Abstract
The evolution and patterns of selection of genes encoding 10 Drosophila odorant receptors (Or) and the sex pheromone receptor Gr68a were investigated by comparing orthologous sequences across five to eight ecologically diverse species of Drosophila. Using maximum likelihood estimates of dN/dS ratios we show that all 11 genes sampled are under purifying selection, indicating functional constraint. Four of these genes (Or33c, Or42a, Or85e, and Gr68a) may be under positive selection, and if so, there is good evidence that 12 specific amino acid sites may be under positive selection. All of these sites are predicted to be located either in loop regions or just inside membrane spanning regions, and interestingly one of the two sites in Gr68a is in a similar position to a previously described polymorphism in Gr5a that causes a shift in sensitivity to its ligand trehalose. For three Ors, possible evidence for positive selection was detected along a lineage. These include Or22a in the lineage leading to D. mauritiana and Or22b in the lineage leading to D. simulans. This is of interest in light of previous data showing a change in ligand response profile for these species in the sensory neuron (ab3A) which expresses both Or22a and Or22b in D. melanogaster. In summary, while the main chemosensory function and/or structural integrity of these 10 Or genes and Gr68a are evolutionarily preserved, positive selection appears to be acting on some of these genes, at specific sites and along certain lineages, and provides testable hypotheses for further functional experimentation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Insects, like other animals, detect a vast array of chemical information in the environment using a specialized olfactory system. In Drosophila melanogaster odors are detected by different functional classes of olfactory receptor neurons (ORNs) in a combinatorial fashion, with most ORNs responding to multiple odors and each odor being detected by multiple classes of ORN (de Bruyne et al. 1999, 2001). Drosophila species exploit many different ecological environments and include generalists that feed on a broad diet of fermenting fruits (D. melanogaster and D. simulans) as well as some specialists, for example, D. sechellia, which feeds exclusively on fruit from the plant Morinda citrifolia (Higa and Fuyama 1993). It is possible that the olfactory systems of these different species have evolved to preferentially detect particular odorants or combinations of odorants that are of ecological relevance. Such differences may manifest as changes in sensitivity and specificity of particular ORNs. For example, electrophysiological recordings from eight ORN functional classes for the entire D. melanogaster subgroup (comprising nine species) have revealed that there are odor sensitivity changes in one ORN type in the simulans clade (Stensmyr et al. 2003).
ORN responses are determined by the odorant receptor protein (Or) that they express. The D. melanogaster Or family comprises 60 genes encoding 62 proteins (Clyne et al. 1999; Gao and Chess 1999; Robertson et al. 2003; Vosshall et al. 1999). Since their discovery in 1999 significant evidence has been obtained showing that these genes represent the critical elements in odor coding and provide a molecular explanation for the variety of response properties of ORNs, with most ORNs expressing only one Or gene, although there are some exceptions (de Bruyne and Warr 2006; Hallem et al. 2006). As with odorant receptors from other organisms, Drosophila Ors are predicted to have seven hydrophobic putative transmembrane domains and have thus been believed to be G protein-coupled receptors (GPCRs). However, the structural conformation of Drosophila Ors is difficult to predict due to the fact that they are extremely divergent in amino acid sequence from each other and from all other known proteins, including Or proteins from other classes of animal. It should also be noted that conclusive evidence that the Drosophila Or proteins activate G protein-coupled signal transduction pathways is yet to be obtained. To further complicate matters, a recent study has suggested that the N terminal domains of two members of the Drosophila Or family are intracellular (Benton et al. 2006), indicating a different protein conformation compared to that found in GPCRs studied to date.
In addition to the uncertainty about protein topology, the lack of sequence homology across paralogues (∼20% identity at the amino acid level, with only a single tryptophan residue conserved across the family [Robertson et al. 2003]) makes it unclear which parts of the proteins are important for their function as odorant receptors, such as in ligand binding or in coupling to intracellular signal transduction proteins. For example, there are no conserved sequences in the third intracellular loop and the C terminal domain, regions that in many GPCRs contain conserved motifs involved in binding G proteins (Probst et al. 1992).
Comparative studies are extremely useful in identifying important structural and functional regions within proteins. However, paralogous comparisons among the Drosophila Ors are not informative due to their ancient splits and low sequence similarity. An alternative approach is to compare orthologous sequences across different species. Such studies have been used to compare chemosensory receptors among species of vertebrates and worms, and have identified selection operating on primate vomeronasal receptors (Mundy and Cook 2003), mammalian Ors (Branscomb et al. 2000), mammalian taste receptors (Fischer et al. 2004), and worm chemosensory receptors of the srz family (Thomas et al. 2005).
With different Drosophila species inhabiting distinct ecological environments we postulated that positive selection may be acting on Or genes, and that this positive selection and the identification of amino acid sites or lineages under selection may be informative with regard to important functional changes in these proteins. We therefore examined the molecular evolution of chemosensory receptors from members of the Drosophila genus. We first identified orthologues of 10 Or genes and of the putative sex pheromone receptor Gr68a from a range of Drosophila species. We then analyzed the evolutionary dynamics and patterns of selective pressures within these genes. Against a common background of purifying selection we found possible instances of positive selection in four of these genes at some amino acid sites and along particular lineages for another three of these genes.
Materials and Methods
Fly Stocks and Genomic DNA Preparation
Stocks of D. sechellia, D. mauritiana, D. erecta, and D. teissieri were obtained from the Tucson Drosophila species stock center. Stock codes for these species are 14021–0248.0, 14021–0241.1, 14021–0224.0, and 14021–0257.0, respectively. A D. simulans stock that was established from Australian field-caught flies was obtained from La Trobe University and used to identify all genes except Or22a and Or22b. D. simulans Or22a and Or22b sequences are from Dobritsa et al. (2003). Stocks of D. serrata, D. birchii, and D. species X (a cryptic member of the D. serrata complex [Schiffer et al. 2004]) were also obtained from La Trobe University. The Canton S strain of D. melanogaster was used for positive controls. Genomic DNA was extracted from 25 adult flies by homogenization in buffer consisting of 2% Triton X-100, 100 mM NaCl, 10 mM Tris-HCl, 1 mM EDTA, and 1% SDS, followed by phenol/chloroform extraction and ethanol precipitation.
Data Mining
D. melanogaster Or and Gr sequences were used to identify orthologous genes from the D. pseudoobscura genome with TBLASTN searches (Altschul et al. 1990) using the program linked to the Web site at Baylor College (http://www.hgsc.bcm.tmc.edu/projects/drosophila/). Homologous Or/Gr sequences for D. yakuba and D. ananassae were identified from 2004 genome assemblies (both Release 1.0). Searches were conducted using D. melanogaster sequences as queries with BLAT through the UCSC genome browser (http://www.genome.ucsc.edu/). Drosophila melanogaster sequences are those available from Flybase (http://www.flybase.bio.indiana.edu). However, two sequences of Or85e of different lengths have been published for the D. melanogaster strains Canton S and Oregon R (as discussed by Goldman et al. 2005). We analyzed only the shorter Canton S sequence.
PCR Cloning and DNA Sequencing
PCR primers (Supplementary Table 1) designed for 10 D. melanogaster Ors—Or22a, Or22b, Or33c, Or42a, Or43a, Or46aa, Or47a, Or71a, Or85d, and Or85e—and for Gr68a were used to amplify orthologous genes. To facilitate amplification primers were designed to the most 5′ and 3′ regions of coding sequence, and consequently gene sequences obtained are missing ∼18–50 base pairs from both the 5′ and the 3′ ends of the coding sequence. If more than 50 bp of sequence could not be obtained, then the gene was not included in the analysis.
Different primer pair combinations were required to amplify gene fragments from the various species due to varying levels of specificity for D. melanogaster primers. Standard PCR amplifications were prepared in a 50-μl reaction volume containing 1 U Taq polymerase, 1× reaction buffer, and 2.5 mM magnesium chloride (Fermentas; supplied by Progen), 0.4 mM dNTP mix, and 1 μM each primer with 1–2 μg genomic DNA template. PCR amplifications were performed in a Thermo Hybaid PCR machine with an initial denaturation step for 5 min at 95°C and amplified for 35 cycles (95°C for 30 s, 48–60°C for 1 min, 72°C for 2 min), then held at 72°C for 10 min. Annealing temperature varied according to primer pairs used and species being amplified. Temperatures for D. simulans and D. sechellia were 55–60°C due to the similarity of these species to D. melanogaster; for all other species annealing temperatures were typically lower (48–55°C). Genes were amplified as two to four PCR fragments with overlap to derive a consensus sequence. To minimize PCR-induced errors the proofreading Expand High Fidelity Taq polymerase (Roche Diagnostics, Germany) was used and PCR products were cloned into the pGEM-T Easy vector so that multiple subclones could be sequenced. At least three independent clones were sequenced for each gene fragment in each species. Additional clones were sequenced when necessary to resolve ambiguities. Cycle sequencing was performed using Big Dye version 3.1 (Applied Biosystems, CA) under standard conditions with gene specific forward, reverse, or internal primers. Sequencing reactions were purified using the Applied Biosystems ethanol/NaAc/EDTA precipitation protocol and resolved on an ABI 3100 automated sequencer.
Sequence Analysis
Nucleotide sequence data were collated using GCG (available from Accelrys, San Diego, CA; used via the Australian National Genomic Information Service) and edited manually to give consensus sequences for the 10 genes from the various Drosophila species. All amplified sequences are available in GenBank under accession numbers EF502092–EF502099. Amino acid sequences were aligned using ClustalW (Thompson et al. 1994) and are supplied as supplementary data. Transmembrane (TM) domains were predicted using the TMHMM transmembrane prediction server (http://www.cbs.dtu.dk/services/TMHMM) for both D. melanogaster and D. pseudoobscura sequences. The predicted domains for both were similar but the D. pseudoobscura predictions appeared more robust (that is, the number of TMs predicted and degree of hydropathy) and these were used for all subsequent analysis.
Coding sequence alignments required for PAML were manually adjusted to reflect the amino acid alignment and any gaps were omitted for the analysis. Parsimony trees were generated using PAUP* version 4 (Swofford 1998) using exhaustive search criteria. Statistical support for the branches was assessed using 1000 bootstrap replicates. Estimation of dN/dS rate ratios (ω) was carried out by maximum likelihood using the codon-based substitution models implemented in codeml with PAML version 3.14 (Wong et al. 2004; Yang 1997). The models used (M0, M3, M7, M8, M8a, and Mfree) are described in detail by Yang and Nielsen (2000), Yang et al. (2000), and Swanson et al. (2003). Briefly, M0 is a simple model of one ω ratio for all sites. M3 has three categories of site, with the ω ratio free to vary for each site class. M7, the “beta” neutral model, has eight categories of site with eight ω ratios in the range 0–1 taken from a discrete approximation of the beta distribution. M8, the “beta plus ω” selection model, has eight categories of site from a beta distribution as in model M7 plus an additional category of site with a ω ratio that is free to vary from 0 to >1. Model M8a is similar to M8 except that the ω1 category is fixed at 1 and, compared with M8, decreases incidences of false-positive detection. Mfree assumes a variable/heterogeneous ω for each lineage, and as such, for a data set to fit this model is a violation of the strictly neutral model M0.
In the cases of paralogous gene duplications, analysis with either copy of the duplications gave similar results (data not shown) and for subsequent analysis we used the copy with highest identity to the D. melanogaster gene.
Results
Identification of Chemosensory Receptor Orthologues from a Range of Drosophila Species
Orthologues of 10 of the D. melanogaster Or genes (Or22a, Or22b, Or33c, Or42a, Or43a, Or46aa, Or47a, Or71a, Or85d, and Or85e) were amplified and sequenced from D. simulans, D. sechellia, D. mauritiana, and D. erecta, and further orthologues from D. yakuba, D. ananassae, and D. pseudoobscura were obtained from genome databases using BLAST searches. These ten genes were chosen as a subset of Ors for which ligand information is available; there are no particular phylogenetic or chromosomal location relationships among the genes. In addition to the 10 Or genes, we also examined the evolution of the gustatory receptor Gr68a. The Gr68a receptor is of interest, as it is a putative sex pheromone receptor in D. melanogaster, suggested to be involved in recognition of species-specific cuticular hydrocarbons (Bray and Amrein 2003). Due to its potential role in mate recognition we hypothesized that this gene may be under positive selection, especially during bouts of speciation. Gr68a orthologues were identified from the same eight species used for the Or genes, together with four additional species, D. teissieri (another member of the melanogaster subgroup) and three montium subgroup species, D. serrata, D. birchii, and D. species X (Schiffer et al. 2004). D. serrata and D. birchii are of particular interest, as they are very closely related species but have different blends of cuticular hydrocarbons important for mate recognition and mate choice (Higgie et al. 2000; Howard et al. 2003).
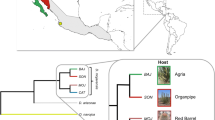
For each gene, PCR amplification was successful for species over a range of evolutionary distances from D. melanogaster (Fig. 1, Supplementary Table 2), except for Or22b, which could not be amplified from D. erecta. This is most likely due to the absence of this gene from the genome rather than PCR failure due to increased primer sequence divergence, as Or22b was not able to be identified from the genome databases for either D. yakuba or D. ananassae.
Phylogenetic relationships and estimated divergence times of lineages in the genus Drosophila used in this study. Species underlined were mined for receptors from their genome database; species with asterisks were only used in Gr68a analysis. Divergence times are based on a linearized Adh molecular clock (Russo et al. 1995; also reviewed by Powell 1997).
The number of exons and introns and the position of introns appear conserved across all orthologous genes in all species. The only exception is Or22a from D. pseudoobscura, which has four introns, while Or22a in all other species has three. However, considerable variation was found in intron size, with orthologous introns varying from −129 to +407 bp compared with those of D. melanogaster. In addition, exons had insertions or deletions of 3 to 18 bp in some Ors in some species. D. pseudoobscura Or proteins are generally one to six amino acids longer (with the exception of Or22a/Or22b). These extra amino acids are in either the N terminus or loop 4 (for definition see below), except for D. pseudoobscura Or42a2 (see below), which has an extra amino acid at the C terminus. For Gr68a two regions of the protein, loop 4 and loop 5, have indels of one to nine amino acids for all species. We found a number of amino acid differences between the Gr68a proteins of the closely related D. birchii and D. serrata (see Supplementary Fig. 1), any of which may be involved in ligand binding changes.
Amino acid identity levels between the D. melanogaster chemosensory receptors and their most divergent orthologues range from 60.3% to 85% (Supplementary Table 2). For most Ors the D. pseudoobscura orthologues are the most divergent. One exception is Or46aa, for which the D. ananassae orthologue is the most diverged.
Pseudogenes and Duplications
Interestingly, although D. melanogaster appears to have no pseudogene members of these multigene families, we were able to identify some pseudogenes in the genomic databases of other Drosophila species. Or22a pseudogenes were identified in both D. ananassae and D. pseudoobscura; in D. ananassae the single Or22a pseudogene has a 4-bp insertion causing a frame shift in the region encoding TM2, and in D. pseudoobscura there are two Or22a pseudogenes which contain multiple frame shifting deletions. D. sechellia has only a partial sequence/pseudogene for Or22b. This confirms the findings of Dekker et al. (2006), who also found that Or22b was a pseudogene in D. sechellia. D. mauritiana appears not to have a functional orthologue of Or85e but, rather, a pseudogene that has a 49-bp deletion in the region encoding TM2, causing a frame shift and a predicted truncated protein. Pseudogene sequences were not used in subsequent analysis.
As well as pseudogenes, we also found some paralogous duplications (nonidentical). D. ananassae has three copies of Or22a (60%–66% amino acid identity), while D. pseudoobscura has two copies of both Or22b (63.5%–63.7%) and Or42a (80%–85%).
Amino Acid Differences Are Not Randomly Distributed
To examine levels of amino acid variation across the receptors we divided them into the following domains: N terminus, transmembrane domains 1–7 (TM1–7), loops 1–6 (L1–L6), and the C terminus. The ratio of amino acid differences relative to domain size, averaged over the 11 receptors, is displayed in Fig. 2. If all regions of the protein contribute equally to function, amino acid changes should occur randomly across the protein. Our analysis of variation, however, shows that TM5, TM7, and the C terminus have less than half the number of expected changes per domain and are much more conserved than other regions of these proteins. Conversely, regions such as the N terminus, L2, and L4 are more variable (Fig. 2). The higher variability in some regions could be a result of selective pressures driving amino acid changes. Conversely, purifying selection could be selecting against amino acid replacements in structurally and functionally important regions of these proteins. To differentiate between these two possibilities we used likelihood-based methods to test for evidence of selection acting on the chemosensory receptor genes.
Ratio of relative amino acid differences per protein domain averaged over 11 chemoreceptors. The ratio value shown for each domain is the average over the 11 receptors of the number of observed changes per domain divided by the number of expected changes. The number of expected changes is calculated as the total number of changes across the protein from the consensus sequences, multiplied by the domain length and divided by the total protein length. If amino acid changes occur equally frequently across the protein, then the ratio is 1. N, N terminus ; TM1–TM7, transmembrane domains 1–7 ; L1–L6, loops 1–6 ; C, C terminus. Error bars represent SEM.
Drosophila Chemosensory Receptors Show Variable Selection Pressures Across Amino Acid Sites
To determine selective pressures on Drosophila chemosensory receptors, the ratio of nonsynonymous-to-synonymous substitution rates of protein coding sequences (ω) was calculated for each set of orthologous genes. In all cases ω was found to be substantially <1, ranging from 0.05 to 0.20 (Table 1). Although the proteins are thus overall under purifying selection, it is common for positive selection to act only on specific domains or residues in a protein sequence (Hughes et al. 1990; Nielsen and Yang 1998). We therefore extended our analysis using codeml (implemented with PAML) to examine ω ratios among sites.
We looked for evidence of variable selective pressures and positive selection among amino acid sites of the Drosophila genes using four site class models: M3 (discrete), M7 (“beta” neutral), M8 (“beta” selection), and M8a (M8 with ω fixed at 1). Likelihood values estimated under the various models indicate which models best fit the data. The significance of these can be tested between nested models M0/M3, M7/M8, and M8/M8a by a likelihood ratio test (LRT). The LRT is twice the log likelihood difference (2Δl), with a χ2 distribution and degrees of freedom equal to the difference in the number of parameters between the models (Yang 1998). The significance of the M8a/M8 comparison was determined using a 50:50 mixture of a point mass at zero and a χ 21 -distribution, as conducted by Swanson et al. (2003). Gene trees, likelihood values, and parameter estimates for the 11 data sets are available as supplementary data (Supplementary Fig. 2, Supplementary Table 3).
For all 11 data sets the discrete model (M3) was a significantly better fit to the data than M0 (M0 vs M3; p < 0.001; Table 2). M0 is a simple model of one ω ratio for all sites, while M3 has three categories of site, with the ω ratio free to vary for each site class. Acceptance of the discrete model indicates that the ω ratio is variable at sites across the coding sequence of these genes. For model M3, five genes, Ors22b, 33c, 42a, 85e, and Gr68a, have an ω value >1 (ω = 1.22–4.59) in 1%–3% of sites. Thus nonsynonymous mutations were fixed at a higher rate than synonymous mutations at these sites, indicating variable selective pressures (p 2 = 0.01–0.03; Supplementary Table 3).
The M0/M3 comparison is a test for variability, but positive selection can be tested specifically by an LRT between M7 and M8 or, more stringently, by an LRT between M8 and M8a (Swanson et al 2003). For four genes, Ors33c, 42a and 85e, and Gr68a, M8 is a significantly better fit to the data than the neutral model M7 (p <0.05; Table 2). For these four data sets, ω = 1.25–2.83 and p 1 = 0.01–0.06 (Supplementary Table 3), indicating possible positive selection at a small proportion (1%–6%) of sites. However, in the M8/M8a comparison only Or33c shows evidence of positive selection at particular sites (Table 2). We also note that the parsimony trees, which were used to estimate the statistics on the different chemoreceptor genes (Supplementary Fig. 2), are not all completely congruent with the classically accepted species tree (Fig. 1), with 4 of 11 having one taxon in an unexpected position. While this is not unexpected for rapidly diverging genes such as these, it does make the results slightly less certain in these cases.
As part of the selection models, a Bayes empirical Bayes computation (Yang et al. 2005) was used to identify the sites potentially under diversifying selection. Under M3 12 sites with posterior probabilities >0.95 were identified across Or33c, Or42a, Or85e, and Gr68a (posterior probabilities = 0.95–0.99; not shown). These same sites were identified in M8 but with lower posterior probabilities (0.74–0.94; Table 3). We acknowledge that likelihood-based methods have been shown to produce high levels of false positives (Suzuki and Nei 2001, 2002). However, the alternative parsimony-based models tend to be very conservative and have low power at detecting true positives, particularly for small data sets such as ours (Suzuki and Gojobori, 1999; Wong et al 2004).
Drosophila Chemosensory Receptors Show Variable Selection Pressures Across Lineages
Drosophila species occupy a range of different habitats and the ability to detect certain odors may be important for one species but not as important for others. Individual lineages or speciation events may have involved changes in odor preferences, thus we looked for evidence of variable selective pressures and positive selection among lineages of these genes using a lineage specific model, Mfree (Yang and Nielsen 2002). For Mfree all codon sites are assumed to be under the same selective pressure within a particular lineage, but the selective pressure can vary among lineages. An LRT comparing M0 and Mfree tests whether lineages are evolving under variable selective pressures.
For eight genes, seven Ors and Gr68a, the Mfree model fit the data significantly better (Table 4), indicating that for these genes different lineages are evolving under different rates of ω. For three of these eight genes (Or22a, Or22b, and Or85e) possible positive selection was detected in a lineage (Table 4, Supplementary Fig. 2): for Or85e in the branch leading to D. yakuba and D. erecta the estimated ω is 3.02 (rate of 16.3 nonsynonymous changes to 1.9 synonymous changes); for Or22a the branch leading to D. mauritiana had a value of ω = 2.90 (27.6 nonsynonymous changes and 3.0 synonymous changes); and for Or22b the branch leading to D. simulans had an estimated ω = 1.25 ( 9.0 nonsynonymous changes and 2.3 synonymous changes). For five of these eight genes, in particular lineages there were no synonymous changes and a number of nonsynonymous changes (0.9–21.1), in which case ω cannot be estimated by this program but it is possible that positive selection is acting in some cases. For Or22a, Or42a, Or71a, and Or85d, for each gene this is the branch leading to D. yakuba and D. erecta. For Gr68a this occurs for both the branch leading to D. sechellia/D. mauritiana and the branch leading to D. ananassae/D. pseudoobscura.
For Or46aa, ω<1 for all lineages and therefore there was no indication of positive selection on any of the branches. Finally, for 3 of the 11 genes (Or33c, Or43a, and Or47a), the free-ratio model did not provide a significantly better fit to the data than M0 (Table 4), indicating no variation in selective pressures among lineages.
Discussion
We have used likelihood-based methods to test for selection acting on 11 chemoreceptors (10 Ors and one Gr) across members of the genus Drosophila. We found that overall these receptors are under purifying selection. However, we did find evidence that four of these genes, Or33c, Or42a, Or85e, and Gr68a, may be under positive selection at a proportion of sites. Furthermore, we have identified 12 amino acid positions within these proteins on which positive selection might be acting. We acknowledge that none of the posterior probabilities under M8 are significant, and as previously mentioned, likelihood-based methods are prone to producing high levels of false positives (Suzuki and Nei 2001, 2002). Nevertheless, the large number of putative positively selected sites that we identified makes it likely that at least some of the 12 sites represent true positives. These results should thus be taken as an indication of possible positive selection, providing testable hypotheses for functionally important regions to be investigated with larger data sets and other analysis methods.
With the above caveats in mind, when considering the possible role of these sites in receptor function we note that many of the amino acid changes at the 12 sites involve charge and polarity changes and could, therefore, affect protein function. In addition, all of these 12 sites either are located in the loop regions of the proteins, which are predicted to protrude from the membrane, or are located just inside transmembrane domains (Fig. 3). Due to uncertainty in the exact position of the TMs in Drosophila Ors and Grs, it is possible that all the sites are in non-membrane-spanning regions. Interestingly, in C. elegans positively selected sites for the srz family of chemoreceptor genes were also found to be located in loop regions (Thomas et al. 2005). Furthermore, it seems more likely that sites on the extracellular surface may play roles in ligand binding, whereas intracellular sites may be involved in signal transduction. At present, however, it is unclear which loops of Drosophila Ors and Grs are extracellular and which are intracellular. Initial topology predictions for Drosophila Ors and Grs gave a structure similar to that of vertebrate Ors and other GPCRs, with the N terminus and even numbered loops on the extracellular surface (Clyne et al. 1999; Gao and Chess 1999; Otaki and Yamamoto 2003; Vosshall et al. 1999). According to this topology, 8 of the 12 sites are on the extracellular surface of the proteins (Fig. 3). However, recent experimental evidence suggests the opposite orientation in the membrane, with the N terminus and even-numbered loops as cytoplasmic (Benton et al. 2006), in which case the other four sites would be extracellular (Fig. 3). We note that the single amino acid change (alanine to threonine at position 218 in loop 3) implicated in ligand binding changes in the Drosophila gustatory receptor Gr5a (Ueno et al. 2001), would be extracellular according to the Benton topology model (Fig. 3).
Predicted topology of an idealized Drosophila chemosensory protein showing positions of sites possibly under positive selection. Transmembrane regions 1–7 (TMs 1–7 from left to right) formed by α-helices pass through the membrane indicated by the double lines. The N-terminal domain (NH2), loops 1–6, and the C-terminal domain (CO2H) fall outside the membrane. A and B refer to the intracellular and extracellular surfaces of the protein: for regular GPCR topology, B is intracellular and A is extracellular; for the Benton et al. (2006) topology, A is intracellular and B is extracellular. Due to differences in the size of chemoreceptor proteins (both overall length and variation in length of loop regions), the numbers of amino acids shown in each domain are representative only, and the positions indicated for the sites possibly under positive selection reflect their approximate location relative to the predicted topology. Only sites selected in M3 with posterior probabilities over >0.95 are indicated. The arrow indicates the approximate site of the Gr5a polymorphism that alters sensitivity to trehalose.
Putative cases of positive selection along a lineage were identified within three Or phylogenies, Or22a, Or22b, and Or85e. The possible selection on Or22a and Or22b is interesting to consider, as some information is available regarding their functional properties in different Drosophila species. Or22a and Or22b are 78% identical at the amino acid level, and in D. melanogaster they are co-expressed in the ab3A neuron, however, only Or22a is functional (Dobritsa et al. 2003). This co-expression means that the ability to link receptor sequence changes to functional changes is complicated by the uncertainty of whether one or both of Or22a and Or22b are functional in the ab3A neuron of particular species. An electrophysiological study by Stensmyr et al. (2003) found that the ab3A neurons of some species within the melanogaster subgroup respond with altered affinities for various similarly structured esters. The most extreme example was found in D. mauritiana, where the ab3A neuron responds more strongly to ethyl butyrate than to D. melanogaster’s preferred ethyl and methyl hexanoate. Interestingly, we found good evidence for positive selection for Or22a along the lineage leading to D. mauritiana (ω = 2.90). As we found that D. mauritiana has an apparently functional Or22b gene, we cannot be sure which gene is responsible for the ab3A response change, but our finding that there is positive selection on Or22a in the lineage leading to D. mauritiana is suggestive of Or22a being responsible for the altered specificity.
In conclusion, our results suggest that positive selection may be acting on some Drosophila chemosensory receptors, on particular amino acid sites and along particular lineages. Sites identified within these receptors, putatively under positive selection, provide testable hypotheses for regions involved in receptor function, particularly in altering odorant specificity. Ultimately the potential role of these sites in receptor function needs to be tested by altering them using site-directed mutagenesis and examining the effect on odor responses.
Supplemental Data
Supplemental data include primers used to amplify receptors, amino acid alignments of all receptor data sets, gene trees, likelihood values, and parameter estimates for site-specific and branch-specific models.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Benton R, Sachse S, Vosshall LB (2006) Atypical membrane topology and heteromeric function of Drosophila odorant receptors in vivo. PLoS Biol 4:1–18
Branscomb A, Seger J, White RL (2000) Evolution of odorant receptors expressed in mammalian testes. Genetics 156:785–797
Bray S, Amrein H (2003) A putative Drosophila pheromone receptor expressed in male-specific taste neurons is required for efficient courtship. Neuron 39:1019–1029
Clyne PJ, Warr CG, Freeman MR, Lessing D, Kim J, Carlson JR (1999) A novel family of divergent seven-transmembrane proteins: candidate odorant receptors in Drosophila. Neuron 22:327–338
de Bruyne M, Warr CG (2006) Molecular and cellular organization of insect chemosensory neurons. BioEssays 28:23–34
de Bruyne M, Clyne PJ, Carlson JR (1999) Odor coding in a model olfactory organ: the Drosophila maxillary palp. J Neurosci 19:4520–4532
de Bruyne M, Foster K, Carlson JR (2001) Odor coding in the Drosophila antenna. Neuron 30:537–552
Dekker T, Ibba I, Siju KP, Stensmyr MC, Hansson BS (2006) Olfactory shifts parallel superspecialism for toxic fruit in Drosophila melanogaster sibling, D. sechellia. Curr Biol 16:101–109
Dobritsa AA, van der Goes Naters W, Warr CG, Steinbrecht RA, Carlson JR (2003) Integrating the molecular and cellular basis of odor coding in the Drosophila antenna. Neuron 37:827–841
Fischer A, Gilad Y, Man O, Paabo S (2004) Evolution of bitter taste receptors in humans and apes. Mol Biol Evol 22:432–436
Gao Q, Chess A (1999) Identification of candidate Drosophila olfactory receptors from genomic DNA sequence. Genomics 60:31–39
Goldman AL, van der Goes Naters W, Lessing D, Warr CG, Carlson JR (2005) Coexpression of two functional odor receptors in one neuron. Neuron 45:661–666
Hallem EA, Dahanukar A, Carlson JR (2006) Insect odor and taste receptors. Annu Rev Entomol 51:113–35
Higa I, Fuyama Y (1993) Genetics of food preference in Drosophila sechellia. Genetica 88:129–136
Higgie M, Chenoworth S, Blows MW (2000) Natural selection and the reinforcement of mate recognition. Science 290:519–520
Howard RW, Jackson LL, Banse H, Blows MW (2003) Cuticular hydrocarbons of Drosophila birchii, D. serrata: Identification and role in mate choice in D. serrata. J Chem Ecol 29:961–976
Hughes AL, Ota T, Nei M (1990) Positive darwinian selection promotes charge profile diversity in the antigen-binding cleft of class i major-histocompatibility-complex molecules. Mol Biol Evol 7:515–524
Kreher SA, Kwon JY, Carlson JR (2005) The molecular basis of odor coding in the Drosophila larva. Neuron 46:445–456
Mundy NI, Cook S (2003) Positive selection during the diversification of class i vomeronasal receptor-like (V1RL) genes, putative pheromone receptor genes, in human and primate evolution. Mol Biol Evol 20:1805–1810
Nielsen R, Yang Z (1998) Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics 148:929–936
Otaki JM, Yamamoto H (2003) Length analyses of Drosophila odorant receptors. J Theor Biol 223:27–37
Powell JR (1997) Progress and prospects in evolutionary biology. The Drosophila model. Oxford University Press, New York
Probst WC, Snyder LA, Schuster DI, Brosius J, Sealfon SC (1992) Sequence alignment of the G-protein coupled receptor superfamily. DNA Cell Biol 11:1–20
Richards S, Liu Y, Bettencourt BR, Hradecky P, Letovsky S, Nielsen R, Thornton K, Hubisz MJ, Chen R, Meisel RP, Couronne O, Hua S, Smith MA, Zhang P, Liu J, Bussemaker HJ, van Batenburg MF, Howells SL, Scherer SE, Sodergren E, Matthews BB, Crosby MA, Schroeder AJ, Ortiz-Barrientos D, Rives CM, Metzker ML, Muzny DM, Scott G, Steffen D, Wheeler DA, Worley KC, Havlak P, Durbin KJ, Egan A, Gill R, Hume J, Morgan MB, Miner G, Hamilton C, Huang Y, Waldron L, Verduzco D, Clerc-Blankenburg KP, Dubchak I, Noor MAF, Anderson W, White KP, Clark AG, Schaeffer SW, Gelbart W, Weinstock GM, Gibbs RA (2005) Comparative genome sequencing of Drosophila pseudoobscura: Chromosomal, gene and cis-element evolution. Genome Res 15:1–18
Robertson HM, Warr CG, Carlson JR (2003) Molecular evolution of the insect chemoreceptor gene superfamily in Drosophila melanogaster. Proc Natl Acad Sci USA 100:14537–14542
Russo CAM, Takezaki N, Nei M (1995) Molecular phylogeny and divergence times of Drosophilid species. Mol Biol Evol 12:391–404
Schiffer M, Carew ME, Hoffmann AA (2004) Molecular, morphological and behavioural data reveal the presence of a cryptic species in the widely studied Drosophila serrata species complex. J Evol Biol 17:430–442
Stensmyr MC, Dekker T, Hansson BS (2003) Evolution of the olfactory code in the Drosophila melanogaster subgroup. Proc R Soc London 270:2333–2340
Suzuki Y, Gojobori T (1999) A method for detecting positive selection at single amino acid sites. Mol Biol Evol 16:1315–1328
Suzuki Y, Nei M (2001) Reliabilities of parsimony-based and likelihood-based methods for detecting positive selection at single amino acid sites. Mol Biol Evol 18:2179–2185
Suzuki Y, Nei M (2002) Simulation study of the reliability and robustness of the statistical methods for detecting positive selection at single amino acid sites. Mol Biol Evol 19:1865–1869
Swanson WJ, Nielson R, Yang Q (2003) Pervasive adaptive evolution in mammalian fertilization proteins. Mol Biol Evol 20:18–20
Swofford D (1998) PAUP*. Phylogenetic Analysis Using Parsimony (*and other methods). Version 4. Sinnauer Associates, Sunderland, MA
Thomas JH, Kelley JL, Robertson HM, Ly K, Swanson WJ (2005) Adaptive evolution in the SRZ chemoreceptor families of Caenorhabditis elegans and Caenorhabditis briggsae. Proc Natl Acad Sci USA 102:4476–4481
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTALW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Ueno K, Ohta M, Morita H, Mikuni Y, Nakajima S, Yamamoto D, Isono K (2001) Trehalose sensitivity in Drosophila correlates with mutations in and expression of the gustatory receptor gene Gr5a. Curr Biol 11:1451–1455
Vosshall LB, Amrein H, Morozov PS, Rzhetsky A, Axel R (1999) A spatial map of olfactory receptor expression in the Drosophila antenna. Cell 96:725–736
Wong WSW, Yang Z, Goldman N, Nielsen R (2004) Accuracy and power of statistical methods for detecting adaptive evolution in protein coding sequences and for identifying positively selected sites. Genetics 168:1041–1051
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. CABIOS 13:555–556
Yang Z (1998) Likelihood ratio tests for detecting positive selection and application to primate lysozyme evolution. Mol Biol Evol 15:568–573
Yang Z, Nielsen R (2000) Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol Biol Evol 17:32–43
Yang Z, Nielsen R (2002) Codon-substitution models for detecting molecular adaptation at individual sites along specific lineages. Mol Biol Evol 19:908–917
Yang Z, Nielsen R, Goldman N, Krabbe Pedersen A-M (2000) Codon-substitution models for heterogeneous selection pressure at amino acid sites. Genetics 155:431–449
Yang Z, Wong WSW, Nielsen R (2005) Bayes empirical Bayes inference of amino acid sites under positive selection. Mol Biol Evol 22:1107–1118
Acknowledgments
The authors would like to thank Dr. M. deBruyne for helpful comments on the manuscript and four anonymous reviewers for constructive suggestions regarding improving the manuscript and the evolutionary analysis. We would also like to thank Dr. Hugh Robertson for sharing unpublished information on the D. pseudoobscura Or genes gathered from the public genome database. This work was supported by grants from the Australian Research Council to C.G.W. and the Marsden Fund (02-HRT-016) to R.D.N.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Reviewing Editor: Dr. David Pollock]
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Tunstall, N.E., Sirey, T., Newcomb, R.D. et al. Selective Pressures on Drosophila Chemosensory Receptor Genes. J Mol Evol 64, 628–636 (2007). https://doi.org/10.1007/s00239-006-0151-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-006-0151-6