Abstract
Bacterial chromosomes are immense polymers whose faithful replication and segregation are crucial to cell survival. The ability of proteins such as FtsK to move unidirectionally toward the replication terminus, and direct DNA translocation into the appropriate daughter cell during cell division, requires that bacterial genomes maintain an architecture for the orderly replication and segregation of chromosomes. We suggest that proteins that locate the replication terminus exploit strand-biased sequences that are overrepresented on one DNA strand, and that selection increases with decreased distance to the replication terminus. We report a generalized method for detecting these architecture imparting sequences (AIMS) and have identified AIMS in nearly all bacterial genomes. Their increased abundance on leading strands and decreased abundance on lagging strands toward replication termini are not the result of changes in mutational bias; rather, they reflect a gradient of long-term positive selection for AIMS. The maintenance of the pattern of AIMS across the genomes of related bacteria independent of their positions within individual genes suggests a well-conserved role in genome biology. The stable gradient of AIMS abundance from replication origin to terminus suggests that the replicore acts as a target of selection, where selection for chromosome architecture results in the maintenance of gene order and in the lack of high-frequency DNA inversion within replicores.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacterial chromosomes are not simply collections of genes; these polymers—up to 100,000 times longer than the cells that contain them—are organized into highly compacted nucleoids (Holmes and Cozzarelli 2000; Wu 2004) as supercoiled domains (Deng et al. 2004; Higgins et al. 1996; Stein et al. 2005), are positioned at defined locations within the cytoplasm (Gitai et al. 2005; Niki et al. 2000; Teleman et al. 1998; Wu and Errington 1998), experience intricately timed replication (Cunningham and Berger 2005), and move through the cytoplasm in precise, choreographed ways (Viollier and Shapiro 2004; Viollier et al. 2004). Beyond encoding thousands of protein and RNA products, as well as signals for their production, DNA molecules must also carry information that controls the tempo and mode of their own replication and segregation into daughter cells. While numerous genetic, molecular biological, and bioinformatic techniques serve to identify DNA sequences that are important because of the products they encode (that is, genes), finding sequences that are important for the maintenance of the DNA molecule itself has proven to be more difficult.
Global chromosome structure is suggested by the nonrandom distribution of genes within replicores. Single replication origins and termini typically apportion bacterial genes nearly symmetrically into two approximately equally sized replicores. The location of some genes relative to the replication origin is known to be important—e.g., the proximity of the Bacillus subtilus spoIIR gene to the replication origin allows its transcription from the newly formed forespore, whereas the origin-distal location of the spoIIAB gene prevents its encapsulation in the forespore, allowing for σF activation there (Dworkin and Losick 2001). Furthermore, dnaA genes are significantly associated with the replication origins. Outside of such special cases, little significance to the positions of other transcription units relative to the replication origin has been postulated beyond the potential for greater gene dosage of origin-proximal genes (Liu and Sanderson 1995b, 1996). Yet we can infer that gene order is constrained, since genetic maps retain order in the face of mechanisms that can rearrange them. More importantly, observed rearrangements are most often symmetrical with respect to replication origins and termini (Eisen et al. 2000; Mackiewicz et al. 2001; Sanderson and Liu 1998; Suyama and Bork 2001; Tillier and Collins 2000), suggesting that inversions that rearrange chromosome structure (i.e., those that move genes from leading to lagging strands) are counterselected.
Beyond the distribution of genes, the nonrandom distribution of certain oligomeric sequences is also consistent with global chromosome structure. One example is the χ recombination signal (Eggleston and West 1997; Kowalczykowski et al. 1994; Kuzminov 1995; Myers and Stahl 1994); this octamer is highly abundant on leading strands (El Karoui et al. 1999; Uno et al. 2000) and serves to disable the RecD exonuclease, allowing the RecBC recombinase to repair double-stranded breaks efficiently via homologous recombination. The overabundance of χ sequences is consistent with their origin by mutational biases and maintenance by selection for function. That is, the signals that mediate global chromosome architecture could arise by mutational bias, where consistent replication from a single origin allows for differences to accumulate between leading and lagging strands (Lobry 1996; Lobry and Louarn 2003; Salzberg et al. 1998). Once placed under selection, differences between strands that arise by chance would be maintained, and disruption of these patterns would be detrimental (Capiaux et al. 2001; Corre et al. 2000). That is, for the overabundance of χ sequences on leading strands to be identifiable, global chromosome structure must exist.
The processes described above—replication termination and DNA segregation—involve the action of proteins at the replication terminus. Therefore, sequences enabling proteins to locate the replication terminus are good candidates for those contributing to chromosome architecture. One may expect these sequences to accumulate near the replication terminus since it is there that selection for their function would be greatest. For example, the FtsK protein translocates along DNA toward the dif site at the replication terminus (Pease et al. 2005) and may mediate segregation of chromosomes across the septum (Lau et al. 2003). FtsK delivers the XerCD recombinase to the dif site (Bigot et al. 2004; Ip et al. 2003; Li et al. 2003; Massey et al. 2004), where it acts to resolve entangled chromosomes during cell division (Blakely et al. 1991; Clerget 1991). The FtsK protein must recognize strand-specific sequences to enable its directional movement toward the replication terminus. Since the frequency at which DNA translocases act is inversely proportional to the distance from the replication terminus, sequences would be under strongest selection—and therefore at highest abundance on their preferred strand—near the terminus. This increase in abundance toward the replication terminus—beyond what would be predicted by changes in mutational bias (Daubin and Perriére 2003)—can be taken as evidence for selection. Here, we describe methods for detecting such sequences and demonstrate that their distributions did not result from mutational biases or chance.
Materials and Methods
Sequence Analysis
Sequences were downloaded from GenBank and analyzed using DNA Master (cobamide2.bio.pitt.edu).
Nucleotide skew was calculated as (G – C)/(G + C) or (A – T)/(A + T) at the third codon positions of protein-coding genes, corrected for the direction of transcription. Global pairwise sequence alignments used the method of Needleman and Wunsch (1970); alignment scores were obtained using the PAM 250 matrix (Altschul 1991), normalized to the average length of the genes being compared. Octamers were classified as matching IUB nondegenerate (GATC) and degenerate (RYMK) bases. Watson strands are defined as the DNA strands—read 5′ to 3′—reported in GenBank files; Crick strands are defined as the complements of Watson strands. Leading strands are defined as Watson strands downstream, and Crick strands upstream, of the replication origin. Skewed octamers were detected as those sequences overrepresented on leading strands; asymmetrically distributed octamers were detected as sequences present in a particular region of a replicore at a significantly higher abundance than predicted from their abundance in the remainder of the replicore as measured by χ2 analysis.
Number of Sequences Defining the Replication Origin or Terminus
Leading strands correspond to the Watson strands on one side of the replication origin or terminus, and to Crick strands on the other side. To locate these positions, a sliding-window analysis was performed, where windows were defined as encompassing 80% of a bacterial genome sequence, centered on a potential “break point.” Strand-biased octamers defining a break point were enumerated as those that were overrepresented on the Watson strand upstream of the break point but overrepresented on the Crick strand downstream of the break point. The numbers of biased octamers would be maximal when the break points lie close to either the replication origin or terminus, where Watson strands change from leading strands to lagging strands.
Detection of Large Inversions and Insertions
Bacterial genomes were divided into segments (typically 10–100 kilobases [kb] in length) that were analyzed independently for strand-biased octamers. The libraries of strand-biased octamers generated for each genome segment were compared to each other. If similar sequences were biased on the Watson strands of both segments, these regions were viewed as being historically replicated in the same direction; if similar sequences were biased on the Watson strand of one segment and the Crick strand of another, these regions were viewed as being historically replicated in opposite directions. Pairwise similarity of octamer libraries was calculated as the Jaccard (1912) coefficient of similarity, SJ. Most genomes could be described as having two large domains, where the Watson strands of segments in one domain were biased in a way similar to the Crick strands of segments in the other domain. Large inversions that did not include the origin or replication terminus were detected as regions where the strand bias of the Crick strand resembled the strand bias of the Watson strand of neighboring segments. Large insertions of foreign DNA—whose strand bias would be different from the remainder of the genome—were detected as regions where the libraries of strand-biased octamers resembled neither the Watson strand nor the Crick strand libraries of any chromosome segment. This pattern would also be reflected in old inversions that had begun to ameliorate their nucleotide composition (Lawrence and Ochman 1997).
Sequences Accumulating Near the Replication Terminus
Octamers that accumulated in abundance toward the replication terminus were initially detected as those that (a) exceeded 100 copies per genome, typically numbering at least one sequence per 10 kb of genomic sequence, (b) were overrepresented on the leading strand, where typically > 70% of the sequences were found on the leading strand, (c) showed abundance in a terminus-proximal window—typically defined as 10 to 25% of the genome length— that exceeded that predicted based on its abundance elsewhere, and (d) showed this pattern on both replicores. Consistent increase in abundance toward the replication terminus was verified by regression of local octamer abundance against distance from the terminus.
Correction for Mutational Bias
The abundances of nucleotides, dinucleotides, trinucleotides, and tetranucleotides were calculated by sliding-window analysis. The expected local abundance of octamers was calculated from the relative abundance of constituent nucleotides, dinucleotides, trinucleotides, or tetranucleotides. For n-mers of length j, where j < 8, the expected frequency of an octamer E j given the abundance of constituent j-mers is defined as
where \( O^{j}_{i} \) is the frequency of the suboligomer of length j at position i within the octamer. Therefore, the expected frequency (E) of an octamer based on the constituent tetramers is calculated as E ABCDEFGH = (P ABCD P BCDE P CDEF P DEFG P EFGH)/(P BCD P CDE P DEF P EFG).
Location of Maximum Octamer Abundance
Sequences that accumulate toward the replication terminus were initially identified via their overabundance in the region adjacent to the terminus. Linear regression was then used to determine if their abundance increased toward the replication terminus. To identify sequences which may accumulate to other, nonterminus locations within the chromosome (either by chance or by selection), we found those that were overabundant in sequence windows away from the terminus. A quadratic regression of the local abundance of octamers against distance of the region from the terminus was then performed. Sequences which reached maximal abundance at a position away from the terminus would show a local maximum (the peak of the parabola) away from the terminus.
Results
Identification of Replication Origins and Termini
To identify sequences imparting chromosome architecture, replication origins and termini must be identified in a robust fashion that is consistent across genomes. Consistent replication initiation and termination at defined points results in strand biases due to the mutational differences between leading and lagging strands. Replication origins and termini can be detected as points of inflection in cumulative nucleotide skew plots (Lobry 1996), where a single strand of DNA is synthesized as a lagging strand upstream, and as a leading strand downstream, of the replication origin. For example, replicore transitions can be identified in the Rhodopseudomonas palustris genome at ∼500 and ∼3100 kb as seen in plots of cumulative GC and AT skew (Fig. 1A). To increase the precision of our assignment, these positions were refined to within ∼5 kb by the identification of highly skewed octamers that were tabulated based on crude localization of the origin and terminus (Fig. 1B). In the absence of a sufficiently strong single-nucleotide bias to make an initial assignment, the change in octamer abundance alone was used to identify the replication origin and terminus by a sliding-window analysis (Fig. 1D). Here, replication origins and termini were identified as those locations maximizing the numbers of octamers that were overrepresented on the Watson strands upstream—and on the Crick strands downstream—of a particular location.
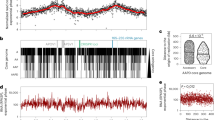
Establishing the locations of the origin and terminus in completely sequenced bacteria: Rhodopseudomonas palustris. A Cumulative third codon-position nucleotide skew. B Positions of five octamers (GAGGAGAG, GAGGAGGG, GAGGGGAG, GAGGGGGG, and GGCGAGGG) are represented as vertical lines on either the Watson (top) or the Crick (bottom) strand. C Cumulative average gene orientation for a 100-gene sliding window, where values are calculated as the proportion of genes transcribed from the Watson strand. The diamond indicates the approximate location of the dnaA gene; arrows indicate the location and orientation of the rRNA cistrons. D Break-point permutation analysis; the numbers of octamers overabundant on the Watson strand upstream of the break point which are also overabundant on the Crick strand downstream of the break point. E Segmental analysis. Black squares denote regions where libraries of Watson strand-biased oligomers are congruent (see Materials and Methods), while white squares denote regions where libraries of Watson strand-biased oligomers of one segment resemble Crick strand-biased oligomer libraries of the other segment. Gray squares denote regions with equivocal data.
Examination of nucleotide skew alone does not identify which inflection point corresponds to the replication origin and which to the terminus. While the dnaA gene is often encoded near the replication origin and rRNA cistrons are often encoded on the leading strands (Fig. 1C), these are not rigorous criteria for localizing origins and termini. To augment these data, we examined gene orientation. Genes are preferentially encoded on the leading strand, perhaps to avoid polymerase collisions at genes under strong selection (Rocha 2004; Rocha and Danchin 2003a, b). Although cumulative gene orientation bias is too crude to identify the replication origin precisely, it may be used to assign the origin and terminus to inflection points identified by mutational bias analysis (Fig. 1B). While more precise localization of the replication origin can be achieved by located dnaA boxes (Mackiewicz et al. 2004), our estimates were sufficiently accurate to enable the identification of strand-biased oligomers.
Single replication origins and termini were established in all large (>1000-kb) bacterial genomes examined, indicating that mutational biases between leading and lagging strands are universal features of bacterial genomes. In most cases, the longest replicore represented between 50% and 55% of the chromosome length (Table 1), suggesting that selection operates to maintain replicores of approximately equal lengths. The positions of the dnaA genes were often, but not always, near the replication origin, and virtually all rRNA cistrons were replicated away from the origin.
In some cases, more than two major inflection points in cumulative nucleotide skew plots were observed; for example, the Pasteurella multocida genome has six major inflection points (Lawrence and Hendrickson 2004). Such patterns could result from inversions, the insertion of foreign DNA with similar strand biases, or the presence of multiple replication origins and termini. In all such cases in Bacteria, we inferred that inversions within replicores had produced regions of the genome with nucleotide skew in the “opposite” direction, because (a) the multiple regions did not reflect more than two symmetrical replicores as was the case in other bacterial genomes and archaeal genomes with likely multiple replication origins (Zhang and Zhang 2003, 2005), and (b) apparent large inversions were common in pathogens with reduced genome sizes where comparisons with the chromosomes of less virulent relatives could delineate the extent of the inverted DNA (Liu and Sanderson 1995a, b; 1996; Parkhill et al. 2003; Read et al. 2000; Suyama and Bork 2001).
Identification of Large Inversions and Insertions
As noted above, the replication history of a DNA segment is reflected in its accumulation of strand bias. Therefore, large insertions and inversions that do not include the replication origin or terminus can be detected by their perturbation of nucleotide-skew and octamer-skew patterns. To identify these regions, a segmental analysis was performed, whereby the local strand biases of individual segments were assessed and compared (Fig. 1E). Here, regions of the chromosome that have historically been replicated in the same direction will have the same sets of octamers biased on their Watson strands. In contrast, large inversions could be identified as regions within well-defined replicores wherein octamers overrepresented on Watson strands were overrepresented on the Crick strands of neighboring segments (Lawrence and Hendrickson 2004). Older large inversions can be recognized as regions where the octamer strand bias is not congruent with either adjacent leading or lagging strands through the process of amelioration; large (>25-kb) insertions will also give this appearance.
Recent large inversions within replicores are typically not evident in bacterial genomes. That is, most genomes showed two large replicores with consistent nucleotide and octamer skew. Exceptions fell into two classes. First, genomes of obligate endosymbionts and intracellular pathogens—typically less than 1000 kb in length—often showed signs of large-scale chromosome rearrangements; examples include the genomes of Buchnera, Wolbachia, Mycoplasma pulmonis, M. genitalium, and Ureaplasma urealyticum. In most cases, chromosomes were sufficiently fragmented to preclude accurate identification of replication origins and termini. Second, pathogens with large genomes also showed rearrangements compared to less virulent relatives with similarly sized genomes. For example, Salmonella enterica serovar Typhi shows substantial rearrangement relative to less virulent salmonellae (Liu and Sanderson 1995b, 1996), and Bordetella pertussis is rearranged relative to B. bronchiseptica (Parkhill et al. 2003). We also detected inversions in E. coli (∼650–740 kb), Fusobacterium nucleatum (∼530–650 kb), Helicobacter pylori (many), and Pasteurella multocida (∼1480–1560 and ∼1880–1960 kb) and inversions shared between Rickettsia prowezeckii and R. connori (∼360–400 and ∼1560–1600 kb).
Identification of Sequences Under Selection
Chromosomes lacking large inversions were examined for octamers that increased in abundance toward the replication terminus only on leading strands. First, sequences that were overrepresented on leading strands on both replicores were identified; this bias in distribution could result solely from the mutational bias inherent in DNA replication and therefore does not in itself suggest that these sequences are under selection. Table 1 reports the number of abundant octamers (found at a frequency of >0.1 per kb) which showed strong strand bias (more than 75%—a 3:1 bias—were located on the leading strand). With these stringent criteria, between 0 and 42,000 sequences were identified in 40 bacterial genomes examined. In genomes that lacked highly abundant oligomers that were skewed to this degree, we identified skewed sequences that were found at least once per 20 kb (Table 1).
Within these sets, we identified sequences under selection as those that increased in abundance on the leading strand toward the replication terminus. These sequences were initially identified as those that were overrepresented on leading strands in the terminus-proximal regions of each replicore. To eliminate sequences which were serendipitously overabundant in these regions—for example, if they were highly abundant in genomic islands integrated near the terminus region—the local abundance of each octamer was calculated for intervals spanning from the replication origin to the terminus. Sequences under selection were identified as those where the slope of the linear regression of abundance vs. position was significantly different from zero (Fig. 2). In most cases, the sequence also significantly decreased in frequency on the lagging strands, thus leading to greater strand bias near the terminus. In other cases, the abundance was extremely low on the lagging strand, precluding accurate assessment of changes in abundance on this strand.
Table 1 shows examples of skewed sequences increasing toward the replication termini in bacterial genomes. For example, there are 312 copies of the GGGCAGGG octamer in the R. palustris genome; 96% of the occurrences are on the leading strand, and twice as many copies are found in the terminus-proximal region of both replicores than would be expected if sequences were distributed randomly (Fig 2). This sequence was found to increase toward the replication terminus in many genomes of proteobacteria (Table 1). We propose that skewed octamers increasing in abundance toward the replication terminus are under selection for maintenance of chromosome structure. Therefore, we term these octamers architecture imparting sequences, or AIMS, to denote their potential involvement in one or more biological processes that use origin to terminus polarity. While the role of each AIMS in cell biology is unknown, it is clear that the distribution of AIMS represents selection operating above the level of the gene and that this selection structures—i.e., provides an architecture to—bacterial chromosomes.
Table 1 presents only a sample of potential AIMS, not a definitive list of all sequences under selection for function. There are many sequences which are less numerous, are less strand-biased, or show a more modest increase in abundance toward the replication terminus which were excluded from this analysis. That is, we chose threshold values so that sequences that met our criteria could not have arisen by chance alone (see below). In genomes where no sequences were found to pass these criteria, we could identify sequences that increased in abundance toward the replication terminus that were less abundant, were less strand-biased, or increased in abundance toward the terminus to a lesser degree. However, this set of sequences includes those whose distributions resulted by chance, thus potentially confounding conclusions drawn from their distributions.
AIMS were found in genomes of bacteria representing every major division, including multiple representatives of Actinobacteria, Chlamydiae, Cyanobacteria, Firmicutes, Proteobacteria, and Spirochetes (Table 1). AIMS were easily identified in genomes of small size (e.g., the TTTAGTTT octamer in the Borrelia burgdorferi genome; 911 kb) and large size (e.g., the AGGAGGGC octamer in the Pseudomonas aeruginosa genome; 6264 kb). We could identify AIMS in genomes with a high GC content (e.g., TGGGGGAG in Streptomyces coelicolor; 72.1% GC), a high AT content (e.g., AAGAAGAT in Clostridium acetobutylicum; 30.9% GC), or a neutral composition (e.g., TGGCTTTG in Prochlorococcus marinus; 50.7% GC). AIMS were often GC-rich, even in genomes with a high AT content (e.g., TAGGG GATG in Enterococcus faecalis; 37.5% GC). AIMS were also found in organisms with linear replicons (e.g., Streptomyces, Borrelia, and Agrobacterium), suggesting that functions utilizing at least some of the AIMS are required for replication and segregation of linear chromosomes. For example, such functions may include DNA translocation across the division septum.
In three instances, multiple, large replicons are found in the same organism: Brucella melitensis, Agrobacterium tumefaciens, and Sinorhizobium meliloti. In Brucella and Agrobacterium, the AIMS identified from one large replicon also appeared to be skewed and increasing in abundance in the other replicon (Table 1); some sequences were less abundant on one replicon and, therefore, are not reported in Table 1. This suggests that they are under selection in both replicons in each organism. Sinorhizobium has three replicons, including the large plasmids pSymA (1354 kb) and pSymB (1683 kb). AIMS found in the Sinorhizobium chromosome (i.e., the largest replicons) were also AIMS in the pSymB sequence. While these sequences are not AIMS in the pSymA plasmid, they are skewed to leading strands. Since both plasmids harbor repABC partitioning operons near their respective replication origins, AIMS may not play as large a role in their maintenance.
As a rule, we did not identify AIMS in genomes <1000 kb in size; most genomes of this size class are found in obligate pathogens and intracellular parasites (e.g., Buchnera, Mycoplasma). We do not interpret this result as a lack of selection for polarity elements in these taxa. Rather, extensive chromosome rearrangements experienced by genomes of pathogens (Mira et al. 2001), coupled with the very small size of these genomes limits the ability to find distributions that are statistically significant. Also, as noted above, replication origins and termini could not be located with confidence in these genomes. As a result, we were not confident of the sequence distributions we could infer.
AIMS Do Not Arise from Changes in Mutational Bias
The underlying mutational biases vary along the chromosome (Daubin and Perrière 2003); that is, GC skew at third codon positions differs between genes that are origin-proximal relative to those that are terminus-proximal. Therefore, one could infer that some octamers may increase in abundance toward the terminus strictly due to changes in mutational bias alone. To correct for changes in mutational bias in the terminus-proximal region, we quantitated the changes in the nucleotide, dinucleotide, trinucleotide, and tetranucleotide frequencies from the origin to the terminus. For octamers under selection, their accumulation near the replication terminus cannot be explained by underlying changes in the distribution of nucleotides, dinucleotides, trinucleotides, or tetranucleotides. For example, Figs. 3A and B show the abundance of the GGGCAGGG octamer in the Rhodopseudomonas palustris genome; toward the replication terminus, it clearly increases in abundance on the leading strand and decreases in abundance on the lagging strand. Yet the predicted abundance of this octamer—as inferred from the abundance of its constituent dinucleotides, trinucleotides and tetranucleotides—does not change appreciably. If any, predicted abundances decrease toward the terminus on the leading strand and increase on the lagging strand. These data suggest that a simple change in mutational bias from the replication origin to the terminus is not responsible for the distribution of the GGGCAGGG octamer in the R. palustris genome. Similar results are seen for the GAAGGGGA octamer in the Bacillus subtilus genome (Figs. 3C and D). We examined the distribution of all potential AIMS listed in Table 1 and concluded that changes in mutational biases alone cannot explain the distribution of any octamer increasing in abundance near a replication terminus.
Distributions of AIMS are not explained by mutational changes from origin to terminus in chromosomes. The accumulation of the GGGCAGGG octamer on the (A) leading strand and (B) lagging strand in the terminus-proximal region of the R. palustris genome, and the accumulation of the GAAGGGGA octamer on the (C) leading strand and (D) lagging strand within the Bacillus subtilus genome. The observed local abundances of these sequences are shown along with the expected abundance predicted from the distributions of the seven constituent dinucleotides, six trinucleotides, or five tetranucleotides as described under Materials and Methods. Sequence abundance is reported as number of AIMS (either observed or predicted) per kilobase.
Sequences Only Accumulate in Abundance Near the Replication Terminus
In identifying potential AIMS in bacterial genomes, we required both a moderately high overall abundance and a strong increase in abundance toward the replication terminus. These criteria were established so that changes in abundance could not be attributed to chance. That is, given 16 million degenerate octamers that are examined, one would expect some to increase in abundance toward the replication terminus strictly by chance; asking for similar increases in both replicores reduces the number of false positives but does not eliminate them. To ascertain how many sequences arise by chance that increase in abundance toward a particular location, we examined genomes for sequences which accumulated at other locations in the genome to the degree shown by AIMS. If the numbers of AIMS merely reflects chance, similar numbers of sequences should be identified that accumulate toward other locations in the genome.
As shown in Fig. 4, more sequences accumulate at the replication terminus than any other location in the genome. Moreover, those sequences appearing to reach maximum abundance outside the terminus region were found in lower copy numbers than AIMS, so their “accumulation” at other chromosomal locations was interpreted as resulting from chance. That is, the numbers of sequences accumulating at a nonterminus location represented the “noise” produced by examining 16 million octamers. Therefore, we interpret AIMS—that is, high-copy-number sequences that accumulate only at the replication terminus in a way unexplained by underlying mutational bias—as sequences under selection for function.
Sequences under selection accumulate at the terminus, not other locations. Sequences that accumulate toward a defined region within each replicore were identified; the total count of individual sequences is plotted (that is, the number of different sequences multiplied by their abundances). There are more sequences that accumulate gradually and have their highest point of abundance at the terminus than other regions of the genome. The gray bars show the numbers of sequences that are overrepresented within the region specified. The black bars show the numbers of sequences that have their maximal abundance within the region specified.
The Sequence Distribution, Not Individual Sequences, Is Under Selection
The accumulation of AIMS toward the replication terminus could result from the nonrandom distribution of genes within genomes. For example, if membrane proteins were located in the terminus proximal region, sequences encoding membrane-spanning domains may be overly abundant in this region. If this were the case, then one would expect individual occurrences of AIMS themselves—not merely their distribution within the replicore—to be conserved among genomes of closely related bacteria. As shown in Table 1, we have identified AIMS in several sets of closely related genomes, including the GGGCAGGG octamer in numerous α-proteobacteria.
To determine if AIMS were under selection for function in their resident proteins, we examined their locations within orthologous genes among closely related taxa. We found that the locations of AIMS within orthologous genes were not conserved; rather, only their distribution—and increase in abundance toward the replication terminus—was shared among these genomes. For example, the distribution of the GGGCAGGG octamer increases in abundance among all α-proteobacteria examined; these genomes range in size from 3.7 to 9.1 MB (Fig. 5B). However, the precise locations of these individual sequences were not conserved among orthologous genes (Fig 5A), and the octamer was found in several reading frames, in both the template and the nontemplate strands, and in intergenic regions. These data support the hypothesis that the distribution itself is under selection, suggesting a unit of selection at the level of the replicore, above the level of the individual gene.
Among closely related bacteria it is the sequence distribution that is conserved, not the absolute positions of the sequences. A The frequency of each octamer—not a cumulative frequency—within genomic regions is plotted as a function of the distance from the terminus of replication. Abundance on the two replicores is averaged. The GGGCAGGG octamer shows comparable distributions in genomes of four species of α-proteobacteria, increasing in abundance on leading strands toward the replication terminus. B AIMS within orthologous genes in the α-proteobacteria do not occur in the same positions. Arrows denote the positions of AIMS; the direction of the arrow denotes orientation. C Orthologues shared between the R. palustris and the S. meliloti genomes. A total of 1666 genes (50% of the S. meliloti gene complement) were reciprocal best matches with adjusted alignment scores of 125 or above, providing a conservative assignment of orthologues. Genes in the same orientation are shown as squares, and those in the opposite orientation as crosses.
More importantly, the cellular functions that require AIMS—candidates include the FtsK protein, which translocates to the dif site in the terminus region during cell division—appear to conserve their choice of AIMS. Table 1 shows several cases of related organisms which share AIMS, even though they share less than 90% sequence identity. For example, many α-proteobacteria share the GGGCAGGG octamer as an AIMS. As shown in Fig. 5C, AIMS may be retained even in the face of extensive chromosomal rearrangements, consistent with strong selection for AIMS.
Discussion
AIMS Are Widespread Among Bacterial Genomes
We have provided evidence that bacterial chromosomes contain sequences whose distributions suggest that they are under selection for a function unrelated to the genes in which they are found. The distributions of these sequences are consistent with their role in specifying strand identity. That is, differential abundance of sequences on leading and lagging strands can be used to locate the terminus; selection for this asymmetry will lead to increased abundance on leading strands, and decreased abundance on lagging strands, that is inversely correlated with distance from the replication terminus. While it is not clear precisely what these functions may be, their distributions are consistent with a role during DNA replication and segregation. We have termed these elements architecture imparting sequences, or AIMS. It does not appear that the specific locations of AIMS with respect to genes or transcripts are under selection as is the case with transcription promoters, ρ-independent transcription terminators, binding sites for regulatory proteins, translation start sites, or translation stop sites. Rather, the distribution of AIMS across the replicore reflects a gradient in selection, where the entire replicore acts as a target of selection, functioning above the level of the individual gene or operon.
We have identified AIMS in nearly every bacterial genome we examined for which the identification of the replication origin and terminus was unambiguous (Table 1); the failure to identify AIMS in some genomes likely reflects the stringency of our search criteria rather than their absence from that genome. This suggests that AIMS are not under selection for a function that is found only in certain organisms, although the proteins that mediate this function may differ among organisms, leading to different AIMS being found in different genomes. For example, ter sites within the E. coli genome—bound by the Tus protein to halt retrograde replication forks—are found in the terminus-proximal region; but the tus gene is not found outside the proteobacteria (Andersen et al. 2000). In other bacteria, sequences like AIMS may contribute to these functions. That is, the function is likely important to all bacteria, but particular sequences (like ter) will not be ubiquitous.
Inspection of Table 1 shows that genomes of closely related organisms often show similar AIMS. For example, the GGGCAGGG octamer—or some closely allied sequence—not only is skewed in proteobacterial genomes, but is increasing in abundance on the leading strand toward the replication terminus—that is, it is an AIMS. Similarly, the AAGAAGAT octamer appears as an AIMS in the genomes of several Firmicutes, and Actinobacteria shared permutations of the YGGGGGAG octamer. As shown in Fig. 5, the common occurrence of AIMS in related genomes is not a result of the sequence being conserved within individual genes; rather, the pattern of increasing abundance toward replication termini is shared. Moreover, AIMS are conserved in the face of extensive rearrangement of these chromosomes (Fig. 5C). The common sets of AIMS among related bacteria are consistent with shared, conserved mechanisms that maintain chromosome architecture in these organisms.
AIMS May Represent Longer, More Degenerate Sequences Under Selection for Function
AIMS do not necessarily indicate the precise sequence acted on by a molecular mechanism; rather, they are only sequences whose distributions must have arisen from selection for their overabundance near the replication terminus. The precise sequences acting as targets of selection could be deduced from the library of AIMS within a genome. First, in many genomes sets of AIMS appear to represent a more degenerate sequence. For example, both permutations of the GGGMAGGG octamer are AIMS in Mesorhizobium loti, as well as both permutations of the GRGCAGGG octamer in Pseudomonas aeruginosa (Table 1). Therefore, the distributions of the nondegenerate sequences may reflect a more degenerate target of selection. Second, AIMS are detected as octamers, while either shorter or longer sequences may actually be under selection. In many genomes, there are AIMS which have overlapping sequences, such as the two octamer permutations of the AGGGCAGGG nonomer in Sinorhizobium meliloti and Brucella melitensis or the three octamer permutations of the YGGGGGAGC nonomer in Mycobacterium tuberculosis (Table 1). Further inspection of these nonomers has not yielded evidence that these longer sequences might be the actual targets of selection; longer sequences do not accumulate toward the terminus to a greater degree than their constituent octamers. More thorough analyses may uncover some examples, but the low abundance of sequences longer than octamers precludes rigorous testing. More importantly, longer sequences may be insufficiently abundant to serve as polarizing elements. For example, the FtsK protein appears to recognize and reorient in response to sequence elements as frequently as once per 2 kb (six times in 12 kb) in the terminus region of the E. coli chromosome (Pease et al. 2005), which may be accommodated by degenerate octamers but likely not by longer sequences.
Selection for Function Does Not Always Lead to Accumulation Near the Replication Terminus
AIMS represent one class of sequences that operates to maintain chromosome architecture; AIMS reflect selection for functions required at or near the replication terminus. Other sequences whose importance is not restricted to this region may show evidence for selection by virtue of their overrepresentation on leading strands throughout the genome. For example, the 8-bp χ sequence (GCTGGTGG) is recognized by the E. coli RecBC helicase/exonuclease/recombinase complex, halting the retrotranslocation of Holliday junctions at these sites and instigating resolution of recombination substrates (Myers and Stahl 1994). It has been noted previously that χ sites are more abundant than would be expected (El Karoui et al. 1999). In the E. coli genome, χ sites are approximately 3.5 times more abundant across the length of the replicore than would be expected given its component tetramers (Fig. 6A). As discussed elsewhere (El Karoui et al. 1999; Uno et al. 2000), this increased abundance is taken as an indication of replicore-wide positive selection for function of this sequence. Here, the χ sequence prevents RecD-mediated degradation of DNA strands, allowing for rapid reestablishment of stalled replication forks. Since selection for the function of χ sequences is independent of genome position, we would not expect the abundance of χ to increase toward the terminus. Unlike AIMS, the abundance of the χ octamer does not increase toward the replication terminus. Also unlike AIMS, selection has not favored the increased abundance of the χ octamer on leading strands and decreased abundance on lagging strands, which would heighten strand bias; χ is actually somewhat more abundant than expected on lagging strands (Fig. 6A).
Distribution of octamers in the E. coli genome. The frequency of each octamer—not a cumulative frequency—within genomic regions is plotted as a function of the distance from the terminus if replication. Abundance on the two replicores is averaged. A Abundance of the χ octamer. B Abundance of the RAGS octamer. C The distributions of octamers that are strand-biased to the same degree as the RAGS oligomer in the E. coli genome were analyzed for positions of maximal abundance; the numbers of oligomers whose distributions were maximal at eight separate intervals (as determined by quadratic regression) are shown. The dashed line denotes the mean of 2.25 oligomers.
All Sequences Accumulating Toward the Terminus Are Not Necessarily Under Selection
The RAG octamer (RGNAGGGS) was identified as a putative polarizing element in the E. coli chromosome, possibly aiding in positioning the dif site at the septum during cell division at the end of bacterial chromosome replication (Bigot et al. 2004; Capiaux et al. 2001; Corre and Louarn 2002). Although the RAG octamer was postulated to act within the terminus-centered 10-kb dif-activity zone (Cornet et al. 1996; Pérals et al. 2000) or the 250-kb FtsK zone (Corre and Louarn 2002, 2005), these boundaries reflect the resolution of the bacteriophage-excision assays used to assess the negative impact of placing AIMS in their non-permissive orientation.
In our analysis, the RAG sequence was not reported as an AIMS in the E. coli genome (Table 1), indicating that its distribution did not satisfy our threshold criteria. The degenerate RAG has 6 bases of information, making it a sufficiently abundant octamer to analyze, and does accumulate somewhat in abundance toward the replication terminus (Fig. 6B). What is not clear is if this increase in abundance is significant. The RAG sequence increases in abundance on leading strands toward the replication terminus 1.5-fold more than would be expected based on the distribution of underlying tetramers (Fig. 6B). Yet AIMS we identified increased to a much greater degree; for example, the degenerate AGGGCRGR octamer increased 3.2-fold in abundance. It is possible that the modest increase in abundance of the RAG octamer—and similar avoidance on the lagging strand—indicates that the RAG octamer is under selection as an AIMS and merely fails to exceed our threshold.
To determine if the degree to which the RAGS sequence accumulates toward the replication terminus is significant, we investigated whether sequences accumulated to this degree at other locations in the E. coli genome (Fig. 6C). We found that sequences that accumulate in abundance to the same degree as the RAG octamer were as likely to be found accumulating toward nonterminus locations (Fig. 6C). Since the “accumulation” of octamers at nonterminus locations reflects baseline noise, one cannot conclude that the apparent increase in abundance of the RAG octamer toward the replication terminus reflects selection for function.
Importantly, a previously identified (Lawrence and Hendrickson 2003), widely distributed (Table 1) AIMS among proteobacteria, GGGCAGGG, has now been implicated as a potential binding site for the FtsK protein in Escherichia coli (Bigot et al. 2005; Levy et al. 2005). This is gratifying, as the FtsK translocase is precisely the sort of protein that would interact with AIMS. Although Levy et al. (2005) point out that the GNGNAGGG octomer is biased in the genomes of several bacteria, strand bias alone does not provide evidence for selection for function. Indeed, strand-biased oligomers may arise by simple differences in mutational proclivities of the DNA polymerases replicating leading and lagging strands (Lobry 1996), and Table 1 shows that genomes may have hundreds or even thousands of octameric sequences that are strand biased. Further analyses, such as those described herein, are required to demonstrate the footprint of natural selection.
Interplay of Mutation and Selection
Although the RAG octamer did not increase in abundance more than one would expect at random (Figs. 6B and 6C), it may still be under selection for function. That is, the strand asymmetry we observe may be sufficient for chromosome polarity to be established. The increase in abundance toward the replication terminus accentuates strand asymmetry, which is also a feature we believe is under selection; if natural mutational biases yield both sufficient sequence abundance and sufficient strand asymmetry, then selection acting on these sequences will not change their distribution in any detectable fashion. The distribution of AIMS within a chromosome reflects a balance of mutation and selection, where a gradient of selection from the replication origin to terminus may increase the abundance of AIMS on leading strands if mutation acts to defeat the required asymmetry. When mutation does not defeat asymmetry, selection is less evident.
In some genomes, strand asymmetry—that is, nucleotide skew reflecting mutational biases—is more evident than in others. For example, Firmicutes show a much larger number of strand-biased oligomers than other taxa (Table 1). The pattern may reflect differences in DNA replication in these taxa; Firmicutes utilize DNA polymerase harboring different subunits to replicate their leading and lagging strands, potentially leading to stronger strand asymmetry (Rocha 2004). In addition, the strong bias of genes to be encoded on leading strands (∼80% in Firmicutes) may lead to stronger strand differences. Similarly, the prevalence of genes being encoded on leading strands will result in transcription-coupled repair processes acting differentially between the strands. As a result, AIMS may be less evident in such highly skewed genomes since mutation does not defeat selected abundance distributions. That is, while the distribution of AIMS reflects selection, the absence of AIMS can not be regarded as an absence of selection.
Impact of AIMS on Genome Evolution
Most genomes show two large replicores with consistent strand asymmetry (Fig. 1E). In using this asymmetry as an indicator of chromosome rearrangement, we could detect inversions without genome comparison and without ambiguity regarding the polarity of the inversion (Darling et al. 2004). Inversions have been described in many genomes that include the replication origin or terminus (Eisen et al. 2000; Mackiewicz et al. 2001); these rearrangements do not disrupt strand asymmetry and are not detected in our analysis. Our findings suggest that most genomes are recalcitrant to inversion within replicores; we found that only the genomes of obligate pathogens or symbionts contained significant numbers of large inversions within replicores. This finding is consistent with published findings for Salmonella typhi (Liu and Sanderson 1995b, 1996), Bordetella pertussis (Parkhill et al. 2003), and Wolbachia (Foster et al. 2005). Therefore, one may ask why large inversions within replicores—that is, those not including the replication origin or terminus—are not found in genomes of free-living, nonpathogenic bacteria.
Selection against some inversions has been demonstrated in the Salmonella enterica genome (Mahan and Roth 1991; Segall et al. 1988). The lack of these “forbidden” inversions does not reflect the inability to form them (Mahan and Roth 1991; Segall et al. 1988). We propose that disruption of the distribution of AIMS—rather than simply placing a gene on the lagging strand or moving its position relative to the replication origin—counterselects organisms which contain large inversions within replicores. Such inversions would place large numbers of AIMS in their nonpermissive orientation and thus confer a fitness defect. For example, if the FtsK protein relies on AIMS to translocate toward the replication terminus, the protein would receive incorrect orientation information within large inversions. It has not escaped our attention that selection would also act to limit the acquisition of genomic islands wherein AIMS were present in large numbers in the nonpermissive orientation.
Just as genomes of pathogens show a large amount of gene loss (Andersson and Andersson 1999a, b; Cole et al. 2001)—reflecting an inability to select for gene retention (Lawrence 2001; Lawrence et al. 2001; Lawrence and Roth 1999)—inversions also accumulate in these genomes. Such inversions would be insufficiently detrimental to prevent the persistence of strains bearing them. Pathogens often have reduced population sizes and reduced rates of recombination, thereby accelerating the fixation of deleterious changes. Yet mispolarized AIMS would still be problematic, and the removal of this DNA may be beneficial. The deletion of inverted DNA would likely not be a strategy employed by most organisms, but it is a likely outcome for organisms experiencing genome reduction (Andersson and Andersson 1999a, b; Cole et al. 2001). The occurrence of large inversions in the genomes of some symbionts (Mira et al. 2001) is consistent with this hypothesis. We speculate that the removal of inverted DNA may provide a selective advantage to DNA loss in organisms experiencing genome reduction. That is, deletion of DNA may not always be neutral or detrimental.
Conclusions
In bacterial genomes, where space is minimal and the DNA is information rich, AIMS represent an elegant solution to the problem of specifying the direction in which landmarks like the replication origin and terminus can be found. The large numbers of AIMS ensure that, even as the tide of random mutation disrupts individual sequences, the overall distribution of these important signaling sequences is maintained. We believe that AIMS are a common feature among bacterial chromosomes and that a previously unrecognized structure plays a role in influencing the evolution of these genomes. Though the mechanism by which most AIMS act has not been determined, it is possible that perturbations of these sequence patterns are sufficiently disruptive to chromosome maintenance that they are having, and have had, a major role to play in the shape and content of bacterial chromosomes as we see them today.
References
Altschul SF (1991) Amino acid substitutions matrices from an information theoretic perspective. J Mol Biol 219:555–565
Andersen PA, Griffiths AA, Duggin IG, Wake RG (2000) Functional specificity of the replication fork-arrest complexes of Bacillus subtilis and Escherichia coli: significant specificity for Tus-Ter functioning in E. coli. Mol Microbiol 36:1327–1335
Andersson JO, Andersson SG (1999a) Genome degradation is an ongoing process in Rickettsia. Mol Biol Evol 16:1178–1191
Andersson JO, Andersson SG (1999b) Insights into the evolutionary process of genome degradation. Curr Opin Genet Dev 9:664–671
Bigot S, Corre J, Louarn JM, Cornet F, Barre FX (2004) FtsK activities in Xer recombination, DNA mobilization and cell division involve overlapping and separate domains of the protein. Mol Microbiol 54:876–886
Bigot S, Saleh OA, Lesterlin C, Pages C, El Karoui M, Dennis C, Grigoriev M, Allemand JF, Barre FX, Cornet F (2005) KOPS: DNA motifs that control E coli chromosome segregation by orienting the FtsK translocase. EMBO J 24:3770–3780
Blakely G, Colloms S, May G, Burke M, Sherratt D (1991) Escherichia coli XerC recombinase is required for chromosomal segregation at cell division. New Biol 3:789–798
Capiaux H, Cornet F, Corre J, Guijo M, Perals K, Rebollo JE, Louarn J (2001) Polarization of the Escherichia coli chromosome. A view from the terminus. Biochimie 83:161–170
Clerget M (1991) Site-specific recombination promoted by a short DNA segment of plasmid R1 and by a homologous segment in the terminus region of the Escherichia coli chromosome. New Biol 3:780–788
Cole ST, Eiglmeier K, Parkhill J, James KD, Thomson NR, Wheeler PR, Honore N, Garnier T, Churcher C, Harris D, Mungall K, Basham D, Brown D, Chillingworth T, Connor R, Davies RM, Devlin K, Duthoy S, Feltwell T, Fraser A, Hamlin N, Holroyd S, Hornsby T, Jagels K, Lacroix C, Maclean J, Moule S, Murphy L, Oliver K, Quail MA, Rajandream MA, Rutherford KM, Rutter S, Seeger K, Simon S, Simmonds M, Skelton J, Squares R, Squares S, Stevens K, Taylor K, Whitehead S, Woodward JR, Barrell BG (2001) Massive gene decay in the leprosy bacillus. Nature 409:1007–1011
Cornet F, Louarn J, Patte J, Louarn JM (1996) Restriction of the activity of the recombination site dif to a small zone of the Escherichia coli chromosome. Genes Dev 10:1152–1161
Corre J, Louarn JM (2002) Evidence from terminal recombination gradients that FtsK uses replichore polarity to control chromosome terminus positioning at division in Escherichia coli. J Bacteriol 184:3801–3807
Corre J, Louarn JM (2005) Extent of the activity domain and possible roles of FtsK in the Escherichia coli chromosome terminus. Mol Microbiol 56:1539–1548
Corre J, Patte J, Louarn JM (2000) Prophage lambda induces terminal recombination in Escherichia coli by inhibiting chromosome dimer resolution. An orientation-dependent cis- effect lending support to bipolarization of the terminus. Genetics 154:39–48
Cunningham EL, Berger JM (2005) Unraveling the early steps of prokaryotic replication. Curr Opin Struct Biol 15:68–76
Darling AC, Mau B, Blattner FR, Perna NT (2004) Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res 14:1394–1403
Daubin V, Perrière G (2003) G+C3 structuring along the genome: a common feature in prokaryotes. Mol Biol Evol 20:471–483
Deng S, Stein RA, Higgins NP (2004) Transcription-induced barriers to supercoil diffusion in the Salmonella typhimurium chromosome. Proc Natl Acad Sci USA 101:3398–3403
Dworkin J, Losick R (2001) Differential gene expression governed by chromosomal spatial asymmetry. Cell 107:339–346
Eggleston AK, West SC (1997) Recombination initiation: Easy as A, B, C, D chi? Curr Biol 7:R745–R749
Eisen JA, Heidelberg JF, White O, Salzberg SL (2000) Evidence for symmetric chromosomal inversions around the replication origin in bacteria. Genome Biol 1:1–11
El Karoui M, Biaudet V, Schbath S, Gruss A (1999) Characteristics of Chi distribution on different bacterial genomes. Res Microbiol 150:579–587
Foster J, Ganatra M, Kamal I, Ware J, Makarova K, Ivanova N, Bhattacharyya A, Kapatral V, Kumar S, Posfai J, Vincze T, Ingram J, Moran L, Lapidus A, Omelchenko M, Kyrpides N, Ghedin E, Wang S, Goltsman E, Joukov V, Ostrovskaya O, Tsukerman K, Mazur M, Comb D, Koonin E, Slatko B (2005) The Wolbachia genome of Brugia malayi: endosymbiont evolution within a human pathogenic nematode. PLoS Biol 3:e121
Gitai Z, Thanbichler M, Shapiro L (2005) The choreographed dynamics of bacterial chromosomes. Trends Microbiol 13:221–228
Higgins NP, Yang X, Fu Q, Roth JR (1996) Surveying a supercoil domain by using the gamma delta resolution system in Salmonella typhimurium. J Bacteriol 178:2825–2835
Holmes VF, Cozzarelli NR (2000) Closing the ring: links between SMC proteins and chromosome partitioning, condensation, and supercoiling. Proc Natl Acad Sci USA 97:1322–1324
Ip SC, Bregu M, Barre FX, Sherratt DJ (2003) Decatenation of DNA circles by FtsK-dependent Xer site-specific recombination. EMBO J 22:6399–6407
Jaccard P (1912) The distribuition of flora in the alpine zone. New Phytol 11:37–50
Kowalczykowski SC, Dixon DA, Eggleston AK, Lauder SD, Rehrauer WM (1994) Biochemistry of homologous recombination in Escherichia coli. Microbiol Rev 58:401–465
Kuzminov A (1995) Collapse and repair of replication forks in Escherichia coli. Mol Microbiol 16:373–384
Lau IF, Filipe SR, Soballe B, Okstad OA, Barre FX, Sherratt DJ (2003) Spatial and temporal organization of replicating Escherichia coli chromosomes. Mol Microbiol 49:731–743
Lawrence JG (2001) Catalyzing bacterial speciation: correlating lateral transfer with genetic headroom. Syst Biol 50:479–496
Lawrence JG, Hendrickson H (2003) Lateral gene transfer: When will adolescence end? Mol Microbiol 50:739–749
Lawrence JG, Hendrickson H (2004) Chromosome structure and constraints on lateral gene transfer. Dev Genet 2004:319–336
Lawrence JG, Ochman H (1997) Amelioration of bacterial genomes: rates of change and exchange. J Mol Evol 44:383–397
Lawrence JG, Roth JR (1999) Genomic flux: genome evolution by gene loss and acquisition. In: Charlebois RL (eds). Organization of the prokaryotic genome. ASM Press, Washington, DC, pp 263–289
Lawrence JG, Hendrix RW, Casjens S (2001) Where are the pseudogenes in bacterial genomes? Trends Microbiol 9:535–540
Levy O, Ptacin JL, Pease PJ, Gore J, Eisen MB, Bustamante C, Cozzarelli NR (2005) Identification of oligonucleotide sequences that direct the movement of the Escherichia coli FtsK translocase. Proc Natl Acad Sci USA 102:17618–17623
Li Y, Youngren B, Sergueev K, Austin S (2003) Segregation of the Escherichia coli chromosome terminus. Mol Microbiol 50:825–834
Liu SL, Sanderson KE (1995a) The chromosome of Salmonella paratyphi A is inverted by recombination between rrnH and rrnG. J Bacteriol 177:6585–6592
Liu SL, Sanderson KE (1995b) Rearrangements in the genome of the bacterium Salmonella typhi. Proc Natl Acad Sci USA 92:1018–1022
Liu SL, Sanderson KE (1996) Highly plastic chromosomal organization in Salmonella typhi. Proc Natl Acad Sci USA 93:10303–10308
Lobry JR (1996) Asymetric substitution patterns in the two DNA strands of bacteria. Mol Biol Evol 13:660–665
Lobry JR, Louarn JM (2003) Polarisation of prokaryotic chromosomes. Curr Opin Microbiol 6:101–108
Mackiewicz P, Mackiewicz D, Kowalczuk M, Cebrat S (2001) Flip-flop around the origin and terminus of replication in prokaryotic genomes. Genome Biol 2:INTERACTIONS1004
Mackiewicz P, Zakrzewska-Czerwinska J, Zawilak A, Dudek MR, Cebrat S (2004) Where does bacterial replication start? Rules for predicting the oriC region. Nucleic Acids Res 32:3781–3791
Mahan MJ, Roth JR (1991) Ability of a bacterial chromosome segment to invert is dictated by included material rather than flanking sequence. Genetics 129:1021–1032
Massey TH, Aussel L, Barre FX, Sherratt DJ (2004) Asymmetric activation of Xer site-specific recombination by FtsK. EMBO Rep 5:399–404
Mira A, Ochman H, Moran NA (2001) Deletional bias and the evolution of bacterial genomes. Trends Genet 17:589–596
Myers RS, Stahl FW (1994) Chi and the RecBC D enzyme of Escherichia coli. Annu Rev Genet 28:49–70
Needleman SB, Wunsch CD (1970) A general method applicable to the search for similarities in the amino acid sequence of two proteins. J Mol Biol 48:443–453
Niki H, Yamaichi Y, Hiraga S (2000) Dynamic organization of chromosomal DNA in Escherichia coli. Genes Dev 14:212–223
Parkhill J, Sebaihia M, Preston A, Murphy LD, Thomson N, Harris DE, Holden MT, Churcher CM, Bentley SD, Mungall KL, Cerdeno-Tarraga AM, Temple L, James K, Harris B, Quail MA, Achtman M, Atkin R, Baker S, Basham D, Bason N, Cherevach I, Chillingworth T, Collins M, Cronin A, Davis P, Doggett J, Feltwell T, Goble A, Hamlin N, Hauser H, Holroyd S, Jagels K, Leather S, Moule S, Norberczak H, O’Neil S, Ormond D, Price C, Rabbinowitsch E, Rutter S, Sanders M, Saunders D, Seeger K, Sharp S, Simmonds M, Skelton J, Squares R, Squares S, Stevens K, Unwin L, Whitehead S, Barrell BG, Maskell DJ (2003) Comparative analysis of the genome sequences of Bordetella pertussis, Bordetella parapertussis and Bordetella bronchiseptica. Nat Genet 35:32–40
Pease PJ, Levy O, Cost GJ, Gore J, Ptacin JL, Sherratt D, Bustamante C, Cozzarelli NR (2005) Sequence-directed DNA translocation by purified FtsK. Science 307:586–590
Pérals K, Cornet F, Merlet Y, Delon I, Louarn JM (2000) Functional polarization of the Escherichia coli chromosome terminus: the dif site acts in chromosome dimer resolution only when located between long stretches of opposite polarity. Mol Microbiol 36:33–43
Read TD, Brunham RC, Shen C, Gill SR, Heidelberg JF, White O, Hickey EK, Peterson J, Utterback T, Berry K, Bass S, Linher K, Weidman J, Khouri H, Craven B, Bowman C, Dodson R, Gwinn M, Nelson W, DeBoy R, Kolonay J, McClarty G, Salzberg SL, Eisen J, Fraser CM (2000) Genome sequences of Chlamydia trachomatis MoPn and Chlamydia pneumoniae AR39. Nucleic Acids Res 28:1397–1406
Rocha EP (2004) The replication-related organization of bacterial genomes. Microbiology 150:1609–1627
Rocha EP, Danchin A (2003a) Essentiality, not expressiveness, drives gene-strand bias in bacteria. Nat Genet 34:377–378
Rocha EP, Danchin A (2003b) Gene essentiality determines chromosome organisation in bacteria. Nucleic Acids Res 31:6570–6577
Salzberg SL, Salzberg AJ, Kerlavage AR, Tomb JF (1998) Skewed oligomers and origins of replication. Gene 217:57–67
Sanderson KE, Liu SL (1998) Chromosomal rearrangements in enteric bacteria. Electrophoresis 19:569–572
Segall A, Mahan MJ, Roth JR (1988) Rearrangement of the bacterial chromosome: forbidden inversions. Science 241:1314–1318
Stein RA, Deng S, Higgins NP (2005) Measuring chromosome dynamics on different time scales using resolvases with varying half-lives. Mol Microbiol 56:1049–1061
Suyama M, Bork P (2001) Evolution of prokaryotic gene order: genome rearrangements in closely related species. Trends Genet 17:10–13
Teleman AA, Graumann PL, Lin DC, Grossman AD, Losick R (1998) Chromosome arrangement within a bacterium. Curr Biol 8:1102–1109
Tillier ER, Collins RA (2000) Genome rearrangement by replication-directed translocation. Nat Genet 26:195–197
Uno R, Nakayama Y, Arakawa K, Tomita M (2000) The orientation bias of Chi sequences is a general tendency of G-rich oligomers. Gene 259:207–215
Viollier PH, Shapiro L (2004) Spatial complexity of mechanisms controlling a bacterial cell cycle. Curr Opin Microbiol 7:572–578
Viollier PH, Thanbichler M, McGrath PT, West L, Meewan M, McAdams HH, Shapiro L (2004) Rapid and sequential movement of individual chromosomal loci to specific subcellular locations during bacterial DNA replication. Proc Natl Acad Sci USA 101:9257–9262
Wu LJ (2004) Structure and segregation of the bacterial nucleoid. Curr Opin Genet Dev 14:126–132
Wu LJ, Errington J (1998) Use of asymmetric cell division and spoIIIE mutants to probe chromosome orientation and organization in Bacillus subtilis. Mol Microbiol 27:777–786
Zhang R, Zhang CT (2003) Multiple replication origins of the archaeon Halobacterium species NRC-1. Biochem Biophys Res Commun 302:728–734
Zhang R, Zhang CT (2005) Identification of replication origins in archaeal genomes based on the Z-curve method. Archaea 1:335–346
Acknowledgments
This work was supported by Grant MCB-0217278 from the National Science Foundation to J.G.L. and a fellowship from the Pennsylvania Space Consortium to H.H. We thank Thomas Murphy VII for assistance with automated global pairwise sequence comparisons.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Reviewing Editor: Dr. Martin Kreitman]
Rights and permissions
About this article
Cite this article
Hendrickson, H., Lawrence, J.G. Selection for Chromosome Architecture in Bacteria. J Mol Evol 62, 615–629 (2006). https://doi.org/10.1007/s00239-005-0192-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-005-0192-2