Abstract
Gene duplications are one of the most important mechanisms for the origin of evolutionary novelties. Even though various models of the fate of duplicated genes have been established, current knowledge about the role of divergent selection after gene duplication is rather limited. In this study, we analyzed sequence divergence in response to neo- and subfunctionalization of segmentally duplicated genes in the genome of Arabidopsis thaliana. We compared the genomes of A. thaliana and the poplar Populus trichocarpa to identify orthologous pairs of genes and their corresponding inparalogs. Maximum-likelihood analyses of the nonsynonymous and synonymous substitution rate ratio \( \left( {\omega = d_{\rm{N}} /d_{\rm{S}} } \right) \) of pairs of A. thaliana inparalogs were used to detect differences in the evolutionary rates of protein coding sequences. We analyzed 1,924 A. thaliana paralogous pairs and our results indicate that around 6.9% show divergent ω values between the lineages for a fraction of sites. We observe an enrichment of regulatory sequences, a reduced level of co-expression and an increased number of substitutions that can be attributed to positive selection based on an McDonald–Kreitman type of analysis. Taken together, these results show that selection after duplication contributes substantially to gene novelties and hence functional divergence in plants.
Similar content being viewed by others

Avoid common mistakes on your manuscript.
Introduction
Gene duplications are one of the most important mechanisms for the origin of evolutionary novelties (Ohno 1970). Duplicated genes are observed from deep to recent levels of evolutionary divergence indicating that gene duplications occurred throughout evolutionary history. There are different types of gene duplications: whole genome, segmental, and tandem duplications. Whole genome duplications occured in yeast, vertebrates, ciliates, and plants (Aury et al. 2006; Scannell et al. 2007; Tang et al. 2008; Kuraku et al. 2009). Tandem gene duplications were observed in nearly every species whose genome was sequenced so far. After duplication, possible evolutionary fates of paralogous genes are nonfunctionalization or neofunctionalization of one duplicate, or subfunctionalization of both copies (Force et al. 1999; Lynch and Force 2000). A complete redundancy over long evolutionary time periods seems unlikely because mutational pressure will ultimately lead to the nonfunctionalization of one of the two genes (Clark 1994; Lynch et al. 2001; O’Hely 2006).
Plant genomes contain high frequencies of duplicated genes (AG Initiative 2000; IRGS Project 2005; Tuskan et al. 2006; Jaillon et al. 2007). One of the most important mechanisms is polyploidization, which has affected more than 50% of all plant species (Blanc and Wolfe 2004b; Soltis and Soltis 2009). In the lineage leading to Arabidopsis thaliana, at least two whole genome duplications occurred at ≈40 million years ago (mya), and probably more than 200 mya (Blanc et al. 2003; Raes et al. 2003). In addition, plant genomes contain a high proportion of duplicated genes that arose by tandem duplications (AG Initiative 2000; Rizzon et al. 2006). Although the basic patterns of genome duplications at different evolutionary levels are now established, little is known about the role of natural selection in the subsequent fate of genes (Hahn 2009). Previous studies of gene duplications in A. thaliana, rice, and other plant species showed that positive selection can drive sequence divergence of both segmentally and tandemly duplicated genes. Examples include pollen-specific oleosins (Schein et al. 2004; Fiebig et al. 2004), genes involved in defense-related secondary metabolism (Benderoth et al. 2006; Mita et al. 2006), and disease resistance genes (Mondragón-Palomino et al. 2002; Kuang et al. 2004; Sun et al. 2006).
Functional analyses and expression patterns suggest that functional divergence of duplicated genes is common. In Arabidopsis, 57% of recent and 73% of older duplicates show divergent expression patterns (Blanc and Wolfe 2004a). Between 31.6 and 85% of pairs of paralogous Arabidopsis genes differ in their tissue-specific expression patterns (Duarte et al. 2006). However, neither functional divergence nor divergence in expression pattern are sufficient to distinguish between neo- and subfunctionalization, unless the ancestral state of expression is known (Lynch and Conery 2000). When the ancestral state of expression was taken into account, only a few of the paralogous pairs were diverged in a way that was fully consistent with either a classic subfunctionalization or neofunctionalization model (Duarte et al. 2006). The majority of duplicated genes apparently underwent both neo- and subfunctionalization (He and Zhang 2005; Rastogi and Liberles 2005).
In the present study, we analyzed rates of sequence evolution to estimate the importance of selection in the divergence of pairs of paralogous genes in A. thaliana. Spillane et al. (2007) described the evolution of the imprinted MEDEA (MEA) gene, which originated by a recent genome duplication and acquired new functions during embryo development. Furthermore, it showed a strong signal of positive Darwinian selection during this period, whereas the sequence and function of its paralog SWINGER (SWN) remained highly conserved. SWN also showed a high level of genetic redundancy with its common ancestor gene, CURLY LEAF (CLF). These results suggested a neofunctionalization of MEA gene but not of its paralog SWN. Here, we analyze how frequently a similar neo- and subfunctionalization of duplicated paralogs can be observed on a genome-wide level. We identify pairs of duplicated genes that were either duplicated as a result of the two whole genome duplications in the past of A. thaliana, or which arose from tandem duplications. By using orthologous genes from the poplar genome as outgroups, we calculated lineage-specific rates of evolution and conducted tests of selection. We found that about 6.9% of A. thaliana paralogous gene pairs exhibit significantly different rates of sequence divergence between duplicated genes.
Materials and Methods
Sequence Data
The genome of A. thaliana was obtained from MIPS (ftp://ftpmips.gsf.de/cress) and TAIR (genomes release 6, ftp://ftp.arabidopsis.org). The Populus trichocarpa genome (version 1.1) was obtained from JGI (http://genome.jpi-psf.org/Poptr1_1/). The segmentally duplicated A. thaliana gene clusters as determined by Blanc et al. (2003) were downloaded from http://wolfe.gen.tcd.ie/athal/all_results. This dataset contains 3,044 pairs of genes in 91 distinct blocks, of which 3,041 genes were consistent with recent genome annotations. The Oryza sativa genome (release 5.0) was obtained from http://rice.tigr.org and the Saccharomyces cerevisiae genome from http://www.yeastgenome.org.
Determining Paralogous and Orthologous Relationships
We follow the established nomenclature for differentiating between orthologs and paralogs (Koonin 2005). Paralogs whose origin predates a speciation event are called outparalogs; they may be misidentified as orthologs if different paralogs are deleted in different lines. Inparalogs originated after a speciation event and are specific to a particular lineage. To identify clusters of inparalogs in the A. thaliana and P. trichocarpa genomes, INPARANOID (version 2.0) was used (Remm et al. 2001), because this program performs well in ortholog classification with a sensitivity and specificity >80% (Chen et al. 2007a). The INPARANOID algorithm identifies inparalogs from two species using BLAST (Altschul et al. 1990) similarity scores between pairs of sequences. The two-way best hits of genes between species are considered as seed orthologs and form a cluster, potential inparalogs are successively added to this seed pair. A BLAST-based clustering assumes equal evolutionary rates among paralogs (Li et al. 2003), but differential levels of selection or differences in the mutation rate among inparalogs may lead to unequal rates. Hence, we changed the default parameters of INPARANOID to allow the inclusion of more divergent inparalogs. The BLAST score cutoff was raised from 50 to 100, which reduces the number of pairwise comparisons used in the clustering step. We also lowered the confidence level for inclusion of inparalogs from 0.5 to −0.5, which increases the number of potential inparalogs for each cluster. The default setting of INPARANOID requires a positive confidence value for a gene to be accepted as an inparalog, but genes evolving under strong positive selection may violate this assumption if they are highly divergent from their paralogs.
After the INPARANOID run, only clusters with exactly two A. thaliana inparalogs were retained for further analysis. For the Arabidopsis inparalogs, we use following nomenclature: The seed ortholog is denoted as At-1 and the added inparalog as At-2. It should be noted that clusters with >2 inparalogs can also be analyzed with appropriate models. Following Remm et al. (2001), we further eliminated all clusters whose BLAST scores were inconsistent with the species phylogeny when S. cerevisiae and O. sativa genomes were used as outgroups.
Refining and Aligning Orthologous Gene Clusters
Orthologous gene clusters identified by INPARANOID were compared with A. thaliana paralogs identified by Blanc et al. (2003) to extract only segmentally duplicated genes. Several paralogs appeared to be located in a segmentally duplicated gene cluster but were not included in the Blanc et al. (2003) dataset due to annotation inconsistencies (i.e., changes of the gene identifier code). To detect such genes, we used a sliding window technique for neighboring genes in segmentally duplicated regions. Groups of inparalogs, which were part of at least six neighboring genes in the same order on both clusters in a window of 20 genes were added to the set of genes from the Blanc et al. (2003) data. The sliding window analysis also identified genes that were present as tandem duplicates within one of two segmentally duplicated regions, but not in the other (Supplementary Figure S1) and they were also included in the analysis. Protein sequences of gene clusters were aligned with CLUSTAL (Higgins 1994) and corresponding gap-free codon-based alignments were generated with PAL2NAL (Suyama et al. 2006). The DNA sequence alignments were used to obtain the tree topology using DNAML from the PHYLIP package (Felsenstein 2005). We obtained 12,573 distinct clusters, of which 3,754 clusters contained >1 and 2,845 clusters exactly two A. thaliana inparalogs (Fig. 1). We also added 203 clusters with >2 A. thaliana inparalogs by considering only A. thaliana inparalogs with a positive confidence value. Among 3,048 INPARANOID clusters with exactly two A. thaliana inparalogs, 2,109 (70%) were identified as segmentally duplicated. Of these, 185 clusters could not be processed by PAL2NAL due to inconsistencies between DNA and protein data that mainly were observed among the poplar sequences and likely result from sequencing errors or wrongly predicted splicing sites; they were excluded from further analysis.
Tests of Sites Under Selection
The ratio of nonsynonymous substitutions per nonsynonymous site (d N) to the synonymous substitutions per synonymous sites (d S), ω = d N/d S, can be used as a test of natural selection (Yang and Bielawski 2000). Positive selection is inferred if ω > 1, purifying selection if ω < 1, and neutral evolution if ω = 1.
We used branch-site models (Forsberg and Christiansen 2003; Bielawski and Yang 2004) to infer ω ratios with the PAML package (Yang 1997). Clade model C (a branch-site model, Forsberg and Christiansen 2003; Bielawski and Yang 2004) was used to detect differences in the proportion of selected sites in the lineages between the two A. thaliana inparalogs. Note that in contrast to branch-site model A of the PAML package the clade model C does not assume a fraction of sites with ω > 1. Tests for significant differences among models were calculated as likelihood ratio tests (LRTs), where the test statistic was 2Δl = 2 × (l 1 − l 2) with l 1 and l 2 as the log of the maximum likelihood (ML) estimated of the two models compared. It is assumed that 2Δl is approximately distributed as χ2 with difference of model parameters as degrees of freedom (d.f.), and critical values were obtained from this distribution. Model C estimates the proportion p 0 of codons with ω0 < 1 and a proportion p 1 of sites with ω1 = 1 for all branches combined and additionally a proportion p 2 of codons which are allowed to differ between the foreground (ω3) and background branches (ω2). We used an extension of clade model C as implemented in PAML version 4.4 which allows for two types of foreground branches (ω3 and ω4). For each cluster, a LRT was carried out with the two A. thaliana paralogs as separate foreground branches (ω3 ≠ ω4) compared with the clade model C for which the two A. thaliana paralogs belonged to the same foreground branch (ω3 = ω4) assuming d.f. = 1 (Fig. 2). We also excluded those clusters for which tree length was larger than the number of branches of the phylogeny and for which the posterior distribution for ω3 and ω4 significantly overlapped (>5%).
Outline of branch-site models used in the study. The phylogeny illustrates the most frequent case with two P. trichocarpa inparalogs (n = 2). The two branches leading to the A. thaliana inparalogs At-1 and At-2 of INPARANOID are labeled with ω3 and ω4, respectively. There are always two inparalogs in A. thaliana, but the number of inparalogs in poplar ranges from 1 to n (resulting in 2n + 1 branches). The duplication events are indicated by black circles. The branches leading to At-1 and At-2 are chosen as foreground branches. The proportions p 0 with ω0, p 1 with ω1, and p 2 of codons with ratio ω2, ω3, and ω4 are estimated. The proportion of codons under purifying selection (0 < ω0 < 1) and neutral codons (ω1 = 1) is estimated together for the whole phylogeny. At Arabidopsis thaliana, Pt Populus trichocarpa
Simulation Studies
The number of sequences in the alignment and the evolutionary distance (i.e., the average number of substitutions in a codon) strongly affect the power of a LRT to detect selection (Anisimova et al. 2001). Our samples are characterized by a low number of sequences and a high level of silent site degeneracy, which both reduce the power to detect lineage-specific evolutionary rates. Therefore, we conducted simulations with the evolver program of the PAML package for three sample clusters. Using the tree topology and the parameter estimates from the codeml branch-site analysis, two sets of 100 alignments each were generated (Table 4). For the first set (Simulation 1), data were simulated using the estimated values of the branch-site test with ω3 ≠ ω4. For the second set (Simulation 2), the estimated values of the branch-site test were used for the case ω3 = ω4. We determined the power and accuracy by conducting the branch-site test on the simulated sequences and counting how often LRTs were significant. We expect the proportion of significant rejected branch-site tests for Simulation 1 to be substantial, while for Simulation 2 the proportion of significant LRTs should be low if our approach is reasonable. However, an important factor is the quality of the alignments as insertions and deletions may produce shorter and less accurate alignments (Fletcher and Yang 2010). Consequently, when gaps are removed from the alignment a proportion of the remaining codons will be incorrectly aligned. These partly misaligned sequences could generate false positive or negative results when the branch-site test is applied. We therefore were interested how insertions and deletions would alter the outcome of the branch-site tests on the simulated sequences. For this analysis, we constructed 100 alignments using the parameter values of the branch-site test with INDELIBLE (Fletcher and Yang 2009). The length distribution of indels and the distribution of indels across the sequences are not known for A. thaliana and P. trichocarpa. We therefore use a scenario with equal rates of insertions and deletions and estimated the parameters as follows: The distribution of indels can be approximated by the Lavalette distribution (Fletcher and Yang 2009) for which the probability P of an indel of size u is given by
where u = 1,2,…,M where M is the maximum indel size. It is not clear which parameter space is reasonable for a and M. However, from the estimate of the mean indel length it is possible to obtain a value for a for a given M. Empirical estimates range from 1.5 to 2 (Zhang and Gerstein 2003; Yamane et al. 2006; Cartwright 2009). We obtain values for a between 2.01 and 2.35 for M = 500 and 1.61 and 1.93 for M = 200 (Supplementary Table S1). We show simulation results for M = 500 only, but results with other parameter values are very similar.
Analysis of Expression Profiles
Arabidopsis thaliana gene expression data were obtained from the Nottingham Arabidopsis Stock Centre microarray database (NASCArray). Hybridization experiments differ by the number of controls, and also the labeling procedures were not consistently standardized. Therefore, we used the raw expression values to calculate Kendall’s τ when comparing expression. Correlations could be calculated for 76% of all pairs (1,463 of 1,924) of inparalogs, because only a subset of A. thaliana genes were included on the microarrays. Owing to the heterogeneity of the type and conditions of experiments, the correlation coefficient should be considered as a rough estimate of co-expression (Table 1).
Comparison of Inparalog Sequence Divergence with Intraspecific Sequence Variation
Resequencing data of 20 A. thaliana accessions obtained with the Perlegene array (Clark et al. 2007) were downloaded from TAIR. The site frequency spectrum was obtained for each locus. We estimated the proportion of adaptive substitutions, α, using an extension of the McDonald–Kreitman (MK) test (McDonald and Kreitman 1991) which takes into account the influence of slightly deleterious mutations (Eyre-Walker and Keightley 2009). Since many genes showed little polymorphism, we split each pair of inparalogs with significantly different ω ratios into two groups and summed data across genes. One group contained genes for which ω3 or ω4 was highest (relaxed group), while the other group harbored the remaining inparalogs (constrained group). Polymorphism data from 225 genes were available for comparison. Lineage-specific divergence was retrieved from the estimates of the free-ratio model from PAML.
Overrepresentation of Gene Ontology terms
Arabidopsis thaliana gene ontology Ashburner et al. (2000) annotations were obtained from the NASCArray. The GO term descriptors were retrieved from the gene ontology website (http://www.geneontology.org). We tested which GO terms are over- or underrepresented among pairs of inparalogs in comparison with all genes, and among inparalog pairs with significant LRTs in comparison with remaining inparalog pairs. A hypergeometrical distribution was assumed, which was approximated with a χ2 distribution for large numbers.
Results
Extraction of Segmentally Duplicated Genes
To extract segmentally duplicated genes from the A. thaliana genome, we obtained pairs of inparalogs with INPARANOID and filtered them with the slightly expanded Blanc et al. (2003) dataset. In the end, 1,924 pairs of A. thaliana inparalogs were analyzed together with their poplar homologs using PAML (Fig. 1), of which 1,588 (82%) are also contained in the Blanc et al. (2003) data. More than 80% of the analyzed clusters consisted of at least four sequences, but 376 clusters consisted of three genes only. A majority of 1,774 clusters (92%) originated in the recent and 57 (3%) in the old duplication event; 93 gene clusters (5%) are tandemly duplicated genes. Among inparalog clusters, functional groups of genes are differentially represented (Supplementary Table S2). GO terms related to terms cellular locations and metabolic processes are under-represented, whereas genes associated with the nucleus, DNA binding, and transcriptional activity are overrepresented.
Pairwise Comparisons of Homologs
The rate of evolutionary divergence between inparalogs and orthologs was estimated as the ratio of nonsynonymous (d N) to synonymous (d S) substitutions, \( \omega = d_{\rm{N}} /d_{\rm{S}} \), in pairwise comparisons of sequences (PAML runmode = −2; Fig. 3). A total of 1,862 (96.7%) A. thaliana inparalog pairs showed d S < 10 and ω < 20 which were used as cutoff values; the median ω value was 0.116. Among 8,712 Arabidopsis-poplar pairwise comparisons, 5,678 (65.2%) showed d S < 10 and ω < 20 and a median ω value of 0.0535. The d S value is assumed to represent the neutral mutation rate since synonymous codon positions are supposed to be largely free from selection. Our data agree with this hypothesis, as pairwise d N values are less variable than d S values. P. trichocarpa is a close relative to Medicago and similar pairwise d S values in comparisons with A. thaliana were obtained for the Populus–Arabidopsis (2.2; Fig. 3) and Medicago–Arabidopsis (2.0–2.2; Blanc et al. 2003) comparisons. In contrast, the median d S value for the A. thaliana inparalog comparison is approximately 1.0, which reflects the high proportion of genes originating from the recent genome duplication. Subsequently, the median ω value of the Arabidopsis-poplar orthologs is smaller than for the Arabidopsis inparalogs because higher d S values decrease the ω ratio.
Tests of Different Selection Pressures After Duplication
We used branch-site models (clade models C) to carry out a test of differences in selection pressures between A. thaliana paralogs (Fig. 2). We find 493 of 1,924 gene clusters (25.6%) resulting in a significant LRT. However, for 299 clusters the posterior distributions of ω3 and ω4 values overlapped significantly and were therefore excluded. Out of the remaining 194 clusters, 62 had unreasonable high tree length estimates in at least one of models and were as well excluded. The remaining set consisted of 132 clusters (6.9%) showed significant differences in ω ratios between the two A. thaliana inparalogs. Out of these 132 clusters, we found for 79 clusters (59.8%) with ω3 or ω4 larger than one. Seed orthologs had smaller ω values than the second inparalog in 114 (86.3%) of 132 significant clusters. This number indicates that INPARANOID uses conserved members of a gene family to identify orthologs in other species, and then adds more divergent paralogs to a cluster. Among the 132 inparalog pairs with significant LRTs, five GO terms were overrepresented (Table 2). They include genes associated to nucleotide binding, protein amino acid phosphorylation as well as response to stress.
Identification of Selection-Driven Genes for Functional Analysis
One goal of this study was the identification of new candidate genes for further functional analyses. Since our study was motivated by the rapid evolution of MEDEA, which controls reproductive development and is likely involved in a genomic conflict, we were interested in genes with elevated ω. Fifteen inparalog pairs which have been identified by the branch-site model and showed ω > 1 for a substantial proportion of sites (n > 100) are shown in Table 3. These clusters contain genes that are involved in stress response (Mao et al. 2006; Sun et al. 2007; Kim et al. 2008), development (Bernhardt et al. 2010) and disease resistance (Kesarwani et al. 2007). Surprisingly, according to the TAIR literature database five out of those 15 gene pairs have yet to be functionally characterized.
Simulation Studies
For three sample clusters (Fig. 4), we checked the power to detect differences in the selective pressure using parameter estimates from the branch-site model (Table 4). We conducted simulations by applying LRTs to 100 simulated sequences for each of the three clusters. In 61–96% of cases, the LRTs were significant (Simulation 1). In contrary, in simulations with equal rates between the lineages (ω3 = ω4), only 7–13% of the LRTs were significant (Simulation 2). We also investigated how misaligned codons would alter the outcome of the branch-site test and modeled insertions and deletions into the simulated alignments using INDELIBLE (Table 4). In 41–91% of cases, the LRTs were significant (Simulation 3). In contrary, 9–14% of LRTs were significant if we model equal evolutionary rates between the lineages (Simulation 4). We therefore conclude that the given parameter values are reasonably well captured by the test statistic and the impact of indels to the outcome of the branch-site test is limited.
Phylogenetic trees of three INPARANOID clusters. a Cluster 745 including protein kinase At1g12460, b cluster 5758 including Rhomboid homolog protein 6 (At1g12750), and c cluster 6450 including carbohydrate binding At1g10150. Tree topologies were obtained with PHYLIP, and branch lengths (substitutions per codon) were calculated with CODEML using the nearly neutral model. Note that the trees are unrooted. ω for sites obtained from the branch-site model are indicated for both Arabidopsis inparalogs in bold
Estimating the Amount of Adaptive Substitutions
We estimated the proportion α of amino acid substitutions that underwent positive selection since the duplication event. For this, we used polymorphism data from a resequencing study of 20 A. thaliana accessions (Clark et al. 2007). A MK type of analysis (see “Materials and Methods” section) was used. Under the assumption that synonymous mutations are neutral, α can be estimated from simple expressions contrasting within-population polymorphism and corresponding levels of between-species divergence at two categories of sites (e.g., synonymous and nonsynonymous sites). Since we were interested in the amount of adaptive substitutions the 132 gene pairs with significantly different ω ratios underwent since the duplication event we used lineage-specific divergence data estimated from the free-ratio model of PAML.
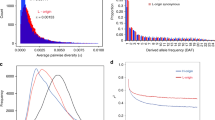
An α value of 0.24 (0.11–0.35) was observed for relaxed inparalogs of the branch-site model which is significantly larger than 0 (Fig. 5a). In contrast, estimates for the constrained inparalogs are −0.33 (−0.59, −0.04) which supports the hypothesis that the relaxed inparalogs have undergone more adaptive evolution. We also estimated the distributions of fitness effects (DFE) for the two sets of genes (Fig. 5b). The results differ substantially between the two groups of genes. For the relaxed group, the proportion of neutral mutations (0 < N e s < 1) is decreased while the proportion of strongly deleterious mutations (N e s > 100) is increased. This indicates that these genes are subject to stronger purifying selection or have a higher effective population size. The DFE of the constrained inparalogs corresponds to a previous genome-wide estimate for A. thaliana (Gossmann et al. 2010) obtained from a different dataset (Nordborg et al. 2005).
Estimates of α and the distribution of fitness effects for the indentified gene pairs. a α, the proportion of fixed amino acid differences since the duplication event driven by positive selection, for pairs of A. thaliana inparalogs with significantly different ω3 and ω4 values. To conduct the test, each gene pair is split into either constrained or relaxed inparalog depending on their ω3 and ω4 values from the branch-site model test. Polymorphism and divergence data are summed across genes. b Estimates of the distribution of fitness effects for the two groups (constrained and relaxed) of genes. Mutations are binned according to their fitness effects
Relationship Between Protein Sequence and Expression Pattern Divergence
Both the neo- and subfunctionalization models accommodate the functional divergence of paralogs in protein function or expression pattern. Genes whose sequences show evidence for positive selection or subfunctionalization after gene duplication may also evolve more rapidly in their expression pattern. We conducted t tests on correlation of co-expression (Table 1), using publicly available microarray experiment data. The average co-expression correlation is increased for pairs of inparalogs than for random pairs of genes (P = 4 × 10−29). Within the set of inparalog pairs, co-expression is significantly reduced for gene pairs identified by the branch-site model (P = 3 × 10−3).
Discussion
The publication of several plant genomes has identified tens of thousands of novel genes over the last decade and the discovery of new genes will continue with the advance of new sequencing technologies (Ellegren 2008). Given the much slower pace with which the function of newly discovered genes can be determined, the evolutionary and bioinformatic characterization of genes is an essential first step in describing genome structure and evolution because it can be automated to a large degree. In the present study, we were interested in estimating the proportion of genes evolving at different evolutionary constraints after gene duplication by calculating the rate of nonsynonymous to synonymous divergence since their origin by duplication. We hypothesized that paralogous genes with significantly different rates of sequence evolution became functionally divergent, because rate differences result either from adaptation to a new function, or from different levels of constraint (i.e., differences in level of purifying selection) after genes acquired new functions.
To investigate the proportion of neo- and subfunctionalized genes in A. thaliana, we constructed a largely automated analysis pipeline based on a whole genome comparison of A. thaliana and P. trichocarpa. In principle, paralogs in P. trichocarpa could be analyzed as well, however, since information about segmental clusters in P. trichocarpa was limited we focused on A. thaliana genes. We generated a dataset of 1,924 pairs of duplicated A. thaliana genes (inparalogs) and their orthologs from the poplar genome with INPARANOID. Genes with annotation inconsistencies and unreasonable high d N, d S, and ω values were excluded because the quality of alignment and of the reconstructed phylogenies are crucial for estimating correct ω values (Wong et al. 2008). As a preliminary analysis, we conducted branch tests to identify differences in selection pressure between the A. thaliana inparalogs (results not shown), of which only 3 out of 1,924 clusters were significant using Bonferroni correction. But a few amino acids may be sufficient for functional divergence of paralogs and there is little power of branch models to detect selection (Anisimova et al. 2001; Studer et al. 2008) especially considering the large divergence since the last duplication event in Arabidopsis. To address this issue, we analyzed our dataset using a branch-site model and determined 132 gene pairs with different evolutionary rates between the two A. thaliana lineages.
Possible explanations for the severe discrepancies between the branch and branch-site tests are (i) a low power of the branch models to detect positive selection, (ii) different levels of purifying selection among paralogs, (iii) subfunctionalization rather than positive selection among paralogs detected with the branch model, (iv) unreasonable parameter estimates of the the branch-site model. To rule out the last possibility, we conducted simulations to estimate the power of PAML to detect positive selection with the branch-site model in alignments with few sequences (Table 4). They indicate that the applied branch-site test is sensitive enough to detect differences in the selective pressure between the lineages using the parameters estimated from the data. On the other hand, the rate of false positives is relatively low. We conclude that models rejected by the branch-site tests could be explained by subfunctionalization of one of the copies resulting in a fraction of sites evolving nearly neutrally. We also investigated by simulations the impact of alignment errors caused by indels on the branch-site model. We observed only a slight reduction in the power to detect significant differences if the impact of indels to the alignment is taken into account. These results suggest that the branch-site test as implemented in our study is fairly robust against alignment errors.
According to GO term descriptors, genes with regulatory activities are enriched among inparalog pairs. After duplication, changes in the regulatory sequence or in the coding sequence can lead to neofunctionalization or subfunctionalization. Our analysis was restricted to coding sites. Genes which are associated with nucleotide binding and amino acid phosphorylation are overrepresented for gene pairs rejected by the branch-site model tests. The enrichment of regulatory genes reflects the hypothesis that changes in regulatory sequences may contribute to the amount of neo- and subfunctionalized genes even though they are not directly covered by our approach. This is also consistent with the observed co-expression pattern of inparalogs as expression values showed a significantly reduced correlation in co-expression for gene pairs identified by the branch-site models. Such a reduction of correlation may be the consequence of a neofunctionalization of one of the copies or of a subfunctionalization of both copies. A previous attempt to distinguish the two possible scenarios by considering an inferred ancestral state of expression (Duarte et al. 2006) revealed that only few gene pairs can be assigned to one of the two categories, instead, a mixture of both models may apply. The ω ratio integrates the selection pressure for a period of about 40 mya for paralogs originating from the recent duplication event (Blanc et al. 2003). To differentiate between the hypotheses that most divergence among inparalogs originated immediately after duplication, or that either one or both inparalogs are evolving rapidly until the present, sequences from additional species covering the phylogenetic distance since duplication are required. Then, variation of ω ratios in the phylogeny of each inparalog can be calculated with greater confidence.
Since genome sequences of close relatives are not yet available, an extension of the MK test was applied to compare lineage-specific divergence and polymorphisms. An application of the MK analysis to paralogs may lead to an overestimate of the amount of adaptive substitutions (α) because polymorphism data provide information only on recent positive or purifying selection but not historical selection (Hahn 2009). Nevertheless, an estimate of α is still meaningful for two reasons. First, it provides an upper boundary of the proportion of fixed adaptions and second, a comparison of the distribution of fitness effects provides a comparable measurement of the recent evolution of both duplicated genes. The comparison of two sets of inparalogs has the advantage that demographic history and breeding system, which both have an effect on the site frequency spectrum and hence on the estimate of α (Eyre-Walker 2006; Foxe et al. 2008), are identical for both groups of inparalogs and do not affect the inference of selection in different ways. Owing to power reasons, we applied each MK analysis to a summed statistic for two groups of genes, dividing each inparalogous pair into either constrained or relaxed categories, according to ω estimates of the branch-site model. Note that this might increase the estimate of α for the relaxed genes. However, differences in the α estimates are largely caused by differences in P N/P S (Fig. 5b). Divergence estimates do not differ significantly between the two groups of genes (P = 0.69 and P = 0.11 for d N and d S, respectively). Therefore, our estimates for genes with a significantly higher ω ratio for a fraction of sites indicate that up to 24% of the divergence since the duplication may be attributed to positive selection.
Conclusions
Our results imply that around 6.9% of the analyzed A. thaliana paralogous gene pairs show different rates of evolution after gene duplication. Asymmetry of selective pressure supports either increased positive selection or relaxation of purifying selection. Other mechanisms such as preservation of duplicate genes by originalization also may be important (Xue and Fu 2009; Tanaka et al. 2009). Our estimate might be a severe underestimate because we had to exclude a substantial number of sequences due to the high degeneracy of A. thaliana paralogs and the relatively highly divergent outgroup. Furthermore, functional differentiation by other mechanisms, such as alternative splicing or gene dosage effects, was not covered by our approach. Studer et al. (2008) have shown that positive selection has been pervasive during vertebrate evolution, but whole genome duplicates had no effect on the prevalence of positive selection. Direct tests for positive selection in yeast and Drosophila are even higher than our estimates (Conant and Wagner 2003), while estimates for Xenopus laevis are lower (Chain and Evans 2006). A recent study in human, macaque, mouse, and rat genomes for young duplicates revealed that about 10% of duplicated gene pairs evolved under positive selection (Han et al. 2009) using a branch-site test. Our result is the highest ever reported value for A. thaliana and shows that selection after duplication contributes substantially to gene novelties and hence functional divergence in plants.
References
AG Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408(6814):796–815
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. doi:10.1006/jmbi.1990.9999
Anisimova M, Bielawski JP, Yang Z (2001) Accuracy and power of the likelihood ratio test in detecting adaptive molecular evolution. Mol Biol Evol 18(8):1585–1592
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25(1):25–29. doi:10.1038/75556
Aury JM, Jaillon O, Duret L, Noel B, Jubin C, Porcel BM, Ségurens B, Daubin V, Anthouard V, Aiach N, Arnaiz O, Billaut A, Beisson J, Blanc I, Bouhouche K, Câmara F, Duharcourt S, Guigo R, Gogendeau D, Katinka M, Keller AM, Kissmehl R, Klotz C, Koll F, Mouël AL, Lepère G, Malinsky S, Nowacki M, Nowak JK, Plattner H, Poulain J, Ruiz F, Serrano V, Zagulski M, Dessen P, Bétermier M, Weissenbach J, Scarpelli C, Schächter V, Sperling L, Meyer E, Cohen J, Wincker P (2006) Global trends of whole-genome duplications revealed by the ciliate Paramecium tetraurelia. Nature 444(7116):171–178. doi:10.1038/nature05230
Benderoth M, Textor S, Windsor AJ, Mitchell-Olds T, Gershenzon J, Kroymann J (2006) Positive selection driving diversification in plant secondary metabolism. Proc Natl Acad Sci USA 103(24):9118–9123. doi:10.1073/pnas.0601738103
Bernhardt A, Mooney S, Hellmann H (2010) Arabidopsis DDB1a and DDB1b are critical for embryo development. Planta 232(3):555–566. doi:10.1007/s00425-010-1195-9
Bielawski JP, Yang Z (2004) A maximum likelihood method for detecting functional divergence at individual codon sites, with application to gene family evolution. J Mol Evol 59(1):121–132. doi:10.1007/s00239-004-2597-8
Blanc G, Wolfe KH (2004a) Functional divergence of duplicated genes formed by polyploidy during Arabidopsis evolution. Plant Cell 16(7):1679–1691. doi:10.1105/tpc.021410
Blanc G, Wolfe KH (2004b) Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. Plant Cell 16(7):1667–1678. doi:10.1105/tpc.021345
Blanc G, Hokamp K, Wolfe KH (2003) A recent polyploidy superimposed on older large-scale duplications in the Arabidopsis genome. Genome Res 13(2):137–144. doi:10.1101/gr.751803
Cartwright RA (2009) Problems and solutions for estimating indel rates and length distributions. Mol Biol Evol 26(2):473–480. doi:10.1093/molbev/msn275
Chain FJJ, Evans BJ (2006) Multiple mechanisms promote the retained expression of gene duplicates in the tetraploid frog Xenopus laevis. PLoS Genet 2(4):e56. doi:10.1371/journal.pgen.0020056
Chen F, Mackey AJ, Vermunt JK, Roos DS (2007a) Assessing performance of orthology detection strategies applied to eukaryotic genomes. PLoS ONE 2(4):e383. doi:10.1371/journal.pone.0000383
Chen Q, Steinhauer L, Hammerlindl J, Keller W, Zou J (2007b) Biosynthesis of phytosterol esters: identification of a sterol o-acyltransferase in Arabidopsis. Plant Physiol 145(3):974–984. doi:10.1104/pp.107.106278
Clark AG (1994) Invasion and maintenance of a gene duplication. Proc Natl Acad Sci USA 91(8):2950–2954
Clark RM, Schweikert G, Toomajian C, Ossowski S, Zeller G, Shinn P, Warthmann N, Hu TT, Fu G, Hinds DA, Chen H, Frazer KA, Huson DH, Schölkopf B, Nordborg M, Rätsch G, Ecker JR, Weigel D (2007) Common sequence polymorphisms shaping genetic diversity in Arabidopsis thaliana. Science 317(5836):338–342. doi:10.1126/science.1138632
Conant GC, Wagner A (2003) Asymmetric sequence divergence of duplicate genes. Genome Res 13(9):2052–2058. doi:10.1101/gr.1252603
Duarte JM, Cui L, Wall PK, Zhang Q, Zhang X, Leebens-Mack J, Ma H, Altman N, dePamphilis CW (2006) Expression pattern shifts following duplication indicative of subfunctionalization and neofunctionalization in regulatory genes of Arabidopsis. Mol Biol Evol 23(2):469–478. doi:10.1093/molbev/msj051
Ellegren H (2008) Sequencing goes 454 and takes large-scale genomics into the wild. Mol Ecol 17(7):1629–1631. doi:10.1111/j.1365-294X.2008.03699.x
Eyre-Walker A (2006) The genomic rate of adaptive evolution. Trends Ecol Evol 21(10):569–575. doi:10.1016/j.tree.2006.06.015
Eyre-Walker A, Keightley PD (2009) Estimating the rate of adaptive molecular evolution in the presence of slightly deleterious mutations and population size change. Mol Biol Evol 26(9):2097–2108. doi:10.1093/molbev/msp119
Felsenstein J (2005) PHYLIP (Phylogeny Inference Package) version 3.6. Distributed by the author. Department of Genome Sciences, University of Washington, Seattle
Fiebig A, Kimport R, Preuss D (2004) Comparisons of pollen coat genes across Brassicaceae species reveal rapid evolution by repeat expansion and diversification. Proc Natl Acad Sci USA 101(9):3286–3291. doi:10.1073/pnas.0305448101
Fletcher W, Yang Z (2009) INDELible: a flexible simulator of biological sequence evolution. Mol Biol Evol 26(8):1879–1888. doi:10.1093/molbev/msp098
Fletcher W, Yang Z (2010) The Effect of insertions, deletions and alignment errors on the branch-site test of positive selection. Mol Biol Evol. doi:10.1093/molbev/msq115
Force A, Lynch M, Pickett FB, Amores A, Yan YL, Postlethwait J (1999) Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151(4):1531–1545
Forsberg R, Christiansen FB (2003) A codon-based model of host-specific selection in parasites, with an application to the influenza A virus. Mol Biol Evol 20(8):1252–1259. doi:10.1093/molbev/msg149
Foxe JP, un Nisa Dar V, Zheng H, Nordborg M, Gaut BS, Wright SI (2008) Selection on amino acid substitutions in Arabidopsis. Mol Biol Evol 25(7):1375–1383. doi:10.1093/molbev/msn079
Gossmann TI, Song BH, Windsor AJ, Mitchell-Olds T, Dixon CJ, Kapralov MV, Filatov DA, Eyre-Walker A (2010) Genome wide analyses reveal little evidence for adaptive evolution in many plant species. Mol Biol Evol. doi:10.1093/molbev/msq079
Hahn MW (2009) Distinguishing among evolutionary models for the maintenance of gene duplicates. J Hered 100(5):605–617. doi:10.1093/jhered/esp047
Han MV, Demuth JP, McGrath CL, Casola C, Hahn MW (2009) Adaptive evolution of young gene duplicates in mammals. Genome Res 19(5):859–867. doi:10.1101/gr.085951.108
He X, Zhang J (2005) Rapid subfunctionalization accompanied by prolonged and substantial neofunctionalization in duplicate gene evolution. Genetics 169(2):1157–1164. doi:10.1534/genetics.104.037051
Higgins DG (1994) CLUSTAL V: multiple alignment of DNA and protein sequences. Methods Mol Biol 25:307–318. doi:10.1385/0-89603-276-0:307
IRGS Project (2005) The map-based sequence of the rice genome. Nature 436(7052):793–800
Jaillon O, Aury JM, Noel B, Policriti A, Clepet C, Casagrande A, Choisne N, Aubourg S, Vitulo N, Jubin C, Vezzi A, Legeai F, Hugueney P, Dasilva C, Horner D, Mica E, Jublot D, Poulain J, Bruyère C, Billault A, Segurens B, Gouyvenoux M, Ugarte E, Cattonaro F, Anthouard V, Vico V, Fabbro CD, Alaux M, Gaspero GD, Dumas V, Felice N, Paillard S, Juman I, Moroldo M, Scalabrin S, Canaguier A, Clainche IL, Malacrida G, Durand E, Pesole G, Laucou V, Chatelet P, Merdinoglu D, Delledonne M, Pezzotti M, Lecharny A, Scarpelli C, Artiguenave F, Pè ME, Valle G, Morgante M, Caboche M, Adam-Blondon AF, Weissenbach J, Quétier F, Wincker P, for Grapevine Genome Characterization FIPC (2007) The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449(7161):463–467
Jakoby M, Weisshaar B, Dröge-Laser W, Vicente-Carbajosa J, Tiedemann J, Kroj T, Parcy F, bZ IP Research Group (2002) bZIP transcription factors in Arabidopsis. Trends Plant Sci 7(3):106–111
Kesarwani M, Yoo J, Dong X (2007) Genetic interactions of TGA transcription factors in the regulation of pathogenesis-related genes and disease resistance in Arabidopsis. Plant Physiol 144(1):336–346. doi:10.1104/pp.106.095299
Kim CY, Bove J, Assmann SM (2008) Overexpression of wound-responsive RNA-binding proteins induces leaf senescence and hypersensitive-like cell death. New Phytol 180(1):57–70. doi:10.1111/j.1469-8137.2008.02557.x
Koonin EV (2005) Orthologs, paralogs, and evolutionary genomics. Annu Rev Genet 39:309–338. doi:10.1146/annurev.genet.39.073003.114725
Kuang H, Woo SS, Meyers BC, Nevo E, Michelmore RW (2004) Multiple genetic processes result in heterogeneous rates of evolution within the major cluster disease resistance genes in lettuce. Plant Cell 16(11):2870–2894. doi:10.1105/tpc.104.025502
Kuraku S, Meyer A, Kuratani S (2009) Timing of genome duplications relative to the origin of the vertebrates: did cyclostomes diverge before or after? Mol Biol Evol 26(1):47–59. doi:10.1093/molbev/msn222
Li L, Stoeckert CJ, Roos DS (2003) OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res 13(9):2178–2189. doi:10.1101/gr.1224503
Lynch M, Conery JS (2000) The evolutionary fate and consequences of duplicate genes. Science 290(5494):1151–1155
Lynch M, Force A (2000) The probability of duplicate gene preservation by subfunctionalization. Genetics 154(1):459–473
Lynch M, O’Hely M, Walsh B, Force A (2001) The probability of preservation of a newly arisen gene duplicate. Genetics 159(4):1789–1804
Mao Y, Pavangadkar KA, Thomashow MF, Triezenberg SJ (2006) Physical and functional interactions of Arabidopsis ADA2 transcriptional coactivator proteins with the acetyltransferase GCN5 and with the cold-induced transcription factor CBF1. Biochim Biophys Acta 1759(1–2):69–79. doi:10.1016/j.bbaexp.2006.02.006
McDonald JH, Kreitman M (1991) Adaptive protein evolution at the Adh locus in Drosophila. Nature 351(6328):652–654. doi:10.1038/351652a0
Mita SD, Santoni S, Hochu I, Ronfort J, Bataillon T (2006) Molecular evolution and positive selection of the symbiotic gene NORK in Medicago truncatula. J Mol Evol 62(2):234–244. doi:10.1007/s00239-004-0367-2
Mondragón-Palomino M, Meyers BC, Michelmore RW, Gaut BS (2002) Patterns of positive selection in the complete NBS-LRR gene family of Arabidopsis thaliana. Genome Res 12(9):1305–1315. doi:10.1101/gr.159402
Nordborg M, Hu TT, Ishino Y, Jhaveri J, Toomajian C, Zheng H, Bakker E, Calabrese P, Gladstone J, Goyal R, Jakobsson M, Kim S, Morozov Y, Padhukasahasram B, Plagnol V, Rosenberg NA, Shah C, Wall JD, Wang J, Zhao K, Kalbfleisch T, Schulz V, Kreitman M, Bergelson J (2005) The pattern of polymorphism in Arabidopsis thaliana. PLoS Biol 3(7):e196, doi:10.1371/journal.pbio.0030196
O’Hely M (2006) A diffusion approach to approximating preservation probabilities for gene duplicates. J Math Biol 53(2):215–230. doi:10.1007/s00285-006-0001-6
Ohno S (1970) Evolution by gene duplication. Springer, Berlin
Raes J, Vandepoele K, Simillion C, Saeys Y, de Peer YV (2003) Investigating ancient duplication events in the Arabidopsis genome. J Struct Funct Genomics 3(1–4):117–129
Rastogi S, Liberles DA (2005) Subfunctionalization of duplicated genes as a transition state to neofunctionalization. BMC Evol Biol 5(1):28. doi:10.1186/1471-2148-5-28
Remm M, Storm CE, Sonnhammer EL (2001) Automatic clustering of orthologs and in-paralogs from pairwise species comparisons. J Mol Biol 314(5):1041–1052. doi:10.1006/jmbi.2000.5197
Rizzon C, Ponger L, Gaut BS (2006) Striking similarities in the genomic distribution of tandemly arrayed genes in Arabidopsis and rice. PLoS Comput Biol 2(9):e115. doi:10.1371/journal.pcbi.0020115
Scannell DR, Butler G, Wolfe KH (2007) Yeast genome evolution–the origin of the species. Yeast 24(11):929–942. doi:10.1002/yea.1515
Schein M, Yang Z, Mitchell-Olds T, Schmid KJ (2004) Rapid evolution of a pollen-specific oleosin-like gene family from Arabidopsis thaliana and closely related species. Mol Biol Evol 21(4):659–669. doi:10.1093/molbev/msh059
Soltis PS, Soltis DE (2009) The role of hybridization in plant speciation. Annu Rev Plant Biol 60:561–588. doi:10.1146/annurev.arplant.043008.092039
Spillane C, Schmid KJ, Laoueillé-Duprat S, Pien S, Escobar-Restrepo JM, Baroux C, Gagliardini V, Page DR, Wolfe KH, Grossniklaus U (2007) Positive darwinian selection at the imprinted MEDEA locus in plants. Nature 448(7151):349–352. doi:10.1038/nature05984
Studer RA, Penel S, Duret L, Robinson-Rechavi M (2008) Pervasive positive selection on duplicated and nonduplicated vertebrate protein coding genes. Genome Res 18(9):1393–1402. doi:10.1101/gr.076992.108
Sun X, Cao Y, Wang S (2006) Point mutations with positive selection were a major force during the evolution of a receptor-kinase resistance gene family of rice. Plant Physiol 140(3):998–1008. doi:10.1104/pp.105.073080
Sun J, Jiang H, Xu Y, Li H, Wu X, Xie Q, Li C (2007) The CCCH-type zinc finger proteins AtSZF1 and AtSZF2 regulate salt stress responses in Arabidopsis. Plant Cell Physiol 48(8):1148–1158. doi:10.1093/pcp/pcm088
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34(Web Server issue):W609–W612. doi:10.1093/nar/gkl315
Tanaka KM, Takahasi KR, Takano-Shimizu T (2009) Enhanced fixation and preservation of a newly arisen duplicate gene by masking deleterious loss-of-function mutations. Genet Res (Camb) 91(4):267–280. doi:10.1017/S0016672309000196
Tang H, Wang X, Bowers JE, Ming R, Alam M, Paterson AH (2008) Unraveling ancient hexaploidy through multiply-aligned angiosperm gene maps. Genome Res 18(12):1944–1954. doi:10.1101/gr.080978.108
Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A, Schein J, Sterck L, Aerts A, Bhalerao RR, Bhalerao RP, Blaudez D, Boerjan W, Brun A, Brunner A, Busov V, Campbell M, Carlson J, Chalot M, Chapman J, Chen GL, Cooper D, Coutinho PM, Couturier J, Covert S, Cronk Q, Cunningham R, Davis J, Degroeve S, Déjardin A, Depamphilis C, Detter J, Dirks B, Dubchak I, Duplessis S, Ehlting J, Ellis B, Gendler K, Goodstein D, Gribskov M, Grimwood J, Groover A, Gunter L, Hamberger B, Heinze B, Helariutta Y, Henrissat B, Holligan D, Holt R, Huang W, Islam-Faridi N, Jones S, Jones-Rhoades M, Jorgensen R, Joshi C, Kangasjärvi J, Karlsson J, Kelleher C, Kirkpatrick R, Kirst M, Kohler A, Kalluri U, Larimer F, Leebens-Mack J, Leplé JC, Locascio P, Lou Y, Lucas S, Martin F, Montanini B, Napoli C, Nelson DR, Nelson C, Nieminen K, Nilsson O, Pereda V, Peter G, Philippe R, Pilate G, Poliakov A, Razumovskaya J, Richardson P, Rinaldi C, Ritland K, Rouzé P, Ryaboy D, Schmutz J, Schrader J, Segerman B, Shin H, Siddiqui A, Sterky F, Terry A, Tsai CJ, Uberbacher E, Unneberg P, Vahala J, Wall K, Wessler S, Yang G, Yin T, Douglas C, Marra M, Sandberg G, de Peer YV, Rokhsar D (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313(5793):1596–1604. doi:10.1126/science.1128691
Wong KM, Suchard MA, Huelsenbeck JP (2008) Alignment uncertainty and genomic analysis. Science 319(5862):473–476. doi:10.1126/science.1151532
Xue C, Fu Y (2009) Preservation of duplicate genes by originalization. Genetica 136(1):69–78. doi:10.1007/s10709-008-9311-5
Yamane K, Yano K, Kawahara T (2006) Pattern and rate of indel evolution inferred from whole chloroplast intergenic regions in sugarcane, maize and rice. DNA Res 13(5):197–204. doi:10.1093/dnares/dsl012
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci 13(5):555–556
Yang Z, Bielawski JP (2000) Statistical methods for detecting molecular adaptation. Trends Ecol Evol 15(12):496–503
Zhang Z, Gerstein M (2003) Patterns of nucleotide substitution, insertion and deletion in the human genome inferred from pseudogenes. Nucleic Acids Res 31(18):5338–5348
Acknowledgments
The authors are grateful to the bioinformatics group of the IPK Gatersleben as well as Christian Kauhaus at University of Jena for access to the computer clusters. Matthias Höffken provided useful hints on Python scripting and contributed Python code for statistical analyses. The authors thank Adam Eyre-Walker for discussion on the MK analysis of inparalogs. The authors also thank D. Tian and two anonymous referees for their valuable comments on the manuscript. The study was supported by an undergraduate scholarship by the Studienstiftung des deutschen Volkes to TG and IPK core funding to KS.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gossmann, T.I., Schmid, K.J. Selection-Driven Divergence After Gene Duplication in Arabidopsis thaliana . J Mol Evol 73, 153–165 (2011). https://doi.org/10.1007/s00239-011-9463-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-011-9463-2