Abstract
Ornithine decarboxylase (ODC) catalyzes the first and rate limiting step in the biosynthesis of polyamines in most eukaryotes. Because polyamines have pleiotropic and often dramatic effects on cellular processes at both high and low concentrations, ODC expression is tightly controlled. ODC is regulated by a family of polyamine-induced proteins, antizymes, which bind to, and inactivate it. In mammals, and apparently most vertebrates, antizymes are in turn antagonized by proteins called antizyme inhibitors. Antizyme inhibitors are homologs of ODC that have lost their decarboxylation activity but have retained their ability to bind antizyme, in most cases even more tightly than ODC. We present a phylogenetic analysis of over 200 eukaryotic homologs of ODC and evaluate their potential to be either true ODCs or catalytically inactive proteins that might be analogs of the previously identified antizyme inhibitors. This analysis yielded several orthologous groups of putative novel antizyme inhibitors each apparently arising independently. In the process we also identify previously unrecognized ODC paralogs in several evolutionary branches, including a previously unrecognized ODC paralog in mammals, and we evaluate their biochemical potential based on their pattern of amino acid conservation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
ODC catalyzes the decarboxylation of ornithine to produce the diamine putrescine. The polyamines spermidine and spermine are then derived from putrescine (Pegg 2006). Increased polyamine concentrations are required in rapidly growing cells, including cancer cells, and indeed targeting ODC activity is being tested for cancer treatment and prevention (Evageliou and Hogarty 2009; Gerner and Meyskens 2009), [as well as in treatment of Trypanosoma brucei infection (Heby et al. 2007; Balasegaram et al. 2009)]. Conversely, cells starved for polyamines accumulate in G1 and are prevented from undergoing DNA replication (Review, Oredsson 2003). The polyamine derived unusual amino acid hypusine is required for activity of eIF5A, which has recently been recognized as an elongation factor (Saini et al. 2009) and inhibitors of hypusine formation are being tested for inhibition of HIV expression (Hoque et al. 2009). Polyamines also have other important physiological effects unrelated directly to cell growth—for instance on certain ion channels (Fleidervish et al. 2008). In light of these effects of polyamines, it is not surprising that ODC activity is tightly regulated in cells. Regulation of ODC is known at the transcriptional, translational, and posttranslational levels. At the posttranslational level, ODC, a homodimer, is primarily regulated by the protein ornithine decarboxylase antizyme (antizyme) (Review, Coffino 2001). Antizyme achieves the down-regulation of ODC activity by binding to it directly and inhibiting it by preventing homodimer formation (both ODC subunits contribute toward the formation of the active site). Antizyme bound to ODC monomers subsequently presents these monomers for ubiquitin-independent degradation by the 26S proteasome (Murakami et al. 1992; Zhang et al. 2003; Takeuchi et al. 2008). In addition antizyme can inhibit the polyamine transporter in cells through an unknown mechanism. Antizyme is encoded by two slightly overlapping reading frames. A +1 frameshifting event is required for ribosomes to access the second and main reading frame and for the synthesis of functional antizyme. This frameshifting occurs at the end of the first reading frame and is induced by polyamines (Review, Ivanov and Matsufuji 2009).
Antizyme is a member of a protein family conserved from human to yeast and protists (Ivanov and Atkins 2007). Several distinct mammalian paralogs exist: the originally cloned antizyme 1 (Miyazaki et al. 1992; Matsufuji et al. 1995) that is expressed in all cells except developing male germ cells; the equally ubiquitously expressed antizyme 2 that is expressed at a lower level (Ivanov et al. 1998; Murai et al. 2009); and the tissue specific antizyme 3 expressed during a discrete stage of spermatogenesis (Ivanov et al. 2000; Tosaka et al. 2000; Snapir et al. 2009). Other vertebrates also have multiple paralogs. For example zebrafish, D. rerio, has four paralogs—two orthologs of mammalian antizyme 1, one ortholog of mammalian antizyme 2 and a fish-specific paralog called antizyme-R (Ivanov et al. 2007; Ivanov and Atkins 2007).
Antizyme itself is down-regulated by a protein called antizyme inhibitor (AZI). AZI is a homolog of ODC which has lost the ability to decarboxylate ornithine (Murakami et al. 1996, 2009; Kahana 2009). Mouse AZI has also lost the ability to form homodimers at physiological conditions (Albeck et al. 2008), but homodimer formation can be restored with the reversion of just 4 amino acid positions (Su et al. 2009). Other evidence shows that AZI does not form heterodimers with ODC and interfere with the catalytic activity of the latter (Murakami et al. 1996). Mouse AZI has also lost the ability to bind to PLP, an essential co-factor in the decarboxylation reaction. AZI, however, binds to antizyme even more tightly than does ODC resulting in antizyme sequestration (Fujita et al. 1982; Kitani and Fujisawa 1989; Nilsson et al. 2000; Cohavi et al. 2009). Cells overexpressing AZI have elevated ODC levels, increased growth and when injected into nude mice, give rise to tumors (Keren-Paz et al. 2006). AZI is involved in the regulation of centrosome duplication (Mangold et al. 2008) and homozygous mutant mice die on the first postnatal day (Tang et al. 2009). AZI was originally discovered in rats and orthologs have been identified in other vertebrates (Hascilowicz et al. 2002).
More recently a second antizyme inhibitor has been identified in mammals. It was initially termed “ODC-like,” later ODCp (for ODC paralog) but currently it is known as AZIN2 (Pitkänen et al. 2001; López-Contreras et al. 2006, 2008; Kanerva et al. 2008; Snapir et al. 2008). Unlike the originally identified AZI (AZIN1) which is ubiquitously expressed, expression of AZIN2 was initially thought to be restricted to brain and testis. However, it is now known to be expressed in sterodogenic cells of human ovaries and testes (Mäkitie et al. 2009) and in mast cells where it influences polyamine regulation of serotonin secretion (Kanerva et al. 2009). Another report suggests that AZIN2 expression could coincide with that of antizyme 3 during mammalian spermatogenesis and interaction between the two could be important for that process (López-Contreras et al. 2009). Homozygous antizyme 3 knockout in mice results in male infertility probably due to easy separation of sperm heads and tails (Tokuhiro et al. 2009). While AZIN1 has been visualized in the nucleus of transfected cells, AZIN2 has been visualized in the ER-Golgi intermediate compartment (ERGIC) and in the cis-Golgi network (López-Contreras et al. 2009). Endogenous AZIN1 changes its subcellular localization during the cell cycle (Mangold et al. 2008; Murakami et al. 2009). As some of its names suggest, AZIN2 is also a homolog of ODC that, like AZIN1, has lost the ability to decarboxylate ornithine but has retained the ability to bind to antizyme. AZIN2 represents a paralogous group independent of AZIN1. Its sequence is much more similar, than its AZIN1 counterpart, to that of ODC.
ODC belongs to the alanine racemase family of pyridoxal-5′-phosphate (PLP) dependant enzymes. In this family, it is most closely related to arginine decarboxylase (ADC) and diaminopimelate decarboxylase (DapDC) (Jansonius 1998). As referenced later, the ODC crystal structure with its ligand, has been solved and the molecular nature of its catalytic site determined—knowledge which has been corroborated by mutational analysis. The crystal structure of AZI is also available for comparison and contrast.
The decarboxylase reaction performed by an ODC dimer begins when the PLP bound Lys69 (using the numbering for human ODC, hODC, throughout) is displaced by L-ornithine followed by decarboxylation of the latter through a nucleophilic attack. It is believed that nucleophilic attack of the substrate is carried out by Cys360. Thus both Lys69 and Cys360 are absolutely essential for the decarboxylation activity and are completely conserved in all known functional decarboxylases, including ODC, ADC and DapDC. Mutation of Lys69 to Ala reduces the k cat by more than 3000-fold and the decarboxylation reaction by 1000-fold (Osterman et al. 1999; Myers et al. 2001). Mutation of Lys69 to Arg reduces the activity of the enzyme to 0.02% of wild type (Tsirka and Coffino 1992). Mutation of Cys360 to Ala completely abolishes activity of the enzyme (Tsirka et al. 1993). Subsequent studies showed that mutation of Cys360 to Ala reduces k cat by 35-fold and causes a decarboxylation-dependent transamination reaction to form pyridoxamine 5-phosphate (PMP) and γ-aminobutyraldehyde, instead of PLP and putrescine (Jackson et al. 2000; Myers et al. 2001). Taken together this evidence indicates that any ODC homolog with alterations in Lys69 or Cys360 is unlikely to be enzymatically active.
Several residues, Asp88, Glu94, Arg154, His197, Ser200, Gly235-Gly237, Glu274, Arg277 Asp332 and Tyr389, in addition to Lys69, either directly bind PLP or otherwise stabilize its coordination (Grishin et al. 1999; Kern et al. 1999). Mutation of Arg277 to Ala increases the K m for PLP by 270-fold compared to wild type, with a 50% drop in k cat of the reaction (Osterman et al. 1997). Mutation of His197 to Ala reduces ODC activity to less than 1% of wild type (Lu et al. 1991). Mutation of Asp88, Glu94 and Asp274 to Ala decreases the k cat of the reaction by 180-, 40- and 50-fold respectively (Osterman et al. 1995). Mutation of Tyr397 to Ala reduces the k cat by 336-fold (Myers et al. 2001). Asp88 and Glu94 additionally interact with Lys69 following displacement of the latter by the formation of putrescine (Jackson et al. 2000). Five of the residues above—Lys69, Gly235–Gly237 and Glu274—are completely conserved in a group of 130 compiled and analyzed ODC, ADC and DapDC sequences (Kidron et al. 2007). The same study identified two additional absolutely conserved amino acids, Gly171 and Gly387, which have structural roles. An earlier study had shown that Gly387 is essential for homodimer formation and Gly387Ala mutation completely abolishes ODC activity (Tobias et al. 1993).
Two amino acids are involved in a salt bridge that forms between the two ODC monomers—Lys169 with Asp364 (Grishin et al. 1999). Mutation of Lys169 to Arg reduced the activity of the enzyme to 0.04% of wild type (Tsirka and Coffino 1992). In addition to their other role in PLP binding described above, Arg277, Asp332, and Tyr389 have also been shown to play an important role in dimer formation (Su et al. 2009). Asp361, adjacent to Cys360, and also Asp332, are believed to interact in a specific manner with the substrate (Grishin et al. 1999; Jackson et al. 2003). Mutating Asp361 to Ala increases the K m for ornithine by 2000-fold (Osterman et al. 1995). Finally, Phe397 is believed to be an integral part of the L-CO2 binding pocket during the decarboxylation (Jackson et al. 2003). Mutation of Phe397 to Ala reduces the steady state rate of product production by 150 folds and the rate of decarboxylation by 2100 folds (Jackson et al. 2003).
In this report we comprehensively compile and investigate sequences of ODC homologs from eukaryotes with the emphasis on species and groups of species, known to harbour antizyme gene(s) (Ivanov and Atkins 2007). Analyzing the identity of the 20 key amino acid residues described above—Lys69, Asp88, Glu94, Arg154, Lys169, Gly171, His197, Ser200, Gly235, Gly236, Gly237, Glu274, Arg277, Asp332, Cys360, Asp361, Asp364, Gly387, Tyr389, and Phe397—is the criteria used in determining whether any given ODC homolog protein is likely to be an active decarboxylase or a catalytically inactive protein similar, at least in this respect, to the known vertebrate AZIs.
Materials and Methods
Compiling ODC Homologs
A list of organisms that have antizyme genes was compiled previously (Ivanov and Atkins 2007). The sequences of ODC homologs from these organisms were then compiled as follows: GenBank was searched with the help of BLAST (http://www.ncbi.nlm.nih.gov/BLAST/) for complete protein sequences (350+ amino acids) corresponding to all ODC homologs in a given organism (irrespective of whether the entry is labeled ODC, ADC, DapDC, AZI, or currently has some other annotation). Another group of sequences was compiled by searching with BLAST and annotated manually using data publically available through Genbank from whole genome sequencing projects in various stages of completion. A third set was compiled by manually assembling EST contigs from the GenBank EST database. ODC homologs from several organisms not known to contain antizyme were also used including: 5 trypanosomal ODCs, 10 plant and algal ODCs, 2 Plasmodium ODCs, and 1 amoeba ODC. We also included 9 ODC homologs from Phytophthora (water molds—Oomycetes). Phytophthora species each appear to have as many as 5 homologs of ODC; some of which group with ODC, some with DapDC, and yet others with ambiguous assignment (Fig. 1). Their sequences were primarily included to help with rooting the phylogenetic tree. We specifically tried to identify and use the three most divergent ADCs and DapDCs (Kidron et al. 2007). In a few instances of special interest, we included less complete sequences (i.e., contigs shorter than 350 amino acids). Some sequences from organisms that are close relatives were excluded to reduce the number of uninformative entries. The number of sequences used for the analysis was 229. All the sequences thus compiled, and used in this study, are listed in Fig. S2, Supplementary Material online.
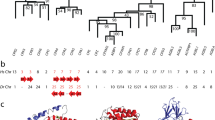
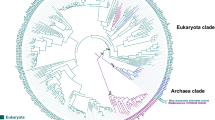
Phylogenetic analysis of the ODC homologs analyzed. The unrooted tree is based on amino acid sequences and was calculated using MrBayes (Huelsenbeck and Ronquist 2001; Ronquist and Huelsenbeck 2003) and plotted with NJ-Plot (Perrière and Gouy 1996). Probabilistic support for key branches is given. The scale bar in the lower left measures the number of amino acid substitutions per site (accounting for multiple substitutions at the same site). Basal branches with dubious support (typically probabilities <0.70 and/or not present in additional realizations of the tree) were collapsed. Putative homologs with impaired ornithine decarboxylation activity are shown in red. Putative functional ODC homologs in Ascomycotal fungi are shown in blue. Putative functional ODC homologs in vertebrates are shown in green. Putative functional ODC homologs in invertebrates are shown in brown. All other functional ODC homologs are shown in black. ODC partial sequences are marked by “*”
Phylogenetic Reconstruction
An amino acid alignment of all compiled sequences was constructed with ClustalW (http://www.clustal.org/). The alignment in Fig. S2 is a hybrid of ClustalW and some manual realignment. A few partial or terminally truncated sequences were excluded from the primary phylogenetic analysis. Alignment columns with gaps in seven or more of the remaining 222 sequences were excluded, leaving 319 amino acid positions. Phylogenetic trees were calculated using the Bayesian likelihood-based method implemented in MrBayes version 3.1.2 (Huelsenbeck and Ronquist 2001; Ronquist and Huelsenbeck 2003), using the amino acid mixture of models with fixed rate matrices and equal rates. Separate trees were calculated for the vertebrate clade, the major fungi ODC1 clade, all other clades combined, and a reduced selection of 80 taxa that sampled all major clades. Each tree was independently calculated at least twice.
The vertebrate tree was also recalculated with the addition of four sequences (Squalus_acanthias_AZIA1, Tetraodon_nigroviridis_ODC1, Tetraodon_nigroviridis_ODC2, Fugu_rubripes_ODC2) that were removed from the initial analysis due to terminal alignment gaps (Fig. S2). The tree in Fig. 1 is a composite of these analyses.
Sequence Analysis
The ratio of nonsynonymous to synonymous substitution rates, Ka/Ks, was calculated using the codeml program from the PAML package (Yang 1997, 2007). AZIB, AZIC, and AZID subgroups were individually realigned and analyzed with the global Ka/Ks value calculated using either the F3x4 model or a table of codon frequencies, with removal of alignment columns that contain gaps.
Results and Discussion
ODC sequences were compiled by searching GenBank with the BLAST algorithm (see “Materials and Methods”). The search identified a total of 319 sufficiently complete sequences. These were then pared down to 229 to reduce the number of sequences from closely related species. The phylogenetic relationship of 224 of them (including 3 known ADCs and 3 known DapDCs used to help root the ODC tree) is presented in Fig. 1. An alignment of the 229 amino acid sequences is shown in Fig. S1. Glu94 is invariable in the set of sequences we analyzed and is thus uninformative and was removed from further consideration. The region surrounding Asp332 is insufficiently conserved to allow reliable alignment of this residue in all homologs. Therefore, it was used for predicting the biochemical properties of select proteins only in the special cases where reliable alignment assessment is possible. Analysis of the 229 sequences identified 68 ODC homologs which have alterations in at least one of the 18 remaining key amino acids described above—henceforth referred to as “the 18 key amino acids.” The alterations in these 68 proteins are shown in Table 1. Three additional proteins have reliable alterations of Asp332 (Fig. S1, and discussion of the cases in the main text below).
One key conclusion presented here is that AZI genes have likely emerged repeatedly during eukaryotic evolution and diversification. Additionally, several AZI genes have undergone branch- (subphylum-) specific duplication. This creates some confusion with the existing nomenclature for naming newly identified AZI genes. To avoid this confusion, we propose an original system of naming newly identified AZI genes and use it for the remainder of this article. A sequential alphabetical letter starting with “A”, and designating members belonging to the same orthologous antizyme inhibitor gene subfamily, is added after the letters “AZI” (the letter “N” is omitted to avoid confusion with one of the existing synonyms of antizyme inhibitor—“AZIN”). Where necessary, a number is also added corresponding to the order of discovery for each additional paralog within an orthologous group (i.e., in an antizyme inhibitor subfamily). Accordingly, the originally identified and cloned antizyme inhibitor from rats is designated AZIA1 while the more recently discovered protein with antizyme inhibitor activity, usually referred to as ODCp or AZIN2, is now designated AZIB1.
ODC Homologues in Vertebrates
Three paralogous groups of vertebrate ODC genes have been described so far. These include ODC itself plus AZIA and AZIB. Most vertebrates have a single true ortholog of ODC though a number of exceptions do exist. In the fish of the Salmonidae family, including salmon and trout, where the entire genome has undergone complete duplication within the last 100 million years (McKay et al. 2004), two ODC paralogs are present. Something similar has occurred in the frog Xenopus laevis which also has two true ODC orthologs, most likely resulting from “a recent” gene duplication (Evans et al. 2004)—the two proteins are 95% identical and 97% similar at the amino acid level. The second ODC ortholog of X. laevis, ODC1-2, described here for the first time, is not to be confused with the gene designated XODC2 (Cao et al. 2001), which is an ortholog of AZIB1 and is discussed below. As should be expected, in all true vertebrate orthologs of ODC, the 18 key residues are identical to their counterparts in hODC suggesting that these genes are functional ODCs.
In addition to the “local” duplications of ODC described above, our analysis identified two paralogous groups of “ODC-like” proteins that, to our knowledge, have never been described before. In the case described first we tentatively designate the encoding gene as ODC2/AZIC. This group of genes is present in three ray-finned fishes—Gasterosteus aculeatus, Tetraodon nigroviridis, and Fugu rubripes. No expressed sequence tags (ESTs) corresponding to these genes can be identified but the genes are maintained without inactivating frameshift mutations and are phylogenetically closer to each other than to ODC1 in the corresponding organism. Application of MLOGD (Firth and Brown 2006), testing a coding model against a noncoding model, resulted in a positive coding signature—indicating that they are likely under purifying selection and therefore expressed. It is worth noting that they are under less stringent purifying selection than the members of the ODC1 group and are diverging at about twice the rate of ODC1 (Fig. 1). Similarly Ka/Ks was calculated to be ~0.30 for the AZIC clade, compared with ~0.17 for AZIA and AZIB (ODCp), and ~0.06 for ODC1. In part because sequence data for this clade is incomplete, it is difficult to predict the biochemical properties of its members.
The second group, whose origin is independent, is present in mammals—monotremes, marsupials, and eutherian mammals—indicating that this paralog diverged from ODC1 prior to the radiation of extant mammals. Like ODC2/AZIC in ray-finned fish, this mammalian gene is rarely expressed, as judged by the low number of known ESTs. This is the most likely reason why the paralogous group has never been noted before. Nonetheless, application of MLOGD again resulted in a positive coding signature indicating that they are likely to be functional, though the few mammalian ESTs available do not provide clues as to specific function. Like the analog in fish, the members of this mammalian orthologous group are diverging faster than their ODC1 counterparts (Fig. 1) and this gene is under less stringent purifying selection than the gene encoding ODC1 (Ka/Ks ~ 0.18 compared to Ka/Ks ~ 0.06 for ODC1) though, here, the degree of purifying selection is not dissimilar to that calculated for AZIA or AZIB. In most members of this subfamily, the region surrounding Asp332 aligns poorly to the same region in ODC1 and only some have an aspartate residue in the vicinity suggesting possibly impaired ODC activity. Additionally, the ortholog in cattle, Bos taurus, has two alterations to the 18 key amino acids—Arg154Trp and Gly235Arg—which are expected to inactivate its decarboxylation activity. In total the evidence suggests that at least some or most members of this subfamily might have biochemical activity other than ornithine decarboxylation—e.g., antizyme inhibition—and are tentatively designated ODC2/AZID. In the ape lineage (humans, chimps, and gorillas) this gene has undergone several frameshift and nonsense mutations that would be predicted to lead to severe truncations though curiously the mRNA continues to be expressed in humans (data not shown).
The root of the phylogenetic branch containing all orthologs of AZIA is located close to the root of the other vertebrate homologs of ODC-indicating that the subfamily evolved in, and is specific to, the vertebrate lineage. Further evidence supporting this hypothesis comes from analysis of the AZIA ortholog from the lamprey (Petromyzon marinus). In lampreys the ortholog of AZIA has remained a functional ODC, as judged by the 18 key amino acids. In fact the two ODC paralogs in lamprey can only be assigned to the ODC and the AZIA orthologous groups, respectively, on the basis of features within their 5′ UTRs which clearly distinguish the two groups (Ivanov et al. 2008). It is noteworthy that this 5′ UTR feature unambiguously puts AZIA members in one orthologous group and ODC1 and AZIB members in another group, independent of the phylogenetic support presented in Fig. 1.
Some vertebrate orthologs of AZIA are present in more than one copy per genome. Like the ODC gene, there are extra copies in X. laevis and Salmonidae as a result of relatively recent whole genome duplications but, in addition all ray-fin fish appear to have a paralogous pair of AZIA genes apparently resulting from a single ancient gene duplication event. These are designated here as AZIA1 and AZIA2. AZIA1 has been cloned and studied previously (Hascilowicz et al. 2002), but to our knowledge AZIA2 is described here for the first time. In individual fish species it appears that the AZIA1 mRNA is more abundant than the mRNA for AZIA2 but specific experiments would be required to untangle the functions of the two forms. As might be expected for ODC homologs that have evolved in a direction independent of decarboxylation activity, AZIA genes have accumulated numerous alterations in the 18 key amino acids. In one ortholog or another, at least 14 of the 18 positions are altered. Individual genes have as many as 11 (T. nigroviridis AZIA1) and as few as 4 (in all mammals) changes. Four residues, Asp88, Arg154, Arg277, and Asp332, involved in binding and coordination of PLP or interacting with the substrate, are altered in all known AZIA orthologs (Table 1; Fig. S1). All other alterations are lineage specific. The most parsimonious analysis suggests that the original inactivating mutation was Arg277Ser.
The first cloned AZIA gene was from rat. At the time, it was noted with curiosity that it has both residues essential for decarboxylation—Lys69 and Cys360. Many AZIA orthologs, however, carry incapacitating alterations in these two essential positions-providing additional evidence that the products of members of this gene subfamily do not perform decarboxylation.
While ODC and AZIA orthologs are apparently present in all vertebrates, including lampreys, the AZIB paralogous group is only found in tetrapods. In Xenopus the AZIB ortholog has been designated XODC2 (Cao et al. 2001) and unlike AZIB proteins in mammals, it is likely an active ornithine decarboxylase as judged by the 18 key amino acids all of which are identical to hODC (Fig. S1). The XODC2 mRNA expression pattern is different from that of XODC1 indicating that their roles are at least partially nonredundant. However, the XODC2 pattern of expression is also markedly different from that of human AZIB1 and its expression is not confined to brain and testes as is the human ortholog. The AZIB orthologs in mammals contain numerous deviations in key amino acids for them to be functional ornithine-, or any other, decarboxylase. To the Cys360Val noted previously in mouse and human AZIB, several other important changes can be added. These include Asp88Ser in all mammals; Arg154Cys in human, macaque, mouse, rat, cattle, and dog; Cys360Ala in dog and cattle; Lys169Arg in mouse and rat; Lys69Gly in mouse; Lys69Arg in rat; Glu274Arg in dog; Glu274Lys and Asp361Ser in cattle; and Asp361Glu in mouse. The chicken, Gallus gallus, AZIB ortholog has some of the same key residues altered as those in eutherian mammals, specifically Arg154 and Cys360; however, none of the changes are the same. Cys360 in the reptile, Anolis carolinensis, shares the same alteration, Cys360Gly, which exists in birds—indicating that perhaps they were inactivated as ODCs in a single event. In platypus (O. anatinus), neither the Cys360 nor the Asp88 are altered but other inactivating alterations exist—e.g., Arg154Ser and Asp364Glu. This pattern is consistent with AZIB evolving from ODC to AZI in at least three separate events. For this reason we propose calling these forms AZIA1 (in marsupials and eutherian mammals), AZIB2 (in monotremes), and AZIB3 (in chicken and gecko). It remains to be seen if all three have similar biochemical properties.
ODC Homologs in Invertebrates
In most invertebrates ODC exists in a single copy. The most fascinating exception involves the ODC homologs in the mosquitoes Aedes aegypti, Culex pipiens, and Anopheles gambiae, each with no less than 6 paralogs. The main ODC paralog in mosquitoes, determined by similarity to other insect ODCs, and designated here as ODC1, is highly transcribed as judged by the number of supporting ESTs. Most of the others are supported by one or zero ESTs, suggesting that they are transcribed infrequently. In the phylogenetic tree of eukaryotic homologs of ODCs, these other paralogs form three distinct clusters. ODC1 orthologs, as indicated above, cluster with the other insect ODCs and not surprisingly appear to be functional ODCs—a conclusion based on the fact that their 18 key residues are identical to hODC. None of the other mosquito ODC homologs cluster with insect ODCs and all have at least one alteration in a key amino acid (see Table 1 and Fig. S1). Curiously, mosquito ODCs2-6, with few exceptions, do not form neat orthologous clusters in the three species examined. Instead, some of these ODC homologs have corresponding orthologs in only one of the two other mosquito species. Several others appear to result from gene duplication events that postdate the divergence of three species from each other, about 150 million years ago (Gaunt and Miles 2002). The possible exception are the homologs designated here as ODC2/AZIE1. The A. gambiae paralog designated ODC3/AZIE8 has no changes in the 18 key amino acids present in hODC but the region that normally contains Asp332 has no identifiable aspartate and is poorly conserved relative to A. gambiae ODC1 (Fig. S1). ODC2/AZIE1 paralogs have two deviations shared in all three species—Ser200Cys and Asp332Arg (in that case the region normally surrounding Asp332 is well conserved relative to A. gambiae ODC1). The first change would be predicted to destabilize PLP binding while the second to affect binding to the substrate and dimer formation, in addition to PLP binding. All other paralogs contain changes in the 18 key residues that are predicted to be even more disruptive of ODC function. Four of them, designated here A. gambiae ODC4/AZIE3 and ODC6/AZIE4, A. aegypti ODC6/AZIE2 and C. pipiens ODC6/AZIE2, on the basis of their amino acid sequences, are predicted to have no decarboxylation activity at all: A. gambiae ODC4/AZIE3 has Lys69Gly while ODC6/AZIE4 has Cys360Met; A. aegypti and C. pipiens ODC6/AZIE2 have Lys69Arg and Lys69Gly, respectively, and both share Cys360Δ. In fact A. aegypti and C. pipiens ODC6/AZIE2 have, respectively, 13 and 11 of the 18 key amino acids altered. Although at this point purely speculative, it seems highly likely that many, or all, of the mosquito ODC homologs with alterations in the 18 key amino acids are analogs of the vertebrate antizyme inhibitors. The surprisingly large number of ODC homologs in mosquitoes suggests that the enzyme could play an unusual role in their physiology and could be a potential target for mosquito control or disease transmission.
Caenorhabditis elegans and related nematodes have two homologs of ODC. They are 27% identical and 45% similar to each other at the amino acid level, indicating an ancient divergence. Members of the first group, here designated ODC1, are diverging from each other at a significantly lower rate than members of the second group. Like the extra ODC homologs in mosquitoes, the second group in C. elegans does not cluster with other animal ODCs but instead forms its own cluster. ODC1 is a functional ODC as shown by recombinant expression (Macrae et al. 1995). Complete disruption of C. elegans ODC1 leads to total loss of ODC activity in the animals indicating that it is the sole ODC (Macrae et al. 1995) and suggesting a nonredundant role for proteins belonging to the second group. The latter, ODC2/AZIF, are described here for the first time. Their lack of ODC activity is entirely consistent with analysis of their amino acid sequence. No less than 12 of the 18 key residues are altered in the protein from C. elegans—most significantly Lys69Ala and Cys360Leu (Table 1). The situation is similar in two other nematode orthologs of this protein, in Caenorhabditis briggsae and Ancylostoma ceylanicum. On the basis of amino acid sequence analysis, members of this group cannot be active decarboxylases; they have completely lost the ability to catalyze decarboxylation, to bind PLP, and also possibly to form homo- and hetero-dimers. Once again the most likely explanation is that they have acquired antizyme inhibitory activity. There is a second curious parallel with the ODC1-AZIA pair in vertebrates. The non-AUG initiated conserved uORF (Ivanov et al. 2008) is present only in the ODC inactive paralog (i.e., in AZIA in vertebrates and ODC2/AZIF in C. elegans). Unlike ODC2/AZIC and ODC2/AZID in vertebrates, C. elegans ODC2/AZIF has very abundant mRNA as judged by the number of ESTs—perhaps a hundred times or more abundant than C. elegans ODC1.
Ciona intestinalis, a chordate, appears to have six ODC homologs. One of the paralogs, designated by us ODC1, clusters with other animal ODCs and has no alterations in the 18 key amino acids. By contrast, the other five paralogs, ODC2/AZIG1 to ODC6/AZIG5, form their own separate cluster which means they are closer to each other than either is to any other eukaryotic ODC homolog (the grouping is supported by 100% probability in our analysis—see Fig. 1). One, ODC4/AZIG3, has His197Phe and Ser200Asp alterations. The other four share an alteration of Asp361 to Ala. Two of them, ODC5/AZIG4 and ODC6/AZIG5, also share Cys360Gly which is predicted to completely abolish decarboxylation activity. There is a distinct possibility that these two, and perhaps all five, have evolved to become antizyme inhibitors.
Another interesting example of an invertebrate organism with multiple homologs of ODC is the lancelet, Branchiostoma floridae, a chordate. This organism also has three homologs of ODC. Two of them have an unaltered set of the 18 key amino acids. One, designated ODC1, clusters with other animal ODCs and is almost certainly a functional enzyme. The same is probably true for the second, ODC2, which is more distantly related to other animal ODCs. Curiously, B. floridae ODC2 clustered with C. intestinalis and the slime mold Dictyostelium discoideum, ODC in our analysis (99% probability—see Fig. 1). The third B. floridae paralog, however, has several very unusual features. In the larger phylogenetic tree, the protein clusters with the DapDCs. Significantly it carries the alteration Asp361Glu which exists naturally in the DapDCs, and is a position known to be important for substrate specificity. Another key feature shared with DapDC is an arginine in helix 11 which is absolutely conserved in these enzymes but is not present in ODCs. In at least one automatically generated database (from the Joint Genome Institute), this protein is already classified as DapDC. We refer to it as ODC3/DapDC pending confirmation of its biochemical activity. To our knowledge this is the first putative DapDC known from metazoans and the gene appears to be a product of horizontal gene transfer.
Several other invertebrates also have more than one ODC paralog. One example is in Drosophila melanogaster which has two copies of the ODC gene (Rom and Kahana 1993), both arising in the Drosophila species subgroup (data not shown). In both proteins all 18 key positions are identical to hODC but the second paralog has an Asp332Tyr alteration that is expected to have a deleterious effect on binding to PLP, the substrate, and for dimer formation. The entire region normally surrounding Asp332 is poorly conserved in D. melanogaster ODC2 but well conserved in ODC1. The biochemical properties of these paralogs have not been investigated, however, based on their sequence, the second is another putative antizyme inhibitor and is here designated ODC2/AZIH. All orthologs of ODC2 in the Drosophila species for which sequence information is available, share the Asp332Tyr alteration (data not shown). The wasp Nasonia vitripennis also has two homologs of ODC. ODC1 groups with the other insect ODCs (Fig. 1) while the second homolog forms a separate unrelated phylogenetic branch (data not shown). Just like ODC2/AZIH in Drosophila, to which it is not related by common descent, Asp332 is altered, in this instance to His. As with Drosophila ODC2/AZIH, this is expected to inactivate the enzyme, and we tentatively designate it ODC2/AZIJ. Hydra magnipapillata, a cnidarian, apparently has as many as four ODC homologs. For only two could we compile sufficient sequence for analysis. In ODC1, the more highly expressed paralog judged by the number of ESTs, all 18 key positions are identical to hODC. In the second homolog three alterations are observed—Ser200Thr, Arg277Lys (Table 1) and Asp332Ser (Fig. S1). These three alterations are expected to completely inactivate ODC activity, and this second homolog is tentatively designated ODC2/AZIK. The flat worm, Schmidtea mediterranea, has at least three ODC homologs. In ODC1 all 18 key positions are identical to hODC. Both the second and third paralogs have a number of alterations in the 18 key amino acids. The second has the alterations Ser200Ile and Glu274Thr (sequence information for the last 6 key amino acid positions is lacking). The third has alterations Ser200Asn, Arg277Thr, and Asp332Glu. The alterations are expected to lead to loss of ODC activity and the proteins are two more potential antizyme inhibitors and designated ODC2/AZIL1 and ODC3/AZIL2.
ODC Homologs in Fungi
Although most fungi have a single homolog of ODC, a number of them have more than one. The ODC paralogs in the Ascomycota subphylum Pezizomycotina are especially interesting. Only about a third of the species in this subphylum show evidence for more than one homolog of ODC per genome. But where more than one copy is present, the paralogous groups usually have deep roots implying divergence for a significant period of time and/or a higher rate of divergence for one of the paralogs. Like the ODC paralogs in mosquitoes mentioned above, wherever sufficient EST data are available, the evidence suggests that one of the two paralogs is transcribed many fold in excess of the other(s). The abundantly transcribed copy is designated here as ODC1. In all the cases the ODC1s are more closely related to the other known fungal ODCs than to any of the extra Pezizomycotina ODCs. From the extra paralogs only the Botryotinia fuckeliana paralog we designate ODC3, which appears to have arisen from ODC1 through a relatively recent gene duplication, groups together with other ODC1 Pezizomycotina paralogs (Fig. 1). The rest of the Pezizomycotina ODC paralogs form three distinct clusters, the members of which we designate ODC2,3/AZIM, ODC2/AZIP, and ODC3/AZIR, respectively. One cluster consists of: Aspergillus clavatus ODC2/AZIM, Neosartorya fischeri ODC2/AZIM1, N. fischeri ODC3/AZIM2 (resulting from a relatively resent duplication of ODC2/AZIM), Aspergillus terreus ODC2/AZIM, and B. fuckeliana ODC2/AZIM. The second cluster consists of: Uncinocarpus reesii ODC2/AZIP, Coccidioides immitis ODC2/AZIP, and Aspergillus fumigatus ODC2/AZIP. The third cluster consists of: Gibberella moniliformis ODC3/AZIR and Fusarium oxysporum ODC3/AZIR. Members of ODC2,3/AZIM do not have any alteration in the 18 key amino acids but they appear to lack Asp332 and, like some other putative AZIs, the region immediately surrounding it is poorly conserved in this group (Fig. S1). This alteration is expected to affect substrate specificity, PLP binding, and dimer formation. Several members of the same cluster also share Ser200Thr (Table 1), expected to affect PLP binding. Proteins in the ODC2/AZIP cluster, like members of the ODC2,3/AZIM cluster, have alteration of Asp332 and again the region surrounding it is poorly conserved. Proteins in the third cluster have diverged from the main ODC paralog more than those in the first two. For example, the proteins corresponding to F. oxysporum ODC1 and ODC3/AZIR are only 21% identical and 40% similar to each other at the amino acid level. In comparison, the corresponding numbers for A. terreus ODC1 and ODC2/AZIM are 46% and 63%, respectively, while the numbers for C. immitis ODC1 and ODC2/AZIP are 37% and 55%, respectively. In addition to the Ser200Thr they share with some of the members of the ODC2/AZIM orthologous group, the members of the ODC3/AZIR group have Asp332Ser and Gly387Ala alterations which are likely to completely inactivate decarboxylation.
At least three of the Pezizomycotina ODC1 orthologs have been shown experimentally to be functional ODCs (Williams et al. 1992; Bailey et al. 2000; Guevara-Olvera et al. 2000). Deleting the gene in Neurospora crassa or Phaeosphaeria nodorum leads to complete loss of ODC activity; however, since both of these organisms lack AZIM/AZIP/AZIR orthologs, the results are not informative regarding the activity of these paralogs in general.
Perspective
Until now, no comprehensive phylogenetic analysis of the antizyme inhibitor genes in eukaryotes has been attempted. There has been some erroneous discussion of orthologs of antizyme inhibitor in invertebrates and attempts to demarcate the extent and spread of either AZIA or AZIB has been limited. There is also significant confusion in the naming and early classification of the protein we here label AZIB. In this study we attempted to clarify all these issues and in the process made several interesting discoveries, the most important of which is the recurrent emergence of apparently enzymatically inactive forms of ODC.
Overall, the structure of the phylogenetic tree for the main paralogs of ODC matches the known phylogenetic relationship of the organisms to which they belong. The most notable exception is the ODC of Trypanosoma brucei which clusters with ODCs in the vertebrate branch of the tree. This clustering has been noted previously and was attributed to horizontal gene transfer (Steglich and Schaeffer 2006). As discussed above, Branchiostoma floridae ODC3/DapDC is almost certainly another example of horizontal gene transfer. The significance of the clustering of flat worm homologs (Dugesia japonica ODC and Schmidtea mediterranea ODC3/AZIL2) with Plasmodium ODC is less clear. Plasmodium ODCs are unique in that they are bifunctional enzyme fusions of S-adenosyl-l-methionine decarboxylase and ornithine decarboxylase (Wrenger et al. 2001). There is no hint of such fusion in flat worms. The future availability of additional flat worm ODC sequences may allow productive revisiting of this unusual clustering.
In all but one of the 68 cases of proteins that vary from human ODC in one or more of the key 18 positions, the same organism has at least one other ODC homolog with a complete set of the key amino acids. The one exception is the ODC homolog in Pichia guilliermondii which is supported by a single high throughput genomic sequence and could well be a sequencing error (Table 1). This strongly implies the possibility that the paralog that deviates from the consensus has evolved into a protein with a function other than decarboxylation of ornithine. We propose that the paralogs with multiple alterations have no decarboxylation activity at all and that some, or all, are instead antizyme inhibitors. Since ODC functions as a dimer and the active site is formed in the interface between the two monomers, one alternative possibility is that some of the inactive paralogs form heterodimers with the enzymatically active paralog and inactivate it—in effect serving as ornithine decarboxylase inhibitors. Currently no ODC homolog is known to possess this activity; however, mutants of mammalian ODC1 can be made which do exhibit this activity (Coleman et al. 1994; Tobias and Kahana 1993). If such activity is demonstrated for any of the newly identified and presumably catalytically dead ODC paralogs, this would add yet another layer of complexity to the already known multitudinous layers of ODC regulation.
References
Albeck S, Dym O, Unger T, Snapir Z, Bercovich Z, Kahana C (2008) Crystallographic and biochemical studies revealing the structural basis for antizyme inhibitor function. Protein Sci 17:793–802
Bailey A, Mueller E, Bowyer P (2000) Ornithine decarboxylase of Stagonospora (Septoria) nodorum is required for virulence toward wheat. J Biol Chem 275:14242–14247
Balasegaram M, Young H, Chappuis F, Priotto G, Raguenaud ME, Checchi F (2009) Effectiveness of melarsoprol and eflornithine as first-line regimens for gambiense sleeping sickness in nine Médecins Sans Frontières programmes. Trans R Soc Trop Med Hyg 103:280–290
Cao Y, Zhao H, Hollemann T, Chen Y, Grunz H (2001) Tissue-specific expression of an ornithine decarboxylase paralogue, XODC2, in Xenopus laevis. Mech Dev 102:243–246
Coffino P (2001) Regulation of cellular polyamines by antizyme. Nat Rev Mol Cell Biol 2:188–194
Cohavi O, Tobi D, Schreiber G (2009) Docking of antizyme to ornithine decarboxylase and antizyme inhibitor using experimental mutant and double-mutant cycle data. J Mol Biol 390:503–515
Coleman CS, Stanley BA, Viswanath R, Pegg AE (1994) Rapid exchange of subunits of mammalian ornithine decarboxylase. J Biol Chem 269:3155–3158
Evageliou NF, Hogarty MD (2009) Disrupting polyamine homeostasis as a therapeutic strategy for neuroblastoma. Clin Cancer Res 15:5956–5961
Evans BJ, Kelley DB, Tinsley RC, Melnick DJ, Cannatella DC (2004) A mitochondrial DNA phylogeny of African clawed frogs: phylogeography and implications for polyploid evolution. Mol Phylogenet Evol 33:197–213
Firth AE, Brown CM (2006) Detecting overlapping coding sequences in virus genomes. BMC Bioinform 7:75
Fleidervish IA, Libman L, Katz E, Gutnick MJ (2008) Endogenous polyamines regulate cortical neuronal excitability by blocking voltage-gated Na+ channels. Proc Natl Acad Sci USA 105:18994–18999
Fujita K, Murakami Y, Hayashi S (1982) A macromolecular inhibitor of the antizyme to ornithine decarboxylase. Biochem J 204:647–652
Gaunt MW, Miles MA (2002) An insect molecular clock dates the origin of the insects and accords with palaeontological and biogeographic landmarks. Mol Biol Evol 19:748–761
Gerner EW, Meyskens FL Jr (2009) Combination chemoprevention for colon cancer targeting polyamine synthesis and inflammation. Clin Cancer Res 15:758–761
Grishin NV, Osterman AL, Brooks HB, Phillips MA, Goldsmith EJ (1999) X-ray structure of ornithine decarboxylase from Trypanosoma brucei: the native structure and the structure in complex with alpha-difluoromethylornithine. Biochemistry 38:15174–15184
Guevara-Olvera L, Hung CY, Yu JJ, Cole GT (2000) Sequence, expression and functional analysis of the Coccidioides immitis ODC (ornithine decarboxylase) gene. Gene 242:437–448
Hascilowicz T, Murai N, Matsufuji S, Murakami Y (2002) Regulation of ornithine decarboxylase by antizymes and antizyme inhibitor in zebrafish (Danio rerio). Biochim Biophys Acta 1578:21–28
Heby O, Persson L, Rentala M (2007) Targeting the polyamine biosynthetic enzymes: a promising approach to therapy of African sleeping sickness, Chagas’ disease, and leishmaniasis. Amino Acids 33:359–366
Hoque M, Hanauske-Abel HM, Palumbo P, Saxena D, D’Alliessi-Gandolfi D, Park MH, Pe’ery T, Mathews MB (2009) Inhibition of HIV-1 gene expression by Ciclopirox and Deferiprone, drugs that prevent hypusination of eukaryotic initiation factor 5A. Retrovirology 6:90
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogeny. Bioinformatics 17:754–755
Ivanov IP, Atkins JF (2007) Ribosomal frameshifting in decoding antizyme mRNAs from yeast and protists to humans: close to 300 cases reveal remarkable diversity despite underlying conservation. Nucleic Acids Res 35:1842–1858
Ivanov IP, Matsufuji S (2009) Autoregulatory frameshifting in antizyme gene expression governs polyamine levels from yeast to mammals. In: Atkins JF, Gesteland RF (eds) Recoding: expansion of decoding rules enriches gene expression. Springer Publishers, New York, pp 281–300
Ivanov IP, Gesteland RF, Atkins JF (1998) A second mammalian antizyme: conservation of programmed ribosomal frameshifting. Genomics 52:119–129
Ivanov IP, Rohrwasser A, Terreros DA, Gesteland RF, Atkins JF (2000) Discovery of a spermatogenesis stage-specific ornithine decarboxylase antizyme: antizyme 3. Proc Natl Acad Sci USA 97:4808–4813
Ivanov IP, Pittman AJ, Chien CB, Gesteland RF, Atkins JF (2007) Novel antizyme gene in Danio rerio expressed in brain and retina. Gene 387:87–92
Ivanov IP, Loughran G, Atkins JF (2008) uORFs with unusual translational start codons autoregulate expression of eukaryotic ornithine decarboxylase homologs. Proc Natl Acad Sci USA 105:10079–10084
Jackson LK, Brooks HB, Osterman AL, Goldsmith EJ, Phillips MA (2000) Altering the reaction specificity of eukaryotic ornithine decarboxylase. Biochemistry 39:11247–11257
Jackson LK, Goldsmith EJ, Phillips MA (2003) X-ray structure determination of Trypanosoma brucei ornithine decarboxylase bound to D-ornithine and to G418: insights into substrate binding and ODC conformational flexibility. J Biol Chem 278:22037–22043
Jansonius JN (1998) Structure, evolution and action of vitamin B6-dependent enzymes. Curr Opin Struct Biol 8:759–769
Kahana C (2009) Regulation of cellular polyamine levels and cellular proliferation by antizyme and antizyme inhibitor. Essays Biochem 46:47–61
Kanerva K, Mäkitie LT, Pelander A, Heiskala M, Andersson LC (2008) Human ornithine decarboxylase paralogue (ODCp) is an antizyme inhibitor but not an arginine decarboxylase. Biochem J 409:187–192
Kanerva K, Lappalainen J, Mäkitie LT, Virolainen S, Kovanen PT, Andersson LC (2009) Expression of antizyme inhibitor 2 in mast cells and role of polyamines as selective regulators of serotonin secretion. PLoS One 4:e6858
Keren-Paz A, Bercovich Z, Porat Z, Erez O, Brener O, Kahana C (2006) Overexpression of antizyme-inhibitor in NIH3T3 fibroblasts provides growth advantage through neutralization of antizyme functions. Oncogene 25:5163–5172
Kern AD, Oliveira MA, Coffino P, Hackert ML (1999) Structure of mammalian ornithine decarboxylase at 1.6 A resolution: stereochemical implications of PLP-dependent amino acid decarboxylases. Structure 7:567–581
Kidron H, Repo S, Johnson MS, Salminen TA (2007) Functional classification of amino acid decarboxylases from the alanine racemase structural family by phylogenetic studies. Mol Biol Evol 24:79–89
Kitani T, Fujisawa H (1989) Purification and characterization of antizyme inhibitor of ornithine decarboxylase from rat liver. Biochim Biophys Acta 991:44–49
López-Contreras AJ, López-Garcia C, Jiménez-Cervantes C, Cremades A, Peñafiel R (2006) Mouse ornithine decarboxylase-like gene encodes an antizyme inhibitor devoid of ornithine and arginine decarboxylating activity. J Biol Chem 281:30896–30906
López-Contreras AJ, Ramos-Molina B, Cremades A, Peñafiel R (2008) Antizyme inhibitor 2 (AZIN2/ODCp) stimulates polyamine uptake in mammalian cells. J Biol Chem 283:20761–20769
López-Contreras AJ, Sánchez-Laorden BL, Ramos-Molina B, de la Morena ME, Cremades A, Peñafiel R (2009) Subcellular localization of antizyme inhibitor 2 in mammalian cells: Influence of intrinsic sequences and interaction with antizymes. J Cell Biochem 107:732–740
Lu L, Stanley BA, Pegg AE (1991) Identification of residues in ornithine decarboxylase essential for enzymic activity and for rapid protein turnover. Biochem J 277(Pt 3):671–675
Macrae M, Plasterk RH, Coffino P (1995) The ornithine decarboxylase gene of Caenorhabditis elegans: cloning, mapping and mutagenesis. Genetics 140:517–525
Mäkitie LT, Kanerva K, Sankila A, Andersson LC (2009) High expression of antizyme inhibitor 2, an activator of ornithine decarboxylase in steroidogenic cells of human gonads. Histochem Cell Biol. E-pub ahead of print
Mangold U, Hayakawa H, Coughlin M, Münger K, Zetter BR (2008) Antizyme, a mediator of ubiquitin-independent proteasomal degradation and its inhibitor localize to centrosomes and modulate centriole amplification. Oncogene 27:604–613
Matsufuji S, Matsufuji T, Miyazaki Y, Murakami Y, Atkins JF, Gesteland RF, Hayashi S (1995) Autoregulatory frameshifting in decoding mammalian ornithine decarboxylase antizyme. Cell 80:51–60
McKay SJ, Trautner J, Smith MJ, Koop BF, Devlin RH (2004) Evolution of duplicated growth hormone genes in autotetraploid salmonid fishes. Genome 47:714–723
Miyazaki Y, Matsufuji S, Hayashi S (1992) Cloning and characterization of a rat gene encoding ornithine decarboxylase antizyme. Gene 113:191–197
Murai N, Shimizu A, Murakami Y, Matsufuji S (2009) Subcellular localization and phosphorylation of antizyme 2. J Cell Biochem 108:1012–1021
Murakami Y, Matsufuji S, Kameji T, Hayashi S, Igarashi K, Tamura T, Tanaka K, Ichihara A (1992) Ornithine decarboxylase is degraded by the 26S proteasome without ubiquitination. Nature 360:597–599
Murakami Y, Ichiba T, Matsufuji S, Hayashi S (1996) Cloning of antizyme inhibitor, a highly homologous protein to ornithine decarboxylase. J Biol Chem 271:3340–3342
Murakami Y, Suzuki J, Samejima K, Kikuchi K, Hascilowicz T, Murai N, Matsufuji S, Oka T (2009) The change of antizyme inhibitor expression and its possible role during mammalian cell cycle. Exp Cell Res 315:2301–2311
Myers DP, Jackson LK, Ipe VG, Murphy GE, Phillips MA (2001) Long-range interactions in the dimer interface of ornithine decarboxylase are important for enzyme function. Biochemistry 40:13230–13236
Nilsson J, Grahn B, Heby O (2000) Antizyme inhibitor is rapidly induced in growth-stimulated mouse fibroblasts and releases ornithine decarboxylase from antizyme suppression. Biochem J 346:699–704
Oredsson S (2003) Polyamine dependence of normal cell-cycle progression. Biochem Soc Trans 31:366–370
Osterman AL, Kinch LN, Grishin NV, Phillips MA (1995) Acidic residues important for substrate binding and cofactor reactivity in eukaryotic ornithine decarboxylase identified by alanine scanning mutagenesis. J Biol Chem 270:11797–11802
Osterman AL, Brooks HB, Rizo J, Phillips MA (1997) Role of Arg-277 in the binding of pyridoxal 5′-phosphate to Trypanosoma brucei ornithine decarboxylase. Biochemistry 36:4558–4567
Osterman AL, Brooks HB, Jackson L, Abbott JJ, Phillips MA (1999) Lysine-69 plays a key role in catalysis by ornithine decarboxylase through acceleration of the Schiff base formation, decarboxylation, and product release steps. Biochemistry 38:11814–11826
Pegg AE (2006) Regulation of ornithine decarboxylase. J Biol Chem 281:14529–14532
Perrière G, Gouy M (1996) WWW-query: an on-line retrieval system for biological sequence banks. Biochimie 78:364–369
Pitkänen LT, Heiskala M, Andersson LC (2001) Expression of a novel human ornithine decarboxylase-like protein in the central nervous system and testes. Biochem Biophys Res Commun 287:1051–1057
Rom E, Kahana C (1993) Isolation and characterization of the Drosophila ornithine decarboxylase locus: evidence for the presence of two transcribed ODC genes in the Drosophila genome. DNA Cell Biol 12:499–508
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Saini P, Eyler DE, Green R, Dever TE (2009) Hypusine-containing protein eIF5A promotes translation elongation. Nature 459:118–121
Snapir Z, Keren-Paz A, Bercovich Z, Kahana C (2008) ODCp, a brain- and testis-specific ornithine decarboxylase ornithine decarboxylase paralogue, functions as an antizyme inhibitor, although less efficiently than AZI1. Biochem J 410:613–619
Snapir Z, Keren-Paz A, Bercovich Z, Kahana C (2009) Antizyme 3 inhibits polyamine uptake and ornithine decarboxylase (ODC) activity, but does not stimulate ODC degradation. Biochem J 419:99–103
Steglich C, Schaeffer SW (2006) The ornithine decarboxylase gene of Trypanosoma brucei: Evidence for horizontal gene transfer from a vertebrate source. Infect Genet Evol 6:205–219
Su K-L, Liao Y-F, Hung H-C, Liu G-Y (2009) Critical factors determining dimerization of human antizyme inhibitor. J Biol Chem 284:26768–26777
Takeuchi J, Chen H, Hoyt MA, Coffino P (2008) Structural elements of the ubiquitin-independent proteasome degron of ornithine decarboxylase. Biochem J 410:401–407
Tang H, Ariki K, Ohkido M, Murakami Y, Matsufuji S, Li Z, Yamamura K (2009) Role of ornithine decarboxylase antizyme inhibitor in vivo. Genes Cells 14:79–87
Tobias KE, Kahana C (1993) Intersubunit location of the active site of mammalian ornithine decarboxylase as determined by hybridization of site-directed mutants. Biochemistry 32:5842–5847
Tobias KE, Mamroud-Kidron E, Kahana C (1993) Gly387 of murine ornithine decarboxylase is essential for the formation of stable homodimers. Eur J Biochem 218:245–250
Tokuhiro K, Isotani A, Yokota S, Yano Y, Oshio S, Hirose M, Wada M, Fujita K, Ogawa Y, Okabe M, Nishimune Y, Tanaka H (2009) OAZ-t/OAZ3 is essential for rigid connection of sperm tails to heads in mouse. PLoS Genet 5:e1000712
Tosaka Y, Tanaka H, Yano Y, Masai K, Nozaki M, Yomogida K, Otani S, Nojima H, Nishimune Y (2000) Identification and characterization of testis specific ornithine decarboxylase antizyme (OAZ-t) gene: expression in haploid germ cells and polyamine-induced frameshifting. Genes Cells 5:265–276
Tsirka S, Coffino P (1992) Dominant negative mutants of ornithine decarboxylase. J Biol Chem 267:23057–23062
Tsirka SE, Turck CW, Coffino P (1993) Multiple active conformers of mouse ornithine decarboxylase. Biochem J 293(Pt 1):289–295
Williams LJ, Barnett GR, Ristow JL, Pitkin J, Perriere M, Davis RH (1992) Ornithine decarboxylase gene of Neurospora crassa: isolation, sequence, and polyamine-mediated regulation of its mRNA. Mol Cell Biol 12:347–359
Wrenger C, Luersen K, Krause T, Muller S, Walter RD (2001) The Plasmodium falciparum bifunctional ornithine decarboxylase, S-adenosyl-L-methionine decarboxylase, enables a well balanced polyamine synthesis without domain-domain interaction. J Biol Chem 276:29651–29656
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl BioSci 13:555–556
Yang Z (2007) PAML 4: a program package for phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Zhang M, Pickart CM, Coffino P (2003) Determinants of proteasome recognition of ornithine decarboxylase, a ubiquitin-independent substrate. EMBO J 22:1488–1496
Acknowledgments
This study was supported by an award from Science Foundation Ireland and National Institutes of Health grant R01 GM079523 to J.F.A.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ivanov, I.P., Firth, A.E. & Atkins, J.F. Recurrent Emergence of Catalytically Inactive Ornithine Decarboxylase Homologous Forms That Likely Have Regulatory Function. J Mol Evol 70, 289–302 (2010). https://doi.org/10.1007/s00239-010-9331-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-010-9331-5