Abstract
We studied the population genetics of two antimicrobial peptide (AMP) loci, called Mytilin B and Mytilus galloprovincialis defensin 2 (MGD2), in the secondary contact mosaic hybrid zone between Mytilus edulis and M. galloprovincialis. The isolation period between the two species was estimated to be ~1 million years (range, 0.5 million to 2 million years) long. During this period, coevolution between microbes and the immune system has likely occurred. The secondary contact, which would date back to ~25,000 (0–200,000) years, recently allowed these coadaptations to be rearranged through hybridization. Distinctive polymorphisms were uncovered in coding sequences of the two AMP loci such as insertion/deletion of codons or bisubstituted codons. Very low levels of differentiation were observed between populations of the two species at both loci, while other nuclear loci often showed marked structure among the same samples. The absence of population differentiation proved to be the consequence of secondary introgression of highly divergent alleles. While only a few recombinants were observed at the Mytilin B locus, the MGD2 locus showed a high intragenic recombination rate, which increased in the exon coding for the mature peptide. In addition, standard neutrality tests revealed significant deviations from the mutation-drift equilibrium at both loci. These results suggest that either balancing or directional selection is likely to play a role in the evolution of the two AMPs and introgression would be adaptive. However, evidence accumulated at the Mytilin B locus allows neither for identification of the direction of selection nor for any conclusions on whether selection acted directly on the antimicrobial peptide itself. At the MGD2 locus, a spatial variation of polymorphism patterns along the sequence suggests that selection was direct, although the precise nature of the selection (directional vs. balancing) remains unclear. This study concurs with previous reports of an effect of slight selection on AMP genes evolution in other invertebrates, although selection does not necessarily act on the mature peptides.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The importance of selection in the evolution of immune system proteins is well recognized. Studies comparing divergence and/or polymorphism between functional categories of genes have clearly identified genes of the immune system as a class of fast-evolving genes (Clark et al. 2003; Schlenke and Begun 2003; Bustamante et al. 2005; Nielsen et al. 2005). Recently, Sackton et al. (2007) compared functional subgroups within the immune system and detected evidence of adaptive evolution for proteins directly involved in the recognition of pathogens. The pattern was less clear for other immune proteins such as antimicrobial peptides (AMPs).
AMPs are an evolutionarily conserved component of the innate immune response and comprise a highly diverse group of small molecules exhibiting antimicrobial activity, isolated from organisms spanning the entire phylogenetic spectrum (review by Zasloff 2002). Even though AMPs play an important role in the immune system, it is not clear whether they can actually take part in tightly coupled host-pathogen coevolution processes such as arms races. AMPs are often involved in first-line immune defenses against microbial attacks and are effective against a broad spectrum of microbes. In addition, the mechanism of action of AMPs does not seem compatible at first with coevolution. AMPs target the bacterial membrane, which is not a structure that can easily engage in an arms race due to extensive functional constraints. Zasloff (2002) suggested that the acquisition of resistance to AMPs by a sensitive microbial strain is too complex to achieve and therefore improbable.
However, some recent studies have provided evidence for signatures of selection in AMP evolution. The molecular diversity of AMPs is characterized by a plethora of multigene families, high nucleotide diversity (Little et al. 2004; Tennessen 2005), and a substantial rate of duplication-subfunctionalization (Nicolas et al. 2003; Quesada et al. 2005). AMPs tend to evolve faster than the genomic average (review by Tennessen 2005). There are reports of an association between genetic variation at AMP loci and bacterial load (Lazzaro et al. 2004) and of bacteria acquiring resistance to AMPs (Yeaman and Yount 2003), sometimes very rapidly (Perron et al. 2005). Finally, the footprint of selection has been identified on the nucleotide variation of some AMP loci (Clark and Wang 1997; Lazzaro and Clark 2003; Semple et al. 2005). On the other hand, while evidence for adaptive evolution has been attested, it does not necessarily mean that selection was driven by host-pathogens interactions. For example, although evidence for adaptive evolution has been detected in the human lineage of the β-defensin 2 gene (Boniotto et al. 2003), the macaque orthologue proved to be more efficient than the human peptide on human pathogens (Antcheva et al. 2004). Sackton et al. (2007) argued that selection may rather act on the efficacy of expression than on the amino acid sequence of AMPs.
Studying the population genetics of AMPs may help to identify more precisely the type of selection that acts on these genes on the short term (Clark and Wang 1997; Lazzaro and Clark 2003; Little et al. 2004). Few studies have tried to incorporate the additional information that can be gained from studying spatially or genetically structured populations. Here, we propose to study the potential effect of selection on AMPs by taking advantage of the population structure observed in secondary contact hybrid zones. Analyzing gene flow between differentiated genomes after secondary contact can provide interesting information on the selective forces at play (Harrison 1990; Martinsen et al. 2001). Coadaptations, evolved during an initial period of divergence in allopatry, can be rearranged in hybrid genotypes and exchanged through subsequent backcrosses. Adaptive introgression of beneficial alleles (Arnold 2000; Weill et al. 2000) or balanced polymorphisms (Castric et al. 2008) into a new genetic background can be detected provided that the barrier to neutral gene flow is sufficiently strong.
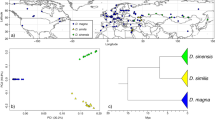
Here we study the mosaic hybrid zone between the marine mussels Mytilus edulis and M. galloprovincialis in Europe (Skibinski et al. 1983; Bierne et al. 2003b). Instead of a single genetic gradient between the two species, successive transition zones delineate enclosed panmictic patches of populations characterized by high frequencies of parental alleles; e.g., a patch of M. edulis-like populations is observed in the Bay of Biscay and a patch of M. galloprovincialis-like populations in Brittany (Fig. 1) (Bierne et al. 2003b). The interspecific barrier to gene flow is due to the synergistic action of various well-characterized pre- and postzygotic isolation mechanisms such as habitat specialization (Gosling and McGrath 1990), spawning asynchrony (Secor et al. 2001), assortative fertilization (Bierne et al. 2002a), and hybrid fitness depression (Bierne et al. 2006). We analyzed DNA sequence polymorphisms at two AMP loci, Mytilus Galloprovincialis Defensin 2 (MGD2) and Mytilin B (Mitta et al. 2000a), in order (i) to characterize amino acid variations in natural populations, (ii) to investigate the level of population differentiation at these loci, and (iii) to search for a possible signature of selection on sequences of these antimicrobial peptide genes. Introgression levels were further analyzed by studying length polymorphisms.
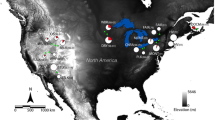
Localities of Mytilus spp. samples. The five samples chosen to represent each panmictic patch are indicated by white and black stars for M. edulis and M. galloprovincialis, respectively. The three hybrid zones described by Bierne et al. (2003) are indicated by dashed lines: HZ1, between M. galloprovincialis of the Atlantic coast of the Iberian Peninsula and M. edulis of the Bay of Biscay; HZ2, between M. edulis of the Bay of Biscay and M. galloprovincialis of Brittany; and HZ3, between M. galloprovincialis of Brittany and M. edulis of the North Sea. Patches of M. galloprovincialis are indicated by solid black lines and patches of M. edulis are indicated by black and white solid lines
Materials and Methods
Samples
Mytilus spp. samples were collected at five localities chosen to represent the previously identified panmictic patches: BA (Banyuls, Pyrénées-Orientales, France), FA (Faro, Algarve, Portugal), and PR (Primel, Finistère, France) for M. galloprovincialis populations of the Mediterranean Sea and the Atlantic coasts of the Iberian Peninsula and Brittany, respectively, and LU (Lupin, Charente-maritime, France) and WS (Wadden Sea, Holland) for M. edulis populations of the Bay of Biscay and the North Sea (Fig. 1). Sample size was 48 individuals per population. These samples have been analyzed previously with three length-polymorphism marker loci (mac-1, EFbis and Glu-5′, Bierne et al. 2003) to verify their genetic composition (Faure et al. 2008).
DNA Polymorphism
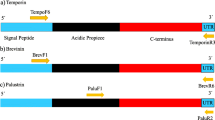
The Mytilin B and MGD2 gene sequences published by Mitta et al. (2000a) were used to design primers that amplify the exon containing the sequence of the mature peptide, together with an intron and an additional exon. The Mytilin B fragment was amplified with the forward primer MytiB-F (5′-ATGTTTCAGTCCATGAGGCAGAGGC-3′) and the reverse primer MytiB-R (5′-ATTGGATCATCTCTTTAAATGCCAG-3′). The amplified fragment includes the third exon that codes for the Mytilin B mature peptide, the third intron, and the coding portion of the fourth exon. The MGD2 fragment was amplified with the forward primer MgD2-F (5′-ATGAAAGCAGTATTCGTCTTGTTGG-3′) and the reverse primer MgD2-R (5′-TACTAGATTCAGAAACGTTCTACTG-3′). The amplified fragment includes the second exon, the second intron, and the third exon that code for the MGD2 mature peptide. Based on the sequence of Mitta et al. (2000a), the predicted length of the Mytilin B fragment was 804 bp and the predicted length of the MGD2 fragment was 1454 bp. However, length polymorphisms are common in mussel introns and sometimes involve long indels so that alleles can be highly variable in length. A standard protocol was used for the PCR reactions, with an annealing temperature of 50°C and 55°C for Mytilin B and MGD2, respectively. We used the Promega GoTaq DNA polymerase (Promega, Madison, WI, USA). To save time and money and to obtain appreciable sample sizes, sequences were cloned following the mark-recapture (MR) cloning protocol (Bierne et al. 2007; Faure et al. 2007, 2008). The principle of MR cloning is to perform a single cloning reaction per pooled population sample. Each individual of a sample was PCR-amplified separately using 5′-tailed primers with small polynucleotide tags. PCR products of similar quantities were mixed together and cloned into a pGEM-T vector using a Promega pGEM-T cloning kit. Clones were sequenced with universal plasmid primers. The individual sequences were identified by the tag sequences upstream of each initial primer. A consequence of the MR-cloning protocol is that sample size cannot be controlled. Within a given number of sequenced clones, the same allele of the same individual (recognized using the nucleotide tag) can be cloned multiple times, while some alleles or individuals remain absent. Therefore, the number of different sequences obtained is less than, although positively correlated with, the number of positive clones sequenced (called the effort of capture by Bierne et al. 2007). However, an advantage of this protocol is that it offers the opportunity to assess the error rate due to mutations during the cloning and amplification process. Singleton mutations (which are important indicators of selective or demographic effects) are particularly sensitive to such artifacts and it thus becomes possible to reduce the potential impacts of these artifacts by restricting the analysis to sequences that were captured twice or more for the same individual.
Length Polymorphism
In both genes we used our sequence datasets to design a new reverse primer in order to study length polymorphism. The new primer was MytiBindel-R2 (5′-TCAACCTGAACATAAAATCCTATCC-3′) for the Mytilin B locus. Together with MytiB-F, this primer allowed us to obtain a biallelic locus with one allele of 268 bp and another allele of 278 bp. The new primer was MgD2indel-R2 (5′-AAAAGTTAACCCTTTACAAAATC-3′) for the MGD2 locus. Together with MgD2-F, this primer allowed us to obtain a biallelic locus with one allele of 105 bp and another allele of 108 bp. The fluorescent dye 5′ end-labeled-primer technique was used, with dye 6-FAM (Sigma Genosys). Polyacrylamide gels were scanned in a FMBIO II fluorescence imaging system (Hitachi Instruments).
Data Analysis
Length polymorphism was analyzed with the Genetix 4.03 software (Belkhir et al. 2002). Estimations of FST were obtained using Weir and Cockerham’s (1984) estimator, and population differentiation tested using the permutation procedure. The method of Foll and Gaggiotti (2008) implemented in the BayeScan 1.0 software was used to assess the probability that a locus depart from the distribution of FST under a neutral model.
Sequence alignment was performed with ClustalW (Thompson et al. 1994) in the Bioedit interface (Hall 1999) and verified by eye. For each sequence, indel composition was registered before excluding alignment gaps from further statistical analyzes. We used DNAsp (Rozas and Rozas 1999) to compute basic population genetic parameters such as number of polymorphic sites (S); number of synonymous (S s ), nonsynonymous (S ns ), and noncoding (S nc ) polymorphisms; levels of nucleotide diversity estimated from the number of polymorphic sites, θW (Watterson 1975), or from pairwise differences, θπ (Nei 1987); Hudson et al.’s (1992) estimator of FST; the minimum number of recombination, RM, estimated by the method of Hudson and Kaplan (1985); and linkage disequilibrium as measured by the ZnS statistic (Kelly 1997). DNAsp was also used to compute Tajima’s D (1989), a well-known statistic that proved efficient in detecting departures of the allele frequency spectrum from the expectation at mutation-drift equilibrium. Departure from the neutral expectation at mutation-drift equilibrium was tested by coalescent simulations conditional on the number of segregating sites. We also used the coalescence-based maximum likelihood method of Galtier et al. (2000). This method is designed to detect a distortion in the shape of gene genealogies generated by a diversity-reducing event (hitchhiking or bottleneck).
Phylogenetic reconstructions were performed differently for the two loci. For the Mytilin B locus, a few recombination breakpoints were easily identified. We first excluded the few recombinant alleles and reconstructed the genealogy with GeneTree (Bahlo and Griffiths 2000). We added recombinants afterward and drew an ancestral recombination graph. For the MGD2 locus the number of recombination breakpoints was far too high to reconstruct a comprehensible genealogy for the entire length of the sequence. To represent an overview of the genetic relationships among alleles, we inferred a network with the NeighborNet algorithm (Bryant and Moulton 2004) in the software SplitsTree4 (Huson and Bryant 2006). As intragenic recombination was very high at this locus, we were able to conduct sliding window analyzes on various statistics. Sliding windows on measures of diversity, departures from mutation-drift equilibrium as measured by Tajima’s D and linkage disequilibrium as measured by ZnS was performed with DNAsp (Rozas and Rozas 1999). Sliding windows on the population recombination rate (4N e r) were performed with the software LDhat that implements the coalescent-based method of McVean et al. (2002).
Genetic data were also fitted to an isolation-with-gene-flow model (Nielsen and Wakeley 2001) by the method of Hey and Nielsen (2007) implemented in the IMa software. The isolation-with-migration (IM) model assumes that two populations of effective size N e1 and N e2 diverged from an ancestral population of effective size N eA at time t and then exchanged migrants at rates m 1 and m 2 . Convergence by the Markov chain Monte Carlo (MCMC) simulations toward the true stationary distribution was checked by monitoring multiple independent chains started at different starting points and by assessing the autocorrelation of the parameter values over the course of the run. Timings of migration events recorded in sampled genealogies at MCMC stationarity were used to generate a posterior distribution of migration times. We used the largest fragment without recombination and we assumed the HKY model of mutation (Hasegawa et al. 1985). We used a burn-in period of 500,000 steps after which 1 million genealogies were saved every 300 steps for parameter estimation. Analyses were conducted separately for each gene and in a combined fashion.
Results
Mark-Recapture Cloning
For the Mytilin B locus we sequenced 132 positive clones and captured 86 sequences (of which 32 were multiple captures) and for the locus MGD2 we sequenced 201 positive clones and captured 104 sequences (of which 51 were multiple captures). Rates of single and multiple captures were in accordance with theoretical and empirical expectations (see Figs. 1 and 2 of Bierne et al. 2007). Final sample sizes are reported in Tables 1 and 2. Alleles resequenced multiple times allowed us to estimate the frequency of artifactual mutations that are inherently produced during PCR, cloning, and sequencing. We found on average one artifactual mutation for ~1000 bp sequenced in both datasets. However, the number of artifactual mutations was not uniformly distributed among sequences; some sequences of multiply captured alleles proved to carry as many as four singleton mutations, which were removed after comparison to the other sequence of the same allele. As previously reported (Faure et al. 2007, 2008) we constructed high-quality datasets (HQ) composed of a subset of sequences that have been captured multiple times, which should be essentially free from artifactual mutations, and compared them to global datasets (G).
Alignment of the amino acid sequences of the 14 different Mytilin B mature peptides observed in the global dataset. Numbers in parentheses are the occurrence of the allele in the dataset. Lines joining cysteines represent disulfide bonds. The structure of the peptide is schematized below the alignment. Amino acid mutations are in boldface and polymorphisms are highlighted within boxes
Amino Acid Sequence Variations
We observed an appreciable level of amino acid polymorphisms at both loci (Tables 1 and 2). Although many amino acid changes were rare variants, some distinctive polymorphisms were segregating at moderate frequencies.
At the Mytilin B locus, distinctive polymorphisms were observed in the mature peptide. Figure 2 presents the 14 different Mytilin B peptides observed in our G dataset. We defined two groups of peptides, B1 and B2, which displayed three major differences: a two-amino acid (VR) indel at the exact N-terminal extremity of the Mytilin B2 peptide and two amino acid substitutions at position 6 (S6L) and 10 (G10A; Fig. 2). The S6L polymorphism was a consequence of two changes at two adjacent nucleotide positions (TCC/CTC). Biochemical isolation of the Mytilin B1 mature peptide (Mitta et al. 2000b) ascertained the cleavage between the signal peptide and the mature peptide to be just after Ala22 for this allele. We verified that the cleavage position remained unchanged despite the VR addition in the Mytilin B2 peptide by using a prediction of the cleavage position. We used the neural network method of Nielsen et al. (1997) implemented in the SignalP 3.0 software (Bendtsen et al. 2004). The prediction retrieved with a very high probability (C-score = 0.92) that the cleavage should occur between position Ala22 of the signal peptide and the valine that we accordingly renumbered position 1 of the mature peptide. A third amino acid mutation at position 24 (L24Q) was also segregating at moderate frequency but was not perfectly associated with the B1/B2 allele definition: although one allele (Brec in Fig. 2) proved to be a likely result of intragenic recombination (see below), two others did not (B2e and B2f).
At the MGD2 locus, the amino acid variation observed in the mature peptide was mainly composed of singleton mutations. However, one amino acid mutation at position 31 (L31R; Fig. 3a) was segregating at moderate frequency. This polymorphism was a consequence of two changes at two adjacent nucleotide positions (TTG/CGG). Distinctive amino acid variations were observed in the MGD2 signal peptide. We present in Fig. 3b the nine different partial signal peptide sequences observed in our G dataset. At positions 16 and 17 an M/T (M16T) and an M/indel (M17-) polymorphism were observed, respectively. At position 21, four different amino acids were segregating (Fig. 3b).
a Alignment of the amino acid sequences of the eight different MGD2 mature peptides observed in the high-quality (multiply captured sequence) dataset. Numbers in parentheses are the occurrence of the allele in the dataset. Lines joining cysteines represent disulfide bonds. The structure of the peptide is schematized below the alignment. Amino acid mutations are in boldface and polymorphisms are highlighted within boxes. b Alignment of the partial amino acid sequences of the nine different signal peptides observed in the global (all sequences) dataset. Numbers in parentheses are the occurrence of the allele in the dataset. Amino acid mutations are in boldface and polymorphisms are highlighted within boxes
Identification of a Repeated Element
A polymorphic 160-bp indel was observed in the MGD2 sequences. The sequence of this indel was submitted to a Blast search in the GenBank database and we found that the sequence displayed high similarities to other Mytilus sequences: a portion (from position 3079 to position 3240) of the second intron of the endo-1,4-β-d-glucanase (AJ308548), a portion (from position 4832 to position 4949) of the fifth intron of the endo-1,4-mannanase gene of (AJ271365), a portion (from position 2069 to position 2206) of the fourth intron of the lysozyme gene (AF334662), a portion (from position 3427 to position 3495) of the 3′ UTR of the hsp70-1 gene (AJ585375), and a portion (from position 3427 to position 3495) of an ISSR sequence (AJ938114). In addition, the sequence also displayed high similarities to untranslated regions of 41 ESTs from a M. galloprovincialis dataset of 5602 ESTs (Tanguy et al. 2008) and 103 ESTs from a M. californianus dataset of 23,808 ESTs. We concluded that the observed indel polymorphism is a consequence of the insertion of a short transposable element.
Population Differentiation
With DNA sequence polymorphism, significant population differentiation was observed at the Mytilin B locus (global FST = 0.22; permutation test, p < 0.0001) but not at the MGD2 locus (global FST = 0.008; p = 0.31). Population differentiation was further analyzed using length polymorphisms. Length polymorphisms allowed us to infer allele frequencies with much bigger sample sizes (48 individuals = 96 chromosomes). Allele frequencies are reported in Fig. 4a, together with those obtained at 11 previously analyzed loci in the same four patches of the French contact zone, 5 allozymes (Coustau et al. 1991) and 6 DNA loci (Bierne et al. 2002b). Loci that were not biallelic were transformed into biallelic loci by pooling alleles according to their frequency in M. edulis and M. galloprovincialis reference populations. The procedure of pooling alleles is often recommended to compare loci with different diversity when the differentiation is strong (McDonald 1994; Hedrick 1999). As was the case with DNA sequence polymorphism, no significant population differentiation was observed at the MGD2 locus (global FST = 0.002, p = 0.33). A significant population structure was observed at the Mytilin B locus (global FST = 0.03, p = 0.01). Two groups of samples were observed for which the homogeneity of genotypic frequencies cannot be rejected: one composed of the two M. galloprovincialis samples from the Mediterranean Sea (BA) and Portugal (FA) and the other composed of M. edulis samples from the North Sea (WS) and the Bay of Biscay (LU) together with the M. galloprovincialis sample from Brittany (PR).
a Frequencies of the B1 allele at the Mytilin B locus, of the –17 allele (gap at position 17 of the signal peptide) at the MGD2 locus, and of the G allele at six length polymorphism loci (Glu-5′, mac-1, EFbis, DAMP1, DAMP2, DAMP3, Bierne et al. 2002b) and five allozyme loci (EST D, LAP, PGI, OCT, MPI, Coustau et al. 1991) in the four patches of panmictic Mytilus spp. populations (see Fig. 1). Some of the 11 loci previously analyzed were not biallelic but were transformed into biallelic loci by pooling alleles according to their frequency in M. edulis and M. galloprovincialis reference populations. b Observed FST values between M. edulis populations of the North Sea and M. galloprovincialis populations of the Atlantic coast of the Iberian Peninsula at 20 loci as a function of the probability to depart from the neutral FST distribution as measured by the decimal logarithm of the Bayes factor computed by the method of Foll and Gaggiotti (2008). Squares, allozymes; dots, DNA loci (black, PAM loci; white, others). The observed FST distribution is depicted at the right
The results obtained with length polymorphisms provided useful additional information to DNA sequence data. Whereas samples BA and FA were exclusively composed of Mytilin B1 alleles in the DNA sequence dataset (Table 1), the length polymorphism survey demonstrated that Mytilin B2 alleles were indeed segregating in these populations, albeit at a low frequency (~10%). At the MGD2 locus, the presence/absence of Met at position 17 of the signal peptide (an indel polymorphism named the ‘M17’ allele; Fig. 3b) was estimated to be 65% with the DNA sequence dataset (Table 2), while it was estimated to be 72% in the analysis of length polymorphism (not significantly different; Fisher’s exact test, p = 0.31). Finally, and most importantly, the analysis of length polymorphisms allowed us to study a parameter that is crucial in population genetics but often unappreciated in DNA sequence surveys, the departure from Hardy-Weinberg equilibrium. Both loci proved never to depart significantly from Hardy-Weinberg expectations. The average FIS value was –0.005 (max value, 0.05 in sample FA) at locus Mytilin B and 0.005 (max value, 0.07 in sample LU) at locus MGD2.
We verified the degrees of differentiation of the Mytilin B and MGD2 loci with respect to other loci by conducting a FST scan analysis between M. edulis populations of the North Sea and M. galloprovincialis populations of the Atlantic coast of the Iberian Peninsula. We compiled data on 11 allozyme loci, 5 from Coustau et al. (1991; EST D, LAP, PGI, OCT, MPI) and 6 from Skibinski et al. (1983; Ap, Pgm, Aap, Idh-1, Aat-1, 6Pgdh), and on 7 nuclear DNA loci, 6 from Bierne et al. (2002b; EFbis, mac-1, Glu5′, DAMP1, DAMP2, DAMP3) and one from Riginos et al. (2006; Lysin M7). Two loci from the study by Skibinski et al. (1983), Aat-2 and Idh-2, were discarded because polymorphism levels were too low. Figure 4b presents the 20 observed FST values (which were different from the locus-population-specific FST coefficients of Foll and Gaggiotti 2008) as a function of the posterior probability that a locus is subject to selection as measured by the decimal logarithm of the Bayes factor (log10[BF]) obtained by the method of Foll and Gaggiotti (2008). MGD2 was the least differentiated locus of the 20 and the only one that was not significantly different from zero. Only one allozyme locus, Pgm, was less differentiated than Mytilin B. According to the method of Foll and Gaggiotti (2008), two loci were “decisively” (log10[BF] > 2) under selection, Glu-5′ and MGD2. However, this result should be taken with caution since FST scan approaches might not be appropriate in the context of a semipermeable hybrid zone (Bierne et al. 2003c; Murray and Hare 2006). In hybrid zones many marker loci may be influenced by indirect pseudoselection, since isolation genes can generate genetic barriers on large genomic scales (Barton and Hewitt 1985). Generating the neutral distribution of FST can be problematic in this context, where a high proportion of loci is affected by selection even though their polymorphisms are intrinsically neutral (Bierne et al. 2003c). Here observed FST values stretch out from 0 to ~1 and the FST distribution does not appear to match with a neutral distribution (Fig. 4b). In any case, outlier tests should not be taken as evidence for the direct action of selection on the identified loci: Glu-5′ might simply be closely linked to an isolation gene, while MGD2 might simply belong to an unimpeded part of the genome.
Fitting an Isolation-with-Migration Model
We compiled a DNA sequence dataset from the two PAM loci obtained in this study (Mytilin B and MGD2), two nuclear loci (EF1α, Faure et al. 2008; Lysin M7 fifth exon, Riginos et al. 2006), and one mitochondrial locus (COI, Gérard et al. 2008). As was the case for the FST outlier test, we were once again confronted with the difficulty that selection is a regular and ubiquitous feature of hybrid zones, while the isolation-with-migration model (Nielsen and Wakeley 2001; Hey and Nielsen 2007) is purely demographic and does not take selection into account. However, it can still be useful to fit this model, as selection might not affect all the parameters of the model. The results we obtained clearly showed that some parameter values were not shared among loci. Estimates of these values are presented in Table 3. First, migration rates were different among loci, which is not surprising across a semipermeable barrier to gene flow and which confirms our previous FST scan analysis. Mytilin B, MGD2, and EF1α displayed a tendency for an asymmetrical gene flow from M. galloprovincialis to M. edulis, while the tendency was in the reverse direction for Lysin M7 and COI. Second, we obtained different estimates of N e for various loci differently affected by selection. For instance, the reduced diversity probably generated by a hitchhiking event at the EF1α locus in M. edulis (Faure et al. 2008) translates into a smaller effective population size (N e ) for this locus in M. edulis. Lysin M7 has also been documented to be influenced by positive selection (Riginos et al. 2006; Springer and Crespi 2007) and the fifth exon indeed exhibited a surprisingly small diversity which translates into a very small N e at this locus. The possible effect of selection on the diversity of the Mytilin B and MGD2 loci is discussed below. However, it is interesting to note that the average Ne is remarkably similar between the two species and the ancestral population (~0.5 × 106).
Considering that the semipermeable nature of the genetic barrier to gene flow was accurately captured by migration parameters and selection by population size parameters, we have some confidence in estimates of the timing of divergence and the timing of migration. Indeed, estimates of the divergence time were similar across loci, ~1 million (range, 0.5 million to 2 million) years, which confirms the divergence time previously estimated from mtDNA (Hilbish et al. 2000). The most probable mean times of migration events were also very similar across loci and very recent, compared to the divergence time, ~25,000 years ago. These results are in accordance with a secondary contact scenario, in which gene flow was recently restored after a long period of isolation.
Nucleotide Variation and Reconstructed Genealogy at the Mytilin B Locus
Basic descriptors of sequence polymorphism at the Mytilin B locus are presented in Table 1. Although intragenic recombination was detected, the number of recombination breakpoints was low enough (Table 1) to reconstruct a single-gene genealogy of the whole sequence (Fig. 5) rather than dividing the sequence into nonrecombining fragments. The genealogy was structured into two highly divergent clades of alleles, one corresponding to the peptide alleles B1 and the other to alleles B2. We observed 36 fixed differences between the two clades (4.5%), together with 7 indels. Two sequences were inferred to be recombinants between alleles of the two clades. The recombination breakpoint of the first interclade recombinant was localized in the 5′ end of the sequence, within exon 3 (that codes for the mature peptide), between nucleotide position 63 and nucleotide position 86. This recombination created a new mature peptide that we named Brec (Fig. 2). The recombination breakpoint of the second interclade recombinant was localized in the middle of the intron between position 506 and position 537. Additionally, intraclade recombination was detected within clade B1. Two recombination events were identified from three intraclade recombinant alleles, of which two were similar alleles. Recombination breakpoints were located between nucleotide position 380 and nucleotide position 496 and between nucleotide position 496 and nucleotide position 594.
Within clades, most alleles coalesced together within star-shaped subgroups of alleles (B1* and B2* in Fig. 5). To infer the state of disequilibrium within the two taxa before secondary introgression occurred, we applied Tajima’s D test within each clade. Tajima’s D tests were mostly significantly negative (Table 1). Since samples BA and FA were exclusively composed of clade B1 alleles (we did not sample any introgressed alleles), we were able to apply neutrality tests more directly. Tajima’s D was significantly negative in sample BA-G but not in samples BA-HQ and FA-G (Table 1). Furthermore, as our datasets were compatible with the infinite-sites mutation model, we were also able to apply the coalescent-based method of Galtier et al. (2000). We found that a model with a recent reduction in effective population size, be it demographic or selective in its origin, was significantly better supported than the mutation-drift equilibrium model (likelihood ratio tests: p = 0.005, p = 0.01, and p = 0.04 for samples BA-G, BA-HQ, and FA-G, respectively).
Nucleotide Variation and Departure from Neutrality at the MGD2 Locus
Basic descriptors of polymorphism at the MGD2 locus are presented in Table 2. Since population differentiation was not detected, we pooled the five geographic samples. A high level of nucleotide diversity was observed (one SNP every 3.6 bp). We found a very high number of recombination breakpoints at this locus, a number that was far too high to reconstruct a comprehensible gene genealogy. A network inferred with the NeighborNet algorithm depicts the genetic relationship among alleles in Fig. 6a. We also present two networks constructed with partial sequences, the 400-bp 5′ end in Fig. 6b and the 400-bp 3′ end in Fig. 6c. A two-clade structure was perceptible at the 5′ end of the locus. Sliding window analyses showed that nucleotide diversity, Tajima’s D, and linkage disequilibrium were stronger at the 5′ than at the 3′ end (Fig. 7a, b). Tajima’s D was negative overall, although it reached positive values at the 5′ end.
a Neighbor network for the entire MGD2 locus. b Neighbor network for the 400 bp of the 5′ end of the fragment. c Neighbor network for the 400 bp of the 3′ end of the fragment. Reticulation of the branches indicates uncertainties due to homoplasia (recombination or multiple hits). Sequences sampled in M. edulis populations are represented by white circles, sequences sampled in M. galloprovincialis populations by black circles, and sequences sampled in both species by gray circles
a Sliding window analysis of the levels of nucleotide diversity estimated from pairwise differences, θπ (Nei 1987), the number of polymorphic sites, θW (Watterson 1975), and Tajima’s (1989) D. b Sliding window analysis of linkage disequilibrium (ZnS) and maximum likelihood estimations of the population recombination rate (4N e r)
The observed pattern was a consequence of a high proportion of mutations segregating at a moderate frequency and in strong linkage disequilibrium while the density of polymorphic sites as measured by θ W did not vary spatially (Fig. 7a). This high density of associated SNPs with a high heterozygosity was located at the beginning of the intron and stopped with a 160-bp indel at position 440. An analysis of the variation of the population recombination rate, 4N e r, showed that recombination increased from the 5′ to the 3′ end of the locus, with a sharp increase in exon 4 (Fig. 7b).
Discussion
Our analysis of the population genetics of two antimicrobial peptides in a Mytilus secondary contact zone yielded intriguing results. The analysis of forces that shape these patterns, however, is far from straightforward. Normally, four hypotheses can be put forward: (1) demography (2) indirect hitchhiking selection (3) direct balancing selection, and (4) weak directional positive selection. We consider the first hypothesis less likely, considering that deviations from demographic equilibrium have not been reported for other loci in Mytilus (Quesada et al. 1998; Riginos et al. 2004, 2006; Springer and Crespi 2007; Faure et al. 2008). Furthermore, the demographic analysis under the isolation-with-migration model suggests that the effective population size averaged over loci has remained fairly constant since the divergence from the ancestral population (Table 3). It is less straightforward to choose between the alternative scenarios invoking selection. Molecular signatures of selection can affect any gene, be they of the immune system or not. Indeed, any neutral locus can be indirectly affected by selection at a linked locus (Barton 2000; Charlesworth et al. 2003) and molecular footprints of positive selection affect the diversity of an appreciable proportion of some genomes (Li and Stephan 2006). Therefore, although candidate amino acid polymorphisms for direct selection have been observed in the two AMPs analyzed here, the accurate interpretation of these results requires a thorough consideration of the patterns observed.
Why Does the Mytilin B Locus Depart from the Standard Coalescent?
For the Mytilin B locus, we propose a scenario of past allopatric differentiation followed by secondary contact and homogenization of allele frequencies by introgression through the interspecific barrier. This scenario is supported both by the divergence time and timing of migration estimated with the immigration-with-isolation model (Table 3) and by the allele genealogy observed at the Mytilin B locus (Fig. 5). Variable levels of differentiation among loci can be quite pronounced in hybrid zones (Harrison 1990), and this also appears to be the case in Mytilus (see Fig. 4). One motivating question, however, is whether gene flow through the interspecific barrier was historical or contemporary. Quesada et al. (1998) provided evidence for both historical and contemporary introgression of the mitochondrial genome between M. edulis and M. galloprovincialis. Evidence for ongoing effective introgression of some nuclear loci is supported by the observation that patches enclosed within the hybrid zone are noticeably more introgressed than peripheral patches (Bierne et al. 2003b) (Fig. 4a). Distinguishing between historical and contemporary introgression is difficult for the Mytilin B locus because the differentiation observed at this locus involves a difference in the frequency of the two highly divergent clades B1 and B2 (Fig. 4). On the one hand, the genetic homogeneity observed between M. edulis populations and the M. galloprovincialis patch enclosed in Brittany (Fig. 4a) suggests that introgression has been effective since the establishment of the present-day mosaic structure of the hybrid zone. On the other hand, the observation of interclade recombinant alleles suggests that a sufficient amount of time has elapsed since alleles of the two clades segregate together within the same populations and genomes for them to recombine and reach detectable frequencies.
Apart from the question of historical versus contemporary introgression, the issue that motivated us was whether we identified a neutral locus located in a chromosome region lacking isolation genes or whether selection has played a role in the introgression process. This is a difficult question to answer. On the one hand, Mytilin B plays a role in the mussel immune system and unusual modifications in the amino acid sequences of the mature peptide (e.g., amino acid indels) have been identified (Fig. 2). Furthermore, the differences in allele frequencies between populations of the two species are surprisingly small compared to previously surveyed loci (Fig. 4). Finally, neutrality tests were significant. On the other hand, the abundance of amino acid polymorphisms suggests that selective constraints on this peptide cannot be too severe. For example, the mutations observed in the cysteine array, which are changes known to be biochemically important, still segregate at a frequency high enough to be detected. The star shape of within-clade genealogies is in accordance with hitchhiking selection. Still, hitchhiking did not completely eliminate all variation and some lineages survived the selective sweeps. This implies recombination events between the locus under study and a neighboring locus under selection (e.g., Fay and Wu 2000). Alternatively, very weak selection is acting directly on the peptide. For this scenario to be plausible, fitness differences between alleles would need to be only very slight, or fluctuating in time or space (i.e., balancing selection), since fixation has obviously not occurred yet. In the end, we have no decisive arguments to conclude that the distinctive amino acid modifications observed between the two Mytilin B alleles are the result of fitness differences, which, at any rate, could only be substantiated with experimental support (Hughes 2007).
Can Selection Explain the Unusual Features Observed at the MGD2 Locus?
The case of the MGD2 locus is somewhat different from the patterns observed at the Mytilin B locus. Even though some loci, of which the Mytilin B locus has just become an additional example, differed very little between M. edulis and M. galloprovincialis (Skibinski et al. 1983) (Fig. 4), the total absence of population differentiation observed at the MGD2 locus was unforeseen. Also unanticipated was the level of intragenic recombination, which contrasts sharply with the rarity of recombination breakpoints observed for the Mytilin B, Lysin M7 (Riginos et al. 2006; Springer and Crespi 2007), and EF1α (Faure et al. 2008) genes, although it is relevant to note that these genes might show a reduced diversity due to hitchhiking. High levels of intragenic recombination have been observed in other species, without the confounding patterns of indirect hitchhiking selection (e.g., Cutter et al. 2006). Despite the high intragenic recombination rates, a high LD block was observed at the 5′ end of the locus. Concurrently, peaks of nucleotide diversity and positive Tajima’s D values were observed, which, together with low population differentiation, is usually taken as evidence for balancing selection (Hudson et al. 1987; Wheat et al. 2006). Considering that the region begins with a distinctive polymorphic amino acid indel (at position 17 of the signal peptide; Fig. 3b) and ends with an insertion of a repeated element, we have at least two candidate positions for balancing selection. Balancing selection, however, would imply selection on some sort of regulating mechanism, as predicted by Sackton et al. (2007), rather than directly on antibacterial activity.
Once again, alternative explanations should be thoroughly considered. Evidence for balancing selection on the ADH gene of Drosophila melanogaster (Hudson et al. 1987), which is one of the most famous examples of balancing selection identified with a peak of diversity around a nonsynonymous polymorphism, proved to be unpersuasive when the same peak was observed in populations monomorphic for this nonsynonymous polymorphism (Begun et al. 1999). Because AMPs are known to be multigene families, we first hypothesized that our data might have been contaminated with paralogous sequences. Three arguments were found against this hypothesis: (i) our peptide sequences were undoubtedly different from the other known defensin of Mytilus, MGD1 (Hubert et al. 1996; Mitta et al. 1999); (ii) we never observed three or more highly divergent alleles when sequences were captured multiple times after the mark-recapture cloning; and (iii) Hardy-Weinberg equilibrium was observed. Therefore, the second hypothesis based on neutral processes is gene conversion. Under this hypothesis, the assemblage of tightly associated SNPs at the beginning of the intron would be the consequence of a single event of conversion from another unknown locus rather than the result of many consecutive mutations. A similar scenario would be that the insertion of the transposon has affected the nucleotide sequence flanking the insertion point. The existence of four perfectly associated adjacent nucleotide substitutions, which are not likely to be four independent mutational events, is in accordance with both hypotheses.
The third and last alternative hypothesis is that the observed pattern for the MGD2 locus is a fingerprint of past allopatric differentiation followed by secondary introgression, as described above for the Mytilin B locus and inferred by the IMa analysis on the 5′ portion (Table 3). Once again, distinguishing between historical and contemporary introgression is not straightforward. In favor of contemporary introgression is the complete absence of significant spatial structure at this locus (Table 2 and Fig. 4). However, there are also arguments in favor of historical introgression. First, we observed small groups of alleles sampled in a single species in the phylogenetic network of Fig. 6. In accordance with this observation is a slightly older mean time of migration events estimated in the IMa analysis compared to other loci (Table 3). Second, recombination has already started to strongly blur the signature of a long period in isolation, which suggests that interspecific alleles segregate together within the same populations for a prolonged period.
Regardless of introgression being contemporary or historical, the fact remains that a modification of the diversity patterns along a sequence (Fig. 7) is usually taken as fair evidence for selection (e.g., Wang et al. 1999; Fay and Wu 2000; Wheat et al. 2006). This would lead us to deduce that the 3′ end instead of the 5′ end of the locus has been impacted by selection. The nature of this selection could be past (in)direct positive selection. Unfortunately recombination has erased most of the signature except for an excess of rare variants (leading to a negative Tajima’s D). It is also possible that we are observing ongoing selection, for which we have two candidate polymorphic sites. One possible site is the L31R polymorphism in the mature peptide, which is a radical change (leucine is a relatively small, nonpolar, neutral amino acid, while arginine is a relatively large, polar, and positively charged amino acid) and is localized in the loop thought to play a key role in the antimicrobial activity (Romestand et al. 2003). In addition, L31R required substituting two neighboring nucleotide positions of the codon. Still, substitutions at two adjacent nucleotide positions were observed in the intron as well. The second candidate position for selection in exon 4 corresponds to the 21st amino acid position of the signal peptide, where no fewer than four different amino acids were found to segregate (phenylalanine, threonine, serine, and asparagine). This is not absolutely unexpected given the nucleotide diversity observed at this locus, but remains a rare observation in polymorphism surveys of proteins. The change toward the relatively large, aromatic and nonpolar phenylalanine is the most radical, while the other three amino acids share conservative physicochemical properties (threonine, serine, asparagine). The modification of the signal peptide at this position together with the indel polymorphism of exon 3 could modify the cleavage frequency, or even specify different modes of targeting (i.e., guiding their passenger proteins through a different pathway) and membrane insertion (Martoglio and Dobberstein 1998).
Conclusion
The population genetics of AMPs in a secondary contact hybrid zone proved to be highly original, although evidence of selection was finally inconclusive. Providing that selection was indeed the explanation for the patterns observed, this selection must have been only very slight and probably of different kinds for the two AMPs investigated here. The spatial variation of polymorphism patterns along the sequence observed at the MGD2 locus was nonetheless a strong argument that selection directly influences a portion of this locus. We now have examples of two loci for which alleles are, or have been, abundantly exchanged between the two species of mussels. The fact that these two loci are immune genes is intriguing. Nonetheless, whether AMPs, or immunity genes in general, tend to cross hybrid zones more easily than other genes remains to be investigated.
References
Antcheva N, Boniotto M, Zelezetsky I, Pacor S, Verga Falzacappa MV, Crovella S, Tossi A (2004) Effects of positively selected sequence variations in humans and Macaca fascicularis b-defensins 2 on antimicrobial activity. Agents Chemother 48:685–688
Arnold ML (2000) Anderson’s paradigm: Louisiana irises and the study of evolutionary phenomena. Mol Ecol 9:1687–1698
Bahlo M, Griffiths RC (2000) Inference from gene trees in a subdivided population. Theor Pop Biol 57:79–95
Barton NH (2000) Genetic hitchhiking. Philos Trans R Soc Lond B Biol Sci 355:1553–1562
Barton NH, Hewitt GM (1985) Analysis of hybrid zones. Annu Rev Ecol Syst 16:113–148
Begun DJ, Betancourt AJ, Langley CH, Stephan W (1999) Is the fast/slow allozyme variation at the Adh locus of Drosophila melanogaster an ancient balanced polymorphism? Mol Biol Evol 16:1816–1819
Belkhir K, Borsa P, Chikhi L, Raufaste N, Bonhomme F (2002) GENETIX, logiciel sous WINDOWSTM pour la génétique des populations. Université de Montpellier 2, Montpellier, France
Bendtsen JD, Nielsen H, von Heijne G, Brunak S (2004) Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 340:783–795
Bierne N, David P, Boudry P, Bonhomme F (2002a) Assortative fertilization and selection at larval stage in the mussels Mytilus edulis and M. galloprovincialis. Evolution 56:292–298
Bierne N, David P, Langlade A, Bonhomme F (2002b) Can habitat specialisation maintain a mosaic hybrid zone in marine bivalves? Mar Ecol Prog Ser 245:157–170
Bierne N, Bonhomme F, David P (2003a) Genetics at larval stage in marine bivalves. In: Fingerman M, Nagabhushanam R (eds) Recent advances in marine biotechnology 10. Science Publishers, Enfield, NH, pp 239–262
Bierne N, Borsa P, Daguin C, Jollivet D, Viard F, Bonhomme F, David P (2003b) Introgression patterns in the mosaic hybrid zone between Mytilus edulis and M. galloprovincialis. Mol Ecol 12:447–462
Bierne N, Daguin C, Bonhomme F, David P, Borsa P (2003c) Direct selection on allozymes is not required to explain heterogeneity among marker loci across a Mytilus hybrid zone. Mol Ecol 12:2505–2510
Bierne N, Bonhomme F, Boudry P, Szulkin M, David P (2006) Fitness landscapes support the dominance theory of post-zygotic isolation in the mussels Mytilus edulis and M. galloprovincialis. Proc R Soc B 273:1253–1260
Bierne N, Tanguy A, Faure M, Faure B, David E, Boutet I, Boon E, Quere N, Plouviez S, Kemppainen P, Jollivet D, Moraga D, Boudry P, David P (2007) Mark-recapture cloning: a straightforward and cost-effective cloning method for population genetics of single-copy nuclear DNA sequences in diploids. Mol Ecol Notes 7:562–566
Boniotto M, Tossi A, DelPero M, Sgubin S, Antcheva N, Santon D, Masters J, Crovella S (2003) Evolution of the beta defensin 2 gene in primates. Genes Immun 4:251–257
Bryant D, Moulton V (2004) NeighborNet: an agglomerative algorithm for the construction of planar phylogenetic networks. Mol Biol Evol 21:255–265
Bustamante CD, Fledel-Alon A, Williamson S, Nielsen R, Hubisz MT, Glanowski S, Tanenbaum DM, White TJ, Sninsky JJ, Hernandez RD, Civello D, Adams MD, Cargill M, Clark AG (2005) Natural selection on protein-coding genes in the human genome. Nature 437:1153–1157
Castric V, Bechsgaard J, Schierup MH, Vekemans X (2008) Adaptive introgression mediated by balancing selection at the self-incompatibility locus between Arabidopsis lyrata and A. halleri. PLoS Genet 4:e1000168
Charlesworth B, Charlesworth D, Barton NH (2003) The effects of genetic and geographic structure on neutral variation. Annu Rev Ecol Syst 34:99–125
Clark AG, Glanowski S, Nielsen R, Thomas PD, Kejariwal A, Todd MA, Tanenbaum DM, Civello D, Lu F, Murphy B, Ferriera S, Wang G, Zheng X, White TJ, Sninsky JJ, Adams MD, Cargill M (2003) Inferring nonneutral evolution from human-chimp-mouse orthologous gene trios. Science 302:1960–1963
Clark AG, Wang L (1997) Molecular population genetics of Drosophila immune system genes. Genetics 147:713–724
Coustau C, Renaud F, Delay B (1991) Genetic characterization of the hybridization between Mytilus edulis and Mytilus galloprovincialis on the Atlantic coast of France. Mar Biol 111:87–93
Cutter AD, Baird SE, Charlesworth D (2006) High nucleotide polymorphism and rapid decay of linkage disequilibrium in wild populations of Caenorhabditis remanei. Genetics 174:901–913
Faure B, Bierne N, Tanguy A, Bonhomme F, Jollivet D (2007) Evidence for a slightly deleterious effect of intron polymorphisms at the EF1alpha gene in the deep-sea hydrothermal vent bivalve Bathymodiolus. Gene 406:99–107
Faure MF, David P, Bonhomme F, Bierne N (2008) Genetic hitchhiking in a subdivided population of Mytilus edulis. BMC Evol Biol 8:164
Fay JC, Wu CI (2000) Hitchhiking under positive Darwinian selection. Genetics 155:1405–1413
Foll M, Gaggiotti O (2008) A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a bayesian perspective. Genetics 180:977–993
Galtier N, Depaulis F, Barton NH (2000) Detecting bottlenecks and selective sweeps from DNA sequence polymorphism. Genetics 155:981–987
Gérard K, Bierne N, Borsa P, Chenuil A, Féral J-P (2008) Pleistocene separation of mitochondrial lineages of Mytilus spp. mussels from Northern and Southern Hemispheres and strong genetic differentiation among southern populations. Mol Phylogenet Evol 49:84–91
Gosling EM, McGrath D (1990) Genetic variability in exposed-shores mussels Mytilus spp. along an environmental gradient. Mar Biol 104:413–418
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Harrison RG (1990) Hybrid zones: windows on evolutionary processes. Oxf Surv Evol Biol 7:69–128
Hasegawa M, Kishino H, Yano TA (1985) Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22:160–174
Hedrick PW (1999) Highly variable loci and their interpretation in evolution and conservation. Evolution 53:313–318
Hey J, Nielsen R (2007) Integration within the Felsenstein equation for improved Markov chain Monte Carlo methods in population genetics. Proc Natl Acad Sci USA 104:2785–2790
Hilbish TJ, Mullinax A, Dolven SI, Meyer A, Koehn RK, Rawson PD (2000) Origin of the antitropical distribution pattern in the marine mussels (Mytilus spp.): routes and timing of transequatorial migration. Mar Biol 136:69–77
Hubert F, Noel T, Roch P (1996) A member of the arthropod defensin family from edible Mediterranean mussels (Mytilus galloprovincialis). Eur J Biochem 240:302–306
Hudson RR, Kaplan NL (1985) Statistical properties of the number of recombination events in the history of a sample of DNA sequences. Genetics 111:147–164
Hudson RR, Kreitman M, Aguade M (1987) A test of neutral molecular evolution based on nucleotide data. Genetics 116:153–159
Hudson RR, Slatkin M, Maddison WP (1992) Estimation of levels of gene flow from DNA sequence data. Genetics 132:583–589
Hughes AL (2007) Looking for Darwin in all the wrong places: the misguided quest for positive selection at the nucleotide sequence level. Heredity 99:364–373
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267
Kelly JK (1997) A test of neutrality based on interlocus associations. Genetics 146:1197–1206
Lazzaro BP, Clark AG (2003) Molecular population genetics of inducible antibacterial peptide genes in Drosophila melanogaster. Mol Biol Evol 20:914–923
Lazzaro BP, Sceurman BK, Clark AG (2004) Genetic basis of natural variation in D. melanogaster antibacterial immunity. Science 303:1873–1876
Li H, Stephan W (2006) Inferring the demographic history and rate of adaptive substitution in Drosophila. PLoS Genet 2:e166
Little TJ, Colbourne JK, Crease TJ (2004) Molecular evolution of daphnia immunity genes: polymorphism in a gram-negative binding protein gene and an alpha-2-macroglobulin gene. J Mol Evol 59:498–506
Martinsen GD, Whitham TG, Turek RJ, Keim P (2001) Hybrid populations selectively filter gene introgression between species. Evolution 55:1325–1335
Martoglio B, Dobberstein B (1998) Signal sequences: more than just greasy peptides. Trends Cell Biol 8:410–415
McDonald JH (1994) Detecting natural selection by comparing geographic variation in protein and DNA polymorphisms. In: Golding B (ed) Non-neutral evolution: theories and molecular data. Chapman and Hall, New York, pp 88–100
McVean G, Awadalla P, Fearnhead P (2002) A coalescent-based method for detecting and estimating recombination from gene sequences. Genetics 160:1231–1241
Mitta G, Vandenbulcke F, Hubert F, Roch P (1999) Mussel defensins are synthesised and processed in granulocytes then released into the plasma after bacterial challenge. J Cell Sci 112:4233–4242
Mitta G, Hubert F, Dyrynda EA, Boudry P, Roch P (2000a) Mytilin B and MGD2, two antimicrobial pep-tides of marine mussels: gene structure and expression analysis. Dev Comp Immunol 24:381–393
Mitta G, Vandenbulcke F, Hubert F, Salzet M, Roch P (2000b) Involvement of mytilins in mussel antimicrobial defense. J Biol Chem 275:12954–12962
Murray MC, Hare MP (2006) A genomic scan for divergent selection in a secondary contact zone between Atlantic and Gulf of Mexico oysters, Crassostrea virginica. Mol Ecol 15:4229–4242
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nicolas P, Vanhoye D, Amiche M (2003) Molecular strategies in biological evolution of antimicrobial peptides. Peptides 24:1669–1680
Nielsen H, Brunak S, Engelbrecht J, von Heijne G (1997) Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng 10:1–6
Nielsen R, Wakeley J (2001) Distinguishing migration from isolation: a Markov chain Monte Carlo approach. Genetics 158:885–896
Nielsen R, Bustamante CD, Clark AG, Glanowski S, Sackton TB, Hubisz MJ, Fledel-Alon A, Tanenbaum DM, Civello D, White TJ, Sninsky J, Adams MD, Cargill M (2005) A scan for positively selected genes in the genomes of humans and chimpanzees. PLoS Biology 3:e170
Perron GG, Zasloff M, Bell G (2005) Experimental evolution of resistance to an antimicrobial peptide. Proc R Soc B 273:251–256
Quesada H, Warren M, Skibinski DOF (1998) Nonneutral evolution and differential mutation rate of gender-associated mitochondrial DNA lineages in the marine mussel Mytilus. Genetics 149:1511–1526
Quesada H, Ramos-Onsins SE, Aguade M (2005) Birth-and-death evolution of the Cecropin multigene family in Drosophila. J Mol Evol 60:1–11
Riginos C, Hickerson MJ, Henzler CM, Cunningham CW (2004) Differential patterns of male and female mtDNA exchange across the Atlantic Ocean in the blue mussel, Mytilus edulis. Evolution 58:2438–2451
Riginos C, Wang D, Abrams AJ (2006) Geographic variation and positive selection on M7 lysin, an acrosomal sperm protein in mussels (Mytilus spp.). Mol Biol Evol 23:1952–1965
Romestand B, Molina F, Richard V, Roch P, Granier C (2003) Key role of the loop connecting the two beta strands of mussel defensin in its antimicrobial activity. Eur J Biochem 270(13):2805–2813
Rozas J, Rozas R (1999) DnaSP version 3: an integrated program for molecular population genetics and molecular evolution analysis. Bioinformatics 15:174–175
Sackton TB, Lazzaro BP, Schlenke TA, Evans JD, Hultmark D, Clark AG (2007) Dynamic evolution of the innate immune system in Drosophila. Nature Genet 39:1461–1468
Schlenke TA, Begun DJ (2003) Natural selection drives drosophila immune system evolution. Genetics 164:1471–1480
Secor CL, Day AJ, Hilbish TJ (2001) Factors influencing differential mortality within a marine mussel (Mytilus spp.) hybrid population in southwestern England: reproductive effort and parasitism. Mar Biol 138:731–739
Semple CA, Maxwell A, Gautier P, Kilanowski FM, Eastwood H, Barran PE, Dorin JR (2005) The complexity of selection at the major primate beta-defensin locus. BMC Evol Biol 5:32
Skibinski DOF, Beardmore JA, Cross TF (1983) Aspects of the population genetics of Mytilus (Mytilidae; mollusca) in the British Isles. Biol J Linn Soc 19:137–183
Springer SA, Crespi BJ (2007) Adaptive gamete-recognition divergence in a hybridizing Mytilus population. Evolution 61:772–783
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tanguy A, Bierne N, Saavedra C, Pina B, Bachère E, Kube M, Bazin E, Bonhomme F, Boudry P, Boulo V, Boutet I, Cancela L, Dossat C, Favrel P, Huvet A, Jarque S, Jollivet D, Klages S, Lapègue S, Leite R, Moal M, Moraga D, Reinhardt R, Samain J-F, Zouros E, Canario A (2008) Increasing genomic information in Bivalves through new EST collections in four species: development of new genetic markers for environmental studies and genome evolution. Gene 408:27–36
Tennessen JA (2005) Molecular evolution of animal antimicrobial peptides: widespread moderate positive selection. J Evol Biol 18:1387–1394
Thompson JD, Higgins DG, Gibson TJ (1994) ClustalW: improving the sensitivity of progressive multiple alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Wang RL, Stec A, Hey J, Lukens L, Doebley J (1999) The limits of selection during maize domestication. Nature 398:236–239
Watterson GA (1975) On the number of segregating sites in genetical models without recombination. Theor Popul Biol 7:256–276
Weill M, Chandre F, Brengues C, Manguin S, Akogbeto M, Pasteur N, Guillet P, Raymond M (2000) The kdr mutation occurs in the Mopti form of Anopheles gambiaes.s. through introgression. Insect Mol Biol 9:451–455
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Wheat CW, Watt WB, Pollock DD, Schulte PM (2006) From DNA to fitness differences: sequences and structures of adaptive variants of Colias phosphoglucose isomerase (PGI). Mol Biol Evol 23:499–512
Yeaman MR, Yount NY (2003) Mechanisms of antimicrobial peptide action and resistance. Pharmacol Rev 55:27–55
Zasloff M (2002) Antimicrobial peptides of multicellular organisms. Nature 415:389–395
Acknowledgments
The authors are grateful to Virginie David and Maeva Fortin for technical assistance and to the National Sequencing Centre (Genoscope at Evry) and the IFR119 “Montpellier Environnement Biodiversité” for providing access to their sequencing platforms. Part of this work was carried out using the resources of the Computational Biology Service Unit at Cornell University, which is partially funded by Microsoft Corporation. We are very much indebted to Patrice David, François Bonhomme, and Grant Pogson for constructive discussions, as well as to Martin Kreitman and three anonymous referees for helpful comments on the manuscript. This work was funded by the French national programme GIS “Génomique Marine” (the Bivalvomix Project; coordinator, N. Bierne), the Marine Genomics Europe NoE (Fish & Shellfish node), and the Agence National de la Recherche (Hi-Flo project ANR-08-BLAN-0334-01; coordinator, N. Bierne). This is article 2009-018 of the Institut des Sciences de l’Evolution de Montpellier.
Author information
Authors and Affiliations
Corresponding author
Additional information
E. Boon and M. F. Faure contributed equally to this work.
Rights and permissions
About this article
Cite this article
Boon, E., Faure, M.F. & Bierne, N. The Flow of Antimicrobial Peptide Genes Through a Genetic Barrier Between Mytilus edulis and M. galloprovincialis . J Mol Evol 68, 461–474 (2009). https://doi.org/10.1007/s00239-009-9211-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-009-9211-z