Abstract
Caspase 1 (CASP-1) inhibitors share sequence similarity to CASP-1 itself and are all mapped to chr11q22.3. Here we show that these inhibitors are all products of a series of gene duplications that occurred at this locus after the divergence between human and mouse. Surprisingly, stop codons originated independently in all duplicated copies to generate CARD-only proteins with inhibitory activity. We discuss this evolutionary model in the context of both neo- and subfunctionalization.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Caspase 1 (CASP-1; also known as interleukin-1B [IL-1B] converting enzyme) is involved in both cytokine maturation and apoptosis of cells infected by pathogens (Cerretti et al. 1992; Thornberry et al. 1992; Jarvelainen et al. 2003; Kanneganti et al. 2006; Mariathasan et al. 2006; Gurcel et al. 2006). In addition to its pro-inflammatory properties, IL-1B contributes to tumorigenesis by influencing angiogenesis, proliferation, and the metastatic capacity of many tumors (Sawai et al. 2003; Apte et al. 2006; Lewis et al. 2006; Elaraj et al. 2006). CASP-1 is synthesized as an inactive zymogen and active caspase-1 is produced by proteolytic cleavage of its pro domain, which contains the CAspase Recruitment Domain (CARD). This CARD domain is localized at the amino end of CASP-1 and serves as a protein-protein interaction module that is important in protein recruitment and proteolytic activation of caspases in the context of apoptosis and inflammation (Lamkanfi et al. 2002; Bouchier-Hayes and Martin 2002). Protein interactions via the CARD domain seem to control the activation of CASP-1 through an “induced proximity” mechanism. According to this model, multimolecular aggregation of the adaptors enforces a locally high concentration of caspase zymogens. The close proximity of the zymogens then allows the immature proteases to self-activate due to their low intrinsic enzymatic activity (Salvesen and Dixit 1999).
CARD domains are found in many caspases and also in proteins known to regulate the activation of caspases. One of these CARD-containing proteins is RIP2, known to induce NF-κB activity (Thome et al. 1998). RIP2, ASC, and Ipaf are adaptor molecules reported to mediate CASP-1 activation. The association of RIP2 with CASP-1 via their homologous CARD domain accelerates the processing of CASP-1 into an active enzyme.
Interestingly, the CARD domain in these proteins is important for caspase regulation by either inducing or suppressing their activation (Srinivasula et al. 1999; Costanzo et al. 1999; Inohara et al. 1999; Reed 2000). One particularly elegant mechanism of caspase regulation involves CARD-only proteins. COP1 (CARD-only protein 1) (Lee et al. 2001; Wang et al. 2006b; Druilhe et al. 2001), INCA (inhibitory CARD) (Lamkanfi et al. 2004), and ICEBERG (Humke et al. 2000; Druilhe et al. 2001) are all CARD-only proteins that use their CARD domain to inhibit CASP-1 activation. COP1 inhibits the RIP2-induced CASP-1 oligomerization and induces NFkB activation (Lee et al. 2001; Lamkanfi et al. 2004). However, neither INCA nor ICEBERG interacts with RIP2 and induces NFkB activation (Lamkanfi et al. 2004). All three inhibitors inhibit the generation of IL-1β in cell culture models (Humke et al. 2000; Lamkanfi et al. 2004; Lee et al. 2001). Interestingly, COP1, INCA, and ICEBERG share significant sequence similarity to CASP-1 and are all mapped to chromosome 11q22, adjacent to the CASP-1 gene. These features, as pointed out by Lee et al. (2001) and Kersse et al. (2007), suggest that these inhibitors originated by gene or exon duplications from CASP-1 during evolution.
In this report, we show that indeed all three inhibitors are products of a series of gene duplications that occurred in the human lineage after the divergence between human and mouse. We provide a detailed map of all the recent duplications events at this locus and map the alterations in gene structure responsible for the emergence of the inhibitory function. Finally, we discuss these data in the context of sub- and neofunctionalization of the duplicated copies.
Materials and Methods
Dot-Plot Analyses
The genome sequences of Homo sapiens (build #36.1), Pan troglodytes (build #2) and Macaca mulatta (build #1) were obtained from the UCSC Genome Browser (http://www.genome.ucsc.edu) and were compared using the dotter program (http://www.pipmaker.bx.psu.edu/pipmaker/).
Alignments and Ka/Ks Analyses
Amino acid sequences for CASP-1 (NP_001214), COP1 (NP_001017534), INCA (NP_001007233), and ICEBERG (NP_067546) were obtained from GenBank using the corresponding accession numbers. For the sequence identified as CB985891.1, we generated an ORF (open reading frame) that encoded a CARD domain using the ORFfinder tool (http://www.ncbi.nlm.nih.gov/gorf). The +2 frame was chosen and the translated sequence used in the following alignments. Amino acid alignments were performed using the “Blast 2 sequences” available at NCBI.
Alignments using the genome sequences from Homo sapiens, Pan troglodytes (chimp), and Macaca mulatta (rhesus) were obtained using three different algorithms: Smith-Waterman (for local alignment), Needleman-Wunsch (for global alignment), and ClustalW (for multiple alignments). For the exon-intron structure characterizations, CASP-1 exons were aligned individually or together with the indicated duplicated sequences using sim4 (Florea et al. 1998) and/or Blat (Kent 2002). When necessary, a manual adjustment of the alignments was performed.
For the analysis involving exon-intron structure and the identification of internal exons that are not present in the inhibitors’ transcripts, each CASP-1 exon was aligned with the duplicated genomic sequence of a given inhibitor using Blast2sequences (NCBI). The Ka/Ks for each exon was calculated using MEGA4 (Nei & Gojobori).
Duplicated Regions
Dot plots were used to determine the duplicated genomic regions. The boundaries (start and end) of each duplicated region were determined through a manual analysis of the BLAT alignments generated by the UCSC Web browser. Manual extension of the alignment was routinely used to precisely define the boundaries of the duplicated regions.
Results and Discussion
COP1, INCA, and ICEBERG are Products of CASP-1 Duplications
CASP-1, COP1, INCA, and ICEBERG are all mapped at 11q22.3 (positions 104,401,452–104,515,016) and share high sequence identity with each other. At the protein level, COP1, INCA, and ICEBERG share 91%, 83%, and 53% identity to CASP-1, respectively (Table 1). These high sequence identities (higher for CASP-1 compared to all other caspases), coupled with their adjacent chromosome mapping, suggest that a series of duplication events from CASP-1 was involved in the origin of these genes.
To gain further insight into these duplications, we performed a dot-plot analysis of the human genomic region spanning CASP-1 and all of its inhibitors (chromosome [chr] 11: 104,399,345–104,516,445). This analysis revealed that, besides the duplications giving rise to COP1, INCA, and ICEBERG, there is an additional duplication block that gave rise to a new gene (or pseudogene) represented by GenBank entry CB985891 (Fig. 1). The characterization of CB985891 as a functional CASP-1 inhibitor remains to be explored. Another feature observed from the dot-plot analysis is that the entire CASP-1 gene and part of its flanking intergenic sequences were found at the genomic regions of the CB985891, INCA, and ICEBERG genes. On the other hand, the duplication that gave rise to COP1 involved a shorter fragment from CASP-1 whose genomic borders are located 6283 bp upstream of CASP-1 start codon and within intron 4 of CASP-1. This sequence analysis involving 11q22 allowed us to define precisely the borders of all duplicated regions, which are shown in Fig. 2.
Dot plot of human genomic region corresponding to chromosome 11: 104,399,345–104,550,000. A 160-kb region corresponding to the genomic sequence spanning CASP-1, COP1, CB, INCA, and ICEBERG was compared to itself. All genes in the region are transcribed from the minus strand. The relative position of each gene is labeled on both axes
To understand the chronological order of these events we also performed global alignments of these duplication regions. Taken together, protein and DNA alignments suggest that the first duplication from CASP-1 gave rise to ICEBERG since the alignments CASP-1/ICEBERG are the most divergent ones (Table 1). Furthermore, ICEBERG is the most distal of all inhibitors (Figs. 1 and 2). As the INCA region is more similar to CASP-1, compared to the CB985891 region, we proposed that a second duplication from CASP-1 originated the region in chromosome 11 from 104,466,665 to 104,480,013, which contains the INCA gene. A third duplication originated in the CB985891 region. Since there are regions specific to both CB985891 and INCA, it is quite likely that CB985891 originated from a duplicated copy of INCA. This is also supported by the dot-plot analysis showing a continuous block of duplication between these two genes (Fig. 1). The last duplication was a partial duplication of CASP-1 involving exons 1 to 4, part of intron 4, and the intergenic region upstream of CASP-1 (chr 11: 104,406,927–104,417,350) giving rise to COP1.
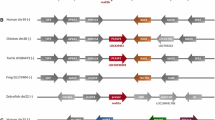
We searched all available vertebrate genomes by the presence of CASP-1 and its inhibitors. Neither CASP-1 nor any of its inhibitors were found in zebrafish, fugu, or frog genomes. On the other hand, CASP-1 was found in the mammalian genome of opossum, and a hybrid CASP-1 and CASP-4 was found in both dog and cat genomes (as also reported by Eckhart et al. 2008). Rodents and primates all have CASP-1. The inhibitors, on the other hand, are only present in primates, suggesting that the duplication events occurred after the rodent/primate divergence.
Interestingly, in the partial genome of Tupaia belangeri (a type of treeshrew, an ancestor of primates), we found a region similar to ICEBERG adjacent to CASP-1 (data not shown), reinforcing that a first duplication from CASP-1 originated ICEBERG. The availability of many other genomes from chordates will certainly shed some light on the exact timing of these duplications.
Independent Origin of New Stop Codons in CASP-1 Duplicates
As shown above, CB985891, INCA, and ICEBERG were originated, directly or indirectly, from an entire CASP-1 duplication. It is striking, however, that they all code for shorter CARD-only proteins. We wondered about the evolutionary mechanisms leading to the generation of shorter proteins at these duplicated regions. To explore this issue, cDNA sequences of CASP-1 (NM_033292.2), CB985891, INCA (NM_001007232.1), and ICEBERG (NM_021571.2) were multiple aligned using ClustalW. As shown in Fig. 3, different stop codons appeared in the duplicated copies generating shorter CARD-only proteins. ICEBERG and INCA, for instance, contain stop codons at exons 2 and 3, respectively. The alignment of ICEBERG exon 2 with CASP-1 exon 2 (Fig. 3) shows a cytosine (C)-to-adenosine (A) substitution (position chr 11: 104,514,751) creating a TAA stop codon. Also in Fig. 3, a TAG stop codon in INCA was created due to a C-to-thymine (T) substitution at position chromosome 11 104,475,302 in INCA’s exon 3.
Exon-intron organization of CASP-1 and its inhibitors. Representation of the exon (rectangles)/intron (horizontal lines) structure for CASP-1 (α isoform), COP1, CB985891, INCA, and ICEBERG. Untranslated and coding regions are labeled in black and gray, respectively. The emergence of new stop codons in all inhibitors is shown in the inserted alignments. The arrow refers to the additional stop codon used in the COP1 variant skipping exon 3
Since the EST CB985891 is not a full-length sequence, we obtained in silico the best ORF containing a CARD domain. Considering the ATG at positions 86 to 88, the first in-frame stop codon is localized at exon 3 (positions 404 to 406; TGA), producing a protein with 106 amino acids. However, this putative transcript would be a target for nonsense-mediated mRNA decay (NMD) since the TAG is located farther than 50 bp upstream of the last exon-exon junction. The alignment showing the emergence of this stop codon is also shown in Fig. 3. An alternative initiation codon at positions 216 to 218 would produce a protein with 135 amino acids, which shares similarities with both the CARD and the caspase C14 domains (data not shown). The problem with this second ORF, however, is the lack of more than half of the CARD domain, which would probably affect the inhibitory activity of this putative protein. It remains to be seen whether any of these ORFs correspond to a bona fide protein.
As we have shown, COP1 arose from a partial duplication of CASP-1 that does not contain an in-frame stop codon. Interestingly, exon 3 of both CASP-1 and COP1 is alternative. As shown in Fig. 3, a C-to-guanine (G) substitution in exon 3 of COP1 originated a new stop codon. The COP1 splicing variant that skips exon 3 produces a longer protein, not yet characterized, only with the CARD domain. The new stop codon has emerged at positions 104,417,337–104,417,339 in chromosome 11 within a 92-bp region that is specific to COP1.
Exon-Intron Organization of CASP1 Inhibitors
The emergence of new stop codons in the CASP-1 inhibitors raises interesting questions about the regulation of splicing in these genes. The INCA gene has four exons, in which exons 1, 2, and 3 are similar to the corresponding CASP-1 exons (Fig. 3). CB985891 has four exons that are similar to CASP-1 exons 1, 2, 6, and 7, respectively. ICEBERG has a three-exon structure in which exons 1 and 2 are similar to the corresponding CASP-1 exons while the third exon has no similarities to any CASP-1 genomic region. Either the similarity of the last exon was lost due to the accumulation of mutations or the exon arose de novo from a specific insertion into the ICEBERG locus.
Although there are few cDNAs reported for the CASP-1 inhibitors, it is striking that all of them seem to exclude some internal exons that code for the caspase catalytic domain. CB985891, for instance, does not use the genomic regions corresponding to CASP-1 exons 3 to 5 (Fig. 3). The same is true for INCA, whose transcripts do not use the genomic regions corresponding to CASP-1 exons 4 to 8.
The emergence of the new stop codons may explain why all duplicated copies (except COP1) have transcripts skipping several downstream exons. If those exons were not skipped, NMD would drive those transcripts to degradation. Alternatively, the possibility exists that transcripts containing those internal exons with stop codons are degraded too fast by NMD to be detected. In accordance with the first above, we found that the regions corresponding to those exons in the duplicated copies are not under negative selection, opposite to the coding exons, especially exons 1 and 2, which code for the CARD domain (Fig. 4).
Ka/Ks ratio for genomic regions corresponding to CASP-1 exons, in all duplicated blocks. CASP-1 exons were aligned with the corresponding genomic regions in all duplicated blocks. Ka/Ks for all exons 1 corresponds to 0. Exon 1, 7 bp; exon 2, 268 bp; exon 3, 64 bp; exon 4, 117 bp; exon 5, 174 bp; exon 6, 235 bp; exon 7, 174 bp; exon 8, 111 bp, exon 9, 131 bp
It is important to emphasize that the low cDNA coverage for most of the inhibitors preclude us from making more confident inferences on exon usage in their transcripts. A better understanding of the evolutionary forces acting on these genes will depend on a higher number of transcript sequences.
Analyses of CASP-1 Inhibitors in Primates
CASP-1 inhibitors are present only in primates with the exception of ICEBERG, which is also present in a primate ancestor (Tupaia belangeri). As discussed, this strongly suggests that all duplications occurred after the human-mouse split, ∼87 million years ago (mya; Springer et al. 2003).
Based on that, we compared the human genomic organization of the CASP-1 locus with the rhesus and chimpanzee genomes. INCA, CB985891, and COP1 are present in both chimpanzee and rhesus, indicating that these duplications occurred before the divergence of Hominoidea and Cercopithecoidea. Interestingly, ICEBERG is also present at the rhesus genome but absent in the chimpanzee genome as observed by Kersse et al (2007). A dot-plot analysis (Fig. 5) shows a deletion at the chimpanzee genome exactly in the ICEBERG region. As the human genome contains ICEBERG, this genomic deletion probably occurred very recently, after the separation of chimps and humans (6 mya).
Dot-plot analyses of the CASP-1 locus in chimpanzee and rhesus. (A) The genomic sequences of rhesus (chr 14: 103,596,394–103,750,052) and human (chr 11:104,399,345–104,558,178) were compared. A clear gap in the dot plot is seen in the intergenic region between INCA and ICEBERG. (B) The genomic sequences of chimpanzee (chr 11: 103,707,584–103,808,312) and human (chr 11:104,399,445–104,558,277) were compared. The loss of ICEBERG is observed in the chimpanzee genome sequence
We asked whether these primates’ duplicates contain the same premature stop codons as observed in humans. We found that all stop codons, except at the rhesus’ ICEBERG, are conserved among these primates. The rhesus ICEBERG seems to use a stop codon downstream from the original one found in humans (Fig. S1). Interestingly, the rhesus stop codon is conserved in the ICEBERG from Tupaia belangeri, suggesting that this is the ancestor stop codon.
CASP-1 Inhibitors: A Case for Neofunctionalization?
New genes originating from duplication of previous ones can undergo either neofunctionalization, in which a new function emerges in the duplicated copy, or subfunctionalization, in which each copy retains one of two functions originally present in the ancestor before the duplication. At first glance, the emergence of the CASP-1 inhibitors from duplicated copies of CASP-1 itself looks like an example of neofunctionalization. However, Wang et al. (2006a) have shown that a CASP-9 splicing variant encodes a protein containing only the CARD domain. Based on experimental evidence, those authors propose that this variant of CASP-9 may function as an apoptotic inhibitor by interfering with CARD-CARD interaction involving the prototype CASP-9. The same is true for CASP-1 itself, which shows two intron retention events in mouse (reported by GenBank entries AK132826 and AK163069) that in theory code for CARD-only proteins. This raises the possibility that an apoptotic inhibitory function is also present in proteic variants of caspases in general. This would make the evolutionary model described here for the CASP-1 inhibitors an example of subfunctionalization. A definitive conclusion on this topic will only be achieved when more cDNAs are available for these genes.
In summary, we show here that CASP-1 inhibitors originated from a series of primary and secondary gene duplications from CASP-1 itself in the human lineage after the human/mouse divergence. Their CARD-only status, and subsequent CASP-1 inhibitory capacity, was achieved either by the emergence of new independent stop codons, generating shorter proteins, or through a partial gene duplication involving only the CARD domain.
Abbreviations
- CASP:
-
Caspase
- COP:
-
CARD-only protein
- INCA:
-
Inhibitory CARD
References
Apte RN, Dotan S, Elkabets M, White MR, Reich E, Carmi Y, Song X, Dvozkin T, Krelin Y, Voronov E (2006) The involvement of IL-1 in tumorigenesis, tumor invasiveness, metastasis and tumor-host interactions. Cancer Metastasis Rev 25:387–408
Bouchier-Hayes L, Martin SJ (2002) CARD games in apoptosis and immunity. EMBO Rep 3:616–621
Cerretti DP, Kozlosky CJ, Mosley B, Nelson N, Van Ness K, Greenstreet TA, March CJ, Kronheim SR, Druck T, Cannizzaro LA (1992) Molecular cloning of the interleukin-1 beta converting enzyme. Science 256:97–100
Costanzo A, Guiet C, Vito P (1999) c-E10 is a caspase-recruiting domain-containing protein that interacts with components of death receptors signaling pathway and activates nuclear factor-kappaB. J Biol Chem 274:20127–20132
Druilhe A, Srinivasula SM, Razmara M, Ahmad M, Alnemri ES (2001) Regulation of IL-1beta generation by Pseudo-ICE and ICEBERG, two dominant negative caspase recruitment domain proteins. Cell Death Diff 8:649–657
Eckhart L, Ballaun C, Hermann M, Vandeberg JL, Sipos W, Uthman A, Fischer H, Tschachler E (2008) Identification of novel mammalian caspases reveals an important role of gene loss in shaping the human caspase repertoire. Mol Biol Evol 25:831–841
Elaraj DM, Weinreich DM, Varghese S, Puhlmann M, Hewitt SM, Carroll NM, Feldman ED, Turner EM, Alexander HR (2006) The role of interleukin 1 in growth and metastasis of human cancer xenografts. Clin Cancer Res 12:1088–1096
Florea L, Hartzell G, Zhang Z, Rubin GM, Miller W (1998) A computer program for aligning a cDNA sequence with a genomic DNA sequence. Genome Res 8:967–974
Gurcel L, Abrami L, Girardin S, Tschopp J, van der Goot FG (2006) Caspase-1 activation of lipid metabolic pathways in response to bacterial pore-forming toxins promotes cell survival. Cell 126:1135–1145
Humke EW, Shriver SK, Starovasnik MA, Fairbrother WJ, Dixit VM (2000) ICEBERG: a novel inhibitor of interleukin-1beta generation. Cell 103:99–111
Inohara N, Koseki T, del Peso L, Hu Y, Yee C, Chen S, Carrio R, Merino J, Liu D, Ni J, Nunez G (1999) Nod1, an Apaf-1-like activator of caspase-9 and nuclear factor-kappaB. J Biol Chem 274:14560–14567
Jarvelainen HA, Galmiche A, Zychlinsky A (2003) Caspase-1 activation by Salmonella. Trends Cell Biol 13:204–209
Kanneganti TD, Body-Malapel M, Amer A, Park JH, Whitfield J, Franchi L, Taraporewala ZF, Miller D, Patton JT, Inohara N, Nunez G (2006) Critical role for Cryopyrin/Nalp3 in activation of caspase-1 in response to viral infection and double-stranded RNA. J Biol Chem 281:36560–36568
Kent WJ (2002) BLAT–the BLAST-like alignment tool. Genome Res 12:656–664
Kersse K, Berghe TV, Lamkanfi M, Vandenabeele P (2007) A phylogenetic and functional overview of inflammatory caspases and caspase-1-related CARD-only proteins. Biochem Soc Trans 35:1508–1511
Lamkanfi M, Declercq W, Kalai M, Saelens X, Vandenabeele P (2002) Alice in caspase land. A phylogenetic analysis of caspases from worm to man. Cell Death Diff 9:358–361
Lamkanfi M, Denecker G, Kalai M, D’hondt K, Meeus A, Declercq W, Saelens X, Vandenabeele P (2004) INCA, a novel human caspase recruitment domain protein that inhibits interleukin-1beta generation. J Biol Chem 279:51729–51738
Lee SH, Stehlik C, Reed JC (2001) Cop, a caspase recruitment domain-containing protein and inhibitor of caspase-1 activation processing. J Biol Chem 276:34495–34500
Lewis AM, Varghese S, Xu H, Alexander HR (2006) Interleukin-1 and cancer progression: the emerging role of interleukin-1 receptor antagonist as a novel therapeutic agent in cancer treatment. J Transl Med 4:48
Mariathasan S, Weiss DS, Newton K, McBride J, O’Rourke K, Roose-Girma M, Lee WP, Weinrauch Y, Monack DM, Dixit VM (2006) Cryopyrin activates the inflammasome in response to toxins and ATP. Nature 440:228–232
Reed JC (2000) Mechanisms of apoptosis. Am J Pathol 157:1415–1430
Salvesen GS, Dixit VM (1999) Caspase activation: the induced-proximity model. Proc Natl Acad Sci USA 96:10964–10967
Sawai H, Funahashi H, Yamamoto M, Okada Y, Hayakawa T, Tanaka M, Takeyama H, Manabe T (2003) Interleukin-1alpha enhances integrin alpha(6)beta(1) expression and metastatic capability of human pancreatic cancer. Oncology 65:167–173
Springer MS, Murphy WJ, Eizirik E, O’Brien SJ (2003) Placental mammal diversification and the Cretaceous-Tertiary boundary. Proc Natl Acad Sci USA 100:1056–1061
Srinivasula SM, Ahmad M, Lin JH, Poyet JL, Fernandes-Alnemri T, Tsichlis PN, Alnemri ES (1999) CLAP, a novel caspase recruitment domain-containing protein in the tumor necrosis factor receptor pathway, regulates NF-kappaB activation and apoptosis. J Biol Chem 274:17946–17954
Thome M, Hofmann K, Burns K, Martinon F, Bodmer JL, Mattmann C, Tschopp J (1998) Identification of CARDIAK, a RIP-like kinase that associates with caspase-1. Curr Biol 8:885–888
Thornberry NA, Bull HG, Calaycay JR, Chapman KT, Howard AD, Kostura MJ, Miller DK, Molineaux SM, Weidner JR, Aunins J (1992) A novel heterodimeric cysteine protease is required for interleukin-1 beta processing in monocytes. Nature 356:768–774
Wang P, Shi T, Ma D (2006a) Cloning of a novel human caspase-9 splice variant containing only the CARD domain. Life Sci 79:934–940
Wang X, Narayanan M, Bruey JM, Rigamonti D, Cattaneo E, Reed JC, Friedlander RM (2006b) Protective role of Cop in Rip2/caspase-1/caspase-4-mediated HeLa cell death. Biochim Biophys Acta 1762:742–754
Acknowledgments
We thank Robson de Souza and Anamaria A. Camargo for discussions about the work, and Daniel O. Vidal, Diogo Meyer, Manyuan Long, and Maria Vibranovski for comments on the manuscript. J.P.C.C. and P.A.F. were supported by a fellowship from FAPESP.
Author information
Authors and Affiliations
Corresponding author
Additional information
Júlia P. C. da Cunha and Pedro A. F. Galante contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
da Cunha, J.P.C., Galante, P.A.F. & de Souza, S.J. Different Evolutionary Strategies for the Origin of Caspase-1 Inhibitors. J Mol Evol 66, 591–597 (2008). https://doi.org/10.1007/s00239-008-9110-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-008-9110-8