Abstract
Asymmetric compositional and mutation bias between the two strands occurs in mitochondrial genomes, and an asymmetric mechanism of mtDNA replication is a potential source of this bias. Some evidence indicates that during replication the heavy strand is subject to a gradient of time spent in a single-stranded state (D ssH) and a gradient of mutational damage. The nucleotide composition bias among genes varies with D ssH. Consequently, partial genome duplications (PGD) will alter the skew for genes located downstream of the duplication, relatively to nascent light strand synthesis, and in the same way, gene rearrangements (GRr) will affect genes by changing their skews. We examined cases where there had been PGD or GRr and determined whether this left a trace in the form of unusual patterns of base composition. We compared the skew of genes differently located on the mtDNA genome of previously published whole mtDNA genomes from amphibians, a group that shows considerable levels of both GRr and PGD. After observing a significant correlation between AT and GC skew with D ssH at fourfold redundant sites, we ran our analysis and detected 31.3% of the species with GRr and/or PGD. By comparing the nucleotide composition at fourfold redundant sites in normal and “abnormal” species, we found that A/C variation occurs and is associated with GRr/PGD. These results show that by analyzing the nucleotide skews of only three genes, it may be possible to predict some mitochondrial GRr and/or PGD without knowing the complete mtDNA genome sequence.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mitochondrial DNA (mtDNA) has been extensively employed as a marker in evolutionary studies. Vertebrate mtDNA is a compact circular DNA molecule about 16.5 kb long, coding for 13 proteins, 22 tRNAs, and 2 rRNAs. These encoding genes have an asymmetric distribution between the two strands. The individual strands of the mtDNA are distinguished by their different buoyant density in a cesium chloride gradient (Radloff et al. 1967) as a result of uneven guanine content: the heavy (H) strand is guanine rich and the light (L) strand is guanine poor (Anderson et al. 1981).
If mutational or selective pressures have equal or no effect within each strand, there are 6 substitution rates instead of 12 (Parity Rule type 1; PR1 [Sueoka 1995]), because the substitution rates for both strands are scored on one strand. Thus, a mutation scored on one strand can be the direct result of that substitution or the indirect result of the complementary mutation on the other strand. In the absence of strand bias the base frequencies within each strand will be in equilibrium: A = T and G = C (Parity Rule type 2; PR2 [Lobry 1995]). Strand bias conditions can be detected by deviations from these rules (Frank and Lobry 1999). Asymmetric compositional and mutation bias between the two strands is known to occur (Raina et al. 2005; Faith and Pollock 2003; Bielawski and Gold 2002; Reyes et al. 1998; Jermiin et al. 1995; Perna and Kocher 1995; Tanaka and Ozawa 1994; Asakawa et al. 1991).
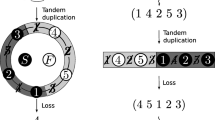
The asymmetric mechanism of mtDNA replication has been referenced as a potential source of strand bias (Faith and Pollock 2003; Reyes et al. 1998). The genome replication process takes about 2 h (Clayton 1982) and starts from the origin of H-strand replication (OH), displacing the parental H-strand. After two-thirds of the H-strand replication, when the origin of L-strand (OL) is displaced, the L-strand synthesis starts in the opposite direction. The parental H-strand is single-stranded until the L-strand replication fork synthesizes the nascent L-strand. Consequentially and depending on the time spent during the single-stranded state (D ssH), the parental H-strand is subject to a gradient of mutational damage, i.e., (i) hydrolytic deamination of cytosine, (ii) hydrolytic deamination of adenine, and (iii) oxidation of guanine. The deamination theory is supported by Reyes et al. (1998), with cytosine-to-uracil mutations occurring more often than adenine-to-hypoxanthine mutations. The observed gradient of bias toward A and C on the complementary L-strand also supports these different mutation rates (Krishnan et al. 2004b; Reyes et al. 1998; Jermiin et al. 1995; Perna and Kocher 1995; Tanaka and Ozawa 1994).
Strand compositional bias (skew) observed in mitochondrial genomes is stronger at fourfold (4×) redundant sites, suggesting the existence of an asymmetric directional mutation pressure. In these sites the action of natural selection is significantly reduced and the frequencies between all nucleotides are presumably a result of mutational patterns. It has been concluded that the nucleotide skew is correlated with the duration of time spent in a single-strand state (Reyes et al. 1998; Jermiin et al. 1995; Perna and Kocher 1995; Tanaka and Ozawa 1994) and is not gene specific (Asakawa et al. 1991). These conclusions are brought into question by recent studies suggesting that the replication mechanism is not asymmetric, because of the double-stranded state of both strands during DNA synthesis (Yang et al. 2002; Holt et al. 2000). Although this is an important issue, there is currently no consensus (Brown et al. 2005).
Our study was based on the widely accepted assumption that nucleotide skew is correlated with D ssH during the replication process. Consequently, PGD would change the skew for genes located downstream of the duplication, relative to nascent L-strand synthesis, because it would change D ssH and the incidence of mutational damage. In the same way, GRr would change the skew of rearranged genes. The majority of organisms currently have only a few gene sequences available, and no method has been developed to detect PGD/GRr without knowing the entire mtDNA genome sequence. The objective of this study was to assess the utility of the AT skew of genes to identify known PGD/GRr. MtDNA protein coding genes can be divided into two different groups based on their specific genome location with respect to the direction of movement of the H-strand replication fork: genes located before (all genes except ND1 and ND2) and after (ND1 and ND2) the OL. It is known that the genome region between the Cyt b and tRNA–Phe genes is the major target for GRr and PGD (Mueller and Boore 2005; Macey et al. 1998). PGD in this region increases D ssH only for ND1 and ND2, because they are located downstream of the PGD in L-strand replication. ND1 or ND2 absolute “abnormal” skew values cannot be directly compared to detect GRr and/or PGD, because these skews are organism-specific even in the absence of GRr and PGD. However, the ratio of skews between two unaffected genes can be used as an organism-specific reference value to compare against the ratio of skews between an unaffected gene and an affected gene. The resultant value should indicate the occurrence of PGD. GRr events affect the involved genes directly, and can also indirectly affect other genes (if the rearrangement alters their positions relative to the OL). Therefore, PGD and GRr can alter nucleotide frequency equilibria by altering relative gene position, order, or both. This should eventually produce detectable deviations from normal patterns of base composition.
Our investigation focused on previously published whole amphibian mtDNA genomes because they have a high occurrence of gene rearrangements relative to other vertebrates, and because there are a large number of complete genome sequences available. We used 4× redundant sites only, to avoid the effect of selection on amino acids. Our aim was to determine if GRr and PGD could be detected solely through comparative skews of compositional bias.
Materials and Methods
The complete genome sequences used in our study were downloaded from GenBank (Table 1). The nucleotide composition of the H-strand coding genes were obtained using the program MEGA version 3.0 (Kumar et al. 2004). The 4× redundant codons included in the analysis were alanine (GCN), proline (CCN), serine (TCN), threonine (ACN), arginine (CGN), glycine (GGN), leucine (CTN), and valine (GTN).
The strand compositional bias is indicated by the GC and AT skew (Perna and Kocher 1995) and calculated according to the formula: AT skew = (A − T)/(A + T), GC skew = (G − C)/(G + C). The standard deviations (SD) of the skews were calculated as (Lobry 1996): SD (AT skew) = 2/(A + T) √[A . T/(A + T)], SD (GC skew) = 2/(G + C) √[G . C/(G + C)]. A, T, C, and G are percentages.
The D ssH was calculated following Faith and Pollock (2003), Reyes et al. (1998), and Tanaka and Ozawa (1994): D ssH = [L − 2 (x − OL)]/L, for ND1 and ND2, the two genes located after the light-strand origin of replication, and D ssH = 2 (OL − x)/L for the remaining genes, where L is the length of the genome, OL is the position of the L-strand origin of replication, and x is the average position of the gene. We assumed that strand synthesis is constant along the genome and equal for both strands. D ssH was calculated for each species and for each gene. In amphibians, the gene order relative to D ssH was the same as in Faith and Pollock (2003) and Reyes et al. (1998): COX1 < COX2 < ATP8 < ATP6 < COX3 < ND3 < ND4L < ND4 < ND1 < ND5 < ND2 < Cyt b. To test the correlation between AT and GC skew with D ssH, we did not use ATP8, ND3 and ND4L because these genes are the smallest protein coding genes and show a low number of 4× redundant sites.
We classified the genomes as “normal” or “abnormal” based on genome organization and length. Species with gene rearrangements relative to the normal vertebrate gene order and/or genome length >17.0 kb were considered “abnormal” (Table 1). We then looked for signs or patterns in nucleotide composition and AT and GC skews that could be indicative of GRr or PGD. To detect nucleotide variation between normal and abnormal species, we compared for each gene the L-strand nucleotide composition at 4× redundant sites.
The ratio of skews of COX1, ND1/ND2, and one of the remaining genes was used to detect varying mitochondrial gene order. A detected species was required to present the following disequilibrium features:
-
(a)
AT skew <0 (for one gene) or ratio of AT skew >2.50.
-
(b)
For the three genes used in each graphical analysis [(A% − C%)/C%] < 0.95. Our reference value was 0.95 instead of 1.00 to reduce the type I error rate.
Results
Skew and D ssH
To test the correlation between the skews with D ssH we plotted the AT and GC skews at 4× redundant sites for each protein coding gene versus D ssH (Fig. 1). AT and GC skew show a significant correlation with D ssH. AT skew is positively correlated (R 2 = 0.724; p = 0.004) and GC skew is negatively correlated (R 2 = 0.603; p = 0.014). Our results with amphibians are in concordance with other studies; Reyes et al. (1998) obtained the same results, although Faith and Pollock (2003) did not find a significant correlation for GC skews with D ssH.
Nucleotide Composition Between Normal and “Abnormal” Species
We compared the nucleotide composition at 4× redundant sites for each gene between normal and abnormal species (Fig. 2). In the two groups T% and G% are similarly represented within each gene. However, genes of abnormal species showed a lower A% and a higher C%. Guanine is the least represented nucleotide in the two groups and adenine is the most represented. In normal mtDNA genes adenine proportions have a tendency to be more than twice the proportions of cytosine (A% > 2C%).
Detecting GRr and PGD
In these analyses the skew of COX1 for each species was used as the denominator for the different ratios, because COX1 is expected to have the lowest skews values within each genome. As a numerator we used the skew of one gene located, like COX1, before OL and the skew of ND1/ND2 (Fig. 3). The analyses using the ratios of GC skew did not show any distinguishable features (analyses not shown). In the analyses using the ratio of AT skews we detected 10 species with GRr/PGD without obtaining any false positives.
Plot of species-specific AT skew rates. (A) Cyt b/COX1 against ND1/COX1; (B) Cyt b/COX1 against ND2/COX1; (C) COX3/COX1 against ND1/COX1; (D) COX3/COX1 against ND2/COX1. (+) Species with “normal” gene order; (Δ) species with “abnormal” genome order. Arrows point to abnormal species detected by the analyses.
Discussion
Fourfold redundant sites are expected to be in the intrastrand equalities A = T and G = C (PR2), because they are less affected by selection. In amphibian mtDNA these sites present a high variability in base composition and, consequently, strong deviations from the nucleotide equilibrium quantities. This violation from PR2 could be due to asymmetric nucleotide substitutions. Previous studies based on mtDNA genes also related nucleotide composition bias with asymmetric substitution matrix (Raina et al. 2005; Krishnan et al. 2004b; Faith and Pollock 2003; Bielawski and Gold 2002; Reyes et al. 1998). One hypothesis to explain these substitution patterns is an asymmetric replication mechanism of mtDNA, since the H-strand is subject to an increase in time spent in a single stranded state. Higher D ssH values are associated with stronger skews and the significant correlations observed between both AT and GC skews with D ssH support the mtDNA asymmetric replication process. However, GC skew correlation with D ssH is lower than AT skew, suggesting that γ-bases (G and C) are less affected by D ssH or that GC skew reaches an early saturation level. This second explanation is more plausible because even COX1, the gene with the lowest D ssH value, shows higher GC skew than AT skews. Reyes et al. (1998) obtained similar results with mammals, but Faith and Pollock (2003), using several vertebrate groups (reptiles, sharks, bony fish, and mammals), reported that the correlation between GC skew and D ssH was not significant. This may indicate GC skew saturation, making it impossible to correlate GC skew with D ssH in distantly related species.
However, the skew values show a high variability between species; in this study the AT skew varies from −0.05 to 0.49 in COX1 and from 0.00 to 0.73 in Cyt b and the GC skew varies from −0.76 to −0.03 in COX1 and from −0.95 to −0.19 in Cyt b.
When we compared the nucleotide composition at 4× redundant sites for each gene between normal and “abnormal” species, all genes indicated that A/C variation occurs and is associated with GRr/PGD. The abnormal species show a lower difference between A% and C%. This suggests the existence of a process directly responsible for the A/C variation. It was hypothesized that the most plausible factor creating biases in single-strand mutation rates is an increased rate of hydrolytic deamination of cytosine and adenine (Reyes et al. 1998). However, these mutations do not explain the direct variation between adenine and cytosine composition. The pattern observed in abnormal species (L-strand C% increase and A% decrease) can be explained by the decreased formation of 8-hydroxyguanine by oxidation, which preferentially pairs with adenine rather than cytosine (Lindahl 1993). In some way, abnormal species could be less affected by this mutation.
The GC skew values are not informative for our analysis as a result of the initial rate of increase and rapid saturation level of the G-to-A transitions (on the L strand). A theoretical maximum rate can be approached rapidly, stabilizing the GC skew (Krishnan et al. 2004a; Faith and Pollock 2003). This sensitivity of GC skew obscured the detection of GRr or PGD.
With the ratio of AT skew values we were able to identify some of the genomes with GRr and PGD. From all these analyses we could detect a minimum of three GRr/PGD (using COX1, ND5, and ND1) and a maximum of seven GRr/PGD (using COX1, ND2, and ATP6). Across all analyses we detected 8 GRr and 2 PGD of 32, representing 31.2% of the “abnormal” species.
Six GRr detected involved tRNA-Leu (before ND5), tRNA-Thr, and tRNA-Pro (after Cyt b). All the species with displaced ND5 and the genome lacking ND5 were detected. Eight species (including the two species that present only PGD) presented PGD in the control region (length longer than 1100 bp). In the mitochondrial genome of Polypedates megacephalus (a detected species) ATP8 and ND5 were absent, and in the Ferjervarya limnocharis (also a detected species) mitochondrial genome there was a tandem duplication of tRNA–Met. One translocation of ND6 and tRNA–Glu was detected (Stereochilus marginatus). Nevertheless, there was no clear pattern for the detected “abnormal” species and new data will be needed to clarify and understand GRr/PGD and its consequences for mitochondrial genome evolution.
With these analyses we conclude not only that GRr/PGD affect displaced genes but that all the genes can be affected. Nine of the species with GRr/PGD were detected by at least two different sets of genes. In fact, GRr/PGD in Bufo melanostictus, Rana nigromaculata, and Rhyacophorus schlegelii were detected by all 12 different analyses. Therefore it could be possible to detect GRr/PGD with just one gene, if the AT skew is negative.
The method developed in this work indicates that some mitochondrial GRr or PGD can be detected by a simple analysis using just three genes. Although in this case applied to amphibians, as more mitochondrial genomes are published, the method can be tested for other groups. By detecting GRr or PGD using only nucleotide skews, unusual mtDNA nucleotides can be predicted without the need for genome sequencing. These “abnormal” mtDNA genomes may cause errors in phylogeny reconstruction if the genes used have atypical skews. By detecting them through the method presented here, such errors may be avoided. In the examples used here, gene order and the position of the OL were known. Similar analyses of compositional bias may be helpful in locating OL where the position is unknown, a common situation in invertebrates (Mizi et al. 2005). When gene rearrangements occur, a gene may be out of equilibrium with its new position. However, to determine this, a well-resolved phylogeny is needed, something that is lacking for most groups that have extensive mtDNA gene rearrangements. There is an acknowledged need for greater understanding of the basic biology of the mtDNA molecule (Ballard and Rand 2005). Further analyses of base composition bias will undoubtedly help in this respect.
References
Anderson S, Bankier AT, Barrell BG, de Bruijn MHL, Coulson AR, Drouin J, Eperon IC, Nierlich DP, Roe BA, Sanger F, Schreier PH, Smith AJH, Staden R, Young IG (1981) Sequence and organization of the human mitochondrial genome. Nature 290:457–465
Asakawa S, Kumazawa Y, Araki T, Himeno H, Miura K, Watanabe K (1991) Strand-specific nucleotide composition bias in echinoderm and vertebrate mitochondrial genomes. J Mol Evol 32:511–520
Ballard JW, Rand DM (2005) The population biology of mitochondrial DNA and its phylogenetics implications. Annu Rev Ecol Syst 36:621–642
Bielawski JP, Gold JR (2002) Mutation patterns of mitochondrial H- and L-strand DNA in closely related Cyprinid fishes. Genetics 161:1589–1597
Brown TA, Cecconi C, Tkachuk AN, Bustamante C, Clayton DA (2005) Replication of mitochondrial DNA occurs by strand displacement with alternative light-strand origins, not via a strand-coupled mechanism. Genes Dev 19:2466–2476
Clayton DA (1982) Replication of animal mitochondrial DNA. Cell 28:693–705
Faith JJ, Pollock DD (2003) Likelihood analysis of asymmetrical mutation bias gradients in vertebrate mitochondrial genomes. Genetics 165:735–745
Frank AC, Lobry JR (1999) Asymmetric substitution patterns: a review of possible underlying mutational or selective mechanisms. Gene 238:65–77
Holt IJ, Lorimer HE, Jacobs HT (2000) Coupled leading- and lagging-strand synthesis of mammalian mitochondrial DNA. Cell 100:515–524
Jermiin LS, Graur D, Crozier H (1995) Evidence from analyses of intergenic regions for strand-specific directional mutation pressure in metazoan mitochondrial DNA. Mol Biol Evol 12:558–563
Krishnan NM, Raina SZ, Pollock DD (2004a) Analysis of among-site variation in substitution patterns. Biol Proced Online 6:180–188
Krishnan NM, Seligmann H, Raina SZ, Pollock DD (2004b) Detecting gradients of asymmetry in site-specific substitutions in mitochondrial genomes. DNA Cell Biol 23:707–714
Kumar S, Tamura K, Nei M (2004) MEGA3: Integrated software for Molecular Evolutionary Genetics Analysis and sequence alignment. Brief Bioinform 5:150–163
Lindahl T (1993) Instability and decay of the primary structure of DNA. Nature 362:709–715
Lobry JR (1995) Properties of a general model of DNA evolution under no-strand-bias conditions. J Mol Evol 40:326–330 (Erratum: J Mol Evol 41:680)
Lobry JR (1996) Asymmetric substitution patterns in the two DNA strands of bacteria. Mol Biol Evol 13:660–665
Macey JR, Shulte II JA, Larson A, Papenfuss TJ (1998) Tandem duplication via light-strand synthesis may provide a precursor for mitochondrial genomic rearrangement. Mol Biol Evol 15:71–75
Mizi A, Zouros E, Moschonas N, Rodakis GC (2005) The complete maternal and paternal mitochondrial genomes of the Mediterranean mussel Mytilus galloprovincialis: implications for the doubly uniparental inheritance mode of mtDNA. Mol Biol Evol 22:952–967
Mueller RL, Boore JL (2005) Molecular mechanisms of extensive mitochondrial gene rearrangement in Plethodontid salamanders. Mol Biol Evol 22:2104–2112
Perna NT, Kocher TD (1995) Patterns of nucleotide composition at fourfold degenerate sites of animal mitochondrial genomes. J Mol Evol 41:353–358
Radloff R, Bauer W, Vinograd J (1967) A dye-buoyant density method for the detection and isolation of closed circular DNA: the closed circular DNA in HeLa cells. Proc Natl Acad Sci USA 57:1514–1521
Raina SZ, Faith JJ, Disotell TR, Seligmann H, Stewart CB, Pollock DD (2005) Evolution of base-substitution gradients in primate mitochondrial genomes. Genome Res 15:665–673
Reyes A, Gissi C, Pesole G, Saccone C (1998) Asymmetrical directional mutation pressure in the mitochondrial genome of mammals. Mol Biol Evol 15:957–966
Sueoka N (1995) Intrastrand parity rules of DNA base composition and usage biases of synonymous codons. J Mol Evol 40:318–325
Tanaka M, Ozawa T (1994) Strand asymmetry in human mitochondrial DNA mutations. Genomics 22:327–335
Yang MY, Bowmaker M, Reyes A, Vergani L, Paolo A, Gringeri E, Jacobs HT, Holt IJ (2002) Biased incorporation of ribonucleotides on the mitochondrial L-strand accounts for apparent strand-asymmetric DNA replication. Cell 111:495–505
Acknowledgments
This work was supported by Fundação para a Ciência e Tecnologia (FCT) grants POCI/61946/2004 (to D.J.H), SFRH/BD/11377/2002 (to E.F.), and POCI/BIA-BDE/61946/2004 (to M.M.F.). We thank the two reviewers for their constructive comments on early drafts of the manuscript and D. Pollock for his extensive editorial assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Reviewing Editor: Dr. David Pollock]
Rights and permissions
About this article
Cite this article
Fonseca, M.M., Froufe, E. & James Harris, D. Mitochondrial Gene Rearrangements and Partial Genome Duplications Detected by Multigene Asymmetric Compositional Bias Analysis. J Mol Evol 63, 654–661 (2006). https://doi.org/10.1007/s00239-005-0242-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-005-0242-9