.
We conclude from all this that the high variability of PGI intron sequences draws on a common pool of fragmented sources which include pieces of native coding regions, transposons, and pathogens. These are presumably moved in and out of introns by deletion, recombination, or transposition, and show some repetitiveness of partial sequences among intron groups.
Are There Phylogenetic Relationships Among PGI Introns?
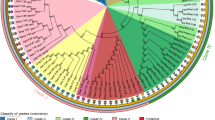
We first used PHYML (Guindon and Gascuel 2003) to evaluate relationships of the coding sequences studied here at the nucleotide level. As found earlier (Wang et al. 2009), there is little support for phylogenetic structure: relationships among these sequences are basically reticulate rather than coalescent (Fig. 7a), and as a result no deep nodes are supported by bootstrap testing. Notable exceptions, also as found earlier, are alleles from two groups of macrostate-4 haplotypes, characterized by significant LD among multiple AA variant combinations. These are named for the (single-letter-abbreviated) AAs defining them: DKMCS and DRVT. Two and three such alleles, respectively, group into shallow-node subclades just as they did with the coding region data including other such alleles earlier (Wang et al. 2009).
Radial (unrooted) plot of phylogenetic relationships estimated by maximum likelihood based on haplotypes’ DNA sequences. a Coding region, b intron 1, c intron 2, d intron 7 (see text for methods). Support for nodes of the phylogeny from 1,000 bootstrap iterations of the estimation is indicated by line width: thin no support at or below 60%, thick support between 60 and 80%, thicker support between 80 and 90%, thickest support between 90 and 100%. For multinode clades, the whole clade is keyed according to the support for the most basal significant (support ≥60%) node of the clade. The two macrostate-4 clades, DKMCS and DRVT, are labeled. Further details are given in the text
For the intronic parts of the gene, we could not analyze phylogeny over all introns because of the fragmentation of alleles into groups with little between-group homology within introns, and inconsistency of those groups among introns. At another level, there are not enough characters in the matrix of those grouping patterns (11 characters for 31 nodes; Fig. 3) on which to base a phylogeny. Therefore, separate analyses were done on sequences of each intron:
-
for introns 1, 2, and 7, which do not form two or more groups, we were able to use all fully sequenced alleles to test phylogenies. The tree topologies are shown in Fig. 7b–d.
-
for other introns, phylogenies were tested within each homologous group. Most clades with strong support at one intron were not supported at other introns (see Supplemental Table 5 for a list of all clades with significant bootstrap support in these analyses).
-
a few shallow clades (other than the AA LD clades) showed strong support in from two to as many as five consecutive introns (Supplemental Table 5). In these cases, the surrounding exons also were closely similar in sequence, suggesting that common blocks of two to several exons and intervening introns were moved by the copy-choice mechanism of IGR into two or more allele copies which we then sampled.
-
no clades were well supported across all the introns except for members of the AA LD clades DKMCS and DRVT. The two DKMCS alleles studied here, 4-66 and 4-37, have almost identical sequences for all 11 introns, and therefore always form a highly supported clade. The same situation is seen between two of the three DRVT alleles, 4-418 and 4-107a, but introns of the third DRVT allele, 4-197, are quite different, never grouping in the same clade for introns 2-10 (for intron 1, 4-197 was not fully sequenced due to amplification failure). The rejection of recombinant AA subset combinations of these sequences by natural selection (Wang et al. 2009) thus extends, but not entirely, to reducing the variation of accompanying introns.
Discussion
Differences Among Introns by Position in Gene
The first intron of a eukaryotic gene is often longer than all downstream introns, as seen in a variety of eukaryotic species (Bradnam and Korf 2008; Zhu et al. 2009). We have found this in Colias, in contrast to the short intron 1 of the Bombyx genome (above). Two hypotheses have been proposed to explain this pattern. First, since introns from the 5′ end of a gene are thought to be “early” introns, they may have had more time to accumulate “junk” DNA (Bradnam and Korf 2008; Zhu et al. 2009). But, our findings of the diversity and apparent rapid turnover, rather than slow accumulation, of intron contents are not consistent with this hypothesis. Another hypothesis for a longer first intron is that it contains more functional elements involved in controlling gene expression. The first introns of eukaryotic genes are indeed often enriched with regulatory elements (Sakurai et al. 2002; Gazave et al. 2007). If so, first introns might be selectively constrained and thus less variable. Comparisons in a variety of vertebrates and Drosophila have shown that first introns are more conserved than later introns (e.g., Marais et al. 2005; Vinogradov 2006). However, Colias PGI’s intron 1, while showing the fourth lowest nucleotide diversity π among the 11 introns (Table 2), has the largest indel diversity and the largest range of length variation of any of the introns. Regulatory functions of intron 1 in Colias PGI, if any, are unclear from study of its variation patterns.
For introns other than the first, we found a significant positive correlation between indel and nucleotide diversities, as seen elsewhere (e.g., Brandström and Ellegren 2007; Zhang et al. 2008). Some Colias PGI introns, e.g., 3, 4, and 5, showing lower diversity of both nucleotides and indels, may be under stronger constraints than others, but it is unclear what these constraints might be. Intron sequences close to the flanking exons are usually more conserved than those in the interior (Jareborg et al. 1999; Hare and Palumbi 2003), which is apparent in our data: in many cases homology between alleles of different groups occurs only very close to the ends. These conserved end-sequences, besides the end dinucleotides GT/AG, may support normal splicing (Burset et al. 2000).
Intron Diversity in Comparative Context
As seen above, the sequence contents of Colias’ PGI introns comprise a wide range of sources, including pieces homologous to subsequences of transposons, of pathogenic viruses, and potentially antisense fragments of “native” protein-coding genes, and clearly “stirred” by spontaneous point mutation, insertion/deletion events which may involve transposable elements, and recombination. It is thus no surprise that PGI introns of Colias and Bombyx have diverged completely in content during tens of millions of years. Moreover, Bombyx mori has undergone 5,000 years of domestication; its adults are flightless and it is dependent on humans for persistence. Thus, it has likely been the subject to initial founder effect and repeated population bottlenecking, maximizing the speed of genetic drift.
The Bombyx genome displays a high level of repetitive sequences, many of which belong to transposable elements (Osanai-Futahashi et al. 2008). Such elements are important players in genome evolution by altering genome structure (Nekrutenko and Li 2001). They contribute to genetic diversity via both insertion site polymorphism and small structural rearrangements (Bennetzen 2000). We found at least traces of these mobile DNA elements both in the PGI introns and in other parts of the Colias transcriptome as revealed by BLAST searches. It will be of interest to study further the dynamics of transposable elements in Colias’ genome evolution.
Recombination and Its Implications in Introns
Recombination may be a common driver of indel formation in non-coding parts of genomes. Carvalho and Clark (1999) found longer introns in regions of low recombination in Drosophila, but this effect was only seen in introns of <80 bp, much shorter than any found in Colias PGI (cf. also Duret 2001). Recombination rates are positively correlated with intron indel density in the chicken genome (Rao et al. 2010). We do not find significant differences in recombination rates among Colias PGI introns, but the longest introns do have a larger range of recombinant event numbers (above).
Watt (1972) showed that IGR could generate new allelic combinations of substitutions at rates far higher than these alleles could arise by primary mutation. Wang et al. (2009) found that IGR was very active in coding regions of Colias PGI, causing mostly reticulate evolution among the haplotypes, with at least one recombination event at each intron position. Studying the introns themselves finds higher levels of IGR, with at least three events per intron and as many as 13 (above). Thus, the introns of the PGI gene increase the potential for recombinative production of new coding alleles even more than recognized by Wang et al. (2009).
Tradeoffs of Selection on Intron Properties?
Comeron and Kreitman (2000) hypothesized selection to maintain intron length to foster recombination, in the face of apparent mutational biases toward deletion events, hence shorter introns. Carvalho and Clark (1999) found a negative correlation in Drosophila between intron length and recombination rate, but no correlation of this kind among “large” (>80 bp) introns. The Colias PGI introns are all much larger than this, with the shortest specimen being a 236 bp Intron 3 (Table 1). It cannot be argued, given the extensive indel variation and extent of recombination found here, that deletion variants are not available to selection for shorter introns if that occurs, e.g., by greater transcription efficiency or splicing accuracy (Lynch 2002). Increases in IGR across the PGI gene as a whole, due to lengthening of the gene by its large introns, would certainly oppose limitations, with respect to any form of selection on PGI, by the “Hill–Robertson effect” (the mutual interference between selection on nearby sites as a result of local LD; cf. Comeron and Kreitman 2000, Duret 2001). Further and in a positive sense, such increases might speed processes of adaptive refinement by interaction of balancing selection and recombination, as we have already suggested in the case of PGI coding sequence variation (Wang et al. 2009).
To this putative balance of pressures we now add the hypothesis of selection for transcription speed, putatively reducing intron % GC and intron length and producing the correlation of these variables found above. This hypothesis is consistent with the lower value of % GC among all introns as compared to their flanking exons, as well as the lower % GC of the longest intron inserts (above). None of the putative selection pressures hypothetically interacting here can be at a comparable order of strength to those known to act on the coding regions. But their interaction, especially the apparent tradeoff between intron length’s increase of recombination and transcription speed’s constraints on intron length, has potential to shape the evolving refinement of complex adaptive genetic combinations, e.g., the AA LD clades DKMCS and DRVT, by interaction of selection and IGR (Wang et al. 2009). Of course, even in the case of these two clades, we find evidence of more rapid turnover and divergence on the part of intron contents than of the synonymous variants in the coding sequences (cf. Wang et al. 2009).
Intron Subgroups and Intron Turnover Dynamics
As seen above, introns 3–6 and 8–11 divide naturally in the alignment process into two or three sequence groups with little inter-relatedness except at their very ends. This may well reflect features of the processes turning over PGI’s intron contents. We note that in general insertions into these introns are longer than deletions from them—e.g., in four introns of varying length (Table 1), average lengths of insertions versus deletions in bp are: intron 1, 127.8 (n = 33) versus 17.2 (n = 30); intron 3, 40.6 (n = 16) versus 16.6 (n = 14); intron 7, 32.4 (n = 16) versus 8.3 (n = 16); intron 10, 71.7 (n = 11) versus 6.6 (n = 11), and similarly for the others. The difference is largely due to a few very long insertions (e.g., in intron 1, two of 1,535 and 1,987 bp and an even longer unsequenced one estimated by size in agarose electrophoresis at ~2,800 bp). Perhaps a balance between such occasional long insertions and erosion of intron contents by equally haphazard smaller deletions tends to lead to the observed subdivided intron groups as a common intermediate stage in the neutrally drifting turnover of intron contents, though constrained by, e.g., selection for PGI transcription speed as already noted.
Evolutionary Interactions of PGI Introns and Their Surrounding Coding Sequences
There seems to be no limitation on the dynamics of Colias intron contents, either by natural variation in the introns themselves or by the range of processes available to rearrange this variation. However, the variation does not extend to intron positions within the coding sequences, which we found to be exactly the same across superfamilies (Bombycoidea, Papilionoidea) of higher Lepidoptera. Besides maintenance of existing introns by selection favoring recombination, this stability of intron position may result from severe fitness consequences of coding sequence alterations accompanying position changes, which risk change in the protein’s reading frame or the number and thus alignment of AAs in protein primary sequence (e.g., Lynch 2002).
Commonality of properties between the introns and their flanking exons seems largely limited to the correlation of their % GC values noted above. For coding sequences, GC-content heterogeneity may result from selectively biased codon usage. Highly expressed genes may be selected for use of codons with less GC (e.g., Hambuch and Parsch 2005; Kotlar and Lavner 2006), again perhaps reflecting pressure for transcription speed. Possible causes of the exon–intron correlation might then include biased repair of mismatches during recombination (Genereux 2002), or issues affecting local stability of mRNA prior to spliceosome processing (Chamary and Hurst 2004). There might also be effects of % GC on secondary structure of the DNA itself and its packing interactions with chromosomal proteins.
Not only did we find no correlation between nucleotide diversities, π, of introns and their flanking exons but also intron π values are seldom quite as high as those of synonymous sites in the coding regions (cf. Wheat et al. 2006, Wang et al. 2009). This may be explained by the fact that the slopes of decay of LD with site separation distance in the introns are, with the exceptions of introns 1 and 2, from 2- to 30-fold larger (Table 7) than those for coding region synonymous sites (Table 3 of Wang et al. 2009). This probably reflects the high rate of IGR seen in the introns, and thus the faster decay of “hitchhiking” of neutral intron variation with the AA variants maintained at multiple sites by balancing selection in the exons. It also emphasizes by contrast the strength of maintenance of LD, in the coding region LD clades, among AA variant sites in distant parts of the primary protein sequence.
On present evidence, then, the evolution of Colias PGI introns is a process of rapidly turning over and diverging sequences, basically neutral but hypothetically channeled by selection based on molecular-genetic functional constraints and/or by adaptive considerations revolving around effects of IGR on the chronic polymorphism of the gene’s coding region. Further study may allow testing for regulatory functions (especially of intron 1) and should clarify further the balance among adaptive, constrained, and neutral variation in these non-coding genetic elements.
References
Batley J, Barker G, O’Sullivan H, Edwards KJ, Edwards D (2003) Mining for single nucleotide polymorphisms and insertions/deletions in maize expressed sequence. Plant Physiol 132:84–91
Belshaw R, Bensasson D (2006) The rise and falls of introns. Heredity 96:208–213
Bennetzen JL (2000) Transposable element contributions to plant gene and genome evolution. Plant Mol Biol 42:251–269
Berger J, Suzuki T, Senti K, Stubbs J, Schaffner G, Dickson BJ (2001) Genetic mapping with SNP markers in Drosophila. Nat Genet 29:475–481
Bergman CM, Kreitman M (2001) Analysis of conserved noncoding DNA in Drosophila reveals similar constraints in intergenic and intronic sequences. Genome Res 11:1335–1345
Bézier A, Annaheim M, Herbinière J, Wetterwald C, Gyapay G, Bernard-Samain S, Wincker P, Roditi I, Heller M, Belghazi M, Pfister-Wilhem R, Periquet G, Dupuy C, Huguet E, Volkoff A-N, Lanzrein B, Drezen J-M (2009) Polydnaviruses of braconid wasps derive from an ancestral nudivirus. Science 323:926–930
Bradnam KR, Korf I (2008) Longer first introns are a general property of eukaryotic gene structure. PLoS ONE 3:e3093. doi:10.1371/journal.pone.0003093
Brandström M, Ellegren H (2007) The genomic landscape of short insertion and deletion polymorphisms in the chicken (Gallus gallus) genome: a high frequency of deletions in tandem duplicates. Genetics 176:1691–1701
Burset M, Seledtsov IA, Solovyev VV (2000) Analysis of canonical and non-canonical splice sites in mammalian genomes. Nucleic Acids Res 28:4364–4375
Carvalho AB, Clark AG (1999) Intron size and natural selection. Nature 401:344
Casillas S, Barbadilla A, Bergman CM (2007) Purifying selection maintains highly conserved noncoding sequences in Drosophila. Mol Biol Evol 24:2222–2234
Chamary JV, Hurst LD (2004) Similar rates but different modes of sequence evolution in introns and at exonic silent sites in rodents: evidence for selectively driven codon usage. Mol Biol Evol 21:1014–1023
Chen J-Q, Wu Y, Yang H, Bergelson J, Kreitman M, Tian D (2009) Variation in the ratio of nucleotide substitution and indel rates across genomes in mammals and bacteria. Mol Biol Evol 26:1523–1531
Comeron JM, Kreitman M (2000) The correlation between intron length and recombination in Drosophila: dynamic equilibrium between mutational and selective forces. Genetics 156:1175–1190
Desjardins CA, Gundersen-Rindal DE, Hostetler JB, Tallon LJ, Fuester RW, Schatz MC, Pedroni MJ, Fadrosh DW, Haas BJ, Toms BS, Chen D, Nene V (2007) Structure and evolution of a proviral locus of Glyptapanteles indiensis bracovirus. BMC Microbiol 7:61
Doolittle WF (1978) Genes in pieces: Were they ever together? Nature 272:581–582
Duan J, Li R, Cheng D, Fan W, Zhu X, Cheng T, Wu Y, Wang J, Mita K, Xiang Z, Xia Q (2010) SilkDB 2.0: a platform for silkworm (Bombyx mori) genome biology. Nucl Acids Res 38:D453–D456. http://silkworm.genomics.org.cn
Duret L (2001) Why do genes have introns? Recombination might add a new piece to the puzzle. Trends Genet 17:172–175
Felsenstein J (2005) PHYLogeny Inference Package, v. 3.63. http://evolution.gs.washington.edu/phylip.html
Feltus FA, Singh HP, Lohithaswa HC, Schulze SR, Silva TD, Paterson AH (2006) A comparative genomics strategy for targeted discovery of single-nucleotide polymorphisms and conserved-noncoding sequences in orphan crops. Plant Physiol 140:1183–1191
Friesen VL (2000) Introns. In: Baker AJ (ed) Molecular methods in ecology. Blackwell, Oxford, pp 274–294
Gazave E, Marqués-Bonet T, Fernando O, Charlesworth B, Navarro A (2007) Patterns and rates of intron divergence between humans and chimpanzees. Genome Biol 8:R21
Genereux DP (2002) Evolution of genomic GC variation. Genome Biology 3:reports0058
Gilbert W (1978) Why genes in pieces? Nature 271:501
Gillespie JH (1991) The causes of molecular evolution. Oxford University Press, New York
Grimaldi D, Engel M (2005) Evolution of the insects. Cambridge University Press, Cambridge
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704. http://atgc.lirmm.fr/phyml/
Haddrill PR, Bachtrog D, Andolfatto P (2008) Positive and negative selection on noncoding DNA in Drosophila simulans. Mol Biol Evol 25:1825–1834
Hall T (2004) Bioedit: biological sequence alignment editor. http://www.mbio.ncsu.edu/BioEdit/bioedit.html
Hambuch TM, Parsch J (2005) Patterns of synonymous codon usage in Drosophila melanogaster genes with sex-biased expression. Genetics 170:1691–1700
Hare MP, Palumbi SR (2003) High intron sequence conservation across three mammalian orders suggests functional constraints. Mol Biol Evol 20:969–978
Hudson R, Kaplan N (1985) Statistical properties of the number of recombination events in the history of a sample of DNA sequences. Genetics 111:147–164
Jareborg N, Birney E, Durbin R (1999) Comparative analysis of noncoding regions of 77 orthologous mouse and human gene pairs. Genome Res 9:815–824
Kotlar D, Lavner Y (2006) The action of selection on codon bias in humans is related to frequency, complexity and chronology of amino acids. BMC Genomics 7:67
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Lynch M (2002) Intron evolution as a population-genetic process. Proc Natl Acad Sci USA 99:6118–6123
Marais G, Nouvellet P, Keightley PD, Charlesworth B (2005) Intron size and exon evolution in Drosophila. Genetics 170:481–485
Morton BR (1993) Chloroplast DNA codon use: evidence for selection at the psb A locus based on tRNA availability. J Mol Evol 37:273–280
Mount SM (1982) A catalogue of splice junction sequences. Nucleic Acids Res 10:459–472
Nekrutenko A, Li WH (2001) Transposable elements are found in a large number of human protein-coding genes. Trends Genet 17:619–621
Osanai-Futahashi M, Suetsugu Y, Mita K, Fujiwara H (2008) Genome-wide screening and characterization of transposable elements and their distribution analysis in the silkworm, Bombyx mori. Insect Biochem Mol Biol 38:1046–1057
Page RDM (2001) TREEVIEW: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Rao YS, Wang ZF, Chai XW, Wu GZ, Nie QH, Zhang XQ (2010) Indel segregating within introns in the chicken genome are positively correlated with the recombination rates. Hereditas 147:53–57
Rogozin IB, Sverdlov AV, Babenko VN, Koonin EV (2005) Analysis of evolution of exon-intron structure of eukaryotic genes. Brief Bioinformatics 6:118–134
Sakurai A, Fujimori S, Kochiwa H, Kitamura-Abe S, Washio T, Saito R, Carninci P, Hayashizaki Y, Tomita M (2002) On biased distribution of introns in various eukaryotes. Gene 300:89–95
Schaeffer SW (2002) Molecular population genetics of sequence length diversity in the Adh region of Drosophila pseudoobscura. Genet Res 80:163–175
Shabalina SA, Spiridonov NA (2004) The mammalian transcriptome and the function of non-coding DNA sequences. Genome Biol 5:105
Siepel A, Bejerano G, Pedersen JS, Hinrichs A, Hou M, Rosenbloom K, Clawson H, Spieth J, Hillier LW, Richards S, Weinstock GM, Wilson RK, Gibbs RA, Kent WJ, Miller W, Haussler D (2005) Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res 15:1034–1050
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting position-specific gap penalties and weight matrix choice. Nucleic Acid Res 22:4673–4680
Urrutia AO, Hurst LD (2003) The signature of selection mediated by expression on human genes. Genome Res 13:2260–2264
Vinogradov AE (2006) “Genome design” model: evidence from conserved intronic sequence in human-mouse comparison. Genome Res 16:347–354
Wang BQ, Watt WB, Aakre C, Hawthorne N (2009) Emergence of complex haplotypes from microevolutionary variation in sequence and structure of Colias phosphoglucose isomerase. J Mol Evol 68:433–447
Wang X, Zhao X, Zhu J, Wu W (2006) Genome-wide Investigation of intron length polymorphisms and their potential as molecular markers in rice (Oryza sativa L.). DNA Res 12:417–427
Watt WB (1972) Intragenic recombination as a source of population genetic variability. Am Nat 106:737–753
Watt WB (2003) Mechanistic studies of butterfly adaptations. In: Boggs CL, Watt WB, Ehrlich PR (eds) Butterflies: ecology and evolution taking flight. University of Chicago Press, Chicago, pp 319–352
Watt WB, Dean AM (2000) Molecular-functional studies of adaptive genetic variation in prokaryotes and eukaryotes. Ann Rev Genet 34:593–622
Wheat CW, Watt WB, Pollock DD, Schulte PM (2006) From DNA to fitness differences: sequences and structures of adaptive variants of Colias phosphoglucose isomerase (PGI). Mol Biol Evol 23:499–512
Wright SI, Andolfatto P (2008) The impact of natural selection on the genome: emerging patterns in Drosophila and Arabidopsis. Annu Rev Ecol Evol Syst 39:193–213
Yang L, Jin G, Zhao X, Zheng Y, Xu Z, Wu W (2007) PIP: a database of potential intron polymorphism markers. Bioinformatics 23:2174–2177
Young ND, Healy J (2003) GapCoder automates the use of indel characters in phylogenetic analysis. BMC Bioinformatics 4:6
Zhang W, Sun X, Yuan H, Araki H, Wang J, Tian D (2008) The pattern of insertion/deletion polymorphism in Arabidopsis thaliana. Mol Genet Genomics 280:351–361
Zhu L, Zhang Y, Zhang W, Yang S, Chen J-Q, Tian D (2009) Patterns of exon–intron architecture variation of genes in eukaryotic genomes. BMC Genomics 10:47–58
Acknowledgments
We thank Carol Boggs, Mike Bramson, Jason Hill, Jen Johnson, Martin Kreitman, Mark Longo, Dmitri Petrov, Steve Palumbi, and Chris Wheat for comments on the paper or other helpful discussions. We also thank Chris Aakre, Will Bassett, Nina Duong, Daniel Herrador, Alejandro Perez, and Eddie Wang for technical assistance. This work was supported by US National Science Foundation grants DEB 05-20315 and MCB 08-46870 to WBW. Our results do not represent official policy of any agency or corporate entity.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, B., Mason DePasse, J. & Watt, W.B. Evolutionary Genomics of Colias Phosphoglucose Isomerase (PGI) Introns. J Mol Evol 74, 96–111 (2012). https://doi.org/10.1007/s00239-012-9492-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-012-9492-5