Abstract
In free-living microorganisms, such as Escherichia coli and Saccharomyces cerevisiae, both synonymous and nonsynonymous substitution frequencies correlate with expression levels. Here, we have tested the hypothesis that the correlation between amino acid substitution rates and expression is a by-product of selection for codon bias and translational efficiency in highly expressed genes. To this end, we have examined the correlation between protein evolutionary rates and expression in the human gastric pathogen Helicobacter pylori, where the absence of selection on synonymous sites enables the two types of substitutions to be uncoupled. The results revealed a statistically significant negative correlation between expression levels and nonsynonymous substitutions in both H. pylori and E. coli. We also found that neighboring genes located on the same, but not on opposite strands, evolve at significantly more similar rates than random gene pairs, as expected by co-expression of genes located in the same operon. However, the two species differ in that synonymous substitutions show a strand-specific pattern in E. coli, whereas the weak similarity in synonymous substitutions for neighbors in H. pylori is independent of gene orientation. These results suggest a direct influence of expression levels on nonsynonymous substitution frequencies independent of codon bias and selective constraints on synonymous sites.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Strains and species diverge in part by accumulating nucleotide substitutions, resulting in a spectrum of substitution frequencies for orthologous genes. Many “universal” factors have been proposed to underlie the different rates at which individual proteins evolve, including expression level (Duret and Mouchiroud 2000; Pal et al. 2001; Krylov et al. 2003; Zhang and Li 2004; Jordan et al. 2004; Nuzdhin et al. 2004; Drummond et al. 2005), metabolic cost (Akashi and Gojobori 2002; Akashi 2003), chromosomal neighborhood (Williams and Hurst 2000; Rison et al. 2002), and essentiality for viability (Hirsh and Fraser 2001; Jordan et al. 2002; Pal et al. 2003; Yang et al. 2003). Additionally, studies involving network analyses have suggested that other, more contextual factors may also be important, such as the number of interacting proteins (in protein interaction networks) (Teichmann 2002; Fraser et al. 2002, 2003) and biochemical centrality (in metabolic networks).
Although the combined action of such factors is likely to drive proteome evolution, gene expression levels stand out as a consistently reported very important contributor. However, because many of the suggested factors are interdependent and because the experimental data are as yet very noisy (Herbeck and Wall 2005), it has proven difficult to assess the relative importance of each factor, as shown by the heated debate about the significance of the correlations observed so far. In particular, contradictory results have been reported on the role of protein–protein interactions (Fraser et al. 2003; Jordan et al. 2003; Bloom and Adami 2003; Fraser and Hirsh 2004; Bloom and Adami 2004) and essentiality in bacteria (Jordan et al. 2002; Rocha and Danchin 2004) and yeast (Hirsh and Fraser 2001; Pal et al. 2003; Hirsh 2003). Recent studies on four species of yeast suggest that essentiality and expression exert independent influences on evolutionary rates, although the relative strength of each still remains undetermined (Wall et al. 2005; Zhang and He 2005).
In the absence of data on protein expression, global analyses of mRNA levels with the aid of microarrays are commonly used as indicators of expression profiles. Codon Adaption Indexes (CAI) provides another set of estimates that are frequently used instead of real expression level data (Rocha and Danchin 2004; Wall et al. 2005). The CAI value reflects the frequency at which a selected subset of codons is being used by an individual gene (Sharp and Li 1987). The use of CAI as a quantifier of gene expression in Escherichia coli (Rocha and Danchin 2004) is justified by the observation that codon usage bias correlates with isoacceptor tRNA (Ikemura 1981, 1982, 1985; Dong et al. 1996) and mRNA concentrations (Kanaya et al. 1999; dos Reis et al. 2003; Goetz and Fuglsang 2005). The more highly expressed the gene, the higher the CAI value and the lower the synonymous substitution rate (Sharp 1991). Likewise, codon usage and CAI values correlate strongly with mRNA abundance in yeast (Coghlan and Wolfe 2000). Also, in Drosophila melanogaster, the frequency of optimal codons is correlated with expression levels (Duret and Mouchiroud 1999).
In free-living unicellular organisms with large population sizes, such as E. coli and Saccharomyces cerevisiae, selection for rapid growth has long been thought to exert a strong influence on codon usage patterns in highly expressed genes (Kurland 1987; Andersson and Kurland 1990) and thereby on fixation rates for synonymous nucleotide substitutions. Because synonymous substitution frequencies often correlate with nonsynonymous substitution frequencies (Li et al. 1985; Mouchiroud et al. 1995), the expression-level effect on amino acid substitutions may be either direct or indirect.
One hypothesis suggests that selection for rapidly translated codons favors certain amino acids and hence determines the frequency at which nucleotide substitutions accumulate at nonsynonymous sites (Lobry and Gautier 1994). The “accuracy hypothesis” postulates the inverse, namely, that the selection is for protein conservation and thereby for codons associated with low missense error rates during protein synthesis (Akashi 1994). Alternatively, protein evolutionary rates may reflect physical constraints, such as the number of active sites and interactions with other proteins (Lobry and Gautier 1994), or be a function of enzymatic performance loss weighted by protein abundance (Rocha and Danchin 2004), and if so, is expected to correlate with expression independently of the rate at which synonymous substitutions accumulate (Rocha and Danchin 2004).
To distinguish between these hypotheses, it is necessary to analyze substitution frequency variation in species that evolve under environmental regimes in which selection on synonymous positions is less effective. Population sizes, environmental conditions, and gene expression levels vary widely among bacteria, as does also the extent of selection on synonymous codon usage (Sharp et al. 2005). Furthermore, recombination (Maynard Smith et al. 1993; Suerbaum et al. 1998), combinatorial gene shuffling, and positive selection (Finlay and Falkow 1997) may drive the divergence of some genes, such as host–interaction genes in pathogenic bacteria, beyond what is achievable by neutral sequence evolution alone.
Thus, what is true for E. coli and yeast is not necessarily true for other microorganisms. Indeed, the absence of translational selection on codon usage has been demonstrated in a broad range of host-associated bacteria (Sharp et al. 2005), such as Rickettsia prowazekii (Andersson and Sharp 1996), Buchnera aphidicola (Rispe et al. 2004), Chlamydia (Lu et al. 2005), and Helicobacter pylori (Lafay et al. 2000). To discern universal trends, it is essential to complement studies of free-living microorganisms with short generation times, such as E. coli, with analyses of substitution frequency variations in species such as these that evolve under fundamentally different selective regimes.
Here, we set out to test whether protein evolutionary rates and expression levels correlate directly (Rocha and Danchin 2004) or indirectly, due to, for example, selection for a subset of codons that are optimal for translation (Lobry and Gautier 1994). To this end, we estimated and analyzed the correlation between expression levels and evolutionary rates in E. coli, an organism with pronounced codon bias, and H. pylori, an organism that lacks codon bias (Lafay et al. 2000). H. pylori establishes long-term persistent infection in the stomach, which can cause gastritis and gastric ulcers and is a leading cause of mucosa-associated gastric lymphoma and gastric adenocarcinoma (Marshall and Warren 1984; Peek and Blaser 2002; Suerbaum and Michetti 2002). Gastric colonization with H. pylori is extremely widespread among humans, with an estimated half of the world’s population being infected (Suerbaum and Michetti 2002; Peek and Blaser 2002).
Despite the absence of codon bias in H. pylori (Lafay et al. 2000), our study revealed a significant correlation between expression levels and nonsynonymous substitution frequencies, as was also observed in E. coli. Hence, codon bias is not the only link between expression and protein evolution; other factors clearly exert an influence on protein evolutionary rates in bacteria.
Materials and Methods
Substitution Frequencies
The genome sequences of H. pylori strains 26995 (NC_000915) and J99 (NC_000921) (Tomb et al. 1997; Alm et al. 1999), E. coli K12 (NC_000913), and Salmonella typhimurium (NC_003197) were downloaded from GenBank (ftp://ftp.ncbi.gov/genomes/Bacteria). H. pylori orthologs were identified by reciprocal best hits using BLAST (Altschul et al. 1997). For E. coli and S. typhimurium, we used the set of orthologs identified by Rocha and Danchin (2004). This process resulted in the identification of 1375 orthologs for H. pylori 26995/J99 and 1980 orthologs for E. coli and S. typhimurium (Table 1).
Protein sequences were aligned using ClustalW v1.81 (Thompson et al. 1994) and then transformed into nucleotide sequence alignments. Nonsynonymous (Ka) and synonymous (Ks) substitution frequencies were estimated by the method of Yang and Nielsen (2000) using PAML (Yang 1997). Note that the term “substitution frequency” and “substitution rate” are used interchangeably in this article, despite the lack of estimated absolute divergence time between compared species. To avoid biased results from a few clusters of genes with extreme K-values, the most highly divergent genes were eliminated in both data sets, resulting in pruned ortholog data sets of 1208 genes in H. pylori and 1908 genes in E. coli (Table 1). Different subsets of the ortholog data sets were used in the various statistical analyses, as detailed in Table 1.
Protein Expression Data
A list of 149 abundant proteins in H. pylori was downloaded from the Proteome 2D-PAGE database (Pleissner et al. 2004; http://www.mpiib-berlin.mpg.de/2D-PAGE/; “Helicobacter pylori 26695 cellular proteins, agar plate, day 3”), representing the detectable cellular proteome of H. pylori and here referred to as the set of highly expressed genes (HEG). A reference set of 168 lowly expressed genes (LEG) was constructed by first selecting all proteins not in HEG, then excluding predicted transmembrane proteins and proteins with predicted biochemical properties outside the detectable range of the two-dimensional polyacrylamide gel electrophoresis (Mw: 10-60 MDa; pI: 5-8) (P. Jungblut, personal communication). The filtering was performed with the TMHMM Server v. 2.0 (Krogh et al. 2001; http://www.cbs.dtu.dk/services/TMHMM-2.0/ ) and ExPASy Compute pI/Mw (Bjellqvist et al. 1993; http://www.expasy.org/tools/pi_tool.html).
mRNA Expression Levels in E. coli
For relative expression levels in E. coli, we used log2-transformed microarray mRNA levels for 1489 genes from E. coli cultured in LB media and grown to OD600 = 0.8 (Bernstein et al. 2002), downloaded from the Stanford Microarray Database (Ball et al. 2005; http://genome-www.stanford.edu/microarray ). These were split into LEG and HEG groups, with 100 genes in each group, using the criteria described in Table 2.
Median ΔK Values for Chromosomal Nearest Neighbors
All 1576 protein-coding H. pylori genes were sorted by the order in which they appear on the chromosome, and adjacent genes in this list were considered nearest neighbors. Neighbor pairs with at least one non-ortholog were excluded (because no ΔK value can be calculated for such a pair, see below) and the remaining data set was divided into two subsets: neighbors located on the same strand and neighbors located on opposite strands, constituting 826 and 193 gene pairs, respectively. For each subset, the positive difference in substitution rates (ΔKa and ΔKs; collectively, “ΔK values”) was calculated for every gene pair and used to compute the median difference in substitution frequency across the dataset. The genes in each true set were permuted to form 10,000 sets of random pairs, and the median ΔK was calculated for each of these random sets. Thus, every random set consisted of equally many gene pairs as the original set and included the exact same subset of genes. The same procedure was followed for E. coli, producing data sets of 1022 nearest-neighbor pairs located on the same strand and 351 nearest-neighbor pairs located on opposite strands.
The proportion of the 10,000 randomized data sets with lower median ΔK values than the original data set is an estimate of the p-value that can be attached to the hypothesis that chromosomal neighbors have more similar substitution frequencies than expected by chance. The complete analysis was repeated for second-nearest, third-nearest, and so on up to 20th-nearest neighbors in both species.
Median ΔExpression Values for Chromosomal Nearest Neighbors and Network Interaction Pairs
In E. coli, where continuous expression data were available, we also tested whether chromosomal neighbors have more similar expression levels than expected by chance, adopting the same strategy as for substitution frequencies outlined above. Neighbor pairs where expression data were lacking for one or both genes were excluded. The median difference in mRNA expression levels was computed for neighbors on the same (n = 532) and opposite (n = 187) strands, and compared to 10,000 corresponding randomized sets.
Using a similar approach, we analyzed adjacent pairs in the protein interaction (n = 362) and metabolic (n = 852) networks (see below), excluding adjacent pairs that were also up to 4th-nearest chromosomal neighbors.
Co-regulation of Genes Inferred from Microarray Data
In order to investigate the degree of co-regulation of gene pairs, expression profile data from numerous microarray experiments were collected from the Stanford Microarray Database (Ball et al. 2005; http://genome-www.stanford.edu/microarray ). A total of 58 and 45 experiments were selected for H. pylori and E. coli, respectively, representing a wide variety of experimental conditions, cellular states, and different stress responses. Before download, data were processed with standard filters (Regression correlation > 0.6 AND [(Channel 1 Mean Intensity/Median Background Intensity > 2.5) OR (Channel 2 Normalized (Mean Intensity/Median Background Intensity) > 2.5)]). For comparative reasons, all arrays were z-normalized (μ = 0, σ = 1). To quantify the level of co-regulation between any two genes, we calculated the correlation between the expression profiles of the two genes (Bhardwaj and Lu 2005), using Spearman correlation.
Essentiality
Putatively essential genes in H. pylori identified by genome-saturating transposon mutagenesis were taken from Salama et al. (2004). All genes with no detectable transposon insertion were included in the essential set, except 11 genes noted as false positives by Salama and colleagues, leaving 324 genes of which 271 have orthologs in both strains. It is estimated that 6% (89 genes) of the total set of genes in H. pylori would have had no transposon by chance given Poisson distribution (Salama et al. 2004). The remaining 937 H. pylori orthologs with transposon insertions were considered nonessential. A set of 250 essential genes in E. coli was downloaded from the PEC database ( http://www.shigen.nig.ac.jp/ecoli/pec/index.jsp;Version 3.6.0), 215 of which have orthologs in S. typhimurium. Excluding orthologs classified as “essential” or “unknown” in the PEC database, the remaining 1321 orthologs were considered nonessential.
Reconstruction of Protein Interaction Networks and Metabolic Networks
Protein interaction networks for H. pylori and E. coli were reconstructed using the highest-confidence data from published yeast two-hybrid studies (Rain et al. 2001; Butland et al. 2005). In these (undirected) networks, each node is a protein, and an edge connects two nodes if the corresponding proteins interact in the two-hybrid screen. These networks comprised 433 interactions among 424 proteins for H. pylori and 829 among 269 proteins for E. coli.
Metabolic networks for H. pylori and E. coli were reconstructed from the KEGG ( http://www.genome.ad.jp/kegg ) LIGAND database (Goto et al. 1998). In these (directed) networks, each node is an enzyme, and an arc connects two nodes if a metabolic product of the first enzyme (the upstream or “from” node) is a substrate for the second enzyme (the downstream or “to” node). Hence, each arc represents a metabolite, and each node–arc–node segment is interpreted to mean that the upstream enzyme makes a metabolite for the downstream enzyme. This method of constructing metabolic networks emphasizes the causal relationships among enzymes, which the method of connecting enzymes simply if they share a metabolite (Jeong et al. 2000) does not. These networks were then “pruned” computationally to remove extremely common metabolites such as water and pyrophosphate; the rationale is that water, for example, is produced and consumed by a large number of enzymes and is not produced expressly by one enzyme for another. These networks comprised 1222 arcs among 254 enzymes for H. pylori and 3944 arcs among 670 enzymes for E. coli. Graph manipulations and calculations were performed using the Graph Template Library (GTL) and in-house implementations of standard algorithms (Newmann 2001).
Median ΔK Values for Gene Pairs in the Protein Interaction and Metabolic Networks
Of the 424 proteins in the protein interaction network, 287 had orthologs between the two strains, and these formed 246 interactions. To avoid interference from chromosomal location effects, all network pairs that were also identified as up to fourth chromosomal neighbors were removed. The remaining set consisted of 274 proteins forming 234 interaction pairs, and it is this set that was analyzed. The network pairs were repeatedly permuted to form 10,000 sets of random pairs, and the median ΔK was calculated for each of these sets. (This is the same as randomizing the protein interaction network.) Each randomized set contained the exact same subset of genes and the same number of pairs as the original set.
The same procedure was repeated for the pruned metabolic network. For this analysis, directionality and multiple arcs between two enzymes were ignored (because we could see no rationale why a bidirectional interaction should be included twice in this analysis and a unidirectional interaction only once), resulting in a network of 234 nodes (enzymes) and 747 edges. As above, the set of network pairs was repeatedly permuted to form 10,000 sets of random pairs, and the median ΔK was calculated for each set.
The analysis was repeated in E. coli, both for the protein interaction network (180 orthologs, 542 interactions) and for the metabolic network (393 orthologs, 1353 interactions).
Results
Variation in Synonymous and Nonsynonymous Substitution Frequencies
We estimated synonymous and nonsynonymous substitution frequencies in H. pylori (which lacks codon bias) and E. coli (which exhibits strong codon bias). Our survey comprised 1375 orthologous genes in H. pylori strains 26995 and J99; this set has a mean Ka value of 0.038 (median Ka = 0.019) and a mean Ks value of 0.22 (median Ks = 0.18). The second set comprised 1980 orthologous genes in E. coli and S. typhimurium; these have a mean Ka value of 0.068 (median Ka = 0.048) and a mean Ks value of 1.24 (median Ks = 1.18). Many of the H. pylori genes with the most extreme K-values (Ka > 0.05; Ks > 0.3; n = 66) are situated in short gene clusters of two to five genes and could potentially bias the results. Hence, in all further analyses we included only orthologs with nonextreme K-values from H. pylori (Ka < 0.05; Ks < 0.3; n = 1208), and to further avoid extreme-value bias we used median rather than average values in the analyses. For ease of comparison, we also pruned the E. coli ortholog dataset to include only nonextreme K-values (Ka < 0.3; Ks < 3.0; n = 1908). Only a weak correlation between synonymous and nonsynonymous substitution frequencies was observed in H. pylori (Spearman’s rho = 0.30, p < 10−4), whereas, as expected (Rocha and Danchin 2004), these correlate strongly in E. coli and S. typhimurium (Spearman’s rho = 0.53; p < 10−4).
Protein Evolutionary Rates Correlate with Expression Independent of Codon Bias
To test whether there is a correlation between expression level and protein evolutionary rates in H. pylori despite the absence of selection on synonymous sites, we compiled a list of 141 highly expressed genes (HEG) identified by 2DE-gel electrophoresis of the H. pylori proteome (Pleissner et al. 2004). These were compared to a corresponding set of 159 lowly expressed genes (LEG), which were not recovered in the same 2DE-gel electrophoresis analysis, although having similar predicted physicochemical properties (10 kDa < Mw < 60 kDa and 5 < pI < 8). We excluded from the LEG data set those genes coding for proteins with predicted transmembrane regions (Krogh et al. 2001), because membrane proteins may not enter the 2DE-gel or may behave unpredictably.
The analysis showed that the HEG set was associated with significantly lower Ka values than the LEG set in H. pylori (Wilcoxon rank sum test, p = 1.2 × 10−4) (Fig. 1a), whereas only a minor difference in Ks-values was observed for the two sets (Wilcoxon rank sum test, p = 0.025) (Fig. 1b). Repeating the analysis replacing LEG with a set of randomly selected genes not found in HEG confirmed these findings (Ka: p = 4.6 × 10−7; Ks: p = 0.013), and thus the result is unlikely to be an artifact of our specific selection of reference genes. We conclude that in H. pylori, the more highly expressed the protein, the lower the fixation rate of nucleotide substitutions at nonsynonymous sites. We return to this finding in the following section.
Substitution frequency distribution for highly and lowly expressed gene sets. In H. pylori, highly expressed genes undergo (a) nonsynonymous substitution at a substantially slower rate than lowly expressed genes, whereas there is a negligible difference for (b) synonymous substitutions. In E. coli, highly expressed genes undergo both (c) nonsynonymous and (d) synonymous substitution at substantially slower rates than lowly expressed genes. The highly expressed gene sets (HEG) are shown in solid lines and the lowly expressed gene sets (LEG) in dashed lines.
How does this compare with E. coli? An examination of mRNA expression levels in E. coli (Bernstein et al. 2002) revealed a substantial and significant inverse correlation with both Ka (Spearman’s rho = −0.41, p < 10−4) and Ks values (Spearman’s rho = −0.46, p < 10−4). To make a direct comparison with the analysis performed in H. pylori, the E. coli proteins were also split into two groups, HEG and LEG, and the analysis was repeated. We noticed the same phenomenon; the HEG set exhibited significantly lower Ka (Wilcoxon rank sum test, p = 5.4 × 10−11) (Fig. 1c) and Ks values (Wilcoxon rank sum test, p = 1.2 × 10−13) (Fig. 1d). This finding is not an artifact of the proteins chosen for the two groups, because the results are robust to multiple different selection criteria for the HEG and LEG sets (Table 2).
Thus, the inverse correlation between Ka and expression is independent of selection on synonymous sites in H. pylori, whereas all three variables (Ka, Ks, and expression) are interdependent in E. coli. Therefore, factors other than codon usage bias and selection for rapid translation have to be invoked to explain protein evolutionary rate variation in H. pylori.
Protein Evolutionary Rates Correlate with Chromosomal Neighborhood
Because expression patterns follow the organization of genes into operons, we expected genes that are close to each other on the chromosome to evolve at similar rates. To determine whether such a dependency is due to co-expression, which presumably would be operonic, or due to some other neighborhood effect, we compared neighbors on the same strand with neighbors on opposite strands.
We made a list of all pairs of orthologs identified as chromosomal nearest neighbors in H. pylori (n = 1019) and E. coli (n = 1373). The H. pylori neighboring pairs were partitioned into two sets: one with 826 pairs located on the same strand (median ΔKa = 0.0093; median ΔKs = 0.042) and the other with 193 pairs located on opposite strands (median ΔKa = 0.011; median ΔKs = 0.041). For each of the two sets, we compared the true median ΔK values to the median ΔK values of 10,000 corresponding sets of random pairs. This analysis showed that chromosomal neighbors on the same strand have more similar Ka values than would be expected by chance (p < 10−4) (Fig. 2a), whereas the median ΔKa value for neighbors on opposite strands was not significantly different from those of the random sets (p = 0.45) (Fig. 2c). In contrast, the effect on Ks values seemed to be independent of the strandedness; same-strand neighbors showed similar ΔKs values (median ΔKs = 0.042, p < 10−4), as neighbors on opposite strands (median ΔKs = 0.041; p = 0.018) (Fig. 2b,d). For ΔKs, the difference in p-value for the two sets seems to be entirely explained by the lower statistical power in the opposite-strand set due to there being fewer opposite-strand pairs than same-strand pairs (Fig 2b,d).
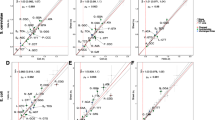
Substitution rates for chromosomal gene neighbors. The median ΔKa (a, c, e, g) and ΔKs (b, d, f, h) values for all pairs of chromosomal nearest neighboring genes located on the same strand (a, b, e, f) and on opposite strands (c, d, g, h) in H. pylori (a, b, c, d) and E. coli (e, f, g, h) are indicated (asterisks). For comparison, the median ΔKa and ΔKs values for 10,000 randomized sets were plotted as histograms. p: permutation p value, n: the number of gene pairs in each data set.
We repeated this analysis for E. coli. For orthologous neighbors on the same strand (n = 1022), the analysis showed a strong difference between the actual and the randomized median ΔKa (median ΔKa = 0.025, p < 10−4) and ΔKs values (median ΔKs = 0.35, p < 10−4) (Fig. 2e, f). For neighbors located on opposite strands (n = 351), the difference between the actual and the randomized data sets was much less significant for both ΔKa (median ΔKa = 0.032, p < 0.067) and ΔKs values (median ΔKs = 0.044, p = 0.064) (Fig. 2g, h).
How close must two genes be in order to see these effects? That is, what are the boundaries on chromosomal neighborhood within which genes evolve at similar rates on the same strand for ΔKa, and on either strand for ΔKs? To address this question, we repeated the above analysis with next-nearest neighbors, third-nearest neighbors, and so on, and found a rapid gradual decay of the proximity effect for ΔKa in both H. pylori and E. coli (SFig. 1a, e), whereas the effect remained for a distance of up to 10 neighbors for ΔKs in E. coli (SFig. 1f).
Hence, nonsynonymous substitution frequencies depend on strandedness. We hypothesize that the effect of chromosomal position on nonsynonymous substitutions is a byproduct of the organization of co-expressed genes into operons in both H. pylori and E. coli.
Chromosomal Neighborhood Correlate withCo-expression Patterns
So far, we have shown that protein evolutionary rates correlate with expression patterns as well as with chromosomal neighborhood, and hypothesized that the effect of chromosomal neighborhood may be secondary to expression.
To test this hypothesis, we analyzed how expression relates to chromosomal neighborhood, using all neighbor pairs with available mRNA expression data. In E. coli, we found that chromosomal neighbors located on the same strand have similar absolute mRNA expression levels in E. coli (n = 532, p < 10−4) (Fig. 3a), which is not the case for neighbors located on opposite strands (n = 187, p = 0.45) (Fig. 3b). In H. pylori, we found that chromosomal neighbors on the same strand are more likely to be both in the same expression group, HEG or LEG (n = 66), than to be in the opposite groups (n = 24) (p < 10−5). It remains to be shown whether expression levels are similar for different-strand neighbors in H. pylori, because the number of cases where both neighbors were found in either HEG or LEG was too small (n = 11) for any statistical conclusion to be drawn.
Absolute mRNA expression levels for chromosomal gene neighbors. The median difference in mRNA expression levels for chromosomal neighbors in E. coli located on the (a) same and (b) opposite strands. For each analysis, 10,000 randomized sets were plotted as histograms. p: permutation p value, n: the number of gene pairs in each data set.
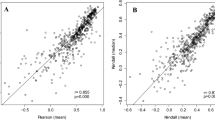
We also tested the co-regulation of neighbors on the same and opposite strands in H. pylori (Fig. 4). For every pair of nearest neighbors, we plotted the change in expression level of one member of the pair against the change in expression level of the other member of the pair under many experimental conditions for H. pylori, using data from 58 microarray experiments (average 56 data points per gene). To analyze coexpression patterns across the entire genome, we compared the distribution of correlations for same-strand nearest neighbors to a distribution of correlations for random pairs. As expected, same-strand nearest neighbors showed strong co-regulation (p < 10−15) (Fig. 4a), whereas the co-regulation of opposite-strand nearest neighbors was only negligibly stronger than would be expected by chance (p = 0.0060) (Fig. 4b). The same analysis was repeated in E. coli using data from 45 microarray experiments (average 34 data points per gene). Co-regulation was seen for both same-strand (p < 10−15) and opposite-strand neighbors (p < 10−15), although the effect in opposite-strand neighbors was less pronounced (SFig. 2).
These results further support the conclusion that the correlation between chromosomal location and protein evolutionary rates is mostly coincidental through transcriptional co-expression of genes located in the same operon.
Protein Evolutionary Rates and Essentiality
It is debated whether gene conservation relates mainly to expression or to essentiality. We compared the distributions of Ka and Ks values for a data set of 271 genes putatively required for viability in H. pylori (Salama et al. 2004) and found that they are not significantly different from the distributions of another set of 937 genes known to be nonessential (Wilcoxon rank sum test, Ka: p = 0.22; Ks: p = 0.092) (Fig. 5a, 5b). In comparison, a significant difference was observed for a set of 215 essential and 1321 nonessential genes in E. coli (Wilcoxon rank sum test, Ka: p < 10−15; Ks: p < 10−15) (Fig. 5c, d), as previously observed (Rocha and Danchin 2004).
Substitution frequency distribution for essential and nonessential gene sets. In H. pylori, there is a negligible difference between essential and nonessential genes with respect to (a) nonsynonymous and (b) synonymous substitutions. In E. coli, essential genes undergo both (c) nonsynonymous and (d) synonymous substitutions at substantially slower rates than essential genes. Essential genes are shown in solid lines and nonessential genes in dashed lines.
Protein Evolutionary Rates Do Not Reflect Network Features
Genes in the same operon typically encode proteins that interact physically (e.g., ribosomal proteins) or via metabolites in enzymatic networks. Hence, the correlation of substitution rates with expression and location in principle may be indirect effects of selective constraints on proteins that interact or have similar number of interacting partners. To explore this possibility, we inferred protein interaction and metabolic networks for H. pylori and E. coli (see Materials and Methods).
First, we tested the hypothesis that chromosomal nearest neighbors are closer to each other (although not necessarily adjacent) in the (undirected) protein interaction network than a randomly selected pair of proteins. For each pair of chromosomal nearest neighbors in H. pylori, we calculated the length of the shortest path between them in the protein interaction network (average path length 7.8; n = 60 shortest paths). Then, we permuted the pairs and repeated the calculation for 10,000 permuted sets. We found that chromosomal neighbors are indeed significantly closer to each other in the network than would be expected by chance (p = 0.011) (SFig. 3a).
We asked the same question for the metabolic network, and found a similar result. Chromosomal nearest neighbors in H. pylori were significantly closer to each other in the metabolic network than would be expected by chance (average path length = 3.3, n = 55, p < 10−4) (SFig. 3b). Likewise, we found that chromosomal nearest neighbors were closer to each other in both the protein interaction (SFig. 3c) and the metabolic network (SFig. 3d) of E. coli than would be anticipated by chance.
Despite the relative closeness of chromosomal neighbors in these networks, we observed no strong correlation between Ka or Ks values and network statistics for either the protein interaction or the enzymatic networks (Table 3). The exceptions were a weak correlation between Ka and degree in the metabolic networks of H. pylori (Kendall’s tau = −0.14, p = 1.2 × 10−3), and E. coli (Kendall’s tau = −0.11, p = 4.0 × 10−4). Additionally, there were weak correlations for Ka and Ks versus node degree and betweeness centrality (Newman 2001) in the E. coli protein interaction network (Table 3). Hence, basic properties of protein interaction network structure, such as the number of interacting partners, do not influence protein evolutionary rates to any major extent.
Finally, to directly investigate whether interacting proteins evolve at more similar rates than noninteracting proteins, we evaluated the differences in substitution frequencies for interacting protein pairs against combinations of randomly paired proteins. To avoid interference from the effect of chromosomal neighborhood, network pairs that were also close on the chromosome (up to fourth nearest neighbors irrespective of strand) were removed from the analysis, yielding 274 proteins with 234 interactions in H. pylori and 180 proteins with 542 interactions in E. coli. The median ΔK values for the interacting protein pairs were well within the range of ΔK values for the random permutations in both H. pylori (ΔKa: p = 0.16; ΔKs: p = 0.13) (SFig. 4a, b) and E. coli (ΔKa: p = 0.47; ΔKs: p = 0.72) (SFig. 4e, f).
We conducted the same analysis on the H. pylori and E. coli metabolic networks to ask whether interacting enzymes in those networks (i.e., enzymes directionally linked by a common metabolite) evolve at more similar rates than would be expected by chance. Again, interacting genes that are also close on the chromosome were removed from the analysis. We found that interactors had significantly more similar ΔKa values than random pairs in H. pylori (p = 0.029) (SFig. 4c), as well as in E. coli (p = 0.0069) (SFig. 4g). Unlike ΔKa values, ΔKs values were not significantly different from random in either species (p = 0.87 and 0.21, respectively) (SFig. 4d, h). Because interacting enzymes do not have more similar expression levels than expected by chance in E. coli (p = 0.27), the effect in ΔKa for the interacting enzymes cannot be directly attributed to expression levels, and may thus reflect a weak additional functional constraint on amino acid substitutions. Thus, although conventionally calculated network parameters, including degree or connectivity (number of interaction partners), do not seem to affect protein evolution, the simple existence of a pairwise interaction does have some effect on the evolutionary rate of the interactors.
Discussion
Although vast amounts of increasingly accurate data on gene expression levels, protein interactions, and metabolic networks continue to be generated, relating substitution frequency variation to the biology and evolution of the species under study remains a major challenge. Here, we have systematically investigated a number of proposed sources, such as expression, physical neighborhood, essentiality, and network-theoretical factors, using the human gastric pathogen H. pylori as our model system. The underlying rationale for selecting this species is that it is one of the most extensively studied human pathogens for which expression and essentiality data are available and that the absence of translationally selected codon usage bias allows us to dissect factors that influence nonsynonymous changes from selective constraints on synonymous sites.
Our results reject theories that try to explain protein evolutionary rates as an effect of selective constraints acting on synonymous sites to increase the translational efficiency (Lobry and Gautier 1994) or the accuracy of protein synthesis (Akashi 1994). The results have shown that the expression-level effect on nonsynonymous substitution frequencies predominates in H. pylori. However, unlike E. coli and S. cerevisiae, the frequency of synonymous substitutions in H. pylori does not correlate with expression levels.
What does this tell us about universal forces that influence protein evolutionary rates? Our results lend further support to the many previous observations made in organisms as diverse as bacteria, yeast, flies, and humans, namely, that protein evolution and expression levels are associated (Duret and Mouchiroud 2000; Pal et al. 2001; Krylov et al. 2003; Zhang and Li 2004; Jordan et al. 2004; Nuzdhin et al. 2004; Drummond et al. 2005). A recurrent, but still unresolved question is whether nonsynonymous substitutions and codon bias are correlated because both are influenced by gene expression levels, or whether the relationship between protein evolutionary rates and expression is a by-product of the association between codon bias and expression. Elaborate attempts have been made previously to dissect the three parameters and determine which of the correlations is the strongest in species that exhibits pronounced codon bias, such as E. coli, S. cerevisiae and Drosophila melanogaster (Rocha and Danchin 2004; Marais et al. 2004; Drummond et al. 2005).
One approach is to remove sites with preferred codons and test whether the Ka-expression correlation remains. Applying this test to yeast genes, Drummond and colleagues (2005) have shown that codon bias alone cannot explain the influence of expression level on protein evolutionary rates. A simpler approach is to test whether the correlation is observed in organisms without codon bias. As in H. pylori, a correlation between protein evolutionary rates and expression was also detected in Homo sapiens, despite the absence of selection on synonymous codon usage in this species (Duret and Mouchiroud 2000). Taken together, the only correlation that seems to hold universally true is between nonsynonymous substitutions and expression. We conclude that factors other than selection for translational efficiency or accuracy, manifested as a strong codon bias and reduced selective constraints on synonymous sites, have to be inferred to explain the inverse correlation between protein evolutionary rates and protein abundance in humans and their intracellular pathogens.
At first glance, these results resemble those obtained in Buchnera aphidicola (Rispe et al. 2004; Schaber et al. 2005) and Wigglesworthia glossinidia (Herbeck et al. 2003). These bacteria are obligate endosymbionts of aphids and tse tse flies, respectively, and like other obligate host-associated bacteria, they are driven by accelerated evolutionary rates and a strong mutational bias for AT. As in H. pylori, there is no observable codon usage bias in these bacteria (Rispe et al. 2004; Herbeck et al. 2003), and nucleotide frequencies at third codon positions reflect primarily strand-specific mutational biases in, for example, B. aphidicola (Rispe et al. 2004) and Blochmannia floridanus (Banerjee et al. 2004). Homologs to genes that are highly expressed in E. coli tend to evolve at slower rates in the endosymbionts, which has been taken as evidence to say that the level of gene expression influences the rate at which amino acids are substituted in B. aphidicola and W. glossinidia.
However, there are several complications with the interpretation of these analyses. First, the conclusions are based on a correlation between protein evolutionary rates in endosymbionts and CAI values of E. coli orthologs (Herbeck et al. 2003, Rispe et al. 2004). The problem is that the CAI value reflects a gene’s relative usage of a set of codons that are optimal for translation in that particular species. The only justification for using the CAI values as a substitute for expression in E. coli is that it has been demonstrated that genes with extreme CAI values tend to be highly expressed (dos Reis et al. 2003; Goetz and Fuglsang 2005). It is questionable whether the CAI value estimated for a gene in a species with a strong codon usage bias is an appropriate measure of the level of expression of the orthologous gene in a species without codon bias.
Another complication is that not only codon usage patterns but also the extent of gene regulation vary dramatically for species that grow in stable versus changing environments of different nutritional quality. For example, the E. coli genome encodes hundreds of transcriptional regulators (Blattner et al. 1997) and responds rapidly to changing conditions, whereas the endosymbiont genomes show an extreme transcriptional inflexibility with only a few transcriptional regulators (Wilcox et al. 2003). Only five genes in total showed differential expression in response to heat stress treatments, in line with the finding that only very few Buchnera genes have retained heat shock promoters (Wilcox et al. 2003). Likewise, only the metE gene, which is involved in methionine biosynthesis and for which the regulator, metR, is retained in B. aphidicola, responds to changes in amino acid concentrations (Moran et al. 2005).
Despite the restricted response in gene expression to environmental changes, a third study exploited data obtained in the heat stress experiment to make inferences about expression levels and protein evolutionary rates (Schaber et al. 2005). In this analysis, the highly and lowly expressed gene sets in B. aphidicola (Sg) were extracted from the 5% upper and lower levels of the normalized signal intensities in the microarray experiment. However, the intensities of spotted polymerase chain reaction products are known to be extremely variable and relative differences cannot be used as quantitative measurements of expression levels, except if the signal is normalized to that of a known reference sample, e.g., genomic DNA from the same species (Wei et al. 2001). Thus, although likely to be true, more convincing data are needed to confirm that protein evolutionary rates are inversely related to expression patterns in aphid endosymbionts, as demonstrated here for H. pylori.
As in a previous study (Jordan et al. 2003), we observed no major correlations between substitution frequency variation and protein network statistics in H. pylori. We also found that interacting proteins did not evolve at significantly more similar rates than expected by chance. However, not all amino acid changes affect the function of the protein. If there are relatively few amino acid changes that affect the function, then a protein evolving under strong functional constraints at these sites could still display an overall high Ka. Thus, our results cannot reject the hypothesis that physical interactions impose constraints on protein evolutionary rates, but even if so, these are unlikely to account for the strong correlation with expression. The concept of a unitary protein interaction network or metabolic network also may be an oversimplification. It is possible that basic network properties, such as the ones that we have calculated for H. pylori, may in fact correlate with substitution rates, but only if a proper network is being considered—a network that includes protein concentrations and kinetic constants, for example. This is an intriguing avenue for future research.
In addition to these factors, we also investigated the influence of essentiality but found no significant effect in H. pylori. The essential gene set comprised some 300 genes with no transposon insertions following random transposon mutagenesis under in vitro laboratory conditions (Salama et al. 2004). About 25% of these may be falsely assigned as essential, as estimated from the expected occurrence by chance of genes with no insertions (Salama et al. 2004). Nevertheless, we anticipate no major effect of essentiality on protein evolutionary rates in H. pylori.
However, just as the “minimal set of genes” required for life (Maniloff 1996; Mushegian and Koonin 1996; Mushegian 1999; Koonin 2000) is meaningful only within a defined environmental context (Klasson and Andersson 2004), the concept of “gene essentiality” may be as well (Papp et al. 2004). The set of essential proteins in a cell depends both on the composition of the interacting nonessential proteins as well as on the environment in which the organism thrives. Knocking out genes under in vitro growth conditions and then testing for cell viability may not tell much about the fitness effects of the deletions under in vivo growth conditions or in other genomic backgrounds. A case in point is the 47 H. pylori genes that have been shown experimentally to be essential for colonization of the gastric mucosa (Kavermann et al. 2003), only 5 of which were identified as essential by transposon mutagenesis (Salama et al. 2004). These include urease enzymes, membrane transport systems, and proteins required for motility and chemotaxis, as well as enzymes involved in the biosynthesis of lipopolysaccharide, all of critical importance for the establishment of the infection and thereby for survival of H. pylori in its natural host, although not necessarily essential under other growth conditions or in other bacterial populations. A systematic study suggested that about half of the genes in yeast that are nonesssential under laboratory conditions are essential in the yeast’s natural environment (Papp et al. 2004). It will be valuable to test this finding in bacteria such as E. coli and H. pylori.
However, the question remains: why do highly expressed proteins evolve slowly? An interesting hypothesis is that selection acts to prevent the misfolding of proteins that contain missense errors, thereby conferring protein robustness (Drummond et al. 2005). Here, the prediction is that evolutionary rates should correlate inversely with the number of folding events, i.e., the number of translational cycles. Because these depend on both the concentration of mRNAs and the rate of translation initiation, they are difficult to estimate solely from mRNA and protein abundance data, particularly in cases where these do not correlate. Further experimental analyses of the rate at which protein synthesis is initiated along with estimates of the rate at which proteins are degraded are required to test the hypothesis.
Finally, species-specific factors may go a long way to explain some of the extreme values in substitution frequencies. For example, positive or diversifying selection on outer membrane proteins in human pathogens may help evade the host immune defense system by combinatorial gene shuffling or by recombination of duplication fragments into expression cassettes. Because immune evasion is essential for bacteria that establish long-term persistent infection, such factors are potent drivers of evolutionary rates (Finlay and Falkow 1997). Only by filtering out the species-specific effects will it be possible to disclose the universality of the many possible factors that influence the rate at which proteins and species evolve.
References
Akashi H (1994) Synonymous codon usage in Drosophila melanogaster: natural selection and translational accuracy. Genetics 136:927–936
Akashi H (2003) Translational selection and yeast proteome evolution. Genetics 164:1291–1303
Akashi H, Gojobori T (2002) Metabolic efficiency and amino acid composition in the proteomes of Escherichia coli and Bacillus subtilis. Proc Natl Acad Sci USA 99:3695–3700
Alm RA, Ling LS, Moir DT, King BL, Brown ED, Doig PC, Smith DR, Noonan B, Guild BC, deJonge BL, Carmel G, Tummino PJ, Caruso A, Uria-Nickelsen M, Mills DM, Ives C, Gibson R, Merberg D, Mills SD, Jiang Q, Taylor DE, Vovis GF, Trust TJ (1999) Genomic-sequence comparison of two unrelated isolates of the human gastric pathogen Helicobacter pylori. Nature 397:176–180
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Andersson SG, Kurland CG (1990) Codon preferences in free-living microorganisms. Microbiol Rev 54:198–210
Andersson SGE, Sharp PM (1996) Codon usage and base composition in Rickettsia prowazekii. J Mol Evol 42:525–536
Ball CA, Awad IA, Demeter J, Gollub J, Hebert JM, Hernandez-Boussard T, Jin H, Matese JC, Nitzberg M, Wymore F, Zachariah ZK, Brown PO, Sherlock G (2005) The Stanford Microarray Database accommodates additional microarray platforms and data formats. Nucleic Acids Res 33:D580–582
Banerjee T, Basak S, Gupta SK, Ghosh TC (2004) Evolutionary forces in shaping the codon and amino acid usages in Blochmannia floridanus. J Biomol Struct Dyn 22:13–23
Bernstein JA, Khodursky AB, Lin PH, Lin-Chao S, Cohen SN (2002) Global analysis of mRNA decay and abundance in Escherichia coli at single-gene resolution using two-color fluorescent DNA microarrays. Proc Natl Acad Sci USA 99:9697–9702
Bhardwaj N, Lu H (2005) Correlation between gene expression profiles and protein-protein interactions within and across genomes. Bioinformatics 21:2730–2738
Bjellqvist B, Hughes GJ, Pasquali C, Paquet N, Ravier F, Sanchez J-C, Frutiger S, Hochstrasser DF (1993) The focusing positions of polypeptides in immobilized pH gradients can be predicted from their amino acid sequences. Electrophoresis 14:1023–1031
Blattner FR, Plunkett III G, Bloch CA, Perna NT, Burland V, Riley M, Collado-Viodes J, Glasner JD, Rode CK, Mayhew GF et al. (1997) The complete genome sequence of Escherichia coli K-12. Science 277:1453–1462
Bloom JD, Adami C (2003) Apparent dependence of protein evolutionary rate on number of interactions is linked to biases in protein-protein interactions data sets. BMC Evol Biol 3:21
Bloom JD, Adami C (2004) Evolutionary rate depends on number of protein-protein interactions independently of gene expression level: response. BMC Evol Biol 4:14
Butland G, Peregrin-Alvarez JM, Li J, Yang W, Yang X, Canadien V, Starostine A, Richards D, Beattie B, Krogan N, Davey M, Parkinson J, Greenblatt J, Emili A (2005) Interaction network containing conserved and essential protein complexes in Escherichia coli. Nature 433:531–537
Coghlan A, Wolfe KH (2000) Relationship of codon bias to mRNA concentration and protein length in Saccharomyces cerevisiae. Yeast 16:1131–1145
Dong H, Nilsson L, Kurland CG (1996) Co-variation of tRNA abundance and codon usage in Escherichia coli at different growth rates. J Mol Biol 260:649–663
Dos Reis, Wernisch L, Saava R (2003) Unexpected correlations between gene expression and codon usage bias from microarray data for the whole Escherichia coli K-12 genome. Nucleic Acids Res 31:6976–6985
Duret L, Mouchiroud D (1999) Expression pattern and, surprisingly, gene length shape codon usage in Caenorhabditis, Drosophila, and Arabidopsis. Proc Natl Acad Sci USA 96:4482–4487
Duret L, Mouchiroud D (2000) Determinants of substitution rates in mammalian genes: expression patterns affect selection intensity but not mutation rate. Mol Biol Evol 17:68–74
Drummond DA, Bloom JD, Adami C, Wilke CO, Arnold FH (2005) Why highly expressed proteins evolve slowly. Proc Natl Acad Sci USA 102:14338–14343
Finlay BB, Falkow S (1997) Common themes in microbial pathogenicity revisited. Microbiol Mol Biol Rev 61:136–169
Fraser HB, Hirsh AE, Steinmetz LM, Scharfe C, Feldman MW (2002) Evolutionary rate in the protein interaction network. Science 296:750–752
Fraser HB, Wall DP, Hirsh AE (2003) A simple dependence between protein evolution rate and the number of protein-protein interactions. BMC Evol Biol 3:11
Fraser HB, Hirsh AE (2004) Evolutionary rate depends on number of protein-protein interactions independently of gene expression level. BMC Evol Biol 4:13
Goetz RM, Fuglsang A (2005) Correlation of codon bias measures with mRNA levels: analysis of transcriptome data from Escherichia coli. Biochem Biophys Acta 327:4–7
Goto S, Nishioka T, Kanehisa M (1998) LIGAND: chemical database for enzyme reactions. Bioinformatics 14:591–599
Herbeck JT, Wall DP, Wernegreen JJ (2003) Gene expression level influences amino acid usage, but not codon usage, in the tsetse fly endosymbiont Wigglesworthia. Microbiology 149:2585–2598
Herbeck JT, Wall DP (2005) Converging on a general model of protein evolution. Trends Biotechnol 23:485–487
Hirsh AE, Fraser HB (2001) Protein dispensability and rate of evolution. Nature 411:1046–1049
Hirsh AE (2003) Rate of evolution and gene dispensability—reply. Nature 421:497–498
Ikemura T (1981) Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes. J Mol Biol 146:1–21
Ikemura T (1982) Correlation between the abundance of yeast transfer RNAs and the occurrence of the respective codons in protein genes. J Mol Biol 158:573–597
Ikemura T (1985) Codon usage and tRNA content in unicellular and multicellular organisms. Mol Biol Evol 2:13–34
Jeong H, Tombor B, Albert R, Oltvai ZN, Barabasi AL (2000) The large-scale organization of metabolic networks. Nature 407:651–654
Jordan IK, Rogozin IB, Wolf YI, Koonin EV (2002) Essential genes are more evolutionarily conserved than are nonessential genes in bacteria. Genome Res 12:962–968
Jordan IK, Wolf YI, Koonin EV (2003) No simple dependence between protein evolution rate and the number of protein-protein interactions: only the most prolific interactors tend to evolve slowly. BMC Evol Biol 3:1
Jordan IK, Marino-Ramirez L, Wolf YI, Koonin EV (2004) Conservation and coevolution in the scale-free human gene coexpression network. Mol Biol Evol 21:2058–2070
Kanaya S, Yamada Y, Kudo Y, Ikemura T (1999) Studies of codon usage and tRNA genes of 18 unicellular organisms and quantification of Bacillus subtilis tRNAs: gene expression level and species-specific diversity of codon usage based on multivariate analysis. Gene 238:143–155
Kavermann H, Burns BP, Angermuller K, Odenbreit S, Fischer W, Haas R (2003) Identification and characterization of Helicobacter pylori genes essential for gastric colonization. J Exp Med 197:813–822
Klasson L, Andersson SGE (2004) Evolution of minimal-gene-sets in host-dependent bacteria. Trends Microbiol 12:37–43
Koonin EV (2000) How many genes can make a cell: the minimal-gene-set concept. Annu Rev Genomics Hum Genet 1:99–116
Krogh A, Larsson B, von Heijne G, Sonnhammer ELL (2001) Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes. J Mol Biol 305:567–580
Krylov DM, Wolf YI, Rogozin IB, Koonin EV (2003) Gene loss, protein sequence divergence, gene dispensability, expression level, and interactivity are correlated in eukaryotic evolution. Genome Res 13:2229–22235
Kurland CG (1987) Strategies for efficiency and accuracy in gene expression. 1. The major codon preference: a growth optimization strategy. Trends Biochem Sci 12:126–128
Lafay B, Atherton JC, Sharp PM (2000) Absence of translationally selected synonymous codon usage bias in Helicobacter pylori. Microbiology 146:851–860
Li W-H, Wu CI, Luo CC (1985) A new method for estimating synonymous and nonsynonymous rates of nucleotide substitutions considering the relative likelihood of nucleotide codon change. Mol Biol Evol 2:150–174
Lobry JR, Gautier C (1994) Hydrophobicity, expressivity and aromaticity are the major trends of amino-acid usage in 999 Escherichia coli chromosome-encoded genes. Nucleic Acids Res 22:3174–3180
Lu H, Zhao WM, Zheng Y, Wang H, Qi M, Yu XP (2005) Analysis of synonymous codon usage bias in Chlamydia. Acta Biochem Biophys Sin 37:1–10
Maniloff J (1996) The minimal cell genome: on being the right size. Proc Natl Acad Sci USA 93:10004–10006
Marais G, Domazet-Loso T, Tautz D, Charlesworth B (2004) Correlated evolution of synonymous and nonsynonymous sites in Drosophila. J Mol Evol 59:771–779
Marshall B, Warren R (1984) Unidentified curved bacilli in the stomach of patients with gastritis and peptic ulceration. Lancet 8390:1311–1315
Moran NA, Dunbar HE, Wilcox JL (2005) Regulation of transcription in a reduced bacterial genome: Nutrient-provisioning genes of obligate symbiont Buchnera aphidicola. J Bacteriol 187:4229–4237
Mouchiroud D, Gautier C, Bernardi G (1995) Frequencies of synonymous substitutions in mammals are gene-specific and correlated with frequencies of nonsynonymous substitutions. J Mol Evol 40:107–113
Mushegian AR, Kooonin EV (1996) A minimal gene set for cellular life derived by comparison of complete bacterial genomes. Proc Natl Acad Sci USA 93:10268–10273
Mushegian AR (1999) The minimal genome concept. Curr Opin Genet Dev 9:709–714
Newman MEJ (2001) Scientific collaboration networks. II. Shortest paths, weighted networks, and centrality. Phys Rev E 64:016132
Nuzhdin SV, Wayne ML, Harmon KL, McIntyre LM (2004) Common pattern of evolution of gene expression level and protein sequence in Drosophila. Mol Biol Evol 21:1308–1317
Pal C, Papp B, Hurst LD (2001) Highly expressed genes in yeast evolve slowly. Genetics 158:927–931
Pal C, Papp B, Hurst LD (2003) Genomic function: Rate of evolution and gene dispensability. Nature 421:496–497
Papp B, Pal C, Hurst LD (2004) Metabolic network analysis of the causes and evolution of enzyme dispensability in yeast. Nature 429:661–664
Peek RM, Blaser MJ (2002) Helicobacter pylori and gastrointestinal tract adenocarcinomas. Nat Rev Camcer 2:28–37
Pleissner K-P, Eifert T, Buettner S, Schmidt F, Boehme M, Meyer TF, Kaufmann SHE, Jungblut PR (2004) Web-accessible proteome databases for microbial research. Proteomics 4:1305–1313
Rain JC, Selig L, De Reuse H, Battaglia V, Reverdy C, Simon S, Lenzen G, Petel F, Wojcik J, Schachter V, Chemama Y, Labigne A, Legrain P (2001) The protein–protein interaction map of Helicobacter pylori. Nature 409:211–215
Rison SCG, Teichmann SA, Thorton JM (2002) Homology, pathway distance and chromosomal localization of the small molecule metabolism enzymes in Escherichia coli. J Mol Biol 318:911–932
Rispe C, Delmotte F, van Ham RCHJ, Moya A (2004) Mutational and selective pressures on codon and amino acid usage in Buchnera, endosymbiotic bacteria of aphids. Genome Res 14:44–53
Rocha EP, Danchin A (2004) An analysis of determinants of amino acids substitution rates in bacterial proteins. Mol Biol Evol 21:108–116
Salama NR, Shepherd B, Falkow S (2004) Global transposon mutagenesis and essential gene analysis of Helicobacter pylori. J Bacteriol 186:7926–7935
Schaber J, Rispe C, Wernegree J, Burness A, Delmotte F, Silva FJ, Moya A (2005) Gene expression levels influence amino acid usage and evolutionary rates in endosymbiotic bacteria. Gene 352:109–117
Sharp PM (1991) Determinants of DNA sequence divergence between Escherichia coli and Salmonella typhimurium: codon usage, map position and concerted evolution. J Mol Evol 33:23–33
Sharp PM, Bailes E, Grocook RJ, Peden JF, Sockett RE (2005) Variation in the strength of selected codon usage bias among genes. Nucl Acids Res 33:1141–1153
Sharp PM, Li W-H (1987) The codon adaptation index—a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res 15:1281–1295
Smith MJ, Smith NH, O’Rourke M, Spratt BG (1993) How clonal are bacteria? Proc Natl Acad Sci USA 90:4384–4388
Suerbaum S, Michetti P (2002) Helicobacter pylori infection. N Engl J Med 347:1175–1186
Suerbaum S, Smith JM, Bapumia K, Morelli G, Smith NH, Kunstmann E, Dyrek I, Achtman M (1998) Free recombination within Helicobacter pylori. Proc Natl Acad Sci USA 95:12619–12624
Teichmann S (2002) The constraints protein-protein interactions place on sequence divergence. J Mol Biol 324:399–407
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tomb JF, White O, Kerlavage AR, Clayton RA, Sutton GG, Fleischmann RD, Ketchum KA, Klenk HP, Gill S, Dougherty BA, Nelson K, Quackenbush J, Zhou L, Kirkness EF, Peterson S, Loftus B, Richardson D, Dodson R, Khalak HG, Glodek A, McKenney K, Fitzegerald LM, Lee N, Adams MD, Hickey EK, Berg DE, Gocayne JD, Utterback TR, Peterson JD, Kelley JM, Cotton MD, Weidman JM, Fujii C, Bowman C, Watthey L, Wallin E, Hayes WS, Borodovsky M, Karp PD, Smith HO, Fraser CM, Venter JC (1997) The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature 388:539–547
Wall DP, Hirsh AE, Fraser HB, Kumm J, Giaever G, Eisen MB, Feldman MW. (2005) Functional genomic analysis of the rates of protein evolution. Proc Natl Acad Sci USA 102:5483–5488
Wei Y, Lee JM, Richmond C, Blattner FR, Rafalski JA, LaRossa RA (2001) High-density microarray-mediated gene expression profiling of Escherichia coli. J Bacteriol 183:545–556
Wilcox JL, Dunbar HE, Wolfinger RD, Moran NA (2003) Consequences of reductive evolution for gene expression in an obligate endosymbiont. Mol Microbiol 48:1491–1500
Williams EJB, Hurst LC (2000) The proteins of linked genes evolve at similar rates. Nature 407:900–903
Yang J, Gu Z, Li WH (2003) Rate of protein evolution versus fitness effect of gene deletion. Mol Biol Evol 20:772–774
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci 13:555–556
Yang Z, Nielsen R (2000) Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol Biol Evol 17:32–43
Zhang J, He X (2005) Significant impact of protein dispensability on the instantaneous rate of protein evolution. Mol Biol Evol 22:1147–1155
Zhang L, Li WH (2004) Mammalian house-keeping genes evolve more slowly than tissue-specific genes. Mol Biol Evol 21:236–239
Acknowledgments
We thank Eduardo P. Rocha for kindly providing data on E. coli/S. typhimarium/Y. pestis orthologs. This work was supported by the Swedish Research Council (VR), the Swedish Foundation for Strategic Research (SSF), the Knut and Alice Wallenberg Foundation (KAW), and the European Union (EU).
Author information
Authors and Affiliations
Corresponding author
Additional information
[Reviewing Editor: Dr. Nicolas Galtier]
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Sällström, B., Arnaout, R.A., Davids, W. et al. Protein Evolutionary Rates Correlate with Expression Independently of Synonymous Substitutions in Helicobacter pylori . J Mol Evol 62, 600–614 (2006). https://doi.org/10.1007/s00239-005-0104-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-005-0104-5