Abstract
Previous studies have reported a positive correlation between the GC content of the double-stranded regions of structural RNAs and the optimal growth temperature (OGT) in prokaryotes. These observations led to the hypothesis that natural selection favors an increase in GC content to ensure the correct folding and the structural stability of the molecule at high temperature. To date these studies have focused mainly on ribosomal and transfer RNAs. Therefore, we addressed the question of the relationship between GC content and OGT in a different and universally conserved structural RNA, the RNA component of the signal recognition particle (SRP). To this end we generated the secondary structures of SRP-RNAs for mesophilic, thermophilic, and hyperthermophilic bacterial and archaeal species. The analysis of the GC content in the stems and loops of the SRP-RNA of these organisms failed to detect a relationship between the GC contents in the stems of this structural RNA and the growth temperature of bacteria. By contrast, we found that in archaea the GC content in the stem regions of SRP-RNA is highest in hyperthermophiles, intermediate in thermophiles, and lower in mesophiles. In these organisms, we demonstrated a clear positive correlation between the GC content of the stem regions of their SRP-RNAs and their OGT. This correlation was confirmed by a phylogenetic nonindependence analysis. Thus we conclude that in archaea the increase in GC content in the stem regions of SRP-RNA is an adaptation response to environmental temperature.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Several studies have shown that the GC content of structural RNAs correlates positively with the optimal growth temperature (OGT) in prokaryotes (Galtier and Lobry 1997; Nakashima et al. 2003; Wang and Hickey 2002; Wang et al. 2006). These studies have established that the increase in GC content concentrates in the double-stranded stem regions of the molecule. Compared to AT pairs, GC pairs are more thermostable because they have an additional hydrogen bond. Thus, it is likely that the increase in the GC content of the stems of a nucleic acid will help to strengthen the stability of its secondary structures. This has led to the proposal of the thermal adaptation hypothesis. This hypothesis suggests that the increase in the GC content in stem regions of structural RNAs results from natural selection acting to favor the thermostability of the molecule. However, to date studies supporting the thermal adaptation hypothesis have focused mainly on ribosomal and transfer RNAs (rRNA and tRNA). The current availability of large genomic datasets offers the possibility of testing this hypothesis more soundly by extending the study of thermal adaptation to other structural RNAs. Therefore, in the present study we addressed the question of the relationship between GC content and OGT in a different and universally conserved structural RNA, the RNA component of the signal recognition particle (SRP).
The SRP is a ribonucleoprotein complex that recognizes nascent polypeptides destined for secretion or membrane insertion. The SRP binds to the signal sequence of the nascent polypeptide emerging from the ribosome and directs the polypeptide toward the SRP receptor (SR) of the targeted membrane (Egea et al. 2004; Halic et al. 2004; Keenan et al. 2001; Gilmore et al. 1982; Walter and Blobel 1981). Both the SRP and the SR contain GTPase domains, and their interaction leads to the reciprocal activation of GTP hydrolysis, allowing delivery of the secretory protein into the nearby translocation channel and recycling of the SRP (Connolly et al. 1991). Experimental studies have shown that the RNA component of the SRP is indispensable for cotranslational protein targeting. SRP-RNA catalytically accelerates the interaction of SRP and SR, which stimulates their GTPase activities. Recently, it has been shown that SRP-RNA accelerates the formation of the complex only when the SRP is bound to a signal sequence (Bradshaw et al. 2009). Thus, SRP-RNA acts as a molecular switch that renders the SRP-SR complex responsive to signal peptide recruitment, coupling GTP hydrolysis to efficient protein targeting.
Genome studies have shown that the SRP-dependent protein targeting mechanism is evolutionarily conserved in the three living domains. However, there are significant differences among the SRPs of Eukaryota, Archaea, and Bacteria. These differences concern both the protein and the RNA components (Zwieb et al. 2005). The mammalian SRP consists of SRP-RNA (7S RNA) and six proteins, called SRP9, SRP14, SRP19, SRP54, SRP68, and SRP72. SRP-RNA (~300 nucleotides long) folds in a secondary structure, which is subdivided in two domains, the Alu and S domains. The Alu domain contains helices 2, 3, and 4 and a portion of helix 5. This domain is associated with the heterodimer SRP9/14 and is thought to modulate the elongation rate of the secretory protein. The S domain also comprises a portion of helix 5 and helices 6–8 and is associated with the proteins SRP19, SRP54, and SRP68/72. This domain is involved in the recognition and capture of the signal peptide (Batey et al. 2000). Archaeal SRP-RNAs (7S RNA) are similar to their eukaryotic homologues. In these organisms SRP-RNA (~300 nucleotides long) is also divided into two domains. The Alu domain comprises helices 1–5 (helix 1 being unique to archaea and some gram-positive bacteria). As for eukaryotic SRP-RNA, the archaeal S domain is composed of a portion of helix 5 and helices 6–8. Despite their similarity to eukaryotic SRPs, only two proteins, SRP19 and SRP54, are believed to be part of the archaeal SRP complex (Eichler and Moll 2001; Zwieb and Eichler 2002). In the majority of bacteria the SRP is composed of a small SRP-RNA (4.5 SRNA) and a homologue of protein SRP54, known as Ffh (Bernstein et al. 1993). Bacterial SRP-RNA (~100 nucleotides long) forms a simple hairpin corresponding to helix 8 and a portion of helix 5. Although most bacterial SRPs lack the Alu domain, some gram-positive bacteria encode a relatively large SRP-RNA (~250 nucleotides). The secondary structure of these SRP-RNAs is similar to that of the archaea, but they lack helix 6 (Larsen and Zwieb 1991; Zwieb et al. 2005). In addition to their secondary structure features, tertiary interactions between the apical loops of helices 3 and 4, involving up to six GC base-pairings, have been described in the SRP-RNAs of eukaryotes, archaea, and some bacteria (Zwieb et al. 1996). Another tertiary interaction involving a hydrogen-bonded A–A pair forms between the loops of helices 6 and 8 in eukaryotic and archaeal SRP-RNAs (Hainzl et al. 2002). These interactions could be involved, respectively, in the early steps of SRP-RNA folding and SRP assemby (Huck et al. 2004).
The crucial role of SRP-RNA in the function of the SRP-dependent translocation mechanism suggests that preservation of the structural features of this RNA is essential. Therefore, we reasoned that in the face of variable environmental parameters such as temperature, natural selection would have acted on this molecule to preserve the stability of its secondary and tertiary structures. Thus, SRP-RNA appeared to be a good candidate to test the thermal adaptation hypothesis. Since the broadest range of growing temperatures is found among prokaryotes, we focused our study on bacteria and archaea. To this end, we generated the secondary structures of the SRP-RNAs of hyperthermophilic, thermophilic, and mesophilic bacteria and archaea, and analyzed the GC content as a function of their OGT. Our study failed to detect a relationship between the GC content of bacterial SRP-RNA and the OGT of these organisms. By contrast, we demonstrated a clear relationship between the growing temperature of the archaea and the GC content of their SRP-RNAs.
Methods
Sequence Data
SRP-RNA sequences were downloaded from the SRP-RNA database, http://rnp.uthct.edu/rnp/SRPDB/SRPDB.html (Rosenblad et al. 2003); from the Integrated Microbial Genomes database, http://img.jgi.doe.gov/cgi-bin/pub/main.cgi (Markowitz et al. 2008); and the NCBI database, http://www.ncbi.nlm.nih.gov. A total of 92 bacterial and 65 archaeal SRP-RNA sequences were retrieved. Although there are more than 1,000 bacterial genomes in the databases, the vast majority of bacterial species are mesophiles, with an OGT below 40°C. Thus, for the purposes of our study we retrieved the SRP-RNA sequences of all the hyperthermopilic and thermophilic bacteria available in the databases. For the mesophiles, we used a sample representative of the different genera present in the databases. For each SRP-RNA sequence, the global GC content was calculated. In addition, we analyzed the secondary structure of these RNAs and calculated the GC content for the stem and loop regions. The secondary structures of the archaeal and bacterial SRP-RNA analyzed in this study were generated using the RNAalifold and RNA fold software from the Vienna RNA package, http://rna.tbi.univie.ac.at (Gruber et al. 2008; Zuker 2003). The RNAalifold software predicts the consensus structure of a set of aligned RNA sequences. It extends standard dynamic programming algorithms for RNA secondary structure prediction by averaging the energy contributions over all sequences and incorporating covariation terms into the energy model to reward compensatory mutations and penalize noncompatible base pairs. The output is a consensus structure for the aligned sequences. Then this consensus structure is used as a constraint to fold each individual sequence using RNAfold. The SRP-RNA sequences were aligned with CLUSTAL W.
Optimal Growth Temperature
OGTs for bacterial (Supplementary Table S1) and archaeal (Supplementary Table S2) species analyzed in this study were retrieved from the Prokaryotic Growth Temperature Database, http://pgtdb.csie.ncu.edu.tw (Huang et al. 2004), and from the German Collection of Microorganisms and Cell Cultures, http://www.dsmz.de. OGT data were carefully checked to ensure that they referred unambiguously to the OGT rather than to a permissive range. Some species are listed in these databases with slightly different OGTs, in such cases we calculated and used the average of the OGTs reported for our study.
Statistical Analyses
Statistical analyses were performed using the statistics software package MATLAB. The nonparametric Kruskal–Wallis rank-sum test was used to compare data among three temperature groups: mesophilic, thermophilic, and hyperthermophilic microorganisms. The Mann–Whitney–Wilcoxon rank-sum test (U-test) was used to compare the average GC content between two temperature groups such as mesophiles and thermophiles. Correlations were evaluated using the nonparametric Spearman’s (1904) rank correlation test. This test assesses how well an arbitrary monotonic function can describe the relationship between two variables, without making any other assumptions about the particular nature of the relationship between the variables. Spearman’s rank correlation coefficient (r s) is used as a measure of the relationship between two sets of ranked data; r s takes a value of −1 and +1. A positive correlation is one in which the ranks of both variables increase together. A correlation of +1 or −1 will arise if the relationship between the two variables is exactly linear. A negative correlation is one in which the ranks of one variable increase as the ranks of the other variable decrease. An r s close to zero indicates no linear relationship between the ranks (Corder and Foreman 2009). In the case of archaea we used the method of comparative analysis by independent contrasts to control for phylogenetic nonindependence of temperature and GC content in stems (Felsenstein 1985). The contrasts were calculated using the PDAP:PDTREE module of the MESQUITE software for phylogenetic analysis (Maddison and Maddison 2009; Midford et al. 2005). The independent contrasts method assumes a Brownian motion model of evolution where variance in a trait increases through evolutionary time and calculates the amount of change between sister taxa at each node of the phylogeny. Independent contrasts are generated by subtracting trait values between all pairs of sister taxa in the tree. Contrasts are standardized by dividing by their standard deviation, calculated as the square root of the sum of branch lengths between the pair of traits. The phylogenetic tree used in this study was kindly provided by Dr. C. Brochier. This tree was generated from the concatenated sequences of ribosomal proteins from 43 archaeal species for which complete genomes are available (Brochier-Armanet et al. 2008).
Results
Analysis of Global GC Content in Bacterial and Archaeal SRP-RNA
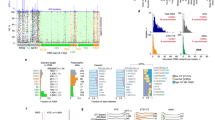
For the purposes of this study bacterial and archaeal species were grouped according to their OGTs. Three groups were defined: the mesophile group contained species with OGTs between 25 and 40°C (52 bacterial species and 22 archaea), the thermophile group included species with OGTs between 40 and 75°C (23 bacterial species and 20 archaea), and the hyperthermophile group contained species with OGTs of 75°C and above (7 bacterial species and 23 archaea). We initiated our study by calculating the global GC content in SRP-RNA sequences from the bacterial and archaeal species in each group (Fig. 1a, b). For bacterial species the average GC levels among the three groups appears to be relatively similar. The mean GC content is 58% for the mesophile group, 62% for the thermophiles, and 65% for the hyperthermophiles. The Kruskal–Wallis test shows that the differences among the three groups are not significant (P = 0.153). In addition, we used the Mann–Whitney–Wilcoxon rank sum test (U-test) to compare the average GC content between two temperature groups such as mesophiles versus thermophiles (P = 0.140), thermophiles versus hyperthermophiles (P = 0.620), and mesophiles versus hyperthermophiles (P = 0.130). Therefore, the analysis of the global GC content in bacterial SRP-RNA does not indicate a relationship with the OGT of these organisms. By contrast, in the case of the archaeal species a trend toward a higher GC content in the thermophilic and hyperthermophilic archaea is observed. The mean GC content is 55% for the mesophile group, 65% for the thermophiles, 65%, and 73% for the hyperthermophiles. The Kruskal–Wallis test shows that these differences are clearly significant (P < 0.001). The Mann–Whitney–Wilcoxon test confirms that the differences among the temperature groups are significant: mesophiles versus thermophiles (P < 0.0001), thermophiles versus hyperthermophiles (P < 0.001), and mesophiles versus hyperthermophiles (P < 0.0001). Thus, the results of the analysis of global sequences prompted us to look in more detail and to investigate the specificity of these compositional differences.
Global GC content in prokaryotic SRP-RNA. a Global GC content in bacterial SRP-RNA (n = 82). b Global GC content in archaeal SRP-RNA (n = 65). Boxes extend from the first quartile to the third quartile, with the median values indicated by the horizontal bar within the box. Whiskers extend to the smallest and largest nonoutlier values. Temperature ranges are as follows: mesophiles (25–40°C), thermophiles (40–75°C), and hyperthermophiles (≥75°C)
Compositional Analysis of Bacterial and Archaeal SRP-RNAs Secondary Structures as a Function of OGT
The secondary structure for SRP-RNA sequences of the 82 bacterial and 65 archaeal species analyzed above were obtained using the RNAalifold and RNA fold software from the Vienna RNA package (Gruber et al. 2008). Then the structures were partitioned into double-stranded stem regions and single-stranded loop regions, and the GC content was calculated for each region. In the case of the bacteria we analyzed separately the SRP-RNAs from species encoding a short SRP-RNA and those encoding a long SRP-RNA. The results of this analysis are summarized in Table 1. For both long and short bacterial SRP-RNA, we did not observe any statistically significant differences in the GC content in stems (or loops) among the three temperature groups. Then we considered the fact that the nucleotide sequences of bacterial SRP-RNAs are highly conserved, thus we performed a more careful analysis by distinguishing between conserved and variable regions (Supplementary Tables S3 and S4). Again, we failed to detect a correlation between the GC content of the stems in short or long bacterial SRP-RNAs and the OGT. However, the GC content in stems of the variable regions of hyperthermophilic short SRP-RNAs is slightly higher than in the other two groups. The mean stem GC content in the SRP-RNA variable regions is 52% ± 11% in mesophilic bacteria, 56% ± 7.3% in thermophilic bacteria, and 71% ± 5.8% in hyperthermophilic bacteria. The differences between the hyperthermophilic organisms and the other two groups are on the cusp of being statistically significant (U-test: mesophiles vs hyperthermophiles, P = 0.061; thermophiles vs hyperthermophiles, P = 0.073). For the archaeal species the compositional analysis of SRP-RNA secondary structures indicates that the GC content of the stem regions is highest in the hyperthermophiles, intermediate in the thermophiles, and lower in the mesophiles. The Kruskal–Wallis rank-sum test indicates that these differences are clearly significant (P < 0.0001). On the other hand, the GC content in the loop regions does not differ among the three archaeal groups, indicating that there is no correlation between the GC content of the loops and the OGT. This result is consistent with previous studies, which have reported that the GC content of stem regions in rRNAs and tRNAs correlates with the OGT, while the loop regions of these molecules show no correlation.
Therefore, we analyzed the secondary structures of archaeal SRP-RNAs versus their OGTs (Fig. 2). A clear linear and positive correlation is observed between the GC content of the stem regions and the OGT (slope = 0.38, r s = 0.821, P < 0.001). On the contrary, no trend is observed for the relationship between GC content of the loop regions and OGT (slope = 0.082, r s = 0.28, P < 0.001).
Phylogenetic Analysis of the Relationship Between OGT and GC Content in SRP-RNA of Archaea
Our results indicated an apparent correlation between the GC content of the stems of archaeal SRP-RNA and the OGT. However, this correlation could be due to shared ancestry rather than to thermal adaptation. Therefore, to control for this possibility we generated phylogenetically independent contrasts for GC content in the stems of SRP-RNA and OGT (Felsenstein 1985). This analysis indicates that a strong positive correlation (r s = 0.821, P < 0.001) persists when the phylogenetic relationships are taken into consideration (Fig. 3). Therefore, it can be concluded that the differences in the stem GC content of the SRP-RNAs of archaea growing at different temperatures are probably due to natural selection.
The Increase in the GC Content of Stems Is More Prominent in the Variable Regions of Archaeal SRP-RNAs
Sequence alignments have shown that archaeal SRP-RNAs contain relatively well-conserved regions as well as variable regions (Larsen and Zwieb 1991). Thus, the sequences of helices 2, 3, and 4, a portion of helix 5, and helix 8 remain relatively well conserved in the different archaeal SRP-RNAs. By contrast, helix 1, the largest portion of helix 5, and helix 6 display more variability in their sequence. To extend our analysis of the adaptation of this RNA molecule to OGT, SRP-RNAs were partitioned into conserved and variable regions, and the GC content in the stems and loops of these regions was analyzed (Table 2). The results show that the differences in the stem GC content of the three temperature groups are highest in the case of the variable regions. However, the Kruskal–Wallis test indicates that the differences are significant in both conserved and variable regions (P < 0.001). Phylogenetically uncontrolled analysis shows a relatively small correlation between the GC content of the conserved regions of SRP-RNA and the OGT (slope = 0.38, r s = 0.65, P < 0.001). A much better correlation with OGT is found when the variable regions are considered (slope = 0.47, r s = 0.79, p < 0.001). These correlations remain robust when phylogenetic nonindependence is analyzed. The contrasts for the conserved and variable regions are shown in Fig. 4. A particularly strong correlation between the GC content of the stem and the OGT was found when helix 6 was considered independently of the rest of the molecule (slope = 0.63, r s = 0.92, P < 0.001). This strong correlation was confirmed by the phylogenetic nonindependence analysis (Fig. 5).
Phylogenetically independent relationship between the GC content in stems of archaeal SRP-RNA and OGT. a Correlation between the GC content in stems of the conserved regions and OGT-independent standardized contrasts (r s = 0.723, P < 0.001). b Correlation between the GC content in stems of the variable regions and OGT-independent standardized contrasts (r s = 0.791, P < 0.001)
Discussion
Previous studies have shown a positive correlation between the GC content in the stems of structural RNAs and the growth temperature of prokaryotes and eukaryotes (Galtier and Lobry 1997; Nakashima et al. 2003; Varriale et al. 2008; Wang and Hickey 2002; Wang et al. 2006). Since GC pairs are more stable than AT, this led to the proposal of the thermal adaptation hypothesis, which states that the GC content in structural nucleic acids should be correlated with the growing temperature of organisms. However, these studies have focused on ribosomal RNAs (16S or 18S rRNA) and transfer RNAs. The current availability of large genomic databases, in particular, for bacteria and archaea, makes it possible to test this hypothesis more rigorously by extending the analysis of thermal adaptation to other structural RNAs. Thus, in the present study we have analyzed the compositional properties of a different structural RNA, SRP-RNA. Our goal was to determine whether there is a positive correlation between the GC content of the double-stranded regions of this universally conserved structural RNA and the growing temperature of bacteria and archaea.
In bacteria we could not detect a correlation between the GC content of SRP-RNA and the growing temperature. Among hyperthermophilic, thermophilic, and mesophilic bacteria, only the hyperthermophiles show a stem GC content slightly higher than that of the other groups. However, in this group the variations in GC content are very important even between organisms growing at similar temperatures. Thus Aquifex aeolicus (OGT = 85°C) shows a GC content in the stems as high as 80%, while Thermotoga petrophila (OGT = 80°C) has a stem GC content of only 65%. Such variations are found also among the thermophilic and mesophilic bacteria. In the bacterial domain we also analyzed 10 psychrophilic species (OGT ≤ 20°C). Interestingly, in this group the mean value for the GC content in the stems of SRP-RNA is 55 ± 6, which is comparable to the mean values found for mesophiles and thermophiles. This suggests that a minimum GC content could be required to ensure the proper folding of SRP-RNA, even at low environmental temperatures. Another interesting observation is that the GC content in the loops remains relatively low and constant among bacteria with different OGTs. This, as discussed below, could be related to their role in ensuring the proper folding of SRP-RNA and its physiological function.
Concerning the archaea this study shows a strong positive linear correlation between the GC content of the SRP-RNA and the OGT of these organisms. In particular, we determined that the GC content in the double-stranded regions of SRP-RNA are higher in hyperthermophiles, intermediate in thermophiles, and lower in mesophiles. Moreover, by using phylogenetic-independence contrast analysis, we demonstrated that this correlation is not an artifact due to phylogenetic relation but more likely reflects an adaptation resulting from natural selection. Also, by distinguishing between the conserved and the variable regions of the archaeal SRP-RNA, we observed that the increase in GC content in stem regions is more significant in helices 5 and 6. In particular, this analysis showed a very robust correlation between the GC content of the stem region of helix 6 and the OGT. This helix is present exclusively in SRP-RNAs of eukaryotes and archaea and displays extensive base-pairing (Zwieb et al. 2005). Helix 6 plays an important role in the assembly of the S-domain of the SRP. Biochemical and crystallographic studies have shown that the conserved adenosine in the apical GNAR (N is for any nucleotide, R is for a purine) tetraloop in helix 6 establishes a tertiary interaction with another adenosine residue in the apical loop of helix 8 (Hainzl et al. 2002; Yin et al. 2004; Zwieb et al. 2005). This interaction is stabilized by SRP19, which binds to the apical tetraloops of helices 6 and 8, clamps them together, and induces extensive interactions between them (Diener and Wilson 2000; Siegel and Walter 1988; Zwieb 1991, 1992, 1994). These interactions trigger an important conformational change in the long strand of the asymmetric loop in helix 8, which enhances the affinity of the RNA for the SRP54 protein (Egea et al. 2008; Hainzl et al. 2005). However, in vitro studies have reported that in some archaea, SRP54 can bind to SRP-RNA in the absence of SRP19 (Bhuiyan et al. 2000; Diener and Wilson 2000; Hainzl et al. 2005; Maeshima et al. 2001; Tozik et al. 2002; Yurist et al. 2007). In these organisms the tertiary interactions between helix 6 and helix 8 occur independently of SRP19 and could suffice to stabilize them in a tertiary structure with an intrinsic affinity for SRP54 (Hainzl et al. 2005; Yin et al. 2004). Thus, the strong positive correlation between the OGT and the GC content of helix 6 in archaeal SRP-RNA could be a reflection of its essential role in the assembly of the S-domain of the SRP. In the absence of a specific adaptation, the high temperatures endured by some archaea could destabilize the stem supporting the GNAR tetraloop and therefore prevent its interaction with the apical tetraloop in helix 8. This would result in failure of the assembly of the SRP. Therefore, it is tempting to speculate that the observed increase in the GC content of helix 6 of thermophilic and hyperthermophilic archaea could be an adaptation to preserve its thermal stability in order to ensure SRP assembly.
Helix 1 was also analyzed independently of the rest of the molecule. This helix pairs the terminal regions of archaea and long bacterial SRP-RNAs, and it has been suggested that it could play a role in preventing the unfolding of the SRP-RNA under the extreme conditions in which archaea thrive (Larsen and Zwieb 1991; Zwieb et al. 2005). In mesophilic and thermophilic archaea the mean GC content in helix 1 is very similar, 63 ± 8.1 and 65 ± 6.3, respectively. In hyperthermophilic archaea the mean GC content in helix 1 is higher, 80 ± 7.0. The difference between the hyperthermophilic group and the other two groups is statistically significant (U-test, P = 0.001). However, we could not find a linear relationship between the GC content of this stem and the OGT. Nevertheless, our data show that in the three temperature groups the GC base-pair content in helix 1 remains relatively high, conferring a great stability to the helix. This suggests that (independently of the growth temperature) the tight closure of the ends of SRP-RNA could be of some importance in the maintenance of the structure of SRP-RNA.
The observed increase in GC content of the stems of archaeal SRP-RNA is consistent with previous studies showing that, in prokaryotes, there is a significant correlation between the GC content of structural RNAs (rRNAs and tRNAs) and the growth temperature, and that the high GC content is concentrated in the double-stranded regions of the molecule. These studies also revealed that in rRNAs the GC content of single-stranded regions remains relatively low and constant. Similarly, our data indicate that the GC content of the single-stranded regions of SRP-RNA remains remarkably constant among the three temperature groups (~40%). It has been proposed that the evolutionary conserved level of guanine (and A) in the unpaired regions of rRNAs could be explained by the fact that these regions may play an essential role in maintaining the secondary and tertiary structure of the molecules (Wang and Hickey 2002). This is most likely the case in SRP-RNAs since phylogenetic comparative sequence analysis shows that there is a high degree of conservation among the nucleotides forming the loops and bulges of these RNAs (Schmitz et al. 1999). This is particularly true for the nucleotides of the loops in the S-domain of archaeal SRP-RNAs (Sauer-Eriksson and Hainzl 2003). Moreover, as mentioned above, structural, biochemical, and mutational studies on prokaryotic SRP-RNAs have shown that the loops of helix 8 of the S-domain are involved in tertiary interactions with the M-domain of protein SRP54, and that such interactions are essential for the physiological function of SRP-RNA (Hainzl et al. 2005). We have also mentioned above the essential role of the tertiary interactions between the apical loops of helices 6 and 8. Finally, mutational studies have shown that tertiary interactions between two distant loops in the Alu domain are necessary to ensure the correct folding and assembly of this region of the SRP-RNA (Huck et al. 2004). Thus, taken together, these results provide evidence that the specific increase in the GC content of the tem regions of archaeal SRP-RNA results from natural selection acting to maintain the folded structure of the molecule at higher temperatures. Moreover, the fact that the composition of the single-stranded regions remains constant and independent of the environmental temperature in this structural RNA indicates that natural selection operates also to ensure that this region remains single-stranded.
A striking finding of this study is that bacterial and archaeal SRP-RNAs seem to have evolved a different response to adapt to environmental temperature. The fact that we could not detect any significant increase in GC content in double-stranded regions of SRP-RNA (both short and long) of thermophilic and hyperthermophilic bacteria suggests that in these organisms this structural RNA could be stabilized by a different mechanism. Possible mechanisms include the use of modified nucleosides and/or the existence of specific RNA chaperones, which could direct the folding of newly transcribed SRP-RNA (Schroeder et al. 2004; Shigi et al. 2006; Wolin and Wurtmann 2006). However, a major problem encountered in the analysis of the thermal adaptation of SRP-RNA in bacteria is that the vast majority of these organisms are mesophiles. Currently, the number of thermophilic and, in particular, hyperthermophilic bacterial species in the genomic databases is very low. Thus, although our study failed to detect a relationship between the OGT of bacteria and the GC content in SRP-RNA stems, the data should not be overinterpreted, as they are not definitive evidence against the thermal adaptation hypothesis. By contrast, our study clearly demonstrates that, as for rRNAs and tRNAs, the SRP-RNAs from archaea have adapted to increasing environmental temperatures by increasing the GC content in their stems.
References
Batey RT, Rambo RP, Lucast L, Rha B, Doudna JA (2000) Crystal structure of the ribonucleoprotein core of the signal recognition particle. Science 287:1232–1239
Bernstein HD, Zopf D, Freymann DM, Walter P (1993) Functional substitution of the signal recognition particle 54-kDa subunit by its Escherichia coli homolog. Proc Natl Acad Sci USA 90:5229–5233
Bhuiyan SH, Gowda K, Hotokezaka H, Zwieb C (2000) Assembly of archaeal signal recognition particle from recombinant components. Nucleic Acids Res 28:1365–1373
Bradshaw N, Neher SB, Booth DS, Walter P (2009) Signal sequences activate the catalytic switch of SRP RNA. Science 323:127–130
Brochier-Armanet C, Boussau B, Gribaldo S, Forterre P (2008) Mesophilic Crenarchaeota: proposal for a third archaeal phylum, the Thaumarchaeota. Nat Rev Microbiol 6:245–252
Connolly T, Rapiejko PJ, Gilmore R (1991) Requirement of GTP hydrolysis for dissociation of the signal recognition particle from its receptor. Science 252:1171–1173
Corder GW, Foreman DI (2009) Nonparametric statistics for non-statisticians: a step-by-step approach. Wiley, New York
Diener JL, Wilson C (2000) Role of SRP19 in assembly of the Archaeoglobus fulgidus signal recognition particle. Biochemistry 39:12862–12874
Egea PF, Shan SO, Napetschnig J, Savage DF, Walter P, Stroud RM (2004) Substrate twinning activates the signal recognition particle and its receptor. Nature 427:215–221
Egea PF, Napetschnig J, Walter P, Stroud RM (2008) Structures of SRP54 and SRP19, the two proteins that organize the ribonucleic core of the signal recognition particle from Pyrococcus furiosus. PLoS One 3:e3528
Eichler J, Moll R (2001) The signal recognition particle of Archaea. Trends Microbiol 9:130–136
Felsenstein J (1985) Phylogenies and the comparative method. Am Nat 125:1–15
Galtier N, Lobry JR (1997) Relationships between genomic G+C content, RNA secondary structures, and optimal growth temperature in prokaryotes. J Mol Evol 44:632–636
Gilmore R, Walter P, Blobel G (1982) Protein translocation across the endoplasmic reticulum. II. Isolation and characterization of the signal recognition particle receptor. J Cell Biol 95:470–477
Gruber AR, Lorenz R, Bernhart SH, Neubock R, Hofacker IL (2008) The Vienna RNA websuite. Nucleic Acids Res 36:W70–W74
Hainzl T, Huang S, Sauer-Eriksson AE (2002) Structure of the SRP19 RNA complex and implications for signal recognition particle assembly. Nature 417:767–771
Hainzl T, Huang S, Sauer-Eriksson AE (2005) Structural insights into SRP RNA: an induced fit mechanism for SRP assembly. RNA 11:1043–1050
Halic M, Becker T, Pool MR, Spahn CM, Grassucci RA, Frank J, Beckmann R (2004) Structure of the signal recognition particle interacting with the elongation-arrested ribosome. Nature 427:808–814
Huang SL, Wu LC, Liang HK, Pan KT, Horng JT, Ko MT (2004) PGTdb: a database providing growth temperatures of prokaryotes. Bioinformatics 20:276–278
Huck L, Scherrer A, Terzi L, Johnson AE, Bernstein HD, Cusack S, Weichenrieder O, Strub K (2004) Conserved tertiary base pairing ensures proper RNA folding and efficient assembly of the signal recognition particle Alu domain. Nucleic Acids Res 32:4915–4924
Keenan RJ, Freymann DM, Stroud RM, Walter P (2001) The signal recognition particle. Annu Rev Biochem 70:755–775
Larsen N, Zwieb C (1991) SRP-RNA sequence alignment and secondary structure. Nucleic Acids Res 19:209–215
Maddison WP, Maddison DR (2009) Mesquite: a modular system for evolutionary analysis. Version 2.6. Available at: http://mesquiteproject.org
Maeshima H, Okuno E, Aimi T, Morinaga T, Itoh T (2001) An archaeal protein homologous to mammalian SRP54 and bacterial Ffh recognizes a highly conserved region of SRP RNA. FEBS Lett 507:336–340
Markowitz VM, Szeto E, Palaniappan K, Grechkin Y, Chu K, Chen IM, Dubchak I, Anderson I, Lykidis A, Mavromatis K, Ivanova NN, Kyrpides NC (2008) The integrated microbial genomes (IMG) system in 2007: data content and analysis tool extensions. Nucleic Acids Res 36:D528–D533
Midford PE, Garland T, Maddison WP (2005) PDAP package of mesquite. Version 1.07. Available at: http://mesquiteproject.org
Nakashima H, Fukuchi S, Nishikawa K (2003) Compositional changes in RNA, DNA and proteins for bacterial adaptation to higher and lower temperatures. J Biochem 133:507–513
Rosenblad MA, Gorodkin J, Knudsen B, Zwieb C, Samuelsson T (2003) SRPDB: Signal Recognition Particle Database. Nucleic Acids Res 31:363–364
Sauer-Eriksson AE, Hainzl T (2003) S-domain assembly of the signal recognition particle. Curr Opin Struct Biol 13:64–70
Schmitz U, Behrens S, Freymann DM, Keenan RJ, Lukavsky P, Walter P, James TL (1999) Structure of the phylogenetically most conserved domain of SRP RNA. RNA 5:1419–1429
Schroeder R, Barta A, Semrad K (2004) Strategies for RNA folding and assembly. Nat Rev Mol Cell Biol 5:908–919
Shigi N, Suzuki T, Terada T, Shirouzu M, Yokoyama S, Watanabe K (2006) Temperature-dependent biosynthesis of 2-thioribothymidine of Thermus thermophilus tRNA. J Biol Chem 281:2104–2113
Siegel V, Walter P (1988) Binding sites of the 19-kDa and 68/72-kDa signal recognition particle (SRP) proteins on SRP RNA as determined in protein-RNA “footprinting”. Proc Natl Acad Sci USA 85:1801–1805
Spearman C (1904) The proof and measurement of association between two things. Am J Psychol 15:72–101
Tozik I, Huang Q, Zwieb C, Eichler J (2002) Reconstitution of the signal recognition particle of the halophilic archaeon Haloferax volcanii. Nucleic Acids Res 30:4166–4175
Varriale A, Torelli G, Bernardi G (2008) Compositional properties and thermal adaptation of 18S rRNA in vertebrates. RNA 14:1492–1500
Walter P, Blobel G (1981) Translocation of proteins across the endoplasmic reticulum III. Signal recognition protein (SRP) causes signal sequence-dependent and site-specific arrest of chain elongation that is released by microsomal membranes. J Cell Biol 91:557–561
Wang HC, Hickey DA (2002) Evidence for strong selective constraint acting on the nucleotide composition of 16S ribosomal RNA genes. Nucleic Acids Res 30:2501–2507
Wang HC, Xia X, Hickey D (2006) Thermal adaptation of the small subunit ribosomal RNA gene: a comparative study. J Mol Evol 63:120–126
Wolin SL, Wurtmann EJ (2006) Molecular chaperones and quality control in noncoding RNA biogenesis. Cold Spring Harb Symp Quant Biol 71:505–511
Yin J, Huang Q, Pakhomova ON, Hinck AP, Zwieb C (2004) The conserved adenosine in helix 6 of Archaeoglobus fulgidus signal recognition particle RNA initiates SRP assembly. Archaea 1:269–275
Yurist S, Dahan I, Eichler J (2007) SRP19 is a dispensable component of the signal recognition particle in Archaea. J Bacteriol 189:276–279
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31:3406–3415
Zwieb C (1991) Interaction of protein SRP19 with signal recognition particle RNA lacking individual RNA-helices. Nucleic Acids Res 19:2955–2960
Zwieb C (1992) Recognition of a tetranucleotide loop of signal recognition particle RNA by protein SRP19. J Biol Chem 267:15650–15656
Zwieb C (1994) Site-directed mutagenesis of signal-recognition particle RNA. Identification of the nucleotides in helix 8 required for interaction with protein SRP19. Eur J Biochem 222:885–890
Zwieb C, Eichler J (2002) Getting on target: the archaeal signal recognition particle. Archaea 1:27–34
Zwieb C, Muller F, Larsen N (1996) Comparative analysis of tertiary structure elements in signal recognition particle RNA. Fold Des 1:315–324
Zwieb C, van Nues RW, Rosenblad MA, Brown JD, Samuelsson T (2005) A nomenclature for all signal recognition particle RNAs. RNA 11:7–13
Acknowledgments
We thank Drs. F. Langa, and J. Y. Zhang for their critical reading of the manuscript and suggestions. We are also grateful to Jean-Christophe Aude for helping with the statistical analysis. Dr. C. Brochier is gratefully acknowledged for providing the phylogenetic tree of the archaea based on concatenated ribosomal proteins. We thank the two anonymous referees for their helpful comments.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Miralles, F. Compositional Properties and Thermal Adaptation of SRP-RNA in Bacteria and Archaea. J Mol Evol 70, 181–189 (2010). https://doi.org/10.1007/s00239-009-9319-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-009-9319-1