Abstract
Most investigations of the forces shaping protein evolution have focussed on protein function. However, cells are typically 50%–75% protein by dry weight, with protein expression levels distributed over five orders of magnitude. Cells may, therefore, be under considerable selection pressure to incorporate amino acids that are cheap to synthesize into proteins that are highly expressed. Such selection pressure has been demonstrated to alter amino acid usage in a few organisms, but whether “cost selection” is a general phenomenon remains unknown. One reason for this is that reliable protein expression level data is not available for most organisms. Accordingly, I have developed a new method for detecting cost selection. This method depends solely on interprotein gradients in amino acid usage. Applying it to an analysis of 43 whole genomes from all three domains of life, I show that selection on the synthesis cost of amino acids is a pervasive force in shaping the composition of proteins. Moreover, some amino acids have different price tags for different organisms—the cost of amino acids is changed for organisms living in hydrothermal vents compared with those living at the sea surface or for organisms that have difficulty acquiring elements such as nitrogen compared with those that do not—so I also investigated whether differences between organisms in amino acid usage might reflect differences in synthesis or acquisition costs. The results suggest that organisms evolve to alter amino acid usage in response to environmental conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
That natural selection acts on protein function is beyond dispute. However, in many proteins only a few sites are under strong functional selection—that is, only a few sites must contain one particular amino acid and no other (Suckow et al. 1996). So what, other than mutation and drift, determines which amino acids make up the rest of the protein? One factor that has been suggested to be important is selection to reduce protein synthesis costs (Richmond 1970; Akashi and Gojobori 2002), or “cost selection.”
A substantial component of the cost of making the protein complement of a cell is the cost of synthesizing, or otherwise obtaining, amino acids. Cells may thus be under pressure to use amino acids that are cheap to make or easy to acquire, particularly in proteins that they mass-produce. Expensive amino acids are thus predicted to be disproportionately common in rarely expressed proteins, and cheap amino acids disproportionately common in highly expressed proteins (Richmond 1970). This effect should be large: protein expression levels are approximately lognormally distributed over five orders of magnitude (Neidhardt et al. 1990; Ghaemmaghami et al. 2003).
Measuring cost selection is not, however, straightforward. For most organisms, direct data on protein expression levels do not exist. To date, therefore, investigations of cost selection on amino acid usage have had to rely on proxies—codon usage patterns (in highly expressed protein genes many genomes avoid all but one of the codons that could encode a given amino acid [Bennetzen and Hall 1982; Ikemura 1982]) or mRNA expression levels. By means of these proxies, it has been shown that per-residue synthesis cost—or in some cases its proxy, molecular weight—decreases with “protein” expression level in a variety of prokaryotes (Lobry and Gautier 1994; Akashi and Gojobori 2002; Palacios and Wernegreen 2002; Zavala et al. 2002; Seligmann 2003; Das et al. 2005; Schaber et al. 2005; Heizer et al. 2006) and eukaryotes (Garat and Musto 2000; Jansen and Gerstein 2000; Seligmann 2003; Urrutia and Hurst 2003; Peixoto et al. 2004; Chanda et al. 2005).
But to demonstrate the existence of cost selection it is not enough just to show a decrease in average per-residue cost with expression. For such a decrease could easily be caused by transcription-induced mutation—the point being that highly expressed genes are transcribed more often, and while being transcribed they are subjected to increased risk of mutation. (This is because the nontemplate strand is left naked and vulnerable to assault by free radicals, and the template strand is touched by various proteins [Francino and Ochman 1997].) Differences in mutation pressure lead to amino acid substitutions (Sueoka 1961; King and Jukes 1969; Lobry 1997; Gu et al. 1998; Nishizawa and Nishizawa 1998; Singer and Hickey 2000), so, a priori, there is a 50% probability that mutation acting on its own will generate a decrease in cost with expression. Another problem is that 3 of the amino acids—tryptophan, phenylalanine, and tyrosine—are much more expensive than the other 17 and, thus, have a disproportionate weight in determining whether a protein is expensive or cheap. But unfortunately these three amino acids share features other than high cost—for example, they are the only three amino acids with phenol rings—and thus a gradient in cost might be caused not by cost selection but by functional selection on some other shared feature of these three costly outliers. To resolve these issues, to convince that cost selection—rather than transcription-induced mutation or functional selection—is really present, it is necessary to go beyond simply showing a decrease in average residue cost with expression. It is necessary to show that the increase (or decrease) in each of the 20 amino acids is proportionate to its synthesis cost. Studies on cost selection have largely ignored this need. The two clear exceptions, Akashi and Gojobori (2002) and Schaber et al. (2005), do provide evidence in favor of cost selection, although in both cases the support is partial only.
An additional problem is with the proxies used for protein expression level. Messenger RNA expression data are available for only a handful of organisms and, in any case, seem typically to show correlation coefficients of ≤0.5 with protein expression levels (Abbott 1999). (In addition to experimental noise, two main factors lead to the poor correlations: the three to four orders of magnitude of variation in protein half-lives [Wagner 2005] and the at least one order of magnitude of variation in the speed at which different codons are translated [Sorensen et al. 1989; Sorensen and Pedersen 1991].) Codon preference data are more widely available, but show even weaker correlations with protein expression.
Furthermore, even if one obtained actual protein expression levels (rather than just a proxy for them), there are several factors that muddy the relationship between protein expression level and cost selection. The strength of cost selection on amino acid usage not only varies positively with protein expression level, but also varies negatively with the degree of functional constraint—that is, with the strength of purifying selection on protein function (Swire 2003): if you need tryptophan at a particular site to survive, then functional selection will maintain tryptophan at that site, with the high cost of tryptophan being utterly irrelevant. But constraint data is, at best, difficult to obtain. More usually, it is unavailable. Worse still, the strength of purifying selection on proteins itself correlates positively with expression level (Sharp and Li 1987; Agrafioti et al. 2005; Drummond et al. 2005). The interplay among these three correlations—negative between cost and expression, positive between cost and purifying selection, and positive between expression and purifying selection—means that analyses of cost purely in terms of protein expression level should be, and indeed are, partially confounded (Swire 2003): such analyses will systematically underestimate the evidence for cost selection.
For all these reasons, any study that seeks to test whether cost selection on amino acid usage is a general phenomenon—a study that requires the analysis of numerous whole genomes—is faced with serious problems. Here, I avoid these problems by developing a new methodology, the “gradient consistency test,” which can detect the signature of cost selection in a genome without reference to expression level or constraint data. Instead, this method depends solely on the analysis of interprotein gradients in amino acid usage. It is motivated by the inference that if cost selection has occurred, then cheap amino acids are expected to be found in proteins mostly composed of other cheap amino acids significantly more often than they are found in proteins mostly composed of expensive amino acids. As a complement to this test, I also apply a second, statistically independent test. This measures whether the overall amino acid usage in a genome differs from that which would be predicted under mutation and drift in such a way that cheap amino acids are favored and expensive ones avoided. Applying both tests to 43 whole genomes shows that selection on the synthesis cost of amino acids strongly influences amino acid usage throughout the eubacteria, archaea, and eukaryota. Further analysis suggests that subtle differences in amino acid usage between genomes may be due, in part, to relative differences in amino acid costs in different environments.
Materials
Data Acquisition
In March 2004, I downloaded 43 whole genomes: roughly a quarter of all the non-organelle genomes then available in GenBank. My choice of genomes was designed principally to maximise breadth of phylogenetic cover (Fig. 1). In addition, I sought to maximize the comparisons between thermal mesophiles, on the one hand, and thermophiles and hyperthermophiles, on the other. Accordingly, my sample of genomes contains more thermophiles and hyperthermophiles than would a selection drawn at random.
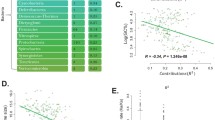
Phylogeny of organisms used in this paper. Organisms appear in the same order as in Table 1. The phylogeny follows that of Madigan et al. (1997), which is based on small subunit rRNA data. Polytomies within the eukaryotes are resolved by placing nematodes with insects in the Ecdysozoa (Aguinaldo et al. 1997) and by treating the microsporidia as the outgroup of the fungi (a position strengthened following the recent sequencing of the genome of Encephalitozoon cuniculi). As the position of Fusobacterium nucleatum is doubtful and unresolved, it is not used in the phylogenetic independent contrasts employed later in this paper.
The whole genome sequences used in this paper are listed below according to their domain and taxonomy. Optimum growth temperatures in degrees Celsius are given (in parentheses) whenever these exceed 40°C. Temperatures were obtained from the online databases of the Deutsche Sammlung von Mikroorganismen und Zellkulturen, http://www.dsmz.de, and the American Type Culture Collection (ATCC), http://www.atcc.org. All sequences were downloaded from GenBank in March 2004, with the exceptions of Candida albicans, downloaded from http://genolist.pasteur.frequency/CandidaDB/ on 26 April 2002, and Neurospora crassa, downloaded from http://www-genome.wi.mit.edu/annotation/fungi/neurospora/ on 13 May 2002.
Bacteria: Thermophilic oxygen-reducing bacteria: Aquifex aeolicus (85); Thermotogales: Thermotoga maritima (70–80); Green non-sulphur bacteria and relatives: Deinococcus radiodurans; Cyanobacteria: Thermosynechococcus elongatus (∼60); Prochlorococcus marinus; Fusobacteria: Fusobacterium nucleatum; Proteobacteria, alpha subdivision: Caulobacter crescentus; Proteobacteria, beta subdivision: Chromobacterium violaceum; Proteobacteria, gamma subdivision: Escherichia coli; Buchnera aphidicola; Proteobacteria, delta subdivision: Bdellovibrio bacteriovorus; Proteobacteria, epsilon subdivision: Helicobacter pylori; Planctomyces and relatives: Chlamydia trachomatis; Pirellula sp.; Spirochetes and relatives; Leptospira interrogans; Treponema pallidum; Green sulphur bacteria: Chlorobium tepidum (45); Bacteroides: Porphyromonas gingivalis; Gram-positives, High G+C: Mycobacterium tuberculosis; Gram-positives, Clostridium and relatives: Clostridium perfringens; Thermoanaerobacter tengcongensis (75); Gram-positives, Mycoplasma and relatives: Mycoplasma pneumoniae; Gram-positives, Bacillus-Lactobacillus-Streptococcus group: Streptococcus pneumoniae; Bacillus subtilis.
Archaea: Crenarchaeota: Aeropyrum pernix (90); Euryarchaeota, Thermococcales: Pyrococcus abyssi (103); Euryarchaeota, Methanococcales: Methanococcus maripaludis: Methanocaldococcus jannaschii (70–85); Euryarchaeota, Methanomicrobacteria and relatives: Thermoplasma volcanium (60); Methanosarcina mazei; Archaeoglobus fulgidus (81).
Eukaryota: Alveolata: Plasmodium falciparum; Viridiplantae: Arabidopsis thaliana; Microsporidia: Encephalitozoon cuniculi; Archiascomycetes: Schizosaccharomyces pombe; Euascomycetes: Neurospora crassa; Hemiascomycetes: Candida albicans; Saccharomyces cerevisiae; Eremothecium gossypii; Deuterostomia: Mus musculus; Homo sapiens; Ecdysozoa: Drosophila melanogaster; Caenorhabditis elegans.
Data Preparation
From each genome, I removed all sequences that were duplicates at the amino acid level. I then prepared a parallel set of genomes from which all proteins under 200 amino acids long had been removed. There are two reasons to suspect that the parallel set might be more reliable. First, short proteins identified by gene detection algorithms are more likely than long ones to be false positives. Second, my analyses involve measuring amino acid frequencies within individual proteins, and sampling noise is louder in short proteins. However, as the greater reliability of the parallel set comes at the expense of discarding some of the data, I performed all analyses on both the full and the parallel genome sets.
Methodology and Results
Gradient Consistency Test for Cost Selection
Procedure
The gradient consistency test measures whether, in a given genome, the relative synthesis costs of the amino acids can explain shifts in amino acid usage in cheap as compared with expensive proteins. This is a three-step procedure. (Note: the synthesis cost of an amino acid is the number of molecules of ATP expended in the biosynthesis of one molecule of that amino acid in Escherichia coli [Craig and Weber 1998].)
Step 1. Calculating the raw shift in usage. For each of the 20 amino acids, I proceeded as follows. (a) I measured the percentage of sites in each protein occupied by the focal amino acid. (b) I calculated the unbiased mean per-residue synthesis cost of each protein—which is to say, I calculated the mean per-residue synthesis cost by only counting those sites not occupied by the focal amino acid. (A failure to exclude the focal amino acid would give a mean cost per residue that is not independent of the occurrence of the focal amino acid.) (c) Having made this pair of calculations for every protein in a genome, I performed a linear regression and estimated the slope. (Slopes are preferable to regression coefficients, as slopes measure the intensity of a shift in usage rather than its consistency, and it is the intensity of the shift that is of interest.)
Figure 2 illustrates this first step for phenylalanine in Saccharomyces cerevisiae.
Step 2. Standardizing the slopes. In order to allow meaningful comparisons between amino acids that are used infrequently and those that are used often, and to compensate for the slight differences in protein mean residue costs that arise when different amino acids are excluded, I standardized each of the 20 slopes as follows:
where y is the mean per residue synthesis cost of all proteins on the relevant 19 amino acid basis, and z is the mean percentage usage of the focal amino acid in all proteins. In the case of phenylalanine in Saccharomyces cerevisiae, the standardized slope is
Step 3. Evaluating the variation in gradients. To evaluate the variation in gradients, I correlated the 20 standardized slopes with the 20 amino acid synthesis costs. I measured the strength of the correlation by Pearson’s r and its statistical significance by a sampled randomization on Kendall’s τ (see Supplementary Information [1] for further details.) A major benefit of using the nonparametric Kendall’s τ to evaluate significance is that this avoids the danger of giving undue weight to outliers, in particular, the three aromatic amino acids, which are far more expensive than other amino acids.
Figure 3 shows the change in usage of amino acids as a function of amino acid synthesis cost in Saccharomyces cerevisiae. The y-axis value for phenylalanine is the one estimated in Step 2, above.
Change in usage of amino acids (with rising protein mean residue cost in Saccharomyces) as a function of amino acid synthesis cost. The 95% confidence intervals of the slope estimates are indicated as vertical lines drawn through the appropriate amino acid—most of these intervals are so small that they are partly obscured by the three-letter amino acid codes.
Results of Gradient Consistency Test
Selection on synthesis cost shapes amino acid usage in all three domains of life
The results for Saccharomyces (shown above) clearly support the cost selection hypothesis: the 20 amino acid synthesis costs provide a consistent explanation of each amino acid’s changing usage along the gradient of protein synthesis cost (Fig. 3; p = 0.0000086 sampled randomization on Kendall’s τ). Moreover, this result remains true even when the analysis is restricted to subsets of proteins that share molecular or cellular function (see Supplementary Information [5].)
More generally, a significant effect of cost selection can be seen in 38 of 43 of the genomes studied (Table 1, column “pval:all”; typical correlations, including that for Homo sapiens, are shown in Fig. 4). Cost selection seems, therefore, to be a pervasive force shaping the amino acid composition of genomes throughout the three domains of life.
Change in usage of amino acids (with rising protein mean residue cost) as a function of amino acid synthesis cost in six representative genomes. The corresponding regression for Saccharomyces cerevisiae is given as Fig. 3. Aeropyrum pernix (d) shows the weakest regression of all 43 genomes: a weak regression is expected, as it is a hyperthermophile.
Mesophiles show a stronger effect than thermophiles
The organisms in the dataset divide into thermal mesophiles, on the one hand, and thermophiles and hyperthermophiles, on the other. Henceforward, I use the term “thermophiles” to refer to both thermophiles and hyperthermophiles; it includes all organisms with optimum growth temperatures of 60°C or higher.
Interestingly, 33 of 34 thermal mesophiles show significant evidence of cost selection (34 of 34 when the analysis is restricted to proteins over 200 amino acids long), but only 5 of 9 thermophiles (Table 2). This difference, which is significant (p = 6.4e−6, one-tailed Mann-Whitney U test), is expected because of differences in amino acid synthesis costs in different environments. Organisms that live in hydrothermal vents are subject to high temperatures, pressures, and reducing environments, all of which alter the Gibbs free energy of formation of amino acids (Amend and Helgeson 1997; Amend and Shock 1998). Thus, an amino acid that is relatively expensive for Saccharomyces to synthesize may be relatively cheap for a hydrothermal vent resident such as Aeropyrum pernix.
The split between mesophiles and thermophiles does not seem to be due simply to a split between archaea (which are disproportionately often thermophilic) and eubacteria, since the split is found within both domains. In both domains the median p-values for cost selection are more significant for mesophiles than for thermophiles (Table 2, with further discussion in Supplementary Information [6]). Moreover, when the entire analysis is repeated but replacing the E. coli amino acid synthesis costs with the synthesis costs estimated for deep sea vent conditions by Amend and Shock (1998), then the pattern in this table reverses: the median eubacterial thermophile correlation is now more positive than the median eubacterial mesophile one and the median archaean thermophile correlation is more positive than the median archaean mesophile one.
Another factor that could, potentially, contribute to thermophiles showing a weaker effect of cost selection is that selection for protein stability at high temperatures may lead to certain amino acids being preferred in proteins (Cambillau and Claverie 2000; Haney et al. 1999; Kreil and Ouzounis 2001; Madigan and Oren 1999; McDonald 2001; McDonald et al. 1999; Menendez-Arias and Argos 1989; Sandberg and Terwilliger 1989; Vogt et al. 1997). Selection for this stability could impose additional functional constraints at sites in thermophile proteins, and therefore these proteins would be less responsive than their mesophile orthologues to cost selection-driven amino acid substitution (Heizer et al. 2006). But although this argument is plausible, it is not obvious how selection for greater thermostability could account for the observation that gradient consistency results are lower in thermophiles. An amino acid that confers greater thermostability should be selected for in all thermophile proteins—regardless, for example, of their expression level. But gradient consistency automatically controls for such blanket selection, as it measures not absolute levels of amino acid usage —all information as to absolute level is erased in the standardisation step—but relative change in amino acid usage within the genome. Thus, although the matter is not definitively settled, differences in synthesis costs—which are predicted to cause relative changes in amino acid usage—seem a more plausible explanation for the lower significance of thermophiles than selection for thermostability per se.
The power of cost selection on mesophiles
The consistency among mesophiles is especially remarkable because the genomes were chosen to give the widest possible phylogenetic cover and because they exhibit considerable variation in mutation pressure: the usage of guanine and cytosine at fourfold degenerate sites varies from 7% to 93% (Table 1). Further analysis confirms that the amino acid usage gradients are not due to mutation pressure; indeed, support for the hypothesis is increased if mutation is taken into account (see Supplementary Information [2]).
Another factor leads to cost selection being underestimated in mesophiles in the foregoing analysis: statistical noise. Each of the regressions in Table 1 is affected by two sources of noise: sampling effects (the finite length of proteins) and interference effects (functional selection is usually far more important than cost in determining the amino acid found at a protein site). We can control for some of this noise by constructing a consensus median mesophile from the data for all 34 mesophiles (Fig. 5). Here, each of the 20 usage slopes (y-axis values in Fig. 5) is the median of the usage slopes of the relevant amino acid among the 34 thermal mesophiles.
Change in usage of amino acids (with rising protein mean residue cost) as a function of amino acid synthesis cost in the consensus median mesophile. In this figure each of the 20 usage slopes (y-axis values) is the median of the usage slopes for the relevant amino acid among the 34 thermal mesophiles.
This consensus mesophile yields a stronger correlation (r = 0.87, p = 0.000098) than 32 of 34 of the individual mesophiles (Table 1). It gives the best overall estimate of the power of cost selection in shaping amino acid usage in thermal mesophiles, and it shows that cost selection is not a trivial force but, rather, one that generates surprisingly strong gradients in amino acid usage: the amino acid that changes the most, tryptophan, increases its usage over the observed range of protein costs by 280%, while the 75th percentile, median, and 25th percentile amino acids change by 90%, 60%, and 20%.
Deviation from Neutrality Test: An Independent Test for Cost Selection
Procedure
The deviation from neutrality test evaluates whether the overall amino acid composition of a genome is consistent with selection pressure for lower cost. This test is statistically independent of the gradient consistency test and, thus, supplies an additional and independent measure of cost selection.
Deviation from neutrality compares the observed amino acid usage with that which would be expected in the absence of selection, the “neutral usage.” If cost selection is sufficiently strong and affects a sufficiently large proportion of proteins in a genome, then the total genomic usage of expensive amino acids should be significantly less than their neutral usage.
Step 1. Obtaining the neutral usage of the 20 amino acids. (a) Eukaryotes and archaea. Neutral usage was estimated for each amino acid from the nucleotide bias at fourfold degenerate sites—sites where all four nucleotides encode the same amino acid—according to the King and Jukes method (King and Jukes 1969; Swire et al. 2005). For example, given nucleotide proportions 0.3 U, 0.2 C, 0.4 A, 0.1 G, the neutral stop codon frequency (codons UAG, UAA, and UGA) is (0.3 × 0.4 × 0.1) + (0.3 × 0.4 × 0.4) + (0.3 × 0.1 × 0.4) = 0.072, and the neutral frequency of cysteine (codons UGU and UGC) is ((0.3 × 0.1 × 0.3) + (0.3 × 0.1 × 0.2))/(1–0.072) = 0.016. (The neutral frequency of stop codons, 0.072, is subtracted from the denominator so as to derive the neutral frequency of cysteine as a function of that of the other amino acids.)
(b) Eubacteria. The King and Jukes method was also used to estimate neutral usage in eubacteria, but this time the method was applied not to fourfold degenerate sites but to intergenic sites, with separate estimates being made for leading and lagging strands, which are under different mutational pressures (Lobry 1996; Francino and Ochman 1997). (None of the results is materially changed if fourfold degenerate sites are used for the eubacteria; however, strand-adjusted intergenic sites are preferable because they give a more accurate neutral estimate, as indicated by the ability to predict the direction of amino acid usage shifts in independent contrasts of genomes [see Table S1 in Supplementary Information (2)].) Strand-adjusted intergenic sites could not be used for eukaryotes or archaea because of the lack of reliable information on the location of origins of replication.
Step 2. Obtaining the observed amino acid usage. Observed usage is the number of times each amino acid is used in the genome divided by the total number of residues, again giving a vector of 20 amino acid usages.
Step 3. Estimating the difference between observed and neutral amino acid usage. The vector of observed usages is divided by the vector of neutral usages, and the resulting vector correlated with the 20-amino acid synthesis costs using Kendall’s τ, with a significant negative correlation being consistent with cost selection.
Results of deviation from neutrality test
All 43 genomes show a negative correlation between cost and (observed_usage/neutral_usage) under Kendall’s τ. The median p-value for the 34 mesophiles is significant, as are the median p-values for the mesophile eubacterium and the mesophile eukaryote (Table 3). The median p-values for the different groups of thermophiles, on the other hand, are never significant. (The difference between all mesophiles and all thermophiles is significant at p = 0.0024 [one-tailed Mann-Whitney U test], that between mesophilic and thermophilic eubacteria at p = 0.018, and that between mesophilic and thermophilic archaea at p = 0.048; for further details see Supplementary Information [6].)
The difference between thermophiles and mesophiles is consistent with the hypothesis, advanced in Mesophiles Show a Stronger Effect Than Thermophiles, above (under Results of Gradient Consistency Test) that because amino acid synthesis costs are different for the two groups, cost selection may have different effects. Repeating the calculations using the synthesis costs that Amend and Shock (1998) estimated for deep sea vent conditions reverses the pattern in the bottom five rows of Table 3: the results for the archaeal and eubacterial thermophiles are significant, whereas those for the mesophiles are not (data not shown).
As in the case of gradient consistency, I also constructed a consensus mesophile to reduce the noise in the expected usage estimates of the individual genomes (Fig. 6). This yields p = 0.0073 under Kendall’s τ—only 5 of 34 individual mesophiles produce more significant correlations.
Correlation between consensus mesophile log(observed amino acid usage/neutral amino acid usage) and amino acid synthesis cost. The y-axis shows the median value for each amino acid among the 34 thermal mesophiles. Compare with Fig. 5.
Note the anomalous position of the three most expensive amino acids, the aromatics tryptophan, phenylalanine, and tyrosine. These appear to be selected against less than would be expected on the basis of their high synthesis costs. Simple neutral explanations of this anomaly in terms either of different mutational effects at the three codon positions or of the proximity of amino acids to stop in the genetic code—which could reduce the substitution rate under mutation pressure—do not work (data not shown), and the anomaly remains mysterious, especially as there is no trace of it under gradient consistency (Fig. 5).
Concordance of the Two Tests
Do gradient consistency and deviation from neutrality reveal the operation of a single force? Two observations suggest they do.
First, despite the disagreement concerning tryptophan, phenylalanine, and tyrosine, the two methods generally concur as to the intensity with which different amino acids respond to cost selection (p = 0.0082, one-tailed Kendall’s τ correlation of the y-axis values in Figs. 5 and 6).
Second, the two methods generally concur as to which genomes show the greater evidence of cost selection. This is revealed by independent contrasts. (Because of the monophyly of life, species cannot simply be treated as independent data points when analyzing correlations between variables. Rather, noncrossing phylogenetically independent contrasts should be drawn: in each contrast the change in the variables between a related pair of species is noted, and the contrasts are then correlated [Burt 1989; Harvey and Pagel 1999]) Independent contrasts show that the two methods generally concur as to which genomes show greater evidence of cost selection (p = 0.0093, one-tailed Kendall’s τ correlation of shiftdvn and shiftgrc in Table 4).
These two observations are independent, giving a Fisher combined probability of p = 0.00080. This consistency argues that the two methods are detecting the same phenomenon.
Selection on the Acquisition Costs of the Components of Amino Acids Probably Modulates Selection on Amino Acid Synthesis Cost
Earlier, I showed that some of the intergenome variation in the apparent strength of cost selection might be explained by the different amino acid synthesis costs for thermophiles versus mesophiles. However, there is another obvious source of potential variation in amino acid costs: the costs of acquiring the components from which amino acids are built. (These costs are not included in the estimation of amino acid synthesis cost, which takes into account only the expenditure of ATP in synthesizing amino acids from components in the cell.)
In principle, intergenome variation in acquisition costs might play an important role in shaping intergenome differences in gradients of amino acid usage. For example, the six amino acids that contain nitrogen atoms in their side-chains—arginine, histidine, lysine, tryptophan, asparagine, and glutamine—might be relatively disfavoured in proteins under strong cost selection in nitrogen-limited organisms. Such disfavor would show itself as positive residuals under gradient consistency for the relevant amino acids and organism—e.g., positive residuals in Fig. 4—the positive residual signifying that the amino acid is behaving as if it were more expensive than expected on the basis of its synthesis cost. Similar expectations may be formed for the two amino acids that contain sulfur in their side-chains—cysteine and methionine—and the seven that contain oxygen—aspartic acid, glutamic acid, tyrosine, asparagine, glutamine, serine, and threonine. (The other two atoms that appear in amino acids, carbon and hydrogen, obviously do not lend themselves to this type of analysis, as they are found in all 20 [hydrogen] or in 19 [carbon] side-chains.)
To test the acquisition cost hypothesis, I once again employ phylogenetically independent contrasts to examine each of the atoms in turn. Thus, for nitrogen, I examine the only two amino acids with multiple side-chain nitrogen atoms—arginine and histidine. If the cost of nitrogen acquisition is shaping intergenome differences in gradients of usage, then the residuals for arginine and histidine should be concordant: that is, the member of the pair of organisms in an independent contrast that shows the more positive residual for arginine should also show the more positive residual for histidine. And, indeed, the two amino acids are concordant in 16 of 21 contrasts (Table 5). This is a higher score than that for any other of the 190 possible amino acid pairs, so an exact randomization test gives a probability of p = 0.0053. The two amino acids with multiple oxygen atoms in their side-chains also give results consistent with the hypothesis, albeit not significantly so: aspartic acid and glutamic acid are concordant in 14 of 21 contrasts (p = 0.10, exact randomization test). The third atom, sulfur, does not conform to the hypothesis: the two amino acids with side-chain sulfur atoms, cysteine and methionine, are concordant in only 9 of 21 contrasts (p = 0.74).
What of those amino acids with single side-chain oxygen or nitrogen atoms? In each case, this involves several amino acids that should be analyzed together, as they are exposed to the same acquisition-cost pressure (lower than that affecting the amino acids with multiple target atoms in their side-chains). The test is similar to that already described for a single pair of amino acids, except that each amino acid is compared with all the others. Thus, in the case of oxygen, where there are five single side-chain oxygen atom amino acids, the result is not a single concordance value but 10 (Table 6a). Sum these 10 concordance values—which gives 124—and compare that with the sum that is obtained on drawing five amino acids at random and subjecting them to the same procedure; 9958 of 10,000 draws of five amino acids yield a total lower than 124, thus the amino acids with a single side-chain oxygen atom show significant concordance, supporting the acquisition cost hypothesis (p = 0.0042, sampled randomization test). An analogous procedure shows that the four amino acids with a single side-chain nitrogen atom also show concordance consistent with the hypothesis, albeit not significantly so (p = 0.11; n = 10,000).
To summarize the evidence for the acquisition cost hypothesis: the test for sulfur shows no tendency, whereas the Fisher combined probability for the two nitrogen tests—multiple and single side-chain atom —is p = 0.0050, and that for the two oxygen tests is p = 0.0037. Both the nitrogen and the oxygen results remain significant at p < 0.012 under sequential Bonferroni correction for three tests, providing modest support in favor of the hypothesis that nutrient availability may affect gradients of amino acid usage in proteins, thereby explaining some of the residual variance remaining from the explanation of those gradients in terms of synthesis cost.
Of course, I have not shown that the organisms in the contrasts that appear to be more limited for a nutrient are so in fact: the data are not available. Quite probably, in many contrasts there is little difference in nutrient limitation, so the tests will have been affected by considerable statistical noise. A more sensitive procedure—once the genomes become available—would be to construct independent contrasts where there are known to be large differences in nutrient limitation: for example, in the case of nitrogen, freshwater versus marine plankton, plants in nitrogen-rich versus nitrogen-poor soils, or nitrogen-fixing versus nonfixing bacteria.
Cysteine Behaves Anomalously
One amino acid, cysteine, conforms to the predictions of the synthesis cost hypothesis considerably more poorly than the other 19. Specifically, under gradient consistency cysteine frequently shows large and inconsistent deviations from the regression of the change in usage of amino acids (with rising protein mean residue cost) on amino acid synthesis cost. This is revealed by the anomalously large standard deviation—taken across the 43 genomes studied—of its residual (Table 7a).
Cysteine is also the only amino acid that is, in some species, the most positive outlier from the gradient consistency regression and, in others, the most negative. Indeed, its residual has a bimodal distribution: in 18 species it falls among the 4 most positive outliers, in 12 species among the 4 most negative, and in only 13 species does it fall among the central 12 (Supplementary Information Table S5).
Some of the anomalous behavior of cysteine is explained by the observation that estimates of the change in usage of cysteine under cost selection are more prone to statistical noise than estimates of the change in usage of the other amino acids. This observation is expected on theoretical grounds. It cannot, however, explain why cysteine is sometimes the most positive and sometimes the most negative outlier: the effects of noise are simply not large enough (Supplementary Information [4]).
About half of the anomalous behavior of cysteine can be attributed to differences between mesophiles and thermophiles, where cysteine is almost always a highly negative outlier (Supplementary Information Table S5). But even within the mesophiles, cysteine shows an anomalously large and outlying standard deviation of its residual (Table 7b). Interestingly, this is not due to phylogenetic constraint: within the mesophiles cysteine shows three phylogenetically independent transitions from being extremely expensive compared with expectation (outlier ranks 1 and 2) to being extremely cheap (ranks 18–20), as can be seen by mapping the last column in Supplementary Information Table S5 onto the phylogeny in Fig. 1.
What might cause these transitions? One possibility is that some taxa are sulfur-limited and therefore disproportionately avoid cysteine in proteins under strong cost selection. However, this hypothesis can be rejected because, as shown in the previous section, the shifts in the residuals of cysteine and those of the other sulfur-containing amino acid methionine are not concordant. Another possibility is that different taxa might use different pathways to synthesize cysteine, expending different amounts of ATP. Cysteine does have more than one biosynthesis pathway (Zubay 1988). But cysteine is not unique in this respect, and the phylogenetic distribution of cysteine biosynthesis pathways does not appear to conform to the distribution of the transitions. The anomalous behavior of cysteine remains, therefore, in part a mystery.
Conclusions
Two independent tests, gradient consistency and deviation from neutrality, agree in showing cost selection to be a pervasive force shaping amino acid usage in all three domains of life. In thermal mesophiles (where the tests were most appropriate), 34 of 34 genomes showed significant evidence of cost selection under gradient consistency, with the consensus mesophile, constructed to reduce statistical noise, showing evidence at p = 0.000098. (Under deviation from neutrality the consensus mesophile showed evidence at p = 0.0073.) There are strong, significant gradients between cheap and expensive proteins in amino acid usage: the median change in usage is 60%; that of the most expensive amino acid, 280%. These effects are not due to mutation (see Supplementary Information [2]).
Gradient consistency and deviation from neutrality are independent tests. Gradient consistency is the more reliable and sensitive test, for two reasons. First, when the amino acid usage expected under neutrality is already cheap, deviation from neutrality cannot readily detect the signature of cost selection (see Supplementary Information [3]). Second, deviation from neutrality is vulnerable to inevitable inaccuracies in the estimation of neutral usage (see Supplementary Information Table S1), whereas gradient consistency is not, as it makes comparisons only between observed usages.
Nevertheless, despite these two disadvantages, there are two reasons for retaining deviation from neutrality as a complement to gradient consistency. First, the fact that two independent methods paint the same picture, both overall and with respect to the differences between thermophiles and mesophiles, greatly strengthens support for the cost selection hypothesis. Second, the deviation from neutrality results confirm that we are indeed seeing selection to reduce protein synthesis cost—for the gradient consistency results considered on their own could have been explained by selection to increase protein synthesis cost, however perverse that might appear.
Two forces appear to modulate the signature of cost selection on different genomes. The first is the general environment. Amino acid usage in thermophiles appears to differ from that of thermal mesophiles owing to different amino acid synthesis costs in the thermophilic environment. The second modulating force is the acquisition cost of the components of amino acids. Where the amino acid usage gradients for a given organism differed from that expected on the basis of amino acid synthesis costs, such differences can potentially be attributed to nutrient limitation increasing the real cost of amino acids containing the limited element.
Finally, we have seen that amino acid synthesis costs estimated from one organism, Escherichia coli, can explain amino acid usage in mesophile organisms in all three domains of life. This suggests that amino acid biosynthesis pathways are highly conserved, and although there are some exceptions—lysine is synthesized in the glutamate pathway in fungi as opposed to the aspartate pathway elsewhere (Zubay 1988)—this is in general doubtless true.
However, even if amino acid synthesis pathways are conserved, a puzzle remains. To wit: not all the mesophiles in this paper synthesize all of their amino acids all of the time. Many are heterotrophs, which acquire several of their amino acids in food—mammals, indeed, can only synthesize about half of their amino acids. This observation raises the question as to why it is possible to observe selection on amino acid synthesis cost in these auxotrophic species: if amino acids are acquired, not synthesized, then a priori, it seems that the amino acid synthesis costs may be poor proxies for the true cost of amino acids to an organism.
There are two mutually compatible resolutions to this conundrum. The first reflects the dependency of heterotrophs upon autotrophs. When a heterotroph eats an autotroph, it obtains amino acids in proportion to their usage in the autotroph’s proteome; this usage will have been shaped by synthesis cost. If the heterotroph is not to have to eat unnecessary amounts of food, the amino acid usage of its proteome must not differ radically from that of the autotroph. Thus amino acid synthesis cost in autotrophs should flow through the food chain to influence amino acid cost for heterotrophs.
The second argument notes that when a heterotroph acquires an amino acid it may either incorporate it into a protein or degrade it to generate ATP and release metabolites for other cellular processes. ATP is generated by inserting the components of degraded amino acids at different points in the glycolytic pathway and citric acid cycle. The number and location of insertion points differs for the different amino acids, and the usable energy released correlates roughly with the different amino acid synthesis costs. Thus, even when an organism does not synthesize an amino acid, it incurs an energetic opportunity cost for incorporating that amino acid into a protein. This opportunity cost is the synthesis cost seen in a distorting mirror.
References
Abbott A (1999) A post-genomic challenge: learning to read patterns of protein synthesis. Nature 402:715–720
Agrafioti I, Swire J, Abbott J, Huntley D, Butcher S, Stumpf MPH (2005) Comparative analysis of the Saccharomyces cerevisiae and Caenorhabditis elegans protein interaction networks. BMC Evol Biol 5(1):article 23
Aguinaldo AMA, Turbeville JM, Linford LS, Rivera MC, Garey JR, Raff RA, Lake JA (1997) Evidence for a clade of nematodes, arthropods and other moulting animals. Nature 387:489–492
Akashi H, Gojobori T (2002) Metabolic efficiency and amino acid composition in the proteomes of Escherichia coli and Bacillus subtilis. Proc Natl Acad Sci USA 99:3695–3700
Amend JP, Helgeson HC (1997) Calculation of the standard molal thermodynamic properties of aqueous biomolecules at elevated temperatures and pressures. Part 1. Amino acids. J Chem Soc 93:1927–1941
Amend JP, Shock EL (1998) Energetics of amino acid synthesis in hydrothermal ecosystems. Science 281:1659–1662
Bennetzen J, Hall B (1982) Codon selection in yeast. J Biol Chem 257:3026–3031
Burt A (1989) Comparative methods using phylogenetically independent contrasts. In: Harvey PH, Partridge L (eds) Oxford surveys in evolutionary biology. Oxford University Press, Oxford, pp 33–53
Cambillau C, Claverie J-M (2000) Structural and genomic correlates of hyperthermostability. J Biol Chem 275:32383–32386
Chanda I, Pan A, Dutta C (2005) Proteome composition in Plasmodium falciparum: higher usage of GC-rich nonsynonymous codons in highly expressed genes. J Mol Evol 61:513–523
Craig CL, Weber RS (1998) Selection costs of amino acid substitutions in ColE1 and ColIa gene clusters harbored by Escherichia coli. Mol Biol Evol 15:774–776
Das S, Ghosh S, Pan A, Dutta C (2005) Compositional variation in bacterial genes and proteins with potential expression level. FEBS Lett 579:5205–5210
Drummond DA, Bloom JD, Adami C, Wilke CO, Arnold FH (2005) Why highly expressed proteins evolve slowly. Proc Natl Acad Sci USA 102:14338–14343
Francino MP, Ochman H (1997) Strand asymmetries in DNA evolution. Trends Genet 13:240–245
Garat B, Musto H (2000) Trends of amino acid usage in the proteins from the unicellular parasite Giardia lamblia. Biochem Biophys Res Commun 279:996–1000
Ghaemmaghami S, Huh W-K, Bower K, Howson RW, Belle A, Dephoure N, O’Shea EK, Weissman JS (2003) Global analysis of protein expression in yeast. Nature 425:737–741
Gu X, Hewett-Emmett D, Li W-H (1998) Directional mutational pressure affects the amino acid composition and hydrophobicity of proteins in bacteria. Genetica 103:383–391
Haney PJ, Badger JH, Buldak GL, Reich CI, Woese CR, Olsen GJ (1999) Thermal adaptation analyzed by comparison of protein sequences from mesophilic and extremely thermophilic Methanococcus species. Proc Natl Acad Sci USA 96:3578–3583
Harvey P, Pagel M (1991) The comparative method in evolutionary biology. Oxford University Press, Oxford
Heizer EMJ, Raiford DW, Raymer ML, Doom TE, Miller RV, Krane DE (2006) Amino acid cost and codon-usage biases in 6 prokaryotic genomes: a whole-genome analysis. Mol Biol Evol 23:1670–1680
Ikemura T (1982) Correlation between the abundance of yeast transfer RNAs and the occurrence of the respective codons in protein genes. J Mol Biol 158:573–597
Jansen R, Gerstein M (2000) Analysis of the yeast transcriptome with structural and functional categories: characterizing highly expressed proteins. Nucleic Acids Res 28:1481–1488
King JL, Jukes TH (1969) Non-Darwinian evolution. Science 164:788–798
Kreil DP, Ouzounis CA (2001) Identification of thermophilic species by the amino acid compositions deduced from their genomes. Nucleic Acids Res 29:1608–1615
Lobry JR (1996) Asymmetric substitution patterns in the two DNA strands of bacteria. Mol Biol Evol 13:660–665
Lobry JR (1997) Influence of genomic G + C content on average amino-acid composition of proteins from 59 bacterial species. Gene 205:309–316
Lobry JR, Gautier C (1994) Hydrophobicity, expressivity and aromaticity are the major trends of amino-acid usage in 999 Escherichia-coli chromosome-encoded genes. Nucleic Acids Res 22:3174–3180
Madigan MT, Oren A (1999) Thermophilic and halophilic extremophiles. Curr Opin Microbiol 2:265–269
Madigan MT, Martinko JM, Parker J (1997) Biol of microorganisms. Prentice-Hall, London
McDonald JH (2001) Patterns of temperature adaptation in proteins from the bacteria Deinococcus radiodurans and Thermus thermophilus. Mol Biol Evol 18:741–749
McDonald JH, Grasso AM, Rejto LK (1999) Patterns of temperature adaptation in proteins from Methanococcus and Bacillus. Mol Biol Evol 16:1785–1790
Menendez-Arias L, Argos P (1989) Engineering protein thermal-stability—sequence statistics point to residue substitutions in alpha-helices. J Mol Biol 206:397–405
Neidhardt FC, Ingraham JL, Schaechter M (1990) Physiology of the bacterial cell: a molecular approach. Sinauer, Sunderland, MA
Nishizawa M, Nishizawa K (1998) Biased usages of arginines and lysines in proteins are correlated with local-scale fluctuations of the G + C content of DNA sequences. J Mol Evol 47:385–393
Palacios C, Wernegreen JJ (2002) A strong effect of AT mutational bias on amino acid usage in Buchnera is mitigated at high-expression genes. Mol Biol Evol 19:1575–1584
Peixoto L, Fernandez V, Musto H (2004) The effect of expression levels on codon usage in Plasmodium falciparum. Parasitology 128:245–251
Richmond RC (1970) Non-Darwinian evolution: a critique. Nature 225:1025–1028
Sandberg WS, Terwilliger TC (1989) Influence of interior packing and hydrophobicity on the stability of a protein. Science 245:54–56
Schaber J, Rispe C, Wernegreen J, Buness A, Delmotte F, Silva FJ, Moya A (2005) Gene expression levels influence amino acid usage and evolutionary rates in endosymbiotic bacteria. Gene 352:109–117
Seligmann H (2003) Cost-minimization of amino acid usage. J Mol Evol 56:151–161
Sharp P, Li W-H (1987) The rate of synonymous substitution in enterobacterial genes is inversely related to codon usage bias. Mol Biol Evol 4:222–230
Singer GAC, Hickey DA (2000) Nucleotide bias causes a genomewide bias in the amino acid composition of proteins. Mol Biol Evol 17:1581–1588
Sorensen MA, Pedersen S (1991) Absolute in vivo translation rates of individual codons in Escherichia coli—the 2 Glutamic acid codons GAA and GAG are translated with a threefold difference in rate. J Mol Biol 222:265–280
Sorensen MA, Kurland CG, Pedersen S (1989) Codon usage determines translation rate in Escherichia coli. J Mol Biol 207:365–377
Suckow J, Markiewicz P, Kleina LG, Miller J, Kisters-Woike B, Muller-Hill B (1996) Genetic studies of the lac repressor XV: 4000 single amino acid substitutions and analysis of the resulting phenotypes on the basis of the protein structure. J Mol Biol 261:509–523
Sueoka N (1961) Correlation between base composition of DNA and the amino acid composition of protein. Proc Natl Acad Sci USA 47:1141–1149
Swire J (2003) Selection on cost as a driver of molecular evolution. PhD thesis, Imperial College London
Swire J, Judson OP, Burt A (2005) Mitochondrial genetic codes evolve to match amino acid requirements of proteins. J Mol Evol 60:128–139
Urrutia AO, Hurst LD (2003) The signature of selection mediated by expression on human genes. Genome Res 13:2260–2264
Vogt G, Woell S, Argos P (1997) Protein thermal stability, hydrogen bonds, and ion pairs. J Mol Biol 269:631–643
Wagner A (2005) Energy constraints on the evolution of gene expression. Mol Biol Evol 22:1365–1374
Zavala A, Naya H, Romero H, Musto H (2002) Trends in codon and amino acid usage in Thermotoga maritima. J Mol Evol 54:563–568
Zubay G (1988) Biochemistry. Macmillan, London
Acknowledgments
Many thanks to Austin Burt, Olivia Judson, Eric de Silva, Michael Stumpf, and two anonymous reviewers for helpful comments, criticisms, and suggestions.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Reviewing Editor: Hector Musto]
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Swire, J. Selection on Synthesis Cost Affects Interprotein Amino Acid Usage in All Three Domains of Life. J Mol Evol 64, 558–571 (2007). https://doi.org/10.1007/s00239-006-0206-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-006-0206-8