Abstract
The transfer and integration of tRNA genes from organellar genomes to the nuclear genome and between organellar genomes occur extensively in flowering plants. The routes of the genetic materials flowing from one genome to another are biased, limited largely by compatibility of DNA replication and repair systems differing among the organelles and nucleus. After thoroughly surveying the tRNA gene transfer among organellar genomes and the nuclear genome of a domesticated rice (Oryza sativa L. ssp. indica), we found that (i) 15 mitochondrial tRNA genes originate from the plastid; (ii) 43 and 80 nuclear tRNA genes are mitochondrion-like and plastid-like, respectively; and (iii) 32 nuclear tRNA genes have both mitochondrial and plastid counterparts. Besides the native (or genuine) tRNA gene sets, the nuclear genome contains organelle-like tRNA genes that make up a complete set of tRNA species capable of transferring all amino acids. More than 97% of these organelle-like nuclear tRNA genes flank organelle-like sequences over 20 bp. Nearly 40% of them colocalize with two or more other organelle-like tRNA genes. Twelve of the 15 plastid-like mitochondrial tRNA genes possess 5′- and 3′-flanking sequences over 20 bp, and they are highly similar to their plastid counterparts. Phylogenetic analyses of the migrated tRNA genes and their original copies suggest that intergenomic tRNA gene transfer is an ongoing process with noticeable discriminatory routes among genomes in flowering plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Among flowering plants, the nuclear genomes often contain significant DNA fractions similar to mitochondrial DNA (mtDNA) and plastid DNA (ptDNA), which are termed nuclear mtDNA (NUMT) and nuclear ptDNA (NUPT), respectively (Lopez et al. 1994; Timmis et al. 2004). Likewise, their mitochondrial genomes also have DNA sequences originated from plastid genomes, designated MTPT (Thorsness and Weber 1996; Timmis et al. 2004). NUMTs, NUPTs, and MTPTs are assumedly generated via nonhomologous recombination among nuclear and organellar genomes and subsequently reintegrated into their final destinies (for reviews, see Noutsos et al. 2005; Richly and Leister 2004a, b; Timmis et al. 2004). Regarding how organellar DNA transfers to nuclear genomes and between organellar genomes, two interpretations—bulk-DNA and cDNA-mediated hypotheses—have been proposed. The bulk-DNA hypothesis suggests that gene transfer is a result of recombination promoted by a large amount of escaped organellar DNA (Henze and Martin 2001; Thorsness and Weber 1996). The cDNA-mediated hypothesis assumes that a cDNA intermediate of organellar mRNAs is a vector transferring genes to the nucleus or interorganelles, based on studies across distant taxa (Adams and Palmer 2003; Adams et al. 2002; Brennicke et al. 1993). As a matter of fact, comparative genomics data in yeast and higher plants suggest that organelle-to-nucleus gene transfer is mostly attributable to the bulk-DNA model. The cDNA-mediated model would be unnecessary for early evolution of mitochondria and plastids because of the few observations of editing and introns in plants, even in free-living α-proteobacteria and cyanobacteria (Timmis et al. 2004).
Ever since the first discovery of plastid sequences in a mitochondrial genome (Stern and Lonsdale 1982), almost all sequenced angiosperm mitochondrial genomes have been demonstrated to contain genetic materials of the plastid counterparts, and mtDNA and ptDNA are also detectable in nuclear genomes. However, no sequence originated from mitochondrial or nuclear genomes has ever been discovered in plastid genomes, and no nuclear sequences in mitochondrial genomes (see Cummings et al. 2003 and references therein). It is reasonable to think the flow of genetic materials among organellar and nuclear genomes is actually biased, although DNA transfer and integration from plastids to mitochondria are ubiquitous and ongoing in flowering plants. As a relic of evolution, mitochondrial tRNA (mt-tRNA) genes show a characteristic of double origin: native and plastid-derived. The different copies of tRNA genes with very significant sequence similarities (95%–100%) to their organellar counterparts are generally referred to as plastid-like (pt-like) or mitochondrion-like (mt-like). In mitochondria, pt-like mt-tRNA genes account for nearly one-third of all tRNA genes in flowering plants. The rest lacking or with sequence similarity of <80% to their plastid counterparts are referred to as native (or genuine) mt-tRNA genes (Hoffmann et al. 2001; Marechal-Drouard et al. 1993). This double-origin feature is considered unique to mt-tRNA genes in flowering plants, because the phenomenon has not yet been identified in liverwort (Marchantia polymorpha) and other lower plants (Oda et al. 1992), even in green alga. The case of mitochondrial pt-like tRNA genes in rice (Oryza sativa, cultivar Nipponbare) has been described recently (Notsu et al. 2002). Nine species (trnC GCA, trnF GGA, trnH GTG, trnM CAT, trnN GTT, trnP TGG, trnR TCT, trnS GGA, and trnW CCA) of all full-length mt-tRNA genes share remarkable sequence similarities (97%–100%) with their plastid counterparts. Compared with the sequences of other higher plants, mitochondrial trnC GCA and trnF GGA are often plastid-originated among monocotyledons (such as rice, maize, and wheat) but are native among dicotyledons, such as Arabidopsis, sugar beet, and tobacco (Veronico et al. 1996).
Moreover, DNA sequence transfer from organellar genomes to the nuclear genome occurs frequently in many eukaryotic cells. NUMTs and NUPTs identifiable with high identity (95%–100%) to their mitochondrial and plastid counterparts have been found in almost all characterized dicotyledons and monocotyledons thus far (Richly and Leister 2004a, b). Investigations of Matsuo and colleagues revealed that rice nuclear DNA with ptDNA integrations has experienced rapid fragmentations and vigorous shuffling, and an equilibrium between frequent integration and rapid elimination of the ptDNA seems to exist among the organellar fragments and the nuclear chromosomes (Matsuo et al. 2005). Furthermore, the transfer of ptDNA to the nucleus has also been detected under experimental conditions in tobacco (Nicotiana tabacum) (Huang et al. 2003; Stegemann et al. 2003). Large blocks of mtDNA sequences are also found in the nuclear chromosomes of Arabidopsis, rice, yeast, and human (Blanchard and Schmidt 1996; Noutsos et al. 2005; Richly and Leister 2004a; Woischnik and Moraes 2002). Therefore, plant nuclear genomes acquire a large number of genes from endosymbiotic organelles, accompanied by frequent and numerous gene gains and losses in organellar genomes, especially in mitochondria (Adams and Palmer 2003; Adams et al. 2002; Martin and Herrmann 1998; Millen et al. 2001; Palmer et al. 2000). For instance, genes encoding ribosomal proteins are frequently lost from mitochondrial genomes, among which the rps10 gene was detected to have been lost over 20 times among 281 angiosperms, as well as the cox2 gene, which has recently been found transferring from mitochondria to the nucleus in legumes (Palmer et al. 2000).
The nuclear genome of flowering plants obtained a variable amount of tRNA genes originated from endosymbiotic organelles, following sequence transfer events during their long history of evolution. It is important to give a detailed documentation of tRNA gene transfer for species whose genome sequence becomes available. Based on the complete genome sequence of a indica rice (Yu et al. 2002; Yu et al. 2005) and those of its mitochondrial (Tian et al. 2006) and plastid (Tang et al. 2004) genomes, here we draw a comprehensive map of the tRNA gene transfer to provide detailed evidence and insights into this phenomenon in flowering plants.
Materials and Methods
Identification of Transferred tRNA Genes in Rice Nuclear and Organellar Genomes
tRNA genes were all predicted with the program tRNAscan-SE (Lowe and Eddy 1997). Potential genes with empirical scores >50 and canonical anticodons were considered to be genuine tRNA genes; and those that had sequence and structure similarities to the genuine tRNA genes but were functionally inactive due to indels (insertions and deletions) or lacked functional promoters were referred to as pseudogenes. To improve the reliability of the prediction, we also performed a BLASTN search against the nonredundant database of GenBank. To further confirm the predicted organellar tRNA genes, we also compared them with as many orthologous sequences as possible from other plants retrieved from databases of OGMP (the Organelle Genome Megasequencing Program; http://www.megasun.bch.umontreal.ca/ogmpproj.html) and PLMItRNA (Rainaldi et al. 2003).
To identify organelle-like tRNA genes from nuclear genomes, we performed similarity searches with BLASTN (Altschul et al. 1990), with an e value of 10 for the parameter of maximum expectation and default values for all other parameters. To search NUPTs, NUMTs, or MTPTs, we performed sequence alignments among rice plastid, mitochondrial, and nuclear genomes with BLASTN under parameter settings similar to those for tRNA searches. The genome data used in this work include (i) the improved whole-genome shotgun sequence assemblies (accession number AAAA02000000) and (ii) the complete genome sequences of the plastid (accession number AY522329) and mitochondrion (accession number DQ167399) of the rice Beijing indica.
Phylogenetic Analysis
Phylogenetic relationships were determined for homologous tRNA genes present in rice plastid, mitochondrial, and/or nuclear genomes, along with relevant orthologous sequences retrieved from GenBank, which include those from other higher and lower plants, as well as green algae and red algae. Multiple sequence alignments were performed using the program CLUSTAL_W version 1.83 (Thompson et al. 1994). Phylogenetic relationships were determined by maximum likelihood analyses using the program PHYML version 2.4.4 (Guindon and Gascuel 2003; Guindon et al. 2005). Parameters of the maximum likelihood model were set as follows: four nucleotide substitution rate categories as the default; the HKY model (Hasegawa et al. 1985) for nucleotide substitution; d alpha, the value of the gamma distribution shape parameter, optimally estimated by maximizing the likelihood of the phylogeny. All these initial parameters were implemented in the heuristic search for estimating maximum likelihood phylogenies. For the final tree, 100 bootstrapped replicates were examined, and a consensus tree was constructed using the program CONSENSE of the PHYLIP package (Felsenstein 1993). All trees were displayed and compiled manually using the program TreeExplorer of MEGA version 3.1 (Kumar et al. 2004).
Results
tRNA Gene Transfer from the Plastid Genome to the Nuclear Genome
The rice plastid genome comprises 31 tRNA genes, including 3 pseudogenes and a complete set of tRNAs for all amino acids. Chromosomal localization of tRNA genes and sequence comparisons between the plastid and the nuclear genomes revealed that 25 of the plastid tRNA (pt-tRNA) genes have excellent sequence similarities (95%–100%) with the nuclear counterparts or poorer sequence similarities but longer NUPTs (>150 bp). The rice nuclear genome possesses 81 pt-like tRNA genes, accounting for 12.2% (564 native nuclear tRNA genes have been identified from indica rice [Wang et al. 2004]) of the tRNA gene content of the nuclear genome (Fig. 1).
Diagram showing the proposed pathways of tRNA gene transfer from organelles to the nucleus and between organelles. Arrows show the directions of tRNA gene transfer from the plastid (green) to the mitochondrion and the nucleus, and from the mitochondrion (blue) to the nucleus. The lines ending in circles indicate nuclear tRNA genes homologous to the connected organellar counterparts but without supporting evidence that transfer from the organelles actually occurred. The dashed lines link uninterpretable pathways of tRNA gene transfer. The numbers are copies of tRNA genes in the nucleus, plastid, and mitochondrion, which are grouped as unique and shared copies among all three genomes.
A full-scale analysis of the pt-like tRNA genes in the rice genome revealed several significant features that are unique to rice in contrast to other flowering plants. pt-like tRNA genes are distributed across all 12 chromosomes (Fig. 2A and Supplementary Fig. S1). Chromosomes 1, 3, 5, and 10, each with 10 pt-like tRNA genes, are the popular targets of pt-tRNA, as opposed to chromosomes 7, 8, and 9, each containing fewer than three pt-like tRNA genes. More than 92% of pt-like tRNA genes are discovered from NUPTs, with a sequence length of >150 bp and identities ranging from 84% to 100% compared with ptDNAs. The trnH GTG gene is the most frequently transferred one among all pt-like tRNA genes. Nine copies of trnH GTG are found on six chromosomes, with transfer fragments (NUPTs) varying from 204 to 2966 bp. The identities between pt-trnH GTG and NUPTs vary from 98% to 100%. In addition, six clusters of cp-like tRNA genes are identified on 11 NUPTs: trnW CCA-trnP TGG on Chr1 and Chr10, trnH GTG-trnI CAT on Chr3 (two copies) and Chr11, trnI CAT-trnL CAA on Chr4 (2 copies), trnN GTT-trnR ACG on Chr5 (2 copies), and trnG GCC-trnS TGA on Chr12 (Supplementary Table S1).
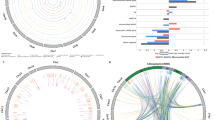
Distributions of mitochondrion-like and plastid-like tRNA genes on nuclear chromosomes (1, 3, 5, 7, 9, and 11; A) and between organellar genomes (B). In A, positions of nuclear chromosomes (Y-axis; color-coded) and plastid (upper X-axis) and mitochondrial (bottom X-axis) genomes are indicated. Colored solid circles and crosses indicate mt-like and pt-like tRNA genes, respectively. The horizontal bars mark repeat regions (>5 kb; blue, cyan, green, red, and brown for different types) of the mitochondrial and plastid genomes, as well as the native (brown) and organelle-like tRNA genes (green). Solid circles with short vertical bars anchored on the X-axis represent tRNA genes with counterparts in the nuclear genome. In B, solid magenta circles indicate the positions of plastid-like tRNA genes in the two organellar genomes. Solid black circles indicate similar DNA segments (with >90% similarity and >100-bp length) between the organellar genomes. The tRNA genes and their pseudogenes (gray) or truncated forms (asterisks) are also indicated. Sequence repeats (>5 kb) are also color-coded (as in A) as vertical or horizontal bars next to the X- and Y-axes.
tRNA Gene Transfer from the Mitochondrial Genome to the Nuclear Genome
In the nuclear genome, 43 intact mt-like tRNA genes are identified from all 12 chromosomes, constituting approximately 6.6% of the tRNA gene content in the rice nuclear genome (Fig. 2A and Supplementary Fig. S1). These mt-like tRNA genes are located on NUMTs, with sequence lengths varying from 82 to 8423 bp. The identities between mt-like nuclear tRNA genes and their mitochondrial counterparts vary from 84% to 100%, while those between the NUMTs and the mitochondrial sequences vary to a slightly larger extent (82% to 100%; Supplementary Table S1).
All mt-like nuclear tRNA genes are homologous to 18 (of 22) of full-length mt-tRNA genes, among which 9 are pt-like (Table 1). In other words, all pt-like mt-tRNA genes have homologous counterparts in the rice nuclear genome (Fig. 1). Did these tRNA genes transfer from mitochondria to the nucleus or from plastids to the nucleus? Our analysis of the corresponding MTPTs, NUMTs, and NUPTs containing these tRNA genes revealed that (i) the NUMTs of seven tRNA genes are larger than their NUPTs in sequence size, suggesting that these tRNA genes may experience a process—originating in the plastid, subsequently migrating to the mitochondrion, and, finally, being integrated into the nuclear chromosomes; and (ii) 19 tRNA genes are located on NUPTs larger than NUMTs, implying that these tRNA genes originated in the plastid and subsequently migrated to the nuclear genome without the mitochondrial intermediates. The organelle-like nuclear tRNA genes, such as trnC GCA, trnN GTT, trnP TCT, and trnW CCA, all have both mitochondrial and plastid homologues that probably migrated via two different routes, direct or mitochondrion-mediated, from the plastid to the nucleus. In addition, four copies of trnM CAT and two copies of trnH GTG are located on NUMTs of the same size as their NUPTs, therefore their transfer routes are uninterpretable based solely on the current information.
The remaining nine native mt-tRNA genes have 11 homologues in the nuclear genome (Supplementary Table S1). Namely, these tRNA genes were mitochondrion-derived, and among them, five mt-like tRNA genes (trnP TGG, trnQ TTG, trnS GCT, trnS TGA, and trnY GTA) coexist with nucleus-native and pt-like homologues in the rice nuclear genome. Although these tRNAs account for identical functions in carrying amino acids, they vary greatly in their DNA sequences in the rice nuclear genome. Therefore, the rice nucleus encompasses trnP TGG, trnQ TTG, trnS GCT, trnS TGA, and trnY GTA with different originations: plastid-derived, mitochondrion-derived, and nucleus vertically inherited.
pt-like tRNA Genes in the Mitochondrial Genome
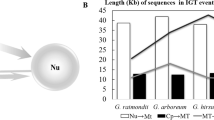
The number and species of pt-like tRNA genes in mitochondrial genomes have been investigated in various flowering plants (Table 1), and here we attempt to address the transfer routes based on the distribution of pt-like mt-tRNA genes and MTPTs. The size and number of MTPTs appear to be larger in rice and maize than in Arabidopsis and tobacco, respectively, in sharp contrast to the number and species of pt-like mt-tRNA genes. Several subtle differences also exist in these four higher plants. For instances, two-thirds of pt-like mt-tRNA genes are located in the repeat regions of the rice mitochondrial genome, as opposed to all their plastid counterparts, which are distributed randomly on the plastid genome (Fig. 2B and Supplementary Table S2). Moreover, in maize almost all pt-like mt-tRNA genes seem to transfer from inverted repeat regions of its plastid genome and be randomly scattered over its mitochondrial genome (Supplementary Fig. S2, A). In Arabidopsis and tobacco, however, the pt-like mt-tRNA genes are distributed in a very similar way but with different patterns, in terms of gene order and interspaces (Supplementary Figs. S2, B and C). Different distribution patterns of pt-like mt-tRNA genes can almost certainly be attributed to tandem duplications, extraneous DNA migrations, and integration-reintegration cycles occurring in mitochondrial genomes of flowering plants, especially after the divergence of dicotyledons and monocotyledons.
tRNA Gene Transfer Through a DNA-Mediated Pathway
Gene transfer from organelles to the nucleus in general follows two different molecular pathways: DNA-mediated (Lopez et al. 1994) and RNA-mediated (Nugent and Palmer 1991). Our study supports the DNA-mediated pathway and the bulk DNA hypothesis. Of all nine pt-like mt-tRNA genes, seven are located on MTPT fragments longer than 150 bp. Likewise, among all mt-like (43 copies) and pt-like (80 copies) nuclear tRNA genes, only 7 copies are located on NUMTs and NUPTs of <150 bp (from 82 to 117 bp) (Supplementary Table S1). After further investigation of tRNA genes with NUMTs, NUPTs, or MTPTs >150 bp and even on their 5′- and 3′-flanking sequences, we reached the result that almost all flanking fragments of transferred tRNA genes contain intergenic regions, noncoding regions, and introns that are similar to their source organellar genomes in gene order and interspaces.
Phylogenetic Analysis of Transferred tRNA Genes
Organelle-like genes are usually considered to be molecular fossils in nuclear genomes for evolutionary relationship analyses (Bensasson et al. 2001; Timmis et al. 2004; Woischnik and Moraes 2002). In phylogenetic analysis of transferred tRNA genes, we aligned sequences of rice nuclear pt-like and/or mt-like tRNA genes with corresponding organellar tRNA genes from rice and several other distant plant species, including lower plants, green alga, and red alga. Subsequently, the phylogenetic trees were constructed based on the maximum likelihood method (detailed under Materials and Methods).
Since the current data collection is not adequate to cover enough variations and to yield fully interpretable topologies, we focus our analysis on a few tRNA genes. First, phylogenetic trees of trnH GTG and trnN GTT show a confinement of all organelle-like nuclear genes, which cluster within branches of flowering plants, suggesting that the relevant transfer events occurred only in these plants. In other words, some of the transferred genes have obvious lineage-specific signatures (Figs. 3A and B). Second, these genes are not simply concentrated around branches of mitochondria or plastids of different origins, but resolved to modules that show organelle specificity. Third, the gene transfer events seem to happen several times, as indicated by other phylogenetic trees for the tRNA genes, such as those for trnR TCT trnW CCA, trnV GAC, trnR TCT, and trnS GCT (Supplementary Fig. S3), together with trnH GTG and trnN GTT, all of which have counterparts in the mitochondrial or plastid genomes (Supplementary Table S1). Fourth, some of the transferred tRNA genes show single organellar origins, such as trnY GTA and trnS TGA (Figs. 3C and D), both of which are native mt-tRNA genes, compared with trnH GTG and trnN GTT, both of which are pt-like mt-tRNA genes.
Phylogenetic trees of trnH GTG (A), trnS TGA (B), trnY GTA (C), and trnS GCT (D) from organellar genomes and their nuclear counterparts. Trees were constructed using the maximum likelihood method. Bootstrap values, calculated from 100 repetitions, are labeled at branching nodes. Rice organelle-like nuclear genes are labeled with their chromosome positions, and ptL, mtL, and un in parentheses depict plastid-like, mitochondrion-like, and uninterpretable tRNA genes, respectively. Organellar genes are labeled with their species names and origins. For mt-tRNA genes, N and ptL indicate the origins as native and plastid-like, respectively. The plastid and mitochondrial tRNA gene sequences retrieved from GenBank and OGMP are as follows: (1) monocotyledons—rice (Oryza sativa; AY522329) and maize (Zea mays; AY506529, NC_001666); (2) dicotyledons—Arabdiposis (Arabidopsis thaliana; NC_001284, NC_000932), tobacco (Nicotiana tabacum; NC_006581, NC_001879), and sugar beet (Beet vulgaris; NC_002511); (3) liverwort (Marchanta polymorpha, NC_001660, NC_001319); (4) Chlorophyta—Chara (Chara vulgaris; AY267353) and Nephroselmis (Nephroselmis olivacea; AF110138, NC_000927); and (5) Rhodophyta—Porphyra (Porphyra purpurea; NC_002007, NC_000925) and Cyanidioschyzon (Cyanidioschyzon merolae; NC_000887, NC_004799).
Discussion
Overview of Flowering Plant tRNA Genes
Flowering plants contain an incomplete set of tRNA genes for protein syntheses inside mitochondria. However , a full set of tRNA is absolutely necessary for carrying all amino acids, so the missing tRNA genes in mitochondria could be supplied by the nucleus (Dietrich et al. 1992; Marechal-Drouard et al. 1993; Veronico et al. 1996). Furthermore, contemporary mt-tRNA genes have a distinct genesis in flowering plants, either from their vertical ancestors or from endosymbiotic gene transfers. The species and the number of pt-like mt-tRNA genes varied widely among flowering plants, showing a clear trend toward losing native tRNAs and continuing transfer events among the organellar and nuclear genomes (Kumar et al. 1996; Small et al. 1999).
The rice mitochondrion contains nine full-length pt-like tRNA genes, accounting for one-third of its tRNA gene content. Likewise, of the rice nuclear tRNA gene content, 91 tRNA genes are cp-like or mt-like, among which 32 copies are homologous to its pt-like mt-tRNA genes. The 5′- and 3′-flanking sequences of these 32 tRNA genes show that (i) the majority (21 copies) of them follow a transfer route from the plastid to the nucleus, rather than from the mitochondrion to the nucleus; (ii) six copies appear to be constantly moving around in the rice endosymbiotic environment, transferring from the plastid to the mitochondrion and subsequently to the nucleus; and (iii) transfer routes of the rest (five copies) are uninterpretable at present.
NUMTs/NUPTs/MTPTs containing tRNA genes indicate a consistent range of identities from 82% to 100% (82 to 8243 bp in sequence length) compared to their source organelles. These data suggest that tRNA genes transfer endosymbiotically under an ongoing process, despite several exceptions in NUMTs and NUPTs of other eukaryotic species (Blanchard and Schmidt 1996; Richly and Leister 2004a, b; Woischnik and Moraes 2002). The high identities of NUMTs/NUPTs/MTPTs to their source organellar DNA suggest that the tRNA genes located on them may have transferred in the recent past. The low identities reflect a higher mutation rate of the nuclear genomes compared with organellar genomes after tRNA genes remained in the nuclear genomes (Palmer et al. 2000; Tang et al. 2004).
Mechanisms and Frequencies of tRNA Gene Transfer
Analyzing the rice data, we discovered noncoding, intronic, and intergenic sequences of the original genomes in the majority of NUPTs, NUMTs, and MTPTs with sequence lengths >150 bp. The information-rich sequences provide evidence supporting the bulk-DNA mechanism, despite a few exceptions where several tRNA genes are located on short transfer fragments (<150 bp). Meanwhile, we could not find sufficient information for the cDNA-mediated transfer model.
The transfer frequency from organelles to the nucleus is rather distinct for different tRNA genes. Several nuclear tRNA genes possess more than five copies of organellar counterparts, such as trnH GTG, trnI CAT, trnL CAA, trnP TGG, and trnR ACG, whereas others barely have a single organellar copy. The higher copy number of an organelle-like tRNA gene in the nuclear genome may be attributed to relatively frequent duplications of nuclear DNA after the migration of organellar DNA and multiple transfer events of an individual tRNA gene. If the higher copy number of an organelle-like tRNA gene is generated by genome duplications (Wolfe and Shields 1997; Yu et al. 2005), the length and the similarity of the corresponding NUMT/NUPT should be nearly identical for a given tRNA gene. Otherwise, the high frequency may be a consequence of continuing transfer events occurring at different stages of evolutionary processes.
The phylogenetic analyses also revealed powerful signatures of tRNA gene transfer, especially when the available data were genome-wide and capable of comparison with other flowering plants, as well as lower plants, green alga, and red alga. In our data, the trees for trnH GTG and trnN GTT genes show that some of the pt-like nuclear tRNA genes are clustered within a branch of the rice plastid genes, whereas several others branched out and were separated into other flowering plants. The clustered group in this case not only suggests the frequency of the transfer events, but also shows the possible timing in an evolutionary context. The branching-out events support the idea that independent transfer events may have occurred at different stages of flowering plant evolution. In addition, we must consider the different mutation spectra and rates of the nuclear, plastid, and mitochondrial genomes. Mutation rates are much higher in the nucleus than in the plastid and mitochondrion of rice and other flowering plants (Cho et al. 2004; Palmer et al. 2000; Tang et al. 2004; Tian et al. 2006; Yu et al. 2002), so the topology of different phylogenetic trees built with data from the different genomes has to be interpreted with caution.
Expression of Transferred tRNA Genes
tRNA genes located on transferred ptDNA provide the only instance where pt-like tRNA genes are actually capable of expression in mitochondria of a flowering plant (Fey et al. 1997). Can all pt-like tRNA genes transcribe and mature to become functional in mitochondria, and can any organelle-like tRNA genes in the nucleus? Miyata and colleagues proved that seven pt-like tRNA genes (trnC, trnF, trnH, trnM, trnN, trnS, and trnW) were transcribed and precisely matured in the rice mitochondrion (Miyata et al. 1998). Likewise, active pt-like mt-tRNA genes have been found in other flowering plants, such as potato mt-tRNA trnH, trnM, trnN, trnS, and trnW (Fey et al. 1997; Marechal-Drouard et al. 1990) and wheat mt-tRNA trnC, trnF, trnM, trnN, trnS and trnW (Ceci et al. 1993; Joyce and Gray 1989). For activation of transferred tRNA genes in a target genome, the following two simple criteria are potentially satisfied: maintaining functional promoters from their original ancestors after transfer and acquiring the regulatory elements from the target genome. We believe that the expressed tRNA genes in the former case are probably those with longer flanking sequences at both ends, and those fitting the latter case must have shorter MTPTs. However, it is difficult to draw reliable conclusions without experimental verification. A complete picture of tRNA gene transfer and its rules among flowering plants and their lower relatives is yet to be painted when more genome sequences become available.
References
Bensasson D, Zhang D, Hartl DL, Hewitt GM (2001) Mitochondrial pseudogenes: evolution’s misplaced witnesses. Trends Ecol Evol 16:314–321
Blanchard JL, Schmidt GW (1996) Mitochondrial DNA migration events in yeast and humans: integration by a common end-joining mechanism and alternative perspectives on nucleotide substitution patterns. Mol Biol Evol 13:893
Ceci LR, Saiardi A, Siculella L, Quagliariello C (1993) A tRNA(Val) (GAC) gene of chloroplast origin in sunflower mitochondria is not transcribed. Plant Mol Biol 23:727–736
Cho Y, Mower JP, Qiu YL, Palmer JD (2004) Mitochondrial substitution rates are extraordinarily elevated and variable in a genus of flowering plants. Proc Natl Acad Sci USA 101:17741–17746
Dietrich A, Weil JH, Marechal-Drouard L (1992) Nuclear-encoded transfer RNAs in plant mitochondria. Annu Rev Cell Biol 8:115–131
Fey J, Dietrich A, Cosset A, Desprez T, Marechal-Drouard L (1997) Evolutionary aspects of “chloroplast-like” trnN and trnH expression in higher-plant mitochondria. Curr Genet 32:358–360
Joyce PB, Gray MW (1989) Chloroplast-like transfer RNA genes expressed in wheat mitochondria. Nucleic Acids Res 17:5461–5476
Kumar R, Marechal-Drouard L, Akama K, Small I (1996) Striking differences in mitochondrial tRNA import between different plant species. Mol Gen Genet 252:404–411
Lopez JV, Yuhki N, Masuda R, Modi W, O’Brien SJ (1994) Numt, a recent transfer and tandem amplification of mitochondrial DNA to the nuclear genome of the domestic cat. J Mol Evol 39:174–190
Marechal-Drouard L, Guillemaut P, Cosset A, Arbogast M, Weber F, Weil JH, Dietrich A (1990) Transfer RNAs of potato (Solanum tuberosum) mitochondria have different genetic origins. Nucleic Acids Res 18:3689–3696
Marechal-Drouard L, Weil JH, Dietrich A (1993) Transfer RNAs and transfer RNA genes in plants. Annu Rev Plant Physiol Plant Mol Biol 44:13–32
Miyata S, Nakazono M, Hirai A (1998) Transcription of plastid-derived tRNA genes in rice mitochondria. Curr Genet 34:216–220
Nugent JM, Palmer JD (1991) RNA-mediated transfer of the gene coxII from the mitochondrion to the nucleus during flowering plant evolution. Cell 66:473–481
Palmer JD, Adams KL, Cho Y, Parkinson CL, Qiu YL, Song K (2000) Dynamic evolution of plant mitochondrial genomes: mobile genes and introns and highly variable mutation rates. Proc Natl Acad Sci USA 97:6960–6966
Richly E, Leister D (2004a) NUMTs in sequenced eukaryotic genomes. Mol Biol Evol 21:1081–1084
Richly E, Leister D (2004b) NUPTs in sequenced eukaryotes and their genomic organization in relation to NUMTs. Mol Biol Evol 21:1972–1980
Small I, Adams KL, Chapron A, Dietrich A, Duchene AM, Lancelin D, Marechal-Drouard L, Menand B, Mireau H, Moudden Y, Ovesna J, Peeters N, Sakamoto W, Souciet G, Wintz H (1999) The strange evolutionary history of plant mitochondiral tRNAs and their aminoacyl-tRNA synthetases. J Hered 90:333–337
Tang J, Xia H, Cao M, Zhang X, Zeng W, Hu S, Tong W, Wang J, Yu J, Yang H, Zhu L (2004) A comparison of rice chloroplast genomes. Plant Physiol 135:412–420
Tian X, Zheng J, Hu S, Yu J (2006) The rice mitochondrial genomes and their variations. Plant Physiol 140:401–410
Timmis JN, Ayliffe MA, Huang CY, Martin W (2004) Endosymbiotic gene transfer: organelle genomes forge eukaryotic chromosomes. Nat Rev Genet 5:123–135
Veronico P, Gallerani R, Ceci LR (1996) Compilation and classification of higher plant mitochondrial tRNA genes. Nucleic Acids Res 24:2199–2203
Woischnik M, Moraes CT (2002) Pattern of organization of human mitochondrial pseudogenes in the nuclear genome. Genome Res 12:885–893
Wolfe KH, Shields DC (1997) Molecular evidence for an ancient duplication of the entire yeast genome. Nature 387:708–713
Yu J, Hu S, Wang J, Wong GK, Li S, Liu B, Deng Y, Dai L, Zhou Y, Zhang X, Cao M, Liu J, Sun J, Tang J, Chen Y, Huang X, Lin W, Ye C, Tong W, Cong L, Geng J, Han Y, Li L, Li W, Hu G, Li J, Liu Z, Qi Q, Li T, Wang X, Lu H, Wu T, Zhu M, Ni P, Han H, Dong W, Ren X, Feng X, Cui P, Li X, Wang H, Xu X, Zhai W, Xu Z, Zhang J, He S, Xu J, Zhang K, Zheng X, Dong J, Zeng W, Tao L, Ye J, Tan J, Chen X, He J, Liu D, Tian W, Tian C, Xia H, Bao Q, Li G, Gao H, Cao T, Zhao W, Li P, Chen W, Zhang Y, Hu J, Liu S, Yang J, Zhang G, Xiong Y, Li Z, Mao L, Zhou C, Zhu Z, Chen R, Hao B, Zheng W, Chen S, Guo W, Tao M, Zhu L, Yuan L, Yang H (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. indica). Science 296:79–92
Yu J, Wang J, Lin W, Li S, Li H, Zhou J, Ni P, Dong W, Hu S, Zeng C, Zhang J, Zhang Y, Li R, Xu Z, Li X, Zheng H, Cong L, Lin L, Yin J, Geng J, Li G, Shi J, Liu J, Lv H, Li J, Deng Y, Ran L, Shi X, Wang X, Wu Q, Li C, Ren X, Li D, Liu D, Zhang X, Ji Z, Zhao W, Sun Y, Zhang Z, Bao J, Han Y, Dong L, Ji J, Chen P, Wu S, Xiao Y, Bu D, Tan J, Yang L, Ye C, Xu J, Zhou Y, Yu Y, Zhang B, Zhuang S, Wei H, Liu B, Lei M, Yu H, Li Y, Xu H, Wei S, He X, Fang L, Huang X, Su Z, Tong W, Tong Z, Ye J, Wang L, Lei T, Chen C, Chen H, Huang H, Zhang F, Li N, Zhao C, Huang Y, Li L, Xi Y, Qi Q, Li W, Hu W, Tian X, Jiao Y, Liang X, Jin J, Gao L, Zheng W, Hao B, Liu S, Wang W, Yuan L, Cao M, McDermott J, Samudrala R, Wong GK, Yang H (2005) The genomes of Oryza sativa: a history of duplications. PLoS Biol 3:e38
Acknowledgments
This work was supported by grants from the Chinese Academy of Sciences (CAS; KSCX1-SW-03), the Ministry of Science and Technology (2004AA231050 and 2005AA235110), and the CAS Hundred Talents Program (to Jun Yu). We thank Dr. Xiyin Wang (Center of Bioinformatics, Peking University, China) for providing information on tRNA genes of the rice 93-11 nuclear genome, and we are grateful to Dr. Chunyuan Huang (School of Molecular and Biomedical Science, The University of Adelaide, Australia) and Dr. Jingui Zhu (Beijing Institute of Genomics, Chinese Academy of Sciences, China) for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Reviewing Editor: Dr. David Guttman
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Tian, X., Zheng, J., Hu, S. et al. The Discriminatory Transfer Routes of tRNA Genes Among Organellar and Nuclear Genomes in Flowering Plants: A Genome-Wide Investigation of indica Rice. J Mol Evol 64, 299–307 (2007). https://doi.org/10.1007/s00239-005-0200-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-005-0200-6