Abstract
The codon assignment of the quasi-universal genetic code can be assumed to have resulted from the evolutionary pressures that prevailed when the code was still evolving. Here, we review studies of the structure of the genetic code based on optimization models. We also review studies that, from the structure of the code, attempt to derive aspects of the primordial circumstances in which the genetic code froze. Different rationales are summarized, compared with experimental data, discussed in the context of the transition from a RNA world to a DNA-protein world, and linked to the emergence of the last universal common ancestor.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the quasi-universal genetic code, 20 amino acids are assigned to 64 codons. This assignment is far from random, suggesting that it is not an accident, i.e., that natural selection shaped the genetic code (Sonneborn 1965; Knight 1999; Judson and Haydon 1999). Indeed, the genetic code possesses a pattern of degeneracies with a large number of symmetries by base substitutions. These have been studied extensively (Rumer 1966; Jestin 2006; Jestin and Kempf 2007). Symmetries by base substitutions have also been noted for codons and their corresponding amino acids associated to the 2′ and 3′ tRNA-aminoacylation classes (Jestin and Soulé 2007). Reasons for why some of these symmetries are present in the genetic code have been uncovered (Jestin 2009). Optimization models have been remarkably successful in providing links between experimental results from molecular biology and genetic code characteristics (Goldberg and Wittes 1966; Jestin and Kempf 1997; Koonin and Novozhilov 2009). Some of these optimization models will be reviewed in the first chapter. In the second chapter, we will summarize studies which analyze the quasi-universal structure of the genetic code for evidence about the evolutionary pressures that shaped the genetic code up until when it froze.
Optimization Models Highlighting Genetic Code Characteristics
These models rely on the hypothesis that the genetic code and the metabolic machineries have been coevolving (Jestin and Kempf 1997; Ardell and Sella 2002; Sella and Ardell 2006).
The observation that hydrophobic amino acids are mainly substituted by amino acids with similar chemical properties in in vitro translation systems suggests, for example, that the genetic code minimizes the effects of amino acid substitutions (Woese 1965).
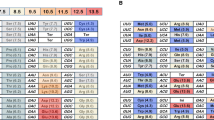
Further, all amino acids encoded by two codons differ by a transition (shown in rectangles in Fig. 1) and not by a transversion (Goldberg and Wittes 1966). This suggests that the code minimizes the effects of the most frequent substitutions which are generally transitions. These substitutions occur during DNA replication or transcription. In addition, most errors occurring during translation in codon–anticodon recognition process at the third codon position (the wobble position) have no effect if a purine is substituted by a purine or a pyrimidine is substituted by a pyrimidine (Crick 1966; Ninio 1971; Lagerkvist 1978).
Amino acids encoded by two codons differ by a transition on the third base which is a pyrimidine or a purine (rectangles). This observation is in accordance with two explanations: (i) as transitions are generally more frequent than transversions, the genetic code can be seen as optimized for the tolerance of the main substitutions (ii) optimal discrimination of bases in codon–anticodon recognition is then achieved. Further, four codons including the two stop codons TAA and TAG are single-base deletion hotspots during in vitro DNA synthesis as the YTRV deletion-prone template sequences are found in their codon (bold) as well as in the reverse complementary sequence (underlined). This observation is consistent with the genetic code’s optimization for frameshift mutation tolerance, as assignment of deletion-prone codons at the end of genes, i.e., stop codons, is most likely to yield functional proteins
The higher the binding energy (i.e., the number of hydrogen bonds in base pairing) for the first two bases, the higher is the probability for the third base to be misread. If the genetic code is optimized for tolerance of this type of errors, high binding energies should be associated to synonymous third codon bases. Hence all (GC)(GC)N codons encode the same amino acid ((GC) means here “G or C”). Conversely, all (AU)(AU)N codons have weaker pairing in the first two bases and can thus be discriminated on the basis of the identity of the third codon position (Lagerkvist 1978).
Our study reported the impact of single-base deletions which are highly deleterious within open reading frames as they cause frameshifts. We observed that the four codons most prone to in vitro single-base deletions during DNA polymerization are 5′ YTRV 3′ sequences and their reverse complementary sequences, the deletion occurring opposite the purine R (Jestin and Kempf 1997). These sequences were mapped on the genetic code codon table and yielded the two stop codons and their complementary codons (shown in bold and underlined in Fig. 1). Assigning the most single-base deletion-prone codons at the end of genes, i.e., stop codons, yield proteins most likely to be functional. Indeed, a single-base within a coding region is associated to truncated proteins most likely to have lost its biological function, whereas a single-base deletion within a stop codon yields full-length proteins fused to a C-terminal peptide, which are most likely to be functional (Fig. 2). This model provides therefore answers for why chain termination signals are encoded within the genetic code as codons (to minimize the deleterious effects of single-base deletions) and for why stop codons have these sequences rather than other sequences (single-base deletions occur mainly in YTRV sequences) (Jestin and Kempf 1997). As shown in Table 1, the codon assignment of stop signals varies depending on the species and on the location within the cell (organites or nucleus) and largely contributes to the diversity of genetic codes found in nature (Lehman and Jukes 1988; Osawa et al. 1992). Analysis of the consensus sequences for single-base deletions within these specific alternative locations may provide further opportunities to test the link between these deleterious consensus sequences and stop codons.
The effects of single-base deletions within a stop codon (1) or within a coding region (2) are compared after transcription and translation. Most single-base deletions occur within 5′ YTRV 3′ sequences or their reverse complementary sequences 5′ BYAR 3′ sequences. Deletions within a coding region yield truncated proteins which are likely to be non functional, whereas deletions within a stop codon yield full-length proteins with a peptide added at their C-termini, which are likely to be functional
In summary, these four optimization models support the theory that the code minimizes the deleterious effects of major mutations and optimizes information preservation (Sonneborn 1965; Goldberg and Wittes 1966; Lagerkvist 1978; Jestin and Kempf 1997; Ardell 1998; Freeland et al. 2000).
It should be noted that the same observations may have another interpretation, considering that the genetic code may have been fixed first and that the metabolic machinery evolved later. Under this assumption, the metabolic machinery evolved to preferentially tolerate certain errors because of the structure of the genetic code. But the model that assumes that the metabolic machinery evolved after the genetic code became fixed, has a major drawback in that it leaves open the question “Why is the genetic code the way it is ?” and is therefore much weaker from a theoretical point of view (Popper 1972).
The study of amino acids’ biosynthesis leads to another “coevolution theory”: it states that in case amino acid biosynthesis pathways coevolved with the genetic code, the introduction of a new amino acid in the code should involve a minimum number of mutations (Wong 1975, 1981, 2005, 2007). Biosynthetically related amino acids (highlighted by a circle on Fig. 3) are then expected to have closely related codons. Only two amino acid pairs (Phe-Leu and Arg-Ser noted in italics in Fig. 3) out of seven do not satisfy these criteria. The statistical significance of this finding has been extensively discussed (Amirnovin 1997; Knight et al. 1999; Di Giulio and Medugno 2000; Di Giulio 2005a).
Approaches to Reconstructing Aspects of the Evolutionary Pressures that Shaped the Genetic Code
The quasi-universal genetic code has been unchanged since at least the last universal common ancestor. It can be assumed, however, that in primordial organisms the genetic code underwent a period of changes under the influence of evolutionary pressures. During this period, any change in the genetic code was equivalent to the systematic introduction of a certain type of mutation all across the genome. The period in which the genetic code evolved therefore had to come to an end. Namely, the genetic code had to “freeze” when the genome started to exceed a certain size, beyond which changes in the genetic code amounted to a generally fatal number of mutations. (Note that this critical genome length has not yet been established.) The genetic code can therefore be viewed as valuable “fossil” (Gutfraind and Kempf 2008) whose study may reveal information about the primordial circumstances under which the genetic code formed (Di Giulio 2000, 2005b, 2005c; Gutfraind and Kempf 2008).
In particular, in (Di Giulio 2000, 2005b, 2005c), the assumption was made that the number of codons per amino acid in the genetic code reflects the frequency of amino acids in ancestral proteins when structuring of the genetic code took place. Thermophilic, barophilic, and acidophilic indexes were constructed for amino acids using protein comparisons from selected species (Picrophilus torridus and Thermoplasma volcanium with intracellular pH of 4.6 and 6.6, respectively, Pyrococcus furiosus and Pyrococcus abyssi isolated at 0.5 or 2000 m, respectively, Bacillus and Methanococcus species being mesophiles and (hyper)thermophiles, respectively). These indexes were then applied to the genetic code by making use of the assumption stated above. It was concluded that the genetic code structuring took place under high hydrostatic pressure, at high temperatures and in an acidic environment (Di Giulio 2000, 2005b, 2005c).
The study of (Gutfraind and Kempf 2008) argues that, very early in evolution, while the genetic code was still evolving, the evolutionary pressure to preserve genetic information can be assumed to have been paramount. Indeed, during the primordial RNA world, in the absence of sophisticated proof-reading mechanisms, organisms can be assumed to have experienced high rates of genetic errors. This implies that there was strong evolutionary pressure to reduce the phenotypical impact of genetic errors. This evolutionary pressure on the still evolving genetic code should have led to a structuring of the codon assignments so as to make the most prevalent types of genetic errors as neutral as possible for the phenotype. Therefore, the relative rates of the various types of genetic errors should have left characteristic imprints in the structure of the genetic code (Gutfraind and Kempf 2008). The authors used the working assumption that the prevalence of each type of genetic error was proportional to the extent to which the genetic code evolved to protect the phenotype from that type of error. It was possible then, to some extent, to reconstruct those relative error rates, as well as the nucleotide frequencies, for the time when the code was fixed.
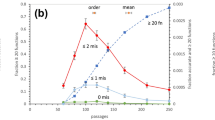
One of the main findings was that the frequencies of G and C in the genome were not elevated at the time when the genetic code froze. Since, for thermodynamic reasons, RNA in thermophiles tends to possess elevated G + C content (Galtier and Lobry 1997), this indicates that the fixation of the genetic code occurred in organisms which were either not thermophiles, i.e., for example not living close to hydrothermal vents, or that the code’s fixation occurred after the rise of DNA. The analysis in (Gutfraind and Kempf 2008) also yielded a reconstruction of the relative sizes of the primordial genetic error rates, see Fig. 2b in (Gutfraind and Kempf 2008). Notice that for any pair of back and forth error rates, say the pair A → G and G → A, only the larger rate of the two is given, since by the method employed only the larger of the two rates could be reconstructed reliably. As the authors pointed out, their results strongly indicate that transitions were more frequent than transversions also in primordial organisms, consistent with expectations.
Let us now attempt a preliminary comparison of the primordial error rates reconstructed in (Gutfraind and Kempf 2008) with experimental data (Fromant et al. 1995). This work used the polymerase chain reaction to amplify DNA fragments in the presence of varying concentrations of magnesium and manganese cations (among other parameters) and studied how these concentrations affect the various mutation rates in the replication process. Recall that manganese cations are known to increase the mutation rate and to extend the substrate specificity of family I DNA-dependent DNA-polymerases, which copy DNA as well as RNA either as wild-type polymerases or after introduction of one or several selected mutations at key positions within the polymerase scaffold (Ricchetti and Buc 1993; Cadwell and Joyce 1994; Jestin et al. 2005; Vichier-Guerre et al. 2006; Jestin and Vichier-Guerre 2005; Bahrami and Jestin 2008). The results that (Fromant et al. 1995) obtained for the mutation rates are listed in their Table 2 and we reproduce these data in Table 2. On the basis of the experimentally obtained data of (Fromant et al. 1995), let us now investigate whether a suitable combination of the concentrations of the two manganese and magnesium cations could account for the reconstructed error rates of (Gutfraind and Kempf 2008). To this end, let us recall first that in (Gutfraind and Kempf 2008) the six error rates were reconstructed only up to an overall constant factor which effectively leaves five independent rates to be matched. If it is not possible to choose these two dications concentrations to match the effectively five independent reconstructed error rates unambiguously, this would indicate that additional factors, such as the DNA polymerase sequence and structure have also significantly affected the primordial error rates. We begin the analysis by considering the reconstructed primordial relative mutation rates obtained in (Gutfraind and Kempf 2008). We recall that (Gutfraind and Kempf 2008) considered a family of scenarios that are parametrized by the unknown overall strength of the evolutionary pressure, labeled by the variable F m. The closest match with the mutation rates of (Fromant et al. 1995) appears to be obtained for the smallest value of F m and we list the corresponding mutation rates in our Table 2. Interestingly (and independently of the value of F m) the reconstructed primordial rates of all transitions are larger than the rates of all transversions, as their Fig. 2b shows. In comparison, the experimental data of (Fromant et al. 1995) in the absence of manganese cations indicate that all transitions are significantly more frequent than all transversions, by a factor of more than at least 5, and up to 52. The experimental data also show that the presence of manganese cations at a concentration of 0.5 mM increases the rate of certain transversions, namely T → A, to the rate of transitions. The elevation of the concentration of magnesium cations, in addition, can increase the rate of A → T transversions even further, far beyond that of some transitions (Fromant et al. 1995). Therefore, if the concentrations of manganese and magnesium cations alone are to account for the reconstructed error rates of (Gutfraind and Kempf 2008), we are led to conclude that manganese cations should have been absent and the concentration of magnesium cations should have been not elevated. This conclusion is also supported by the observation that the ratio, p, of the rates for AG versus TC transitions (Table 2), indicates that the reconstructed data are closer to the in vitro data in the absence of manganese ions.
Let us now consider also the ratios, r, of the overall rates of transitions versus the overall rates of transversions (Table 2). According to this criterion we see that, if the concentrations of manganese and magnesium cations alone are to account for the reconstructed error rates of (Gutfraind and Kempf 2008), then these concentrations should have been high. (Other ratios between the various error rates including transversions, in particular, could be considered but might be subject to significant uncertainties as rates for transversions are typically very low.)
The conclusion we have to draw, therefore, is that no combination of concentrations of manganese and magnesium dications could unambiguously account for the reconstructed error rates. The results of (Gutfraind and Kempf 2008) and (Fromant et al. 1995) together therefore indicate that, beyond the magnesium and manganese cations concentrations, there should have existed additional factors that significantly affected the primordial error rates. It should be interesting to try to identify these factors through detailed experimental analyses of parameters that affect the error rates.
Finally, we note that the concentration of manganese cations that was found necessary in (Fromant et al. 1995) to increase the rate of some transversions to the rate of some transitions is in fact comparable to the concentrations of manganese ions that have been used in studies on bacterial growth in media mimicking hydrothermal vents. In this scenario, one of the primordial organisms’ first sources of energy could have been the oxidation of manganese from basalt (Templeton et al. 2005). Note that for primordial organisms one may be able to assume that the concentration of manganese ions in the cell was comparable to that in the environment, as the maintenance of a significant concentration gradient would have been energetically expensive. We remark that the question of whether hyperthermophilic organisms might be at the origin of life is indeed still a matter of active debate (Islas et al. 2003; Schwartzman and Lineweaver 2004). The emergence of life at the interface between hot and cold environments is an attractive hypothesis as strong gradients in complex dynamical systems can be linked to self-organization processes (Hoelzer et al. 2006).
The study of the genetic code as a fossil whose structure can reveal information about the primordial circumstances under which the genetic code formed (Di Giulio 2000, 2005b, 2005c; Gutfraind and Kempf 2008) is a fascinating emerging field. Among the issues that wait to be addressed from this perspective let us here only mention the following: experimental evidence for a code linking RNAs and amino acids was obtained from studies of RNA helices which mimick tRNAs and their acceptor stems in particular, which are without anticodons and which are amino-acylated by aminoacyl-tRNA synthetases (Thiebe et al. 1972; Schimmel et al. 1993). This code has been called the operational code (Schimmel et al. 1993; Rodin et al. 1996). This code may have been relevant during the transition between a RNA world with its ribozymes (Maynard-Smith and Szathmary 1995) and the RNA-protein world endowed with a genetic code (Penny et al. 2003; Rodin and Rodin 2006). The last universal common ancestor (LUCA) was suggested to derive from the RNA-protein world (Becerra et al. 2007) and to have a RNA genome (Glansdorff et al. 2008), whereas earlier reports mentioned that LUCA emerged from the DNA-protein world (Penny et al. 2003).
The field is still in its early stages and a consensus has not yet been reached on what exactly the structure of the genetic code can tell about the evolution of the primordial organisms and their environment. More work in this direction is needed, as this approach is one of only very few ways in which any evidence about the circumstances during the early stages of life can be obtained.
Bibliography
Amirnovin R (1997) An analysis of the metabolic theory of the origin of the genetic code. J Mol Evol 44:473–476
Ardell DH (1998) On error minimization in a sequential origin of the standard genetic code. J Mol Evol 47:1–13
Ardell DH, Sella G (2002) No accident: genetic code freezes in error-correcting patterns of the standard genetic code. Phil Trans R Soc Lond B 357:1625–1642
Bahrami F, Jestin JL (2008) Streptococcus agalactiae DNA polymerase I is an efficient reverse transcriptase. Biochimie 90:1796–1799
Becerra A, Delaye L, Islas S, Lazcano A (2007) The very early stages of biological evolution and the nature of the last common ancestor of the three major cell domains. Ann Rev Ecol Evol Syst 38:361–379
Cadwell RC, Joyce GF (1994) Mutagenic PCR. PCR Methods Appl 3:S136–S140
Crick FH (1966) Codon-anticodon pairing: the wobble hypothesis. J Mol Biol 19:548–555
Di Giulio M (2000) The late stage of genetic code structuring took place at a high temperature. Gene 261:189–195
Di Giulio M (2005a) The origin of the genetic code: theories and their relationships, a review. Biosystems 80:175–184
Di Giulio M (2005b) Structuring of the genetic code took place at acidic pH. J Theor Biol 237:219–226
Di Giulio M (2005c) The ocean abysses witnessed the origin of the genetic code. Gene 346:7–12
Di Giulio M, Medugno M (2000) The robust statistical bases of the coevolution theory of genetic code origin. J Mol Evol 50:258–263
Freeland SJ, Knight RD, Landweber LF, Hurst LD (2000) Early fixation of an optimal genetic code. Mol Biol Evol 17:511–518
Fromant M, Blanquet S, Plateau P (1995) Direct random mutagenesis of gene-sized DNA fragments using polymerase chain reaction. Anal Biochem 224:347–353
Galtier N, Lobry JR (1997) Relationships between genomic G + C content, RNA secondary structures and optimal growth temperature in prokaryotes. J Mol Evol 44:632–636
Glansdorff N, Xu Y, Labedan B (2008) The last universal common ancestor: emergence, constitution and genetic legacy of an elusive forerunner. Biol Direct 3:29
Goldberg AL, Wittes RE (1966) Genetic code: aspects of organization. Science 153:420–424
Gutfraind A, Kempf A (2008) Error-reducing structure of the genetic code indicates code origin in non-thermophile organisms. Orig Life Evol Biosph 38:75–85
Hoelzer GA, Smith E, Pepper JW (2006) On the logical relationship between natural selection and self-organization. J Evol Biol 19:1785–1794
Islas S, Velasco AM, Becerra A, Delaye L, Lazcano A (2003) Hyperthermophily and the origin and earliest evolution of life. Internat Microbiol 6:87–94
Jestin JL (2006) Degeneracy in the genetic code and its symmetries by base substitutions. Comp Rend Biol 329:168–171
Jestin JL (2009) A rationale for the symmetries by base substitutions of degeneracy in the genetic code. Biosystems (in press)
Jestin JL, Kempf A (1997) Chain-termination codons and polymerase-induced frameshift mutations. FEBS Lett 419:153–156
Jestin JL, Kempf A (2007) Degeneracy in the genetic code: how and why? Genes. Genomes and Genomics 1:100–103
Jestin JL, Soulé C (2007) Symmetries by base substitutions in the genetic code predict 2′ or 3′ aminoacylation of tRNAs. J Theor Biol 247:391–394
Jestin JL, Vichier-Guerre S (2005) How to broaden enzyme substrate specificity: strategies, implications and applications. Res Microbiol 156:961–966
Jestin JL, Vichier-Guerre S, Ferris S (2005) Methods for obtaining thermostable enzymes. Patent application WO2005IB00734 20050225
Judson OP, Haydon D (1999) The genetic code: what is it good for? An analysis of the effects of selection pressures on genetic codes. J Mol Evol 49:539–550
Knight RD, Freeland SJ, Landweber LF (1999) Selection, history and chemistry: the three faces of the genetic code. Trends Biochem Sci 24:241–247
Koonin EV, Novozhilov AS (2009) Origin and evolution of the genetic code: the universal enigma. IUBMB Life 61:99–111
Lagerkvist U (1978) “Two out of three”: an alternative method for codon reading. Proc Natl Acad Sci USA 75:1759–1762
Lehman N, Jukes TH (1988) Genetic code development by stop codon takeover. J Theor Biol 135:203–214
Maynard-Smith J, Szathmary E (1995) The major transitions in evolution. Oxford Freeman, pp 81–95
Ninio J (1971) Codon-anticodon recognition: the missing triplet hypothesis. J Mol Biol 56:63–74
Osawa S, Jukes TH, Watanabe K, Muto A (1992) Recent evidence for evolution of the genetic code. Microbiol Rev 56:229–264
Penny D, Hendy MD, Poole AM (2003) Testing fundamental evolutionary hypotheses. J Theor Biol 223:377–385
Popper K (1972) Objective knowledge; an evolutionary approach. Oxford University Press, New York
Ricchetti M, Buc H (1993) E. coli DNA polymerase I as a reverse transcriptase. EMBO J 12:387–396
Rodin S, Rodin A (2006) Origin of the genetic code: first aminoacyl-tRNA synthetases could replace isofunctional ribozymes when only the second base of codons was established. DNA Cell Biol 25:365–375
Rodin S, Rodin A, Ohno S (1996) The presence of codon-anticodon pairs in the acceptor stem of tRNAs. Proc Natl Acad Sci USA 93:4537–4542
Rumer Y (1966) About the codon’s systematization in the genetic code (in Russian). Proc Acad Sci USSR (dokljady) 167:1393–1395
Schimmel P, Giege R, Moras D, Yokoyama S (1993) An operational RNA code for amino acids and possible relationship to genetic code. Proc Natl Acad Sci USA 90:8763–8768
Schwartzman DW, Lineweaver CH (2004) The hyperthermophilic origin of life revisited. Biochem Soc Trans 32:168–171
Sella G, Ardell DH (2006) The coevolution of genes and genetic codes: Crick’s frozen accident revisited. J Mol Evol 63:297–313
Sonneborn TM (1965) Degeneracy of the genetic code: extent, nature and genetic implications. In: Bryson V, Vogel HJ (eds) Evolving genes and proteins. Academic Press, New York, pp 377–397
Templeton AS, Staudigel H, Tebo BM (2005) Diverse Mn(II)-oxidizing bacteria isolated from submarine basalts at Loihi Seamount. Geomicrobiol J 22:127–139
Thiebe R, Zachau HG, Harbers K (1972) Aminoacylation of fragment combinations from yeast tRNA Phe. Eur J Biochem 26:144–152
Vichier-Guerre S, Ferris S, Auberger N, Mahiddine K, Jestin JL (2006) A population of thermostable reverse transcriptases evolved from Thermus aquaticus DNA polymerase I by phage display. Angew Chem Int Ed Engl 45:6133–6137
Woese CR (1965) On the evolution of the genetic code. Proc Natl Acad Sci USA 54:1546–1552
Wong JT (1975) A co-evolution theory of the genetic code. Proc Natl Acad Sci USA 72:1909–1912
Wong JT (1981) Coevolution of genetic code and amino acid biosynthesis. TIBS 6:33–36
Wong JT (2005) Coevolution theory of the genetic code at age thirty. Bioessays 27:416–425
Wong JT (2007) Coevolution theory of the genetic code: a proven theory. Orig Life Evol Biosph 37:403–408
Acknowledgments
AK has been supported by the Discovery and Canada Research Chair Programs of the National Science and Engineering Research Council (NSERC) of Canada.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jestin, J.L., Kempf, A. Optimization Models and the Structure of the Genetic Code. J Mol Evol 69, 452–457 (2009). https://doi.org/10.1007/s00239-009-9287-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-009-9287-5