Abstract
Analysis of cDNAs from Macrovipera lebetina transmediterranea (Mlt) and Echis ocellatus (Eo) venom gland libraries encoding disintegrins argued strongly for a common ancestry of the messengers of short disintegrins and those for precursors of dimeric disintegrin chains. We now report the sequence analysis of disintegrin-coding genes from these two vipers. Genomic DNAs for dimeric disintegrin subunits Ml_G1 and Ml_G2 (Mlt) and Eo_D3 (Eo) contain single 1-kb introns exhibiting the 5′-GTAAG (donor)/3′-AG (acceptor) consensus intron splicing signature. On the other hand, the short RTS-disintegrins Ml_G3 (Mlt) and Eo_RTS (Eo) and the short RGD-disintegrin ocellatusin (Eo) are transcribed from intronless genomic DNA sequences, indicating that the evolutionary pathway leading to the emergence of short disintegrins involved the removal of all intronic sequences. The insertion position of the intron within Ml_G1, Ml_G2, and Eo_D3 is conserved in the genes for vertebrate ADAM (A disintegrin and metalloproteinase) protein disintegrin-like domains and within the gene for the medium-size snake disintegrins halystatins 2 and 3. However, a comparative analysis of currently available disintegrin(-like) genes outlines the view that a minimization of both the gene organization and the protein structure underlies the evolution of the snake venom disintegrin family.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
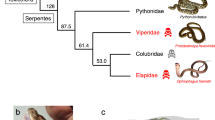
Venom represents a key innovation in ophidian evolution that allowed advanced snakes to transition from a mechanical (constriction) to a chemical (venom) means of subduing and digesting prey larger than themselves. Venom toxins likely evolved from endogenous proteins with normal physiological functions that were recruited into the venom proteome before the radiation of the advanced snakes at the base of the Colubroidea radiation (Fry and Wüster 2004; Fry 2005; Fry et al. 2006). The superfamily Colubroidea comprises >80% of the approximately 2900 species of snake currently described (Vidal et al. 2002). Within this taxon, venoms from the Viperinae (vipers) and Crotalinae (pitvipers) subfamilies of Viperidae snakes contain proteins that interfere with the coagulation cascade, the normal hemostatic system, and tissue repair, and human envenomations are often characterized by clotting disorders, hypofibrinogenemia, and local tissue necrosis (Markland 1998; Fox and Serrano 2005a). Despite the fact that viperid venoms are complex mixtures of protein components (Fox et al. 2006), venom proteins comprise only a few major protein families, including enzymes (serine proteinases, Zn2+-metalloproteases, L-amino acid oxidase, group II PLA2) and proteins without enzymatic activity (disintegrins, C-type lectins, natriuretic peptides, myotoxins, CRISP toxins, nerve and vascular endothelium growth factors, cystatin, and Kunitz-type protease inhibitors) (Fry and Wüster 2004; Fry 2005; Markland 1998; Fox et al. 2006; Juárez et al. 2004; Bazaa et al. 2005). Notably, most venom toxins are extensively cross-linked by disulfide bonds and have flourished into functionally diverse, toxin multigene families that exhibit interfamily, intergenus, interspecies, and intraspecific variability. The existence in the same venom of functionally diverse isoforms of the same protein family reflects accelerated Darwinian evolution (Moura da Silva et al. 1996; Ménez 2002; Tani et al. 2002; Ohno et al. 2003). The evolutionary pressure acting to promote high levels of variation in venom proteins may be part of a predator-prey arms race that allows the snake to adapt to a variety of different prey, each of which is most efficiently subdued with a different venom formulation (Daltry et al. 1996). This evolutionary pattern parallels the “birth-and-death” model of protein evolution proposed to underpin the evolution of resistance genes in plants (Michelmore and Meyers 1998), the vertebrate´s adaptative immune responses to protect the organism from a wide range of foreign antigens (Nei et al. 1997; Nei and Rooney 2005), and the evolution of elapid three-finger toxins (Fry et al. 2003). However, details of the molecular events leading to snake venom toxin diversification remain unclear.
Venom Zn2+-dependent metalloproteinases (SVMPs) represent Serpentes-specific hemorrhagic toxins derived from cellular ADAM (A disintegrin and metalloproteinase) proteins (Fry et al. 2006; Moura da Silva et al. 1996; Calvete et al. 2003). Snake venom hemorrhagins have been classified according to their domain structure (Jia et al. 1996; Lu et al. 2005; Fox and Serrano 2005b). The PIII class comprises the closest homologues of cellular ADAMs, which are large multidomain toxins (60–100 kDa) built up by an N-terminal metalloproteinase domain and C-terminal disintegrin-like and cysteine-rich domains. The class PII metalloproteinases (30–60 kDa) contain a disintegrin domain at the carboxyl terminus of the metalloproteinase domain. PI metalloproteinases (20–30 kDa) are single-domain proteins. Disintegrins, a broad family of small (40–100 amino acids), cysteine-rich polypeptides isolated from venoms of vipers and rattlesnakes (Calvete et al. 2005), are released in viper venoms by proteolytic processing of PII SVMP precursors (Shimokawa et al. 1996) or synthesized from short-coding mRNAs (Okuda et al. 2002), and selectively block the function of cell surface adhesive receptors of the integrin family (Calvete et al. 2005; Sanz et al. 2006). Currently, disintegrins can be classified according to their length and number of disulfide bonds (Calvete et al. 2003). The first group includes short disintegrins, composed of 41–51 residues and four disulfide bonds. The second group is formed by the medium-sized disintegrins, which contain about 70 amino acids and six cystine bonds. The third group includes long disintegrins, with an ∼84-residue polypeptide cross-linked by seven disulfide bridges. The fourth group is composed of homo- and heterodimers. Dimeric disintegrins contain subunits of about 67 residues with 10 cysteines involved in the formation of four intrachain disulfides and two interchain cystine linkages (Calvete et al. 2000; Bilgrami et al. 2004, 2005). Like many other venom toxins, the integrin inhibitory activity of disintegrins depends on the appropriate pairing of cysteines, which determines the conformation of the inhibitory loop that harbors an active tripeptide located at the apex of a mobile loop protruding 14–17 Å from the protein core (Calvete et al. 2005; Monleón et al. 2003, 2005). The current view is that functional diversification among disintegrins has been achieved during ophidian evolution by amino acid substitutions within the active loop, whereas structural diversification was driven through a disulfide bond engineering mechanism involving the selective loss of pairs of cysteine residues engaged in the formation of disulfide bonds (Calvete et al. 2003). The great sequence and structural diversity exhibited by the different subfamilies strongly suggests that disintegrins, like toxins from other venoms (Duda and Palumbi 1999; Kordis et al. 2002; Ohno et al. 2002), have evolved rapidly by adaptative evolution.
Our earlier studies on venom gland cDNAs encoding disintegrins of Cerastes vipera, Macrovipera lebetina transmediterranea (Sanz et al. 2006), and Echis ocellatus (Juárez et al. 2006a) provided compelling data indicating a common ancestry of the messengers coding for precursors of dimeric disintegrin chains and short disintegrins. In this study we sought to investigate the molecular mechanism underlying the structural diversification of these disintegrins through analysis of the genomic organization of their genes.
Materials and Methods
Extraction of Genomic DNA
Genomic DNA was extracted from fresh tissues of M. l. transmediterranea captured in the rocky mountains of northern Tunisia and kept in captivity at the serpentarium of the Institute Pasteur de Tunis (Tunisia) until sacrificed, and from fresh liver of E. ocellatus (Kaltungo, Nigeria), of different ages and of both sexes, maintained at the herpetarium of the Liverpool School of Tropical Medicine. The M. l. transmediterranea tissues were homogenized in 400 μl of sterile salt buffer (0.4 M NaCl, 110 mM Tris-HCl, pH 8.0, containing 2 mM EDTA) using a Polytron tissue homogenizer for 10–15 s. Echis ocellatus liver was ground to a fine powder under liquid nitrogen and the genomic DNA extracted using a Roche DNA isolation kit for cells and tissue containing sodium dodecyl sulfate (SDS; 2% final concentration) and proteinase K (400 μg/ml final concentration). The homogenates were incubated at 55°C overnight. Thereafter, 300 μl of 6 M NaCl (NaCl-saturated H2O) was added to each sample, and the mixture was vortexed for 30 s at maximum speed and centrifuged for 30 min at 10,000g. An equal volume of isopropanol was added to each supernatant, and the sample mixed, incubated at −20°C for 1 h, and centrifuged for 20 min at 4°C and 10,000g. The resulting pellets were washed with 70% ethanol, dried, and, finally, resuspended in 300–500 μl sterile distilled H2O.
DNA Amplification and Sequencing
Disintegrin-encoding DNAs from M. l. transmediterranea were amplified by PCR from genomic DNAs using the following pairs of primers, whose sequences are listed in Table 1. For Ml_G1 and Ml_G2, the forward primer was F1, corresponding to the nucleotide sequence coding for the N-terminal amino acid sequence (MNSANPC) of dimeric disintegrin ML-(2,8,15) (SwissProt/TrEMBL accession code AM114016) (Sanz et al. 2006). The reverse primer was R1. For Ml_G3, the forward primer was F2, which contains the sequence coding for the conserved first six residues of the mature short disintegrins jerdostatin (Protobothrops jerdoni) (Sanz et al. 2005) (SwissProt/TrEMBL accession code AY262730), CV-short (Cerastes vipera) (Q3BK17) (Sanz et al. 2006), and lebestatin (Q3BK14) (M. l. transmediterranea) (Sanz et al. 2006) and the CCATGG NcoI restriction site. The reverse primer was R2, which includes a STOP codon (TTA), the CTCGAG restriction site for XhoI, and the last six C-terminal residues of jerdostatin/CV-short/lebestatin. The Touchdown 60°C/50°C PCR protocol included an initial denaturation step at 95°C for 10 min followed by 4 cycles of denaturation (30 s at 94°C), annealing (30 s at 60°C), and extension (120 s at 72°C); 21 cycles starting with the above conditions and, in subsequent cycles, decreasing the annealing temperature by 0.5°C (reaching 50°C in cycle 21); 10 cycles of denaturation (30 s at 94°C), annealing (30 s at 50°C), and extension (120 s at 72°C); and a final extension for 10 min at 72°C.
Genomic DNA fragments of E. ocellatus encoding disintegrins were amplified using primers listed in Table 1. The dimeric disintegrin (Eo_D3) was amplified using the forward primer F3 and the reverse primer R3 complementary to the highly conserved open reading frame (ORF). Amplification of the genomic DNA coding for ocellatusin precursor (Eo_C3) was achieved using the primer F4 designed from the N-terminal of the disintegrin domain and the reverse primer R3. Amplification of the short RTS-containing disintegrin (Eo_RTS) was achieved with the primers F5 (forward) and R5 (reverse). PCR amplification of the E. ocellatus genomic DNAs was performed in a 25-μl volume containing 0.34 μg of genomic DNA, dNTP (each at 10 mM), 0.6 μl of 5′ and 3′ primers (10 μM each), 0.25 μl of Ampli Taq Gold polymerase, and buffer provided by the supplier (Roche). PCR amplification was performed using the Touchdown 60°C/50°C PCR protocol as above. The resulting PCR products were directly ligated into a pTOPO vector using the TA cloning kit (Invitrogen) for sequencing.
PCR-amplified genomic DNA fragments from M. l. transmediterranea were separated by 1% agarose gel electrophoresis, purified using the Perfect Pre Gel Clean Up kit (Eppendorf, Hamburg, Germany), and cloned in a pGEM-T vector (Promega), which was then used to transform Escherichia coli DH5α cells (Novagen, Madison, WI, USA) by electroporation using an Eppendorf 2510 electroporator following the manufacturer’s instructions. Positive clones, selected by growing the transformed cells in Luria broth (LB) medium containing 10 μg/ml ampicilin, were confirmed by PCR amplification using the above primers and the sequences of the inserts were subjected to sequencing on an Applied Biosystems Model 377 DNA sequencing system.
DNA Sequence Analysis
The nucleotide sequences of Ml_G1, Ml_G2, Ml_G3, Eo_D3, and Eo_C3 were compared to all sequences in the GenBank database (http://www.ncbi.nlm.nih.gov/blast/) (GenBank + EMBL + DDBJ + PDB sequences) using the BLASTN 2.2.13 program (Altschul et al. 1997). Multiple sequence alignment was done with CLUSTALW (Thompson et al. 1994) through the online facility of the Kyoto University Bioinformatics Center (http://clustalw.genome.jp) using default parameters.
Phylogenetic Reconstruction
We have used three methods for phylogeny reconstruction and assessment of the inferred evolutionary relationships based on amino acid sequences. First, we used maximum-likelihood inference as implemented in MEGA 3.1 (Kumar et al. 2001), which optimizes the likelihood function by simultaneously adjusting the topology and branch lengths. We used the JTT (Jones, Taylor, and Thornton 1992) substitution matrix, with a discrete gamma function with eight categories plus invariant sites to account for substitution rate heterogeneity among sites and empirically estimated amino acid frequencies. Second, we employed Bayesian inference with MrBayes 3.1.2. (Ronquist and Huelsenbeck 2003) using the same substitution model (JTT) as in MEGA. The method uses Markov chain Monte Carlo methods to generate posterior probabilities for each clade represented in the tree. The analysis was performed by running 106 generations in four chains, using burn-in at 50,000 generations, and saving every 100th tree. The final tree was rooted using the branch of the human ADAM 7 and 28 molecules as the outgroup.
Accession Numbers
The DNA sequences of clones Ml_G1, Ml_G2, and Ml_G3 from M. l. transmediterranea and Eo_D3, Eo_C3, and Eo_RTS from E. ocellatus are accessible from the SwissProt/TrEMBL data bank (http://us.expasy.org) under accession codes AM261811, AM261812, AM261813, AM286800, AM286799, and AM286798, respectively.
Results and Discussion
Genomic Organization of Dimeric Disintegrin Subunits
Disintegrin-coding DNAs were PCR-amplified using genomic DNAs as template and a pair of primers complementary for the nucleotide sequences coding for the N- and C-terminal amino acid sequences of dimeric disintegrin subunits from M. l. transmediterranea (Mlt) (Sanz et al. 2006) and E. ocellatus (Eo) (Juárez et al. 2006a). The 1.2-kb sequences of two such Mlt genes, termed Ml_G1 and Ml_G2 (Fig. 1A), revealed that their ORFs were each interrupted by single introns of 1052 and 1043 bases, respectively (Figs. 2A and 2B). The 1.7-kb sequence of Echis ocellatus Eo_D3 (Fig. 1C) coded for a dimeric disintegrin subunit and also contained a single 1006-nucleotide intron (Fig. 3A). The exon-intron-exon structure of the disintegrin domain would support the hypothesis of exon shuffling as a putative mechanism for structural diversification of toxins suggested by Junqueira-de-Azevedo and colleagues (2006) in a recent report on the transcriptome analysis of Lachesis muta. The medium-sized disintegrins halystatins 2 and 3 from Gloydius halys are produced by alternative splicing of a single gene (GenBank accession code D28871). Addressing this point requires detailed genomic and transcriptomic comparative analyses.
PCR amplification of disintegrin genes of M. l. transmediterranea. One percent agarose gel electrophoretic analysis of the PCR-amplified genomic DNAs from M. l. transmediterranea (A) Ml_G1 (lane G1) and Ml_G2 (lane G2) and (B) Ml_G3 (lane G3). C, D PCR-amplified genomic DNAs from E. ocellatus coding for dimeric disintegrin subunit Eo_D3, short RGD-disintegrin (Eo_C3), and short RTS-disintegrin (Eo_RTS). STD, the “1 kb plus” DNA ladder from Invitrogen was used as a reference for estimating DNA fragment sizes.
Disintegrin genes of M. l. transmediterranea. Nucleotide sequences of the genomic DNA forMl_G1 (A), Ml_G2 (B), and Ml_G3 (C). The deduced amino acid sequences of exons are shown in the one-letter code. The nucleotide sequences complementary to the primers used for PCR amplification are underlined. In A and B, the 5′-GTAAG (donor)/3′-AG (acceptor) consensus intron splice site signature is in italics and double-underlined, and the intronic sequences overlapping in the 5′→3′ and the 3′→5′ sequencing directions are underlined. The VGD (A), MLD (B), and RTS (C) integrin binding motifs are depicted on a gray background.
Disintegrin genes of E. ocellatus. Nucleotide sequences of the genomic DNA for Eo_D3 (A), Eo_C3 (B), and Eo_RTS (C). The deduced amino acid sequences of exons are shown in the one-letter code and in boldface. The nucleotide sequences complementary to those of the primers used for PCR amplification are underlined. In A the 5′-GTAAG (donor)/3′-AG (acceptor) consensus intron splice site signature is in italics and double-underlined. Intronic sequences overlapping in the 5′→3′ and the 3→5′ sequencing directions are underlined. The MLD (A), RGD (B), and RTS (C) integrin binding motifs are depicted on a gray background.
The intragenic regions of Ml_G1, Ml_G2, and Eo_D3 exhibit ∼88% sequence identity among themselves and display approximately 90% sequence identity with the full-length (999-base) intron 2 of the gene for prepro-halystatins 2 and 3. The exon-intron organization of prepro-halystatins 2 and 3 and the partial intron sequences (139 nucleotides) of a disintegrin gene from several Agkistrodon contortrix subspecies (Soto et al. 2006), which show 92% identity with the 5′ regions of the M. l. transmediterranea and the E. ocellatus introns (comprising nucleotides 12–155) are the only reported sequences of disintegrin genes (Fig. 4). The partial exon sequences of the Agkistrodon contortrix isogenes show the highest similarity with cDNA sequences for the α-subunit of acostatin and for the α- and β-subunits of piscivostatin, two dimeric disintegrins from A. c. contortrix and A. piscivorus piscivorus, respectively (Okuda et al. 2002).
Comparison of intronic sequences. Alignment of the nucleotide sequences of the introns of Ml_G1, Ml_G2, and Eo_D3 with that of intron 2 of prepro-halystatins 2 and 3 from the Chinese water mocassin (G. halys pallas) (GenBank accession code D28871) and the partial intronic sequence of a disintegrin gene from several Agkistrodon contortrix subspecies (Soto et al. 2006). Identical nucleotides are labeled with an asterisk below the multiple alignment. The 5′-GTAAG (donor)/3′-AG (acceptor) consensus intron splice site signature is in italics and double-underlined.
The M. l. transmediterranea, E. ocellatus, and G. halys introns all display the 5′-GTAAG (donor)/3′-AG (acceptor) consensus intron splice site signature (Vicens and Cech 2006) (Figs. 2A and B, 3A, and 4), whereas the introns from the Agkistrodon contortrix subspecies genes lack the first 11 nucleotides, which are absolutely conserved in the 5′ end of the Macrovipera and the Gloydius introns (Fig. 4). Nonetheless, all these introns share equivalent topology within their respective disintegrin genes (Fig. 5). On the other hand, comparison of the exon-intron organization of the gene for the medium-sized disintegrins halystatins 2 and 3 and the genomic DNA coding for the dimeric disintegrin subunits Ml_G1 and Ml_G2 revealed that intron 1 of the former (G. halys) disintegrins is absent from the M. l. transmediterranea G1 and G2 genes (Fig. 5). Accumulating evidence suggests that the subunits of dimeric disintegrins arose from duplicated medium-sized disintegrin genes (Calvete et al. 2003). Deletions and mutations involving, among others, the codons of the first two cysteine residues, yielded polypeptides with 10 cysteines. Cys-6 and Cys-7, which in monomeric medium-sized disintegrins are disulfide-bonded to the lost cysteines, form interchain disulfide bonds with homologous cysteines from another 10-cysteine-containing disintegrin chain (Calvete et al. 2000; Bilgrami et al. 2004, 2005), giving rise to homo- and heterodimers (Marcinkiewicz et al. 1999a,b; Zhou et al. 2000; Gasmi et al. 2001; Calvete et al. 2002). Beside mutations affecting the exon-coded sequences of disintegrin genes, sequence analysis of dimeric disintegrin subunit genes from different vipers indicates that the loss of intron 1 may represent a conserved feature in the transition from a medium-sized to a subunit of dimeric disintegrin.
Comparison of amino acid sequences and organization of disintegrin genes. Alignment of the genomic DNA-deduced amino acid sequences of dimeric disintegrin subunits Ml_G1, Ml_G2, and Eo_D3 and the short disintegrins Ml_G3 and Eo_C3 from M. l. transmediterranea (“Ml_X”) and E. ocellatus (“Eo_X”) with representative disintegrin-like domains from ADAM molecules of other vertebrate taxa (hu, human; ck, chicken; mm, mouse; zf, zebra fish) and snake venom disintegrins whose gene organization has been reported: Halys-2 and Halys-3, disintegrin domains of halystatins 2 and 3 from the Chinese water mocassin (Gloydius halys pallas; GenBank accession code D28871); Contortrix, partial sequence of a disintegrin gene from several Agkistrodon contortrix subspecies (Soto et al. 2006). The amino acid residues preceding the conserved insertion position of introns 1, 2, and 3 (marked with arrows) in the respective disintegrin/disintegrin-like genes are underlined and boxed. Cysteine residues are depicted on a gray background. Accession codes: hu_ADAM-11 (NC_000017.9), hu_ADAM-19 (NC_000005.8), hu_ADAM-22 (NC_000007.12), ck_ADAM-22 (http://www.ensembl.org; ENSGALP00000014587), mm_ADAM-19 (AK147217), zf_ADAM-22 (NC_007128.1).
Intronless Genomic Sequences Coding for Short Disintegrins
A 133-bp genomic DNA fragment from M. l. transmediterranea (Ml_G3) was amplified (Fig. 1B) using primers complementary to conserved regions of the short disintegrins lebestatin (M. l. transmediterranea), CV-short (Cerastes vipera) (Sanz et al. 2006), and jerdostatin (Protobothrops jerdoni) (Sanz et al. 2005). This DNA fragment corresponded to a small intronless genomic sequence coding for a full-length RTS-disintegrin (Fig. 2C) identical to mature CV-short and jerdostatin. Ml-G3 and lebestatin show 89% amino acid sequence identity. An identical RTS-disintegrin sequence (Eo_RTS) was amplified from genomic DNA of E. ocellatus using a different set of primers (Figs. 1D and 3C). This striking finding was consistently confirmed in more than 20 DNA sequence experiments from independent clones. Thus, using the same set of primers, we (Sanz et al., in preparation) have amplified intronless jerdostatin-like genes from genomic DNAs of a number of species classified into very diverse genera from the subfamilies Viperinae (pitless vipers: Daboia russelli, Bitis arietans) and Crotalinae (pit vipers: P. jerdoni, G. halys, Sistrurus catenatus catenatus, Crotalus viridis, P. mucrosquamatus) (http://www.embl-heidelberg.de/∼uetz/families/Viperidae.html).
The short RGD-disintegrin ocellatusin (E. ocellatus) is also transcribed from an intronless genomic 258-bp DNA sequence (Eo_C3) (Figs. 1D and 3B). The nucleotide sequence of Eo_C3 is identical to the region comprising nucleotides 1228–1485 of the long precursor (PII metalloprotease) of ocellatusin (clone Eo_00006; Juárez et al. 2006a). Analysis of cDNAs from M. l. transmediterranea and E. ocellatus venom gland libraries encoding disintegrins argued strongly for a common ancestry of the messengers of short disintegrins and those for precursors of dimeric disintegrin chains (Juárez et al. 2006a; Sanz et al. 2006). In line with this evidence, our current findings indicate that the evolutionary pathway leading to the emergence of short disintegrins involved the removal of all intronic sequences (Fig. 6).
Minimization of protein structure and gene organization. A Scheme of the domain organization, disulfide bond patterns, and proposed evolutionary pathway from the PIII disintegrin/cysteine-rich proteins to short disintegrins (Calvete et al. 2003, 2005; Calvete 2005; Juárez et al. 2006a, b). Structural features (the cysteine-rich domain of PIII disintegrin-like molecules, and class-specific disulfides) lost along the disintegrin diversification pathway are highlighted with thick lines. B Scheme of the conserved exon-intron organization of known disintegrin-like domains of vertebrate ADAM proteins and of the medium-sized disintegrins halystatins 2 and 3, the dimeric disintegrin subunits Ml_G1, Ml_G2, and Eo_D3, and the short-disintegrins Ml_G3 and Eo_C3. GT...AG, denotes the 5′-GTAAG (donor)/3′-AG (acceptor) consensus intron splice site signature conserved in all known disintegrin-like and disintegrin genomic DNAs. The concept of minimization of both the protein and the gene structures along the diversification pathway of disintegrins is highlighted.
Loss of Introns Along the Diversification Pathway of Disintegrins
A survey of complete genome sequences (http://www.ensembl.org;http://www.ncbi.nlm.nih.gov/BLAST) reveal- ed that the genes for ADAM proteins from a number of species, including zebra fish (Danio rerio), mouse (Mus musculus), rat (Rattus norvegicus), chicken (Gallus gallus), and human (Homo sapiens), contain three topologically conserved introns within their disintegrin-like domains (Fig. 5). ADAMs represent the closest homologues of PIII snake venom proteins, which in turn are likely to represent the ancestors of the PII disintegrin family (Moura da Silva et al. 1996; Calvete et al. 2003, 2005; Calvete 2005). Noteworthily, all known ADAM and disintegrin introns share the 5′-GTAAG (donor)/3′-AG (acceptor) consensus splice site signature (Figs. 5 and 6). Moreover, the insertion sites of introns 1 and 2 are conserved in the genes for ADAMs and for the medium-sized snake disintegrins halystatins 2 and 3, and in addition, the insertion position of intron 2 is also conserved in the genes for dimeric disintegrin subunits Ml_G1, Ml_G2, and Eo_D3 (Figs. 5 and 6). On the other hand, intron 3 has been removed from the halystatin gene; introns 1 and 3 are not present in the genes for Ml_G1, Ml_G2, and Eo_D3; and the short disintegrins Ml_G3 and Eo_C3 (ocellatusin) are encoded by intronless genes (Figs. 5 and 6).
Phylogenetic relationships among disintegrins inferred using maximum likelihood and Bayesian analysis (Fig. 7) indicate that the disintegrin-like domains of PIII snake venom metalloproteinases and those of cellular ADAMs are the closest homologues, and are in line with the view that PIII SVMPs diverged from an ADAM precursor (Moura da Silva et al. 1996). PIII SVMPs are modular proteins containing N-terminal Zn2+-metalloproteinase, disintegrin-like, and C-terminal cysteine-rich domains. Gene duplication and removal of the region encoding the cysteine-rich domain generated the PII SVMP genes (Moura da Silva et al. 1996; Juárez et al. 2006b). The phylogenetic tree inferred for the disintegrin family (Fig. 7) also supports our hypothesis that the structural diversity of PII disintegrins has been achieved during evolution through mutations causing the loss of pairs of cysteine residues engaged in the formation of disulfide bonds, generating successively the precursors of long, medium-sized, dimeric, and short disintegrins as depicted schematically in Fig. 6A (Calvete et al. 2003; Juárez et al. 2006a). Our current results showing the sequential loss of introns along the diversification pathway of disintegrins provide additional evidence in favor of our hypothesis that a minimization of both the gene organization and the protein structure underpins the evolution of the snake venom disintegrin family (Fig. 6). Challenges for future investigations are to identify the molecular machinery responsible for, and to dissect the individual steps along, the transformation pathway of disintegrins from ADAM PIII disintegrin-like domains to snake venom short PII disintegrins.
Phylogenetic tree for the disintegrin family. Evolutionary relationships among disintegrins were inferred using maximum-likelihood and Bayesian methods. The analyses were based on the amino acid sequences of PIII disintegrin-like and PII long, medium-sized, dimeric, and short disintegrin domains displayed in Fig. 3 of Calvete et al. (2003). When available, database accession numbers are indicated. The sequences of the short disintegrins multisquamatin, pyramidinA, and leucogastin are from Okuda et al. (2001). The primary structures of the dimeric disintegrin subunits EMS11A, VLO4, VLO5A, VLO5B, VA6, VB7A, and VB7B are reported by Calvete et al. (2003). Bilitoxin-1 is from Nikai et al. (2000), and the sequence of the disintegrin-like domain of graminelysin is from Wu et al. (2001). For rooting the tree, the branch of disintegrin-like domains of the human ADAM-7 and ADAM-28 molecules was used as the outgroup. Nodes with confidence values >60% are indicated.
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Bazaa A, Marrakchi N, El Ayeb M, Sanz L, Calvete JJ (2005) Snake venomics: comparative analysis of the venom proteomes of the Tunisian snakes Cerastes cerastes, Cerastes vipera and Macrovipera lebetina. Proteomics 5:4223–4235
Bilgrami S, Tomar S, Yadav S, Kaur P, Kumar J, Jabeen T, Sharma S, Singh TP (2004) Crystal structure of schistatin, a disintegrin homodimer from saw-scaled viper (Echis carinatus) at 25 Å resolution. J Mol Biol 341:829–837
Bilgrami S, Yadav S, Sharma S, Perbandt M, Betzel C, Singh TP (2005) Crystal structure of the disintegrin heterodimer from saw-scaled viper (Echis carinatus) at 19 Å resolution. Biochemistry 44:11058–11066
Calvete JJ (2005) Structure-function correlations of snake venom disintegrins. Curr Pharm Des 11:829–835
Calvete JJ, Jürgens M, Marcinkiewicz C, Romero A, Schrader M, Niewiarowski S (2000) Disulfide bond pattern and molecular modelling of the dimeric disintegrin EMF-10, a potent and selective integrin α5β1 antagonist from Eristocophis macmahoni venom. Biochem J 345:573–581
Calvete JJ, Fox JW, Agelan A, Niewiarowski S, Marcinkiewicz C (2002) The presence of the WGD motif in CC8 heterodimeric disintegrin increases its inhibitory effect on αIIbβ3, αvβ3, and α5β1 integrins. Biochemistry 41:2014–2021
Calvete JJ, Moreno-Murciano MP, Theakston RDG, Kisiel DG, Marcinkiewicz C (2003) Snake venom disintegrins: novel dimeric disintegrins and structural diversification by disulphide bond engineering. Biochem J 372:725–734
Calvete JJ, Marcinkiewicz C, Monleón D, Esteve V, Celda B, Juárez P, Sanz L (2005) Snake venom disintegrins: evolution of structure and function. Toxicon 45:1063–1074
Daltry JC, Wüster W, Thorpe RS (1996) Diet and snake venom evolution. Nature 379:537–540
Duda TF Jr, Palumbi SR (1999) Developmental shifts and species selection in gastropods. Proc Natl Acad Sci USA 96:6820–6823
Fox JW, Serrano SMT (eds) (2005a) Special Issue: Snake Toxins and Hemostasis. Toxicon 45:951–1181
Fox JW, Serrano SMT (2005b) Structural considerations of the snake venom metalloproteinases, key members of the M12 reprolysin family of metalloproteinases. Toxicon 45:969–985
Fox JW, Ma L, Nelson K, Sherman NE, Serrano SMT (2006) Comparison of indirect and direct approaches using ion-trap and Fourier transform ion cyclotron resonance mass spectrometry for exploring viperid venom proteomes. Toxicon 47:700–714
Fry BG (2005) From genome to “venome”: molecular origin and evolution of the snake venom proteome inferred from phylogenetic analysis of toxin sequences and related body proteins. Genome Res 15:403–420
Fry BG, Wüster W (2004) Assembling an arsenal: origin and evolution of the snake venom proteome inferred from phylogenetic analysis of toxin sequences. Mol Biol Evol 21:870–883
Fry BG, Wüster W, Kini RM, Brusic V, Khan A, Venkataraman D, Rooney AP (2003) Molecular evolution and phylogeny of elapid snake venom three-finger toxins. J Mol Evol 57:110–129
Fry BG, Vidal N, Norman JA, Vonk FJ, Scheib H, Ramjan SF, Kuruppu S, Fung K, Hedges SB, Richardson MK, Hodgson WC, Ignjatovic V, Summerhayes R, Kochva E (2006) Early evolution of the venom system in lizards and snakes. Nature 439:584–588
Gasmi A, Srairi N, Guermazi S, Dkhil H, Karoui H, El Ayeb M (2001) Amino acid structure and characterization of a heterodimeric disintegrin from Vipera lebetina venom. Biochim Biophys Acta 1547:51–56
Jia L-G, Shimokawa K-i, Bjarnason JB, Fox JW (1996) Snake venom metalloproteinases: structure, function and relationship to the ADAMs family of proteins. Toxicon 34:1269–1276
Jones DT, Taylor WR, Thornton JM (1992) The rapid generation of mutation data matrices from protein sequences. Comput Appl Biosci 8:275–282
Juárez P, Sanz L, Calvete JJ (2004) Snake venomics: characterization of protein families in Sistrurus barbouri venom by cysteine mapping, N-terminal sequencing, and tandem mass spectrometry analysis. Proteomics 4:327–338
Juárez P, Wagstaff SC, Sanz L, Harrison RA, Calvete JJ (2006a) Molecular cloning of Echis ocellatus disintegrins reveals non-venom-secreted proteins and a pathway for the evolution of ocellatusin. J Mol Evol 63:183–193
Juárez P, Wagstaff SC, Oliver J, Sanz L, Harrison RA, Calvete JJ (2006b) Molecular cloning of disintegrin-like transcript BA-5A from a Bitis arietans venom gland cDNA library: a putative intermediate in the evolution of the long chain disintegrin bitistatin. J Mol Evol 63:142–152
Junqueira-de-Azevedo ILM, Ching ATC, Carvalho E, Faria F, Nishiyama NY, Ho PL, Diniz MRV (2006) Lachesis muta (Viperidae) cDNAs reveal diverging pit viper molecules and scaffolds typical of cobra (Elapidae) venoms: implications for snake toxin repertoire evolution. Genetics 173:877–889
Kordis D, Krizaj I, Gubensek F (2002) Functional diversification of animal toxins by adaptative evolution. In: Ménez A (ed) Perspectives in molecular toxinology. John Wiley & Sons, Chichester, UK, pp 401–419
Kumar S, Tamura K, Jacobsen IB, Nei M (2001) Mega2: Molecular Evolutionary Genetic Analysis software. Bioinformatics 17:1244–1245
Lu X, Lu D, Scully M.F, Kakkar VV (2005) Snake venom metalloproteinase containing a disintegrin-like domain, its structure-activity relationships at interacting with integrins. Curr Med Chem Cardiovasc Hematol Agents 3:249–260
Marcinkiewicz C, Calvete JJ, Marcinkiewicz MM, Raida M, Vijay-Kumar S, Huang Z, Lobb RR, Niewiarowski S (1999a) EC3, a novel heterodimeric disintegrin from Echis carinatus venom, inhibits α4 and α5 integrins in an RGD-independent manner. J Biol Chem 274:12468–12473
Marcinkiewicz C, Calvete JJ, Vijay-Kumar S, Marcinkiewicz MM, Raida M, Schick P, Lobb RR, Niewiarowski S (1999b) Structural and functional characterization of EMF10, a heterodimeric disintegrin from Eristocophis macmahoni venom that selectively inhibits α5β1 integrin. Biochemistry 38:13302–13309
Markland FS (1998) Snake venoms and the hemostatic system. Toxicon 36:1749–1800
Ménez A (2002) Perspectives in molecular toxinology. John Wiley & Sons, Chichester, UK
Michelmore RW, Meyers BC (1998) Clusters of resistance genes in plants evolve by divergent selection and a birth-and-death process. Genome Res 8:1113–1130
Monleón D, Moreno-Murciano MP, Kovacs H, Marcinkiewicz C, Calvete JJ, Celda B (2003) Concerted motions of the integrin-binding loop and the C-terminal tail of the non-RGD disintegrin obtustatin. J Biol Chem 278:45570–45576
Monleón D, Esteve V, Kovacs H, Calvete JJ, Celda B (2005) Conformation and concerted dynamics of the integrin-binding site and the C-terminal region of echistatin revealed by homonuclear NMR. Biochem J 387:57–66
Moura-Da-Silva AM, Theakston RDG, Crampton JM (1996) Evolution of disintegrin cysteine-rich and mammalian matrix-degrading metalloproteinases: gene duplication and divergence of a common ancestor rather than convergent evolution. J Mol Evol 43:263–269
Nei M, Rooney AP (2005) Concerted and birth-and-death evolution of multigene families. Annu Rev Genet 39:121–152
Nei M, Gu X, Sitnikova T (1997) Evolution by the birth-and-death process in multigene families of the vertebrate immune system. Proc Natl Acad Sci USA 94:7799–7806
Nikai T, Taniguchi K, Komori Y, Masuda K, Fox JW, Sugihara H (2000) Primary structure and functional characterization of bilitoxin-1, a novel dimeric P–II snake venom metalloproteinase from Agkistrodon bilineatus venom. Arch Biochem Biophys 378:6–15
Ohno M, Ogawa T, Oda–Ueda N, Chijiwa T, Hattori S (2002) Accelerated and regional evolution of snake venom gland isozymes, In: Ménez A (ed) Perspectives in molecular toxinology. John Wiley & Sons, Chichester, UK, pp 387–401
Ohno M, Chijiwa T, Oda-Ueda N, Ogawa T, Hattori S (2003) Molecular evolution of myotoxin phospholipases A2 from snake venom. Toxicon 42:841–854
Okuda D, Nozaki C, Sekiya F, Morita T (2001) Comparative biochemistry of disintegrins isolated from snake venom: consideration of the taxonomy and geographical distribution of snakes in the genus Echis. J. Biochem 129:615–620
Okuda D, Koike H, Morita T (2002) A new gene structure of the disintegrin family: a subunit of dimeric disintegrin has a short coding region. Biochemistry 41:14248–14254
Ronquist F, Huelsenbeck JP (2003) MrBayes3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Sanz L, Chen RQ, Pérez A, Hilario R, Juárez P, Marcinkiewicz C, Monleón D, Celda B, Xiong YL, Pérez-Payá E, Calvete JJ (2005) cDNA cloning and functional expression of jerdostatin, a novel RTS-disintegrin from Protobothrops jerdoni and a specific antagonist of the α1β1 integrin. J Biol Chem 280:40714–40722
Sanz L, Bazaa A, Marrakchi N, Pérez A, Chenik M, Bel Lasfer Z, El Ayeb M, Calvete JJ (2006) Molecular cloning of disintegrins from Cerastes vipera and Macrovipera lebetina transmediterranea venom gland cDNA libraries: insight into the evolution of the snake venom integrin-inhibition system. Biochem J 395:385–392
Shimokawa K, Jai L-G, Wang X-M, Fox JW (1996) Expression, activation and sequencing of the recombinant snake venom metalloproteinase, pro-atrolysin E. Arch Biochem Biophys 335:283–294
Soto JG, Powell RL, Reyes SR, Wolana L, Swanson LJ, Sanchez EE, Perez JC (2006) Genetic variation of a disintegrin gene found in the American copperhead snake (Agkistrodon contortrix). Gene 373:1–7
Tani A, Ogawa T, Nose T, Nikandrov NN, Deshimaru M, Chijiwa T, Chang CC, Fukumaki Y, Ohno M (2002) Characterization, primary structure and molecular evolution of anticoagulant protein from Agkistrodon actus venom. Toxicon 40:803–813
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Vicens Q, Cech TR (2006) Atomic level architecture of group I introns revealed. Trends Biochem Sci 31:41–51
Vidal N (2002) Colubroid systematics: evidence for an early appearance of the venom apparatus followed by extensive evolutionary tinkering. J Toxicol. Toxin Rev 21:21–41
Wu WB, Chang SC, Liau MY, Huang TF (2001) Purification, molecular cloning and mechanism of action of graminelysin I, a snake-venom-derived metalloproteinase that induces apoptosis of human endothelial cells. Biochem J 357:719–728
Zhou Q, Hu P, Ritter MR, Swenson SD, Argounova S, Epstein AL, Markland FS (2000) Molecular cloning and functional expression of contortrostatin, a homodimeric disintegrin from southern copperhead snake venom. Arch Biochem Biophys 375:278–288
Acknowledgments
This study was partly financed by Grant BFU2004-01432/BMC from the Ministerio de Educación y Ciencia, Madrid, Spain, and by Spanish-Tunisian Cooperation Programme 29p/02 of the Agencia Española de Cooperación Internacional (AECI). P.J. is the recipient of a predoctoral fellowship from the Formación de Personal Investigador (FPI) program from the Ministerio de Educación y Ciencia, Madrid. The authors thank Iñaki Comas (Instituto Cavanilles de Biodiversidad y Biología Evolutiva, Universidad de Valencia) for helping with phylogenetic analysis.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Reviewing Editor: Dr. Bryan Fry]
Amine Bazaa and Paula Juárez contributed equally to this work and may both be considered first authors.
Rights and permissions
About this article
Cite this article
Bazaa, A., Juárez, P., Marrakchi, N. et al. Loss of Introns Along the Evolutionary Diversification Pathway of Snake Venom Disintegrins Evidenced by Sequence Analysis of Genomic DNA from Macrovipera lebetina transmediterranea and Echis ocellatus . J Mol Evol 64, 261–271 (2007). https://doi.org/10.1007/s00239-006-0161-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-006-0161-4