Abstract
The “cognate bias hypothesis” states that early in evolutionary history the biosynthetic enzymes for amino acid x gradually lost residues of x, thereby reducing the threshold for deleterious effects of x scarcity. The resulting reduction in cognate amino acid composition of the enzymes comprising a particular amino acid biosynthetic pathway is predicted to confer a selective growth advantage on cells. Bioinformatic evidence from protein-sequence data of two bacterial species previously demonstrated reduced cognate bias in amino acid biosynthetic pathways. Here we show that cognate bias in amino acid biosynthesis is present in the other domains of life—Archaebacteria and Eukaryota. We also observe evolutionarily conserved underrepresentations (e.g., glycine in methionine biosynthesis) and overrepresentations (e.g., tryptophan in asparagine biosynthesis) of amino acids in noncognate biosynthetic pathways, which can be explained by secondary amino acid metabolism. Additionally, we experimentally validate the cognate bias hypothesis using the yeast Saccharomyces cerevisiae. Specifically, we show that the degree to which growth declines following amino acid deprivation is negatively correlated with the degree to which an amino acid is underrepresented in the enzymes that comprise its cognate biosynthetic pathway. Moreover, we demonstrate that cognate fold representation is more predictive of growth advantage than a host of other potential growth-limiting factors, including an amino acid’s metabolic cost or its intracellular concentration and compartmental distribution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Proteins are polymers constructed from linear combinations of 20 amino acid monomers. Before protein synthesis can occur, amino acids must be imported whole from the external environment by cells or manufactured de novo by the appropriate metabolic pathways. Different groups of enzymes are involved in the production (i.e., biosynthesis) of one or several chemically related amino acid(s) from simpler nitrogen-containing raw materials, like ammonia (Umbarger 1978). These biosynthetic enzymes are generally expressed or catalytically active in cells when supplies of a given amino acid have been exhausted. Feedback inhibition, or the allosteric inhibition of the amino acid biosynthetic enzymes by their end products, which may be the amino acid itself or a precursor metabolite thereof, is one of several regulatory mechanisms that both prokaryotic and eukaryotic cells use to modulate the protein levels and anabolic activity of these enzymes (Szentirmai and Horvath 1976; Bhattacharjee 1985; Kovach et al., 1969; Chasin and Magasanik 1968). By and large, amino acid metabolism has remained largely unchanged over evolution, as eukaryotic amino acid biosynthetic genes exhibit sequence and catalytic homology to their bacterial orthologues (Bono et al., 1998).
However, eukaryotic regulatory control of amino acid metabolism differs from prokaryotic regulatory control of amino acid metabolism in several ways. First, eukaryotic cells are compartmentalized. Accordingly, amino acids display nonuniform intracellular distributions, with most free amino acid compartmentalized as reserves in the vacuole (in fungi) or the lysosome (in metazoa). Second, eukaryotes possess both specific and general (i.e., suprapathway) genetic regulatory controls of amino acid metabolism (Messenguy and Cooper 1977). For example, in S. cerevisiae upon amino acid starvation, among other stresses, the protein encoded by the gene GCN4 activates multiple amino acid biosynthetic pathways, as well as aminoacyl-tRNA synthetases and protein-degrading genes (Hinnebusch 2005). On the other hand, bacteria have limited general regulatory controls that affect induction of multiple amino acid biosynthesis, such as the stringent response (Jain et al., 2006).
It has been estimated that the energy stored in 20 billion to 60 billion phosphate bonds is consumed in the production of a typical microbial cell (Stouthamer 1973). One of the costliest cellular activities is amino acid biosynthesis. Recently, the “cognate bias hypothesis” was postulated and then tested using protein-sequence data from two bacterial species (Alves and Savageau 2005). However, this hypothesis falls under a more general hypothesis of metabolic cost minimization (Dufton 1997), and several studies have provided evidence for the sequence optimization of metabolic pathways whose activity depends on their own product. For example, a pre-genomic-era study of a marine bacterium showed that the sulfur-containing amino acids methionine and cysteine are selectively eliminated from the sequences of abundantly expressed proteins during sulfur starvation (Mazel and Marliere 1989). This work was later extended in yeast and bacteria, where it was shown that the number of sulfur and carbon atoms is reduced in sulfur and carbon assimilatory pathways, respectively (Baudouin-Cornu et al., 2001). Sequence optimization has been seen in the depletion of metabolically costly amino acids in the sequences of highly expressed proteins in E. coli and B. subtilis (Akashi and Gojobori 2002), and this work was later extended to four additional bacterial species (Heizer et al., 2006). Optimization of amino acid usage with respect to amino acid molecular weight has also been shown to be a general, adaptive metabolic strategy in a variety of organisms and ecological contexts (Seligmann 2003). With the precedent of these examples, we build and expand on the principles of constrained sequence optimization by combining multigenomic computational and experimental approaches to demonstrate the universality of cognate bias, and the selective growth advantage conferred by it to the model eukaryote Saccharomyces cerevisiae. As predicted under a metabolic cost minimization framework, we correlated cognate bias with experimental measures of S. cerevisiae growth in limiting amounts of amino acid. We also generalize the notion of cognate bias by determining the under- and overrepresentation of each amino acid in noncognate biosynthetic pathways.
Methods
Amino Acid Biosynthetic Pathway Sequence Retrieval
Genome-wide GO annotations were obtained from the European Bioinformatics Institute Gene Ontology Annotation Database (http://www.ebi.ac.uk/GOA) (Harris et al., 2004). It should be noted that the biosynthetic pathways for four amino acids—alanine, phenylalanine, tyrosine, and valine—were not included because of an insufficient number of annotated enzymes attributed to them. Also, due to incomplete functional annotation or their actual absence, some biosynthetic pathways in one or more of the five organisms examined in this study are not present.
Yeast and Media
MY1384 was obtained from the American Tissue Culture Collection (ATCC). Synthetic media contained 2% glucose, 6.7 g/L yeast nitrogen base (YNB), and 0.05% ammonium sulfate (AS). All 20 amino acids were purchased as powder stocks (Sunrise Science Products, USA) and dissolved in water to a final concentration of 85.6 mg/L (except leucine, which was at 173.4 mg/L) before use. The concentration of each amino acid in the culture media is in accordance with specifications for Synthetic Complete Hopkins Mixture.
Amino Acid Limitation Experiments
Parallel liquid cultures consisting of synthetic media lacking a single amino acid were inoculated with log-phase cells pregrown in synthetic medium supplemented with all 20 amino acids but washed with distilled water to remove residual amino acid. These cultures were grown at constant temperature (30°C) in a humidified incubator in NUNC 384-well, clear-bottom, untreated, sterile plates (VWR; 62409-604). A given amino acid was titrated in six replicates across a plate. As a control, we titrated water lacking amino acid in six replicates to normalize growth and allow comparison between each amino acid limitation condition; each amino acid limitation experiment was performed in duplicate. At three to five 90-min intervals during log phase growth plates were vortexed on a standard tabletop vortexer (VWR) for 10–30 s prior to measurement in a Varioskan plate reader (Thermo Electron Corp.) set to 600-nm emission. OD values of each well were normalized on a well-by-well basis using position-equivalent water-titrated culture ODs. The growth rate of cultures at a particular amino acid calculation was ultimately calculated as the average fold-change in normalized OD across two to four 90-min observation intervals, 6× replicate wells, and 2× replicate plates (yielding 24 < n < 96).
Statistical Analysis
In addition to determining the relationship between amino acid parameters and the experimental growth metric using the Spearman rank correlation, we also used a variance-dependent unequal weighting regression analysis. In particular, if X i is the property of amino acid i, and Y i are growth metrics of that amino acid across experimental replicates, then
follows the χ2 distribution with n – 2 degrees of freedom. This statistic weights more heavily those observations with lower experimental noise (i.e., when var[Y i ] is low). Moreover, it yields the variance-adjusted degree of deviation from the null hypothesis of a linear relationship between X and Y, when a and b are chosen to minimize ε.
Results
Amino Acid Cognate Bias Observed in All Three Domains of Life
We began by compiling a list of all enzymes belonging to each amino acid biosynthetic pathway (as defined by the Gene Ontology [GO] “Amino Acid Biosynthesis” annotation [GO:0008652]) or one of its subannotations (Gene Ontology Consortium 2001) for the following species: Escherichia coli (Eubacteria; Gram-negative), Bacillus subtilis (Eubacteria; Gram-positive), Methanococcus jannaschii (Archaebacteria), Saccharomyces cerevisiae (Eukaryote; Fungus), and Homo sapiens (Eukaryote; Metazoan). For each amino acid, we counted the number of times it appears in its cognate biosynthetic pathway and then compared this observed number to an estimate of the a priori expected number determined by multiplying the size of the cognate biosynthetic pathway by the average frequency of the particular amino acid within all amino acid biosynthetic pathways. We call the ratio of the expected number to the observed number “cognate fold-representation.” Importantly, and in contrast to previous studies (Alves and Savageau 2005), we chose not to use the entire proteome to determine the expected number of each amino acid because of the possibility of systematic amino acid frequency biases in the sequences of amino acid biosynthetic enzymes. However, this choice of baseline expectation did not qualitatively affect our results; using an amino acid’s proteomic observed frequency instead of our baseline yields essentially the same qualitative results (Supplementary Fig. 1). In related work, we found that amino acid composition varies strongly as a function of GO function (de Bivort and Perlstein, unpublished data). Previous studies have used genomic amino acid frequencies to calculate amino acid biases within biosynthetic pathways, but this could introduce systematic errors if biosynthetic enzymes, as a group, exhibit amino acid biases with respect to genomic averages. Therefore, the best control for detecting cognate bias in a particular amino acid biosynthetic pathway is to use a baseline control composed of the average observed frequency across all amino acid biosynthetic pathways. This choice of baseline frequency furthermore controls for other physicochemical properties common to biosynthetic enzymes, such as length, hydrophobicity, and total metabolic cost.
We calculated cognate fold-representation of each amino acid (Table 1). Two striking features are immediately apparent. First, validating the study by Alves and Savageau, we show that many amino acids are underrepresented in their cognate biosynthetic pathways (bold rows). Specifically, 10 of 15 amino acid pathways have fewer cognate residues than expected in yeast. Within E. coli, B. subtilis, M. jannaschii, and H. sapiens: 13 of 14, 8 of 13, 5 of 10, and 6 of 10 pathways show underrepresentation, respectively. The percentage of underrepresentation attains a minimum in the case of the cysteine biosynthetic pathway in S. cerevisiae, in which 12 cysteine residues are expected but only 2 exist (0.167-fold underrepresentation). Cysteine is a rare amino acid, and its frequent underrepresentation across species illustrates the second observation: there is an evolutionarily conserved trend between the rarity of an amino acid and its cognate representation. Rare amino acids (e.g., cysteine and tryptophan) and the very most common amino acids (e.g., leucine), but not amino acids of intermediate abundance, exhibit greater conserved underrepresentation than expected by chance. This is depicted in Fig. 1 as a plot of an amino acid’s cognate fold-representation (the ratio of observed to expected) versus its expected frequency for all analyzed pathways in E. coli, B. subtilis, and S. cerevisiae.
Amino acid fold-representation in its cognate biosynthetic enzymes versus frequency. Points falling below the unity line indicate pathways in which the cognate amino acid is underrepresented. S. cerevisiae pathways are indicated by squares; B. subtilis, by circles; E. coli, by triangles. The degrees to which there were monotonically increasing trends in the -fold versus frequency relationship was tested for each organism using the Spearman rank correlation: S. cerevisiae, r = 0.289, p = 0.296; B. subtilis, r = 0.291, p = 0.334; E. coli, r = 0.635, p = 0.014; aggregated data, r = 0.357, p = 0.0203
For example, there are 2413, 2424, and 4984 total amino acids in the tryptophan biosynthetic pathways of E. coli, B. subtilis, and S. cerevisiae, respectively; one would expect to find 22, 18, and 39 tryptophan residues, but there are in fact only 8, 9, and 20 tryptophan residues, respectively, a ~50% reduction in each case. However, cysteine, which is underrepresented in its cognate biosynthetic pathway in E. coli and S. cerevisiae, is curiously overrepresented in its cognate biosynthetic pathway in B. subtilis. Cognate overrepresentation of cysteine in B. subtilis was not observed by Alves and Savageau because they analyzed only two members of the cysteine biosynthetic pathway, cysK (cysteine synthase) and cysE (serine acetyltransferase), while we included nine enzymes, including those involved in sulfur assimilation, an indispensable component of de novo cysteine production. The cysteine biosynthetic pathway in B. subtilis does not appear to be substantially different from that of E. coli, so cognate overrepresentation of cysteine may reflect local environmental conditions of B. subtilis during its evolution history (Albanesi et al., 2005). For example, E. coli is gut-dwelling, whereas B. subtilis is soil-dwelling, and this difference may have affected the elemental sulfur availability during the evolutionary optimization of cognate cysteine representation. Nevertheless, the positive correlation between fold and frequency is statistically significant in the case of B. subtilis (p = 0.014 by Spearman rank correlation) and in the aggregated case of the pathways from all three organisms (p = 0.020). The correlation between fold and frequency in yeast and B. subtilis is diminished by two outliers, histidine (in S. cerevisiae) and cysteine (in B. subtilis), both of which are relatively rare amino acids that are overrepresented in their species-specific cognate biosynthetic pathways.
Evolutionarily Conserved Amino Acid Noncognate Bias Also Observed
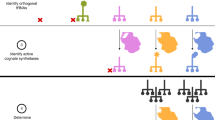
In order to explore amino acids’ compositional bias further both in cognate and in noncognate amino acid biosynthetic pathways, as well as to visualize better the evolutionary conservation of these relationships across E. coli, B. subtilis, M. jannaschii, S. cerevisiae, and H. sapiens, we assembled networks for the 16 annotated amino acids and for 3 higher-level amino acid groups based on shared biophysical properties (aromatic, branched-chain, and sulfur-containing) (Fig. 2). These higher-level groups include enzymes that are involved in the production of more than one related amino acid. For example, the yeast gene ILV3, dihydroxyacid dehydratase, catalyzes the third biosynthetic step in the pathway that eventually yields the branched-chain amino acids valine, leucine, and isoleucine (Velasco et al., 1993). An arrow connecting amino acid (or higher-level amino acid group) A to B indicates that there is an underrepresentation of amino acid A (or that group of amino acids) within the biosynthetic enzymes that produce B. Underrepresentation of an amino acid in its cognate biosynthetic pathway is depicted as a self-loop. We observe several instances of underrepresentation of an amino acid in a noncognate biosynthetic pathway (Fig. 2).
Networks of amino acid underrepresentation in cognate and noncognate biosynthetic pathways. An arrow connecting an amino acid or higher-level amino acid group to another indicates that the source amino acid (or group) is underrepresented in the sequence of the biosynthetic enzymes of the destination amino acid (or group), with a χ2 significance of p < 0.05. Icons on the arrows indicate conserved relationship across species: circles on arrows indicate identical conservation of that relationship in another species. Triangles indicate a related relationship in which the underrepresentation of amino acid A in pathway B in one species is conserved in another species as either (1) an underrepresentation of A with in the higher-level pathway of enzymes that produce B (as well as the other amino acids in B’s synthetic family) or (2) an underrepresentation of the amino acids in A’s higher-level family within the B biosynthetic pathway. The color of a triangle or circle indicates the organism in which the relationship is also found: brown, E. coli; green, B. subtilis; cyan, M. jannaschii; orange, S. cerevisiae; and purple, H. sapiens
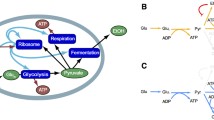
The most conserved noncognate underrepresentation is glycine in the methionine biosynthetic pathways of E. coli, B. subtilis, and H. sapiens and in the biosynthetic pathways responsible for producing sulfur-containing amino acids (methionine and cysteine) in S. cerevisiae (Fig. 2). This observation is consistent with the secondary metabolism of glycine, which can serve as an intermediate in the production of methionine via single-carbon metabolism. The following is an elucidation of the metabolic steps linking glycine and methionine. Methionine is produced by a series of enzymatic transformations starting with the metabolite homoserine, which itself is derived from the amino acid aspartate. The immediate precursor of methionine is homocysteine, which is methylated by the methyl-group donor 5-methyltetrahydrofolate (methyl-THF [Thomas and Surdin-Kerjan 1997]). Catabolism of glycine (as well as serine and formate) fuels the THF cycle, which is an important source of single-carbon units for many downstream cellular processes, including methionine production (Newman and Magasanik 1963; Christensen and MacKenzie 2006; Piper et al., 2000). Therefore one can envision the following scenario: methionine starvation increases demands on the THF cycle; free glycine is cannibalized by single-carbon metabolism in order to meet these demands; glycine that is no longer required to assemble the enzymes of the methionine biosynthetic pathway may instead be used to increase methionine production through single-carbon metabolism. As serine can also be a catabolic precursor of the THF cycle, we appropriately find that it, like glycine, is also underrepresented in sulfur amino acid synthesizing enzymes in yeast and humans (Fig. 2).
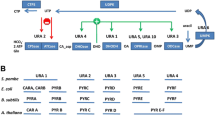
We also visualized the evolutionary conservation of amino acid overrepresentation both in cognate and in noncognate amino acid biosynthetic pathways (Fig. 3). Strikingly, tryptophan is overrepresented in the asparagine biosynthetic pathway in all organisms except H. sapiens, which lacks an identifiable tryptophan biosynthetic pathway. In fact, aromatic amino acids as a whole are overrepresented in asparagine biosynthetic pathways (Fig. 3). Secondary metabolism of tryptophan is not an obvious explanation for this observation, because tryptophan is not an intermediate in asparagine production. In fact, tryptophan is derived from the glycolytic intermediate erythrose-4-phosphate, while asparagine is derived from the tricarboxylic acid cycle intermediate oxaloacetate. However, the strong evolutionary conservation of this observation suggests that asparagine starvation correlates, or co-occurs, with tryptophan abundance. Therefore, the asparagine biosynthetic pathway may serve as a “sink” for excess tryptophan, but further experiments are needed to verify this claim.
Networks of amino acid overrepresentation in cognate and noncognate biosynthetic pathways. As Fig. 1 except that an arrow connecting amino acid A to amino acid B indicates that there is an overrepresentation of A residues in the B biosynthetic enzymes
Experimental Validation of the Cognate Bias Hypothesis Using Yeast
Using S. cerevisiae, we sought to verify experimentally whether the fold change in cognate underrepresentation of a given amino acid causes a detectable change in the actual final yield of a population of cells challenged with amino acid limitation, as had been previously predicted (Alves and Savageau 2005). According to the cognate bias hypothesis, eliminating a cognate amino acid from an enzyme within its biosynthetic pathway would confer a selective growth advantage on cells whenever that amino acid is scarce; fewer occurrences of the cognate amino acid in its biosynthetic pathway will engender greater growth advantage. We used a prototrophic yeast strain MY1384 (isogenic to Σ1278b), which is competent for growth on synthetic media lacking all 20 amino acids. Most commonly used laboratory strains are auxotrophic for growth in media lacking particular amino acids, making them suitable for crossing, but unsuitable for our studies of amino acid limitation. Amino acid biosynthetic pathways in yeast are subject to varying degrees of feedback inhibition, and these pathways are generally derepressed upon starvation of their cognate amino acid (Jones and Fink 1982). In total we screened all 15 amino acids for which there are annotated amino acid biosynthetic enzymes in S. cerevisiae. The data for three representative amino acids—cysteine, asparagine, and histidine—are discussed below in detail, while the remaining data may be found in Supplementary Materials.
Cysteine is a highly underrepresented amino acid, with only 2 cysteine residues observed in its cognate biosynthetic pathway, while 12 are expected (0.17). During cysteine starvation, cells displayed reduced final yields (Fig. 4A). As a function of cysteine concentration, final yield declined roughly linearly from its baseline levels to 85% of baseline (see Supplementary Materials) as cysteine was diluted across the cultures to 2−4 of its original concentration. This was the greatest decline in final yield seen after any amino acid depletion. Further dilutions did not further decrease final yield. Asparagine is a moderately underrepresented amino acid, with 79 asparagine residues observed in its cognate biosynthetic pathway, while 105 are expected (0.75) (Fig. 4B). During asparagine starvation, cells exhibited their lowest final yield in the 2−8 dilution, but only a 0.7% decline in total final yield compared to baseline.
Amino acid limitation experiments in S. cerevisiae. Normalized growth rate of wild-type yeast in media progressively dilute in (A) cysteine, (B) asparagine, and (C) histidine. Average growth rates (see Methods) were standardized for comparability across depleted amino acids by scaling the maximal growth rate to 1 and minimum growth rate to 0. Error bars are the standard error of the mean (12 < n < 36). (E–P) Growth metric comparing growth in intermediately depleted media and strongly diluted media (average of left gray region minus average of right gray region) versus a suite of amino acid parameters: (E) mass, (F) van der Waals volume, (G) hydrophobicity, (H) pK1, (I) metabolic phosphate cost, (J) total metabolic cost, (K) pK2, (L) metabolic hydrogen cost, (M) size of biosynthetic pathway (in amino acids), (N) isoelectric point (pI), (O) amino acid frequency, and (P) fold-representation. Plots are ordered from least to most correlated (Spearman rank correlation); see text for r values
On the other hand, histidine is an example of an overrepresented amino acid, with 156 histidine residues in its cognate biosynthetic pathway, while only 139 are expected (1.14) (Fig. 4C). This represents a potential metabolic cost to cells experiencing histidine starvation. Indeed, during histidine starvation, cells experience a 1.25% decrease in final yield following 1:1000 or higher dilution of histidine. Interestingly, the final yield following histidine depletion is lowest in the 2−4 and 2−5 dilutions. Cells experiencing complete depletion of a given amino acid will have significant difficulty synthesizing any new proteins, let alone the biosynthetic enzymes that can synthesize that amino acid de novo and, therefore, must rely on compensatory mechanisms such as releasing that amino acid from the vacuole or regenerating it by protein catabolism.
Differences in the declines in final yield following elimination of an amino acid from the culture medium presumably reflect varying degrees to which the cells can acquire this amino acid by compensatory mechanisms. Therefore we would predict that amino acids with the weakest compensatory mechanisms (such as cysteine) would receive the greatest benefit following an evolutionary underrepresentation in their cognate synthetic enzymes. We found that the difference in final yield between baseline and highly diluted media (measured as the average growth rates of the 2−11 through 2−16 dilutions) was negatively correlated with fold-representation in cognate pathways. This trend was not statistically significant across all amino acids, the notable outlier being glutamate, which exhibited a 13% reduction in growth compared to baseline, but exhibits 2% overrepresentation. Excluding glutamate, the negative correlation is significant, with a p-value of 0.028 (Fig. 5). Glutamate is the amino-group donor in transamination reactions vital to the biosynthesis of many amino acids from glycolytic and citric acid cycle intermediates, and its role as such may confound the effects of its cognate overrepresentation.
Fold representation in cognate pathways versus growth rate reduction following amino acid depletion. Fold representation is negatively correlated with the percentage difference in culture growth rate between wild-type yeast in full media and in media highly depleted for each amino acid (average growth rate of the 2−11 through 2−16 dilutions). Gray line is the linear best fit with Spearman rank correlation; r = 0.42 and p = 0.11 for all amino acids. Excluding the glutamate data point (orange) gives r = 0.59, p = 0.02, and the red best-fit line
Amino Acid Cognate Bias Correlates with Selective Growth Advantage
Underrepresentation of an amino acid might bring about its strongest selective advantage in cases of moderate amino acid deprivation, during which incorporation of the amino acid into the biosynthetic enzymes begins to become rate-limiting, but the cell is not yet relying entirely on any compensatory mechanisms as sources of the amino acid. To gauge the cultures’ behavior in these conditions, we extracted a metric that captures the growth advantage afforded cells under moderate amino acid limitation, when underrepresentation is predicted to confer the most energy cost savings. To do so, we averaged the final yields of the 2−1 through 2−5 dilution cultures and subtracted the growth rates in highly diluted cultures (2−11 through 2−16 dilutions). We correlated the growth metric derived from experiments on all 15 amino acids to a battery of physical parameters (Nelson and Cox 2000) in an effort to discern what property of amino acids best predicts how well a culture would grow following moderate depletion of a single amino acid (Figs. 4E–P) (Karlin and Bucher 1992). We found that the following parameters are essentially nonpredictive: total number of amino acids in the cognate biosynthetic pathway (r = –0.02), amino acid mass (r = 0.05), metabolic synthetic cost in units of phosphate charges (r = –0.068) (Akashi and Gojobori 2002), metabolic phosphate cost plus cost in reducing hydrogens (r = 0.074) (Akashi and Gojobori 2002), amino acid volume (r = –0.13), metabolic cost in hydrogens (r = 0.167) (Akashi and Gojobori 2002), intracellular amino acid concentration (r = 0.18) (Jones and Fink 1982), amino acid hydrophobicity (Kyte and Doolittle 1982) (r = 0.22), and pK1 (COOH) (r = 0.23). Because the measures of metabolic cost were determined in E. coli, we excluded lysine from their correlation analyses, as lysine biosynthesis in S. cerevisiae uses different enzymes (Umbarger 1978). The following parameters are mildly predictive: the vacuolar fraction of intracellular amino acid pools (r = –0.27) (Jones and Fink 1982), pK2 (NH2) (r = –0.29), and isoelectric point (pI) (r = –0.35). The two most predictive parameters are an amino acid’s expected frequency (Fig. 1) (r = –0.45) and the fold-representation of an amino acid in its cognate biosynthetic pathway (r = –0.51, p = 0.052). This result was reiterated using a variance-weighted regression analysis in which the growth metric of amino acids were more heavily weighted if their experimental replicates showed less variance (Table 2, Methods). This second method reiterates the result attained using correlation analysis, particularly that the fold-cognate representation is the most predictive factor of the experimental growth metric.
Discussion
This study of amino acid metabolism offers a striking example of the thrift and efficiency of natural selection in response to the complex crosscurrent of selective pressures that have arisen during evolution. For example, the unique ecological history of B. subtilis, a soil-dwelling bacterium, in contrast to that of E. coli, a human gut-dwelling symbiont, may have overridden the tendency of a rare amino acid like cysteine to exhibit cognate underrepresentation. More to the point, significant differences in amino acid usage between proteomes of different species, let alone between related pathways in the proteome of a single species, have often been observed but, due to the intrinsic complexity underlying the observation, have not yet been fully explained (Gerstein and Hegyi 1998; Pascal et al., 2006; Bogatyreva et al., 2006). It is likely that no single parameter can fully explain our observations, which are almost assuredly the products of a complex integration of many variables, such as protein thermostability, codon-usage bias, metabolic flux, and ecological conditions.
However, in an unbiased fashion, we evaluated the potential predictive role of many of those variables within the well-established metabolic cost minimization framework. We found that the physicochemical properties of amino acids, such as mass and hydrophobicity, do not predict final yield in a regime of moderately limiting amino acid levels. Therefore the structure of biosynthetic enzymes per se plays a negligible role in the sequence optimization of amino acid biosynthetic pathways. This interpretation is supported by the fact that there are no known amino acid compositional biases of biosynthetic enzymes as a class, which may be cytoplasmic or localize to intracellular compartments (e.g., mitochondria), and exhibit a diverse array of enzymatic activities. However, such compositional biases are theoretically detectable, because hydrophobic amino acids are known to be overrepresented in integral membrane proteins as a class (Pascal et al., 2005). The metabolic cost of an amino acid is also a poor predictor. On the other hand, cognate underrepresentation, which is especially significant for rare amino acids, is the strongest predictor. We can reconcile those two observations as follows. Akashi and Gojobori demonstrated that the sequences of abundantly expressed proteins, including amino acid biosynthetic pathways, contain fewer metabolically expensive amino acids (e.g., tryptophan). Therefore, genome-wide sequence optimization on the basis of amino acid metabolic cost may dampen the signal of specific cognate underrepresentation in amino acid biosynthetic pathways.
The variation in final yield between baseline and highly diluted amino acid media is suggestive of different compensatory mechanisms associated with each amino acid. For example, yeast cells might be able to more easily convert or degrade related metabolites to histidine, which does not exhibit cognate underrepresentation, following depletion compared to cysteine, which exhibits significant cognate underrepresentation. Alternatively, steady-state concentrations of amino acids in the cytosol and/or vacuole may vary such that depletion of some amino acids has more or less of an effect on growth because the cell is capable of drawing down on smaller or larger intracellular pools (Messenguy et al., 1980; Kitamoto et al., 1988). This interpretation in consistent with the observation that amino acid vacuolar compartmentalization is a modest predictor of growth advantage during amino acid limitation. Appropriately, 90% of intracellular histidine, which is overrepresented in its cognate biosynthetic pathway in S. cerevisiae, is present in the vacuole, and this reservoir of histidine may relax the selective pressure to reduce its cognate bias (Jones and Fink 1982). Regardless of the mechanism that explains the differences between baseline final yield and final yield in highly depleted media, if a cell is unable to generate quickly the amino acid by secondary metabolism or by tapping its intracellular pools, any reduction in the amount of that amino acid needed to create its biosynthetic enzymes will provide selective advantage.
Future computational studies and experiments may assess the significance of amino acid noncognate overrepresentation. Until then, we speculate that directed overrepresentation might arise as a consequence of the zero-sum constraint of switching residue identities away from the cognate amino acid. Overrepresentation of metabolically unrelated amino acids (such as tryptophan in the asparagine pathway) may be the least disadvantageous because environmental depletion of tryptophan may be essentially uncorrelated with the need to synthesize asparagine biosynthetic enzymes. The optimization strategy we observe in amino acid biosynthetic pathways may also manifest itself in amino acid composition in the enzymes that comprise other metabolic (e.g., glycolysis) or nonmetabolic (e.g., signal transduction) pathways.
References
Akashi H, Gojobori T (2002) Metabolic efficiency and amino acid composition in the proteomes of Escherichia coli and Bacillus subtilis. Proc Natl Acad Sci USA 99:3695–3700
Albanesi D, Mansilla MC, Schujman GE, de Mendoza D (2005) Bacillus subtilis cysteine synthetase is a global regulator of the expression of genes involved in sulfur assimilation. J Bacteriol 187:7631–7638
Alves R, Savageau MA (2005) Evidence of selection for low cognate amino acid bias in amino acid biosynthetic enzymes. Mol Microbiol 56:1017–1034
Baudouin-Cornu P, Surdin-Kerjan Y, Marliere P, Thomas D (2001) Molecular evolution of protein atomic composition. Science 293:297–300
Bhattacharjee JK (1985) Alpha-aminoadipate pathway for the biosynthesis of lysine in lower eukaryotes. Crit Rev Microbiol 12:131–151
Bogatyreva NS, Finkelstein AV, Galzitskaya OV (2006) Trend of amino acid composition of proteins of different taxa. J Bioinform Comput Biol 4:597–608
Bono H, Ogata H, Goto S, Kanehisa M (1998) Reconstruction of amino acid biosynthesis pathways from the complete genome sequence. Genome Res 8:203–210
Chasin LA, Magasanik B (1968) Induction and repression of the histidine-degrading enzymes of Bacillus subtilis. J Biol Chem 243:5165–5178
Christensen KE, MacKenzie RE (2006) Mitochondrial one-carbon metabolism is adapted to the specific needs of yeast, plants and mammals. Bioessays 28:595–605
Dufton MJ (1997) Genetic code synonym quotas and amino acid complexity: Cutting the cost of proteins? J Theor Biol 187:165–173
Gene Ontology Consortium (2001) Creating the gene ontology resource: design and implementation. Genome Res 11:1425–1433
Gerstein M, Hegyi H (1998) Comparing genomes in terms of protein structure: surveys of a finite parts list. FEMS Microbiol Rev 22:277–304
Harris MA, et al. (2004) The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res 32[Database Issue]:D258–D261
Heizer EM, Raiford DW, Raymer ML, Doom TE, Miller RV, Krane DE (2006) Amino acid cost and codon-usage biases in 6 prokaryotic genomes: a whole-genome analysis. Mol Biol Evol 23:1670–1680
Hinnebusch AG (2005) Translational regulation of GCN4 and the general amino acid control of yeast. Annu Rev Microbiol 59:407–450
Jain V, Kumar M, Chatterji D (2006) ppGpp: stringent response and survival. J Microbiol 44:1–10
Jones EW, Fink GR (1982) The molecular biology of the yeast Saccharomyces. Metabolism and gene expression. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Karlin S, Bucher P (1992) Correlation analysis of amino acid usage in protein classes. Proc Natl Acad Sci USA 89:12165–12169
Kitamoto K, Yoshizawa K, Ohsumi Y, Anraku Y (1988) Dynamic aspects of vacuolar and cytosolic amino acid pools of Saccharomyces cerevisiae. J Bacteriol 170:2683–2686
Kovach JS, Berberich MA, Venetianer P, Goldberger RF (1969) Repression of the histidine operon: effect of the first enzyme on the kinetics of repression. J Bacteriol 97:1283–1290
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132
Mazel D, Marliere P (1989) Adaptive eradication of methionine and cysteine from cyanobacterial light-harvesting proteins. Nature 341:245–248
Messenguy F, Colin D, ten Have JP (1980) Regulation of compartmentation of amino acid pools in Saccharomyces cerevisiae and its effects on metabolic control. Eur J Biochem 108:439–447
Messenguy F, Cooper TG (1977) Evidence that specific and “general” control of ornithine carbamoyltransferease production occurs at the level of transcription in Saccharomyces cerevisiae. J Bacteriol 130:1253–1261
Nelson DL, Cox MM (2000) Lehninger principles of biochemistry, 3rd ed. Worth, New York
Newman EB, Magasanik B (1963) The relation of serine-glycine metabolism to the formation of single-carbon units. Biochim Biophys Acta 78:437–448
Pascal G, Medique C, Danchin A (2006) Persistent biases in the amino acid composition of prokaryotic proteins. Bioessays 28:726–738
Piper MD, Hong SP, Ball GE, Dawes IW (2000) Regulation of the balance of one-carbon metabolism in Saccharomyces cerevisiae. J Biol Chem 275:30987–30995
Seligmann H (2003) Cost-minimization of amino acid usage. J Mol Evol 56:151–161
Stouthamer AH (1973) A theoretical study on the amount of ATP required for synthesis of microbial cell material. Antonie Van Leeuwenhoek 39:545–565
Szentirmai A, Horvath I (1976) Regulation of branched-chain amino acid biosynthesis. Acta Microbiol Acad Sci Hung 23:137–149
Thomas D, Surdin-Kerjan Y (1997) Metabolism of sulfur amino acids in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 61:503–532
Umbarger HE (1978) Amino acid biosynthesis and its regulation. Annu Rev Biochem 47:532–606
Velasco JA, Cansado J, Pena MC, Kawakami T, Laborda J, Notario V (1993) Cloning of the dihydroxyacid dehydratase-encoding gene (ILV3) from Saccharomyces cerevisiae. Gene 137:179–185
Acknowledgments
We are indebted to Drs. Boris Magasanik and Finny Kuruvilla for their constructive discussions. This work was supported by a Merck-Wiley Fellowship (B.L.d.) and by the National Institute of General Medicine Sciences (S. L. S.). S. L. S. is an Investigator at the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Additional information
Reviewing Editor: Dr. Niles Lehman
Ethan O. Perlstein and Benjamin L. de Bivort contributed equally to this work.
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Perlstein, E.O., de Bivort, B.L., Kunes, S. et al. Evolutionarily Conserved Optimization of Amino Acid Biosynthesis. J Mol Evol 65, 186–196 (2007). https://doi.org/10.1007/s00239-007-0013-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-007-0013-x