Abstract
The pentose phosphate cycle is considered as a major source of NADPH and pentose needed for nucleic acid biosynthesis. 6-Phosphogluconate dehydrogenase (6PGD), an enzyme participating in this cycle, catalyzes the oxidative decarboxylation of 6PGD to ribulose 5-phosphate with the subsequent release of CO2 and the reduction of NADP. We have determined the amino acid sequence of 6PGD of Bactrocera oleae and constructed a three-dimensional model based on the homologous known sheep structure. In a comparative study of 6PGD sequences from numerous species, all the conserved and variable regions of the enzyme were analyzed and the regions of functional importance were localized, in an attempt promoted also by the direct involvement of the enzyme in various human diseases. Thus, analysis of amino acid variability of 37 6PGD sequences revealed that all regions important for the catalytic activity, such as those forming the substrate and coenzyme binding sites, are highly conserved in all species examined. Moreover, several amino acid residues responsible for substrate and coenzyme specificity were also found to be identical in all species examined. The higher percentage of protein divergence is observed at two regions that accumulate mutations, located at the distant parts of the two domains of the enzyme with respect to their interface. These peripheral regions of nonfunctional importance are highly variable and are predicted as antigenic, thus reflecting possible regions for antibody recognition. Furthermore, locating the differences between diptera 6PGD sequences on the three-dimensional model suggests probable positions of different amino acid residues appearing at B. oleae fast, intermediate, and slow allozymic variants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The enzyme 6-phosphogluconate dehydrogenase (EC 1.1.1.44; 6PGD) is involved in the third step in the pentose phosphate pathway. It catalyzes the oxidative decarboxylation of 6-phosphogluconate to ribulose 5-phosphate, with the release of CO2 and the reduction of NADP (Novello and McClean 1968). Prokaryotic and eukaryotic 6PGDs are proteins of about 470 amino acids whose sequences are highly conserved, and this enzyme is dimeric and NADP dependent for almost all species (Rosemeyer 1987) and not metal ion dependent. The subunit molecular weight was found to be 52,000 Da and the crystal structure of the sheep liver enzyme with 5′-ADP has been determined at 2.5-Å resolution (Adams et al. 1991). The enzyme crystallizes in space group C2221 and the dimer has crystallographic twofold symmetry. The monomer has three domains: the amino-terminal β–α–β domain (residues 1–176), with a typical dinucleotide-binding fold (Rossmann et al. 1975), followed by a short helix and an additional, antiparallel to the first domain, β–α–β unit; the second helical domain (residues 177–434), which is the largest; and the third, the carboxy-terminal tail (residues 435–482), which burrows through the second subunit. In addition, a tightly bound sulfate ion was identified in the apoenzyme crystals which were grown from ammonium sulfate at pH 6.5. The role of this sulfate was proved to be very important since it makes intersubunit contacts and is bound by strictly conserved amino acid residues in the helical domain of one subunit and in the tail of the twofold-related subunit (Adams et al. 1994).
So far, it is generally recognized that the two main functions of the 6PGD pathway are to provide NADPH for fat synthesis and ribose for nucleic acid synthesis (Wood 1985). The amino acid sequences of more than 40 different 6PGDs have been reported, including sheep (Carne and Walker 1983), pig (Harbitz et al. 1990), human (Tsui et al. 1996), Bacillus subtilis (Fujita et al. 1986), and Escherichia coli (Nasoff et al. 1984). Nevertheless, the sheep liver enzyme is the only 6PGD whose three-dimensional structure has been determined (at 2.5-Å resolution) so far (Adams et al. 1991) (PDB code 2PGD).
The functional importance of the enzyme and its involvement in various human diseases was pointed out in several studies performed during the last 25 years. Thus, in a rare case of hereditary erythrocyte enzymopathy, namely, 6PGD deficiency, the activity of the enzyme was reduced to 35% in the affected members of a family under investigation (Caprari et al. 2001). The 6PGD activity was found to be decreased in aged human erythrocyte populations, with the aged enzyme having 11 fewer lysine residues than the young enzyme. Oxidation of 6PGD may be also considered an important process that takes place during aging of human erythrocytes (Gordillo and Machado 1991). Jonges et al. (1995) revealed that V max values of 6PGD for phosphogluconate decreased after partial hepatectomy in intermediate and pericentral zones of liver lobules (V max represents a maximum rate of a reaction that is reached when essentially all of the enzyme will be bound to substrate at steady state). In contrast, a selective increase in the activity of 6PGD was monitored in the inferior temporal cortex (Palmer 1999) as well as the neocortex (Martins et al. 1986; Balazs and Leon 1994) of patients with Alzheimer’s disease. Simultaneously, the activity of the enzyme was significantly increased as a result of the chronic neurogenic lesion of the muscle fibers in patients with neurogenic atrophy (Langohr 1980). Moreover, the specific activity of the enzyme was increased in a monozygotic twin that presented acute myeloblastic leukemia (Barretto et al. 1986). Renewed interest concerning the pentose phosphate pathway has been expressed since the pathway was shown to play a central role in the tumor proliferation process, providing ribose 5-phosphate in nucleic acids biosynthesis (Boros et al. 1997). Moreover, the antineoplastic effect of 6-aminonicotinamide has been considered a consequence of the inhibition of 6PGD and the secondary strong decrease in glycolysis, mediated by 6-phosphogluconate inhibition of phosphoglucose isomerase (Street et al. 1997). In addition, it has been suggested that 6PGD may be a target for chemotherapy against parasitic infections such as trypanosomiasis (Hanau et al. 1996). All these findings promote for further analysis of the 6Pgd locus from the molecular point of view. Furthermore, more information dealing with the putative implications of 6PGD in medical treatments and human diseases might be provided upon a better understanding of the crystal structure of this enzyme.
Our knowledge of 6PGD in insects is limited and is mostly concentrated on three Drosophila species, D. melanogaster (Scott and Lucchesi 1991), D. yakuba, and D. simulans (Begun and Aquadro 1994), and one Tephritidae species, the medfly Ceratitis capitata (Scott et al. 1993). In Bactrocera oleae biochemical studies have shown the existence of an allozyme polymorphism with three major alleles (plus a null allele appearing at a very low frequency), the fast (F), the intermediate (I), and the slow (S), mapping at a single locus (Bush and Kitto 1979; Loukas et al. 1985). Population genetics studies performed in natural populations of B. oleae as well as in laboratory colonies revealed striking allele frequency differences dealing with 6PGD. Under laboratory conditions the 6PGD polymorphism is driven to a new equilibrium state within only four generations. These changes in allele frequencies leave little doubt that the polymorphism is under the influence of strong selection pressure. In addition, artificial rearing of the insect in the laboratory results in drastic changes in its physiology and behavior, which can be correlated with an allozyme frequency change (Economopoulos 1980). Further insight into the evolutionary and adaptational significance of the 6Pgd system of this insect required the cloning of the 6PGD gene.
We report here the successful cloning and sequence analysis of the 6Pgd locus of B. oleae. The aim of this study is threefold: first, to develop a three-dimensional model of B. oleae 6PGD based on the sheep structure; second, to analyze all the conserved and variable regions of 6PGD from over 37 species and localize the regions of functional importance; and, third, to correlate the located regions with the enzyme’s function.
Materials and Methods
Fly Species
The Bactrocera oleae species used is the colony kept in our laboratory (Agricultural University of Athens, Greece) for about 20 years. The origin, extraction, and maintenance of the stock have been described in detail by Cosmides et al. (1997).
Amplification of the Genomic 6Pgd Gene
Preparation of genomic DNA from each of the species was done according to the protocol described by Holmes and Bonner (1973). The cDNA 6Pgd sequence of C. capitata (GenBank accession No. S67873) was used to design primers for PCR amplification of the corresponding genomic fragment of B. oleae The gene-specific 5′-GCAAAGCAATAACAAAATAATCATGTC-3′ and 5′-GGCATATTTTAGAAATACAAGCAAAGC-3′ upstream primers and the 5′-GTACATAACGACCTTACGCCTGGTATG-3′ downstream primer were used to generate the 6Pgd product of B. oleae. The amplification was carried out using high-fidelity conditions (Kwiatowski et al. 1991). To this end, Pwo polymerase (a proofreading enzyme) (Boehringer–Mannheim) was used to get amplification products of high fidelity. A hot start was used, with initial heating at 94°C for 5 min, followed by the addition of the polymerase and then 35 cycles of denaturation (at 94°C for 1 min), annealing (at 52°C for 1 min), and chain extension (at 72°C for 1.5 min), followed by a final extension step at 72°C for 10 min.
Cloning and Sequencing of the 6Pgd Gene
The resulting PCR product was cloned into the plasmid vector pGEM (Promega). Restriction and DNA modification enzymes were provided by MINOTECH and New England Biolabs. Agarose gel electrophoresis and other recombinant DNA methods were performed essentially as described by Sambrook et al. (1989). For the PCR amplified region, both strands were completely sequenced and the consensus sequence was obtained for two different clones, generated from two independent PCR reactions. Sequencing of the double-stranded plasmids was carried out according to the dideoxy-chain termination method, using either vector-specific (T7, SP6) or custom gene-specific (internal) primers. A LiCor 4200L sequencer of the Laboratory of Microchemistry (IMBB-FORTH, Crete, Greece) was used as well.
DNA and Protein Sequence Analysis
The DNA and protein sequences were analyzed with the GCG Sequence Analysis Software Computer Package. The alignment of the sequences was done using the Clustal X program (Thompson et al. 1997). The FASTA (Pearson and Lipman 1988) and SWISSPROT (2000) programs were also used for the alignment of the amino acid sequences. The nucleotide sequence reported in this study and the deduced amino acid sequence have been deposited in the EMBL Database (accession number: B. oleae 6Pgd, AJ517226).
Prediction of Antigenic Sites on the Protein
Analysis of data from experimentally determined antigenic sites on proteins has revealed that the hydrophobic residues Cys, Leu, and Val, if they occur on the surface of a protein, are more likely to be a part of antigenic sites. To predict antigenic determinants on proteins, a semiempirical method which makes use of physicochemical properties of amino acid residues and their frequencies of occurrence in experimentally known segmental epitopes was used (Kolaskar and Tongaonkar 1990). Prediction of antigenic sites was performed using EMBOSS’s (European Molecular Biology Open Software Suite) Antigenic option (Rice et al. 2000).
Construction of a B. oleae 6PGD Three-Dimensional (3D) Model
In order to construct a three-dimensional model for 6PGD of B. oleae and because of its high similarity to the sheep protein (69% sequence identity), two automated protein structure homology-modeling servers have been used: the Swiss-model via the ExPASy web server (Peitsch 1995) and the 3D-PSSM web server at Imperial College of Science, Technology and Medicine (Fischer et al. 1999). The derived models were checked for folding and packing errors using the QUANTA-CHARMm program (Brooks et al. 1983; Quanta 1998) to arrive at a combined model with no bad atom contacts and optimal side-chain conformation.
Results
The 6Pgd Locus in B. oleae
The use of degenerate primers based on the cDNA sequences of the 6Pgd of C. capitata produced an ∼2-kb fragment when 200 ng of genomic DNA from the species B. oleae was used as template. Nested PCR that was performed using a series of degenerate primers based on internal sequences of the C. capitata 6Pgd produced an initial positive evidence for the authenticity of the 6Pgd-like sequence. Therefore, we proceeded in the cloning and sequencing of the PCR products. In order to define the intron/exon splice junctions, the generally approved (C/A)AGGTAAGTA and YYYNYAGG consensus sequences were considered, thus enabling the extraction of the coding sequence of 6Pgd (Breathnach et al. 1978). The 6Pgd-coding region consists of 1443 nucleotides that correspond to 481 amino acid residues.
The consensus model of 6PGD of B. oleae
Among 143 positions where there are changes between the mammalian and the insect 6PGD amino acid sequences (140 mutations and three deletions in incects), we have selected 36 variable amino acid positions from diptera and mammalian sequences. This selection was made after locating these positions on the sheep 6PGD experimental structure and identifying that, of the 140 mutation positions, 135 are on or near the surface of the molecule and 4 (A286S, A328C, C401G, and C421A) are internal in secondary structure element contact points. Since on the internal mutations the changes between Gly–Ala and Ser–Cys amino acid side chains produced no major volume changes, we have adopted the following two criteria for the selection of variable positions: (1) variation of electronic charge (positive to negative, and vice versa, or loss of charge) and (2) variation of hydrophobicity/polarity hydrogen bond forming potential. We have located the 36 positions that follow these criteria on the 3D model of the dimer. These are shown together with the heavily conserved residues of 37 different species at the NADP binding site and 6PG active site in Fig. 1. The dimer was formed by application of the crystallographic symmetry as described in the PDB entries (2PGD, 1PGO).
Three-dimensional model structure of the B. oleae 6PGD dimer. The first domain of the two subunits is shown in dark colors and the second in light colors. Green indicates binding site amino acid residues (shown only in one subunit for clarity) that are conserved in all 6PGD molecules, and blue indicates the nonconserved amino acid residues of B. oleae with respect to the amino acid residues of sheep (shown only in one subunit for clarity). All molecular diagrams were performed using Molscript 2 (Kraulis 1991) and Raster3D (Merritt and Bacon 1997).
Comparison of 6PGD Amino Acid Sequences from Various Species
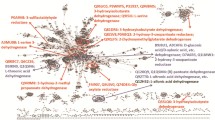
Thirty-seven different 6PGD sequences have been used in the comparison done, using FASTA (Pearson and Lipman 1988) on SWISSPROT, v 3.3t09, May 18, 2001. A consensus pattern for the 6PGD family can be found in the PROSITE database [LIVM]–x–D–x(2)–[GA]–[NQS]–K–G–T–G–x–W (PROSITE Release 40.7). In total, 279 positions on 6PGD sequences of various lengths (sequence length, 445 to 517, with a mean of 480) are highly conserved (Fig. 2). In particular, 69 totally conserved positions and 46 totally character conserved positions (charged, polar, large hydrophobic, and small hydrophobic) were found, with 53, 56, and 6 of them distributed in domains 1, 2, and 3, respectively. Moreover, 111 sites were found exhibiting more than 80% conservation; 33 are located on domain 1, 70 on domain 2, and 8 on domain 3. Finally, 53 sites showing conservation ranging between 60 and 80% were identified; 21 of these sites are located on domain 1, 31 on domain 2, and 1 on domain 3. The totally conserved positions have also been placed on the 3D model (Fig. 5).
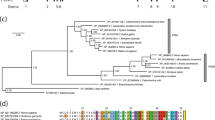
Multiple alignment of the 6PGD amino acid sequences from 37 species. All numbering is based on the numbering of the sheep 6PGD sequence. Black background indicates totally conserved (100%) amino acid residues, dark gray indicates more than 80% conservation, and light gray more than 60% conservation. Below the multiple alignment, a line with characters and numbers characterizes conservation: capital letters indicate totally conserved residues among all species, lowercase letters indicate the dominant residue on highly conserved positions, and numbers indicate total conservation of character (1: D, N; 3: S, T; 5: F,Y,W; 6: L, I, V). Domain allocation and helix names are given on line „domain/helix.” Active-site residues and associated coenzymes are indicated by capital A and C, respectively, and lowercase c indicates residues indirectly associated (e.g., through a water molecule) with the enzyme. The nucleotide sequences used in this study have the following GenBank accession numbers: human, P52209; sheep, P00349; Bactrocera oleae, AJ517226; Ceratitis capitata, P41570; Drosophila melanogaster, P41572; Drosophila simulans, P41573; Saccharomyces cerevisiae, P53319; Schizosaccharomyces pombe, P78812; Trypanosoma brucei brucei, P31072; Bacillus subtilis, P12013; Bacillus licheniformis, P52207; Cunninghamella elegans, O60037; Actinobacillus actinomycetem, P70718; Chlamydia muridarum, Q9PKX7; Chlamydia trachomatis, O84066; Citrobacter amalonaticus, P41581; Citrobacter koseri, P41582; Citrobacter freundii, P41583; Candida albicans,O13287; Chlamydophila pneumoniae, Q9Z813; Buchnera aphidocola, P57208; Escherichia coli, P37754; Escherichia vulneris, P41574; Lactococcus lactis, P96789; Klebsiella pneumoniae, P41576; Raoultella terrigena, P41577; Raoultella planticola, P41575; Shigella boydii, P41578; Shigella flexneri, P37756; Shigella dysenteriae, P41579; Shigella sonnei, P41580; Salmonella typhimurium, P14062; Synechocystis sp. PCC 6803, P52208; Synechococcus sp., P21577; Treponema pallidum, O83351; Schizaphis gominum, Q9ZHD9; Haemophilus influenzae, P43774.
Location of Highly Sequence Conserved Regions on the 3D Model
All amino acid residues that come into contact with either the substrate (15 amino acids for the reduced NADPH) or the ligand (10 amino acids for 6-phosphogluconate) or both (2 amino acids) (Adams et al. 1994) on the active site are totally conserved (Table 1). In addition, 162 amino acids that form the core of the two domains (the ß–α–ß dinucleotide-binding domain and the entirely helical active site-containing second domain) are also heavily conserved. In particular, in the first dinucleotide-binding domain, all the key features of the domain are conserved.
In the second domain, helices αh, αi, part of αj and αk, αl, αn, αo, αp, αq, and αr (Adams et al. 1991) are very well conserved. Finally, high conservation is observed in helix αs, in the third domain (amino acids 438–454) of the twofold-related subunit of the dimer that is also involved in the formation of the active site. Regions of the dimer interface formed by helices αk, αl, αq, and αr are also highly conserved.
As can be seen from Fig. 2, all residues that are responsible for the specificity of the coenzyme as well as substrate binding (see Discussion for details) are highly conserved. For reasons of consistency, we follow the position numbering used in the literature for the sheep 6PGD. Thus, Asn32, Arg33, and Thr34, all involved in NADP specificity, are identical or highly conserved in all species aligned. This is also true for amino acid residues Lys183 and Gly130. Moreover, residues Tyr191, Lys260, Ser128, and Glu190, which confer substrate specificity, are identical in all species examined. The turn βA–αa has been recognized as a conserved fingerprint „Gly–X–Gly–X–X–Ala” for some NADP enzymes (Hanukoglu and Gutfinger 1989; Scrutton et al. 1990). The corresponding residues in 6PGD are „Gly–X–Ala–X–X–Gly” (amino acid residues 9–14) and they were found to be totally conserved as well. Therefore, the adenine is bound in a pocket, one side of which is formed by side chains of conserved hydrophobic residues in the loop between βD and αa (Val174 and Ala79) and in αa (Phe83). Phe83 defines the end of this binding pocket (Adams et al. 1994).
Adams et al. (1994) suggested that the phosphate group of 6-phosphogluconate makes close contacts with two conserved arginines in the binding site. These arginine residues, which are located in positions 287 and 446 of the second subunit of the dimer and are crucial for balancing the phosphate charge and providing substrate specificity, were found conserved in the species examined (Figs. 2 and 3). Furthermore, there are also two conserved histidines in the substrate-binding site, His186 and His452, of the twofold-related subunit but neither histidine is in a suitable position to act as base or acid in the enzymic mechanism (Adams et al. 1994). Both histidines were found to be conserved in the present study (Figs. 2 and 3).
Multiple alignment of the sheep, human, C. capitata, B. oleae, and D. melanogaster 6PGD amino acid sequences. The shading pattern, numbering, domain allocation, helix names, active site residues, and coenzyme-associated residues are defined as in Fig. 2. Secondary structure elements are shown below the sequence alignment. Symbols used are H for helix, E for β-strand, and T, S for turn or coil. The secondary structure assignment is as on the X-ray structure (PDB code 2PGD).
Location of High Sequence VariabilityRegions in the 3D Model
The highly variable amino acid positions have been located on the 3D model (Fig. 1) based on the above criteria, and we observed several noteworthy findings. All positions are located on the surface of the molecule and on the accessible sides of otherwise extensively conserved regular secondary structural elements (α-helix and β-strand) forming the two domains. There are two main regions of the protein molecule where a high accumulation of mutations is observed. Both are located in the distant parts of the two domains with respect to their interface and on either side of the cavity accommodating the active site of the enzyme, where high amino acid conservation has been found. On domain 1 (amino acid positions 1–176), as shown in Figs. 3 and 4, the major changes occur on the external sides of helices aA (20–26), aB (37–49), and aE (111–119). Helix aB neighbors the putative NADP-binding site (defined by superposition of the model on the 2PGO [Berman et al. 2002] crystal structure, i.e., sheep 6PGD + reduced NADPH) where high conservation is observed (i.e., positions 12, 13, 14, 15, 32, 33, 34). On this helix a comparison of the amino acid sequences of dipteran vs. mammalian species (Fig. 3) shows the introduction of two extra negative charges (Asp39 and Asp40) for the mammalian species and the loss of a negative (Glu36) and a positive (Lys43) charges in diptera. A charged interaction between Asp39 and His55 stabilizes the helix–loop strand region 37–57 as regards the mammalian protein. Region 119–121, consisting of the C-terminal part of a helix plus a turn, is predicted to be antigenic and its high variability might reflect antibody recognition (Kolaskar and Tongaonkar 1990) (Table 2).
On domain 2 (amino acid positions 177–434), as shown in Fig. 5, major changes occur on the external loop 306–313 and on the packing region of helices (positions 321 and 407) of the otherwise highly conserved helices aQ (392–415) and aN (314–347), respectively. In the second region, proximal to the binding site, consisting of the external side of helices aI (212–224) and of loops 244–245 and 250–251, the introduction of additional charged residues has been revealed (Lys213, Lys217) in the sheep protein as well as a loss of the negatively charged residue Asp237. Region 308–320, which consists of an exposed loop together with region 405–413, the C-terminal part of an otherwise conserved helix is predicted to be antigenic and the high variability in these regions might also reflect antibody recognition (Kolaskar and Tongaonkar 1990) (Table 2).
Comparison of Dipteran 6PGD Amino Acid Sequences
Between diptera there are 59 positions exhibiting sequence variation, of which 22 fell under the criteria used earlier (Fig. 3). Of these, 11 (positions 38, 41, 57, 60, 119, 121, 209, 213, 244, 310, 320) are in the regions of high variability described earlier, 6 (positions 118, 120, 220, 242, 350, 380) are in positions where Drosophilidae diversify from the other animal (Tephritidae and mammalian) sequences, and 4 (positions 225, 240, 399, 414) differ between Drosophilidae and Tephritidae in external nonconserved regions of the protein chain. The latter are possible positions where the F, I, and S variants of B. oleae might diversify, since on positions 225 and 413 there is the introduction of a negative charge, while on positions 240 and 399 there is the introduction of a hydrophobic residue (Val) or a polar one (Asn), respectively, instead of a positively charged one (Arg and Lys).
Discussion
The nicotinamide adenine dinucleotide phosphate (NADP)-dependent oxidative decarboxylase 6PGD has been reported to be involved in several human diseases as pointed out in various studies performed over the last three decades (Caprari et al. 2001; Martins et al. 1986; Barretto et al. 1986). Thus, the putative implication of 6PGD in medical treatments (i.e., developing of selective inhibitors for therapeutic reasons) and human diseases made necessary a focus on the structural features of the enzyme from many species, aiming at a better understanding of its function. Alignment of the amino acid sequence of D. melanogaster 6PGD with those of the Bacillus subtilis, Escherichia coli, porcine, and ovine enzymes revealed remarkable conservation. In particular, a strikingly high degree of similarity was found in two regions as shown by Scott and Lucchesi (1991). In our case, also taking into consideration the 6PGD sequences of B. oleae and C. capitata, we observed that the 6PGD amino acid sequences of the two species show 96.7% identity. The sheep liver enzyme is the only 6PGD whose 3D structure has been determined experimentally and refined (at 2.5-Å resolution) so far (Adams et al. 1991). The phylogenetic distance between the sheep and the tephritid species under examination cannot be considered a prohibiting factor for further comparisons and analyses since the enzyme shows a notable conservation. When the amino acid sequences of sheep and human or E. coli 6PGD were aligned, 94.2 and 50% identity between these proteins was found, respectively (Tsui et al. 1996; Nasoff et al. 1984). Furthermore, the Drosophila and medfly protein coding sequences were found to be 73.5% identical (Scott et al. 1993).
We have attempted to relate the conservation of the amino acids to the catalytic role of the enzyme as far as (a) the functionality of the binding and active site and (b) the core of the protein forming the binding and active site. The first domain (residues 1–177) contains both the parallel β pleated sheet with strands βA to βF and α helices αa to αe forming the β–α–β fold, known as a „dinucleotide-binding fold” (Rossman et al. 1975). The second domain (residues 178–436) provides much of the dimer interface (Adams et al. 1991) and is entirely helical. Both form a solid, highly conserved base for the formation of the binding site structure.
It is known that there are two ways of stabilizing the nicotinamide of the coenzyme in this protein, depending on its redox state (Adams et al. 1994). Substrate specificity is achieved through binding of the carboxyl group to Ser128 and Glu190 and the phosphate moiety to Tyr191, Lys260, Arg287, and Arg446 of the second subunit. Lys183 and an ordered water molecule bonded to the main-chain nitrogen of the conserved Gly130 have been shown to be the most likely catalytic groups. Dehydrogenases participating in the pentose phosphate pathway contribute to its metabolic role by providing reduced nicotinamide adenine dinucleotide phosphate (NADPH) for synthesis. Their NADP specificity is apparently very crucial, and as detected previously (Adams et al. 1994), amino acids Asn32, Arg33, Thr34, Ala11, and Lys183 contribute to the NADP specificity of the enzyme. Thus, from the structural and functional point of view the importance of the conservation of all these amino acid residues is obvious. The phosphate group of 6-phosphogluconate (6PG) is in contact with the two binding site conserved arginines (R287 and R446) that balance the phosphate charge and provide specificity. The bound 6PG is stabilized by hydrogen bonds from Arg287 through water 528 to Glu190 and the 1-carboxyl of 6PG (Adams et al. 1994).
A series of Lys charged residues is also involved in interactions with the cofactor. Four lysine residues (Lys 39, Lys75, Lys 183, and Lys260), all at a distance of about 5 Å from the charged groups of NADP, are heavily associated with the enzymatic activity of 6PGD and surround the cofactor. Lys 260 is in the active site, while Lys 264, on the subunit contact interface, is actively involved in the formation of the dimer. Oxidation of 6PGD may be considered an important process that takes place during aging of human erythrocytes (Gordillo and Machado 1991). In an attempt to determine the presence of posttranslational modified 6PGD in aged human erythrocytes (as had been observed previously in rat liver cells, with the „old” enzyme being about 26% less active than the „young” enzyme [Gordillo et al. 1989]), it was found that indeed there is a modification of lysine residues, probably caused by oxidation. This modification could imply the loss of enzymatic activity of 6PGD reported, a situation resembling the oxidation of histidines in malic enzyme of rat, which is also accompanied by the loss of enzymatic activity (Gordillo et al. 1989). 6PGD, as many other members of the β-hydroxyacid dehydrogenase family, appears to be readily inactivated by chemical modifiers of lysine residues, such as 2,4,6-trinitrobenzene sulfonate (TNBS) (Gordillo and Machado 1991). This may be due to either the inactivation of substrate binding (Lys 39, 75, 183, and 260) or the malformation of the homodimer (Lys 264 and 406).
The substrate binding site is formed at the inner subunit interface with many highly conserved residues within 4 Å of 6PG (Table 2). The substrate makes hydrogen bonds to residues in the βF–αf loop of the first domain and to residues in the second domain of one subunit: one face of helix αh, the end of the long αj–αk loop, and Arg287 in αl. Hydrogen bonds are also made to residues in the tail of the second subunit of the dimer, which wraps around the other two of the first subunit, completing the formation of the substrate site (Adams et al. 1994).
Peripheral regions of nonstructural importance are highly variable and candidates for antigenicity. Thus, according to Kolaskar and Tongaonkar (1990) these regions exhibit antigenic properties (i.e., regions 308–320, 406–414, 394–400, etc.). In B. oleae 6PGD 34 nonconserved amino acid positions were detected. A percentage of them apparently belong in predicted antigenic regions (Nos. 4, 119, 121, 155, 166, 308–320, 406). In an attempt to classify them into categories we come to the following conclusions. In domain 1 (residues 1–177) we find most nonconserved positions located on solvent accessible areas of amphipathic α-helices (residues 24, 41, 42, 45, 57, 60, 115, 119, 121, 155, and 156) and very few (4, 138, 148, and 166) on external loops. In domain 2 nearly all are located on external loops (213, 244, 245, 250, 307–312, 379, 383, and 414) and very few (237, 319, 320, and 406; the latter three forming a cluster) on the external sides of helices. These findings are in line with the general observations that conservation occurs in important elements of secondary structure when rigidity is of absolute importance, restricting species variation on external loop regions. When variation occurs on the external side of otherwise conserved elements of secondary structure the trend is that this element is involved in molecular recognition of some sort.
Site-directed mutagenesis was used to change K183 of sheep liver 6PGD to A, E, H, C, Q, R, and M to probe its possible role as a general base catalyst (Zhang et al. 1999). The only mutant enzyme that gives a significant amount of catalysis is the K183R mutant, and the extent of catalysis is decreased by about three orders of magnitude. All other mutant enzymes exhibit rates that are decreased by about four orders of magnitude compared to that of the wild-type enzyme, and these data are consistent with the general base function of K183. In all sequences examined in this study, K183 is totally conserved. It is worthwhile noticing the multiple functional role of glutamate 190 (Glu190), another amino acid residue found to be conserved in the present study. Site-directed mutagenesis was used to change E190 of sheep liver 6PGD to A, D, H, K, Q, and R in order to probe its possible role as a general acid catalyst (Karsten et al. 1998). Each of the mutant proteins was characterized with respect to the pH dependence of kinetic parameters. Moreover, the data were also consistent with some participation of E190 in an isomerization required to form the active Michaelis complex. Glu190 was found to be totally conserved in this study, also, in accordance with the proposed role of this amino acid residue.
In this study the high conservation of 6PGD in numerous species, ranging from bacteria and yeasts to insects and mammals, has been placed in a structural framework. By doing so certain biochemical and clinical findings associated with the irregular action of the enzyme in various human diseases can be directly associated with regions around the active site or linked to the cofactor. Variability is observed mainly in peripheral regions of the structure, not associated directly with its function as an enzyme. These regions have been associated with antigenic sites on the molecular structure.
References
MJ Adams S Gover R Leaback C Phillips DO’N Somers (1991) ArticleTitleThe structure of 6-phosphogluconate dehydrogenase refined at 2.5 A resolution Acta Crystallogr B 47 817–820 Occurrence Handle10.1107/S0108768191010315 Occurrence Handle1793548
MJ Adams GH Ellis S Gover CE Naylor C Phillips (1994) ArticleTitleCrystallographic study of coenzyme, coenzyme analogue and substrate binding in 6-phosphogluconate dehydrogenase: Implications for NADP specificity and the enzyme mechanism Structure 2 651–668 Occurrence Handle10.1016/S0969-2126(00)00066-6 Occurrence Handle1:CAS:528:DyaK2MXot1ertg%3D%3D Occurrence Handle7922042
L Balazs M Leon (1994) ArticleTitleEvidence of an oxidative challenge in the Alzheimer’s brain Neurochem Res 19 1131–1137 Occurrence Handle10.1007/BF00965146 Occurrence Handle1:CAS:528:DyaK2cXlslKkurg%3D Occurrence Handle7824065
OC Barretto K Nonoyama GM Colletto (1986) ArticleTitleAcquired erythroenzymopathy in a monozygotic twin with acute myeloid leukemia Braz J Med Biol Res 19 63–67 Occurrence Handle1:STN:280:BiiC3c7msVA%3D Occurrence Handle2948602
DJ Begun CF Aquadro (1994) ArticleTitleEvolutionary inferences from DNA variation at the 6-phosphogluconate dehydrogenase locus in natural populations of Drosophila: selection and geographic differentiation Genetics 136 155–171 Occurrence Handle1:CAS:528:DyaK2cXmtVWhtL8%3D Occurrence Handle8138153
HM Berman T Battistuz TN Bhat WF Bluhm PE Bourne K Burkhardt Z Feng GL Gilliland L Iype S Jain P Fagan J Marvin D Padilla V Ravichandran B Schneider N Thanki H Weissig JD Westbrook C Zardecki (2002) ArticleTitleThe Protein Data Bank Acta Crystallogr D 58 IssueID6:1 899–908 Occurrence Handle10.1107/S0907444902003451 Occurrence Handle12037327
LG Boros J Puigjaner M Cascante WN Lee JL Brandes S Bassilian FI Yusuf RD Williams P Muscarella WS Melvin WJ Schirmer (1997) ArticleTitleOxythiamine and dehydroepiandrosterone inhibit the nonoxidative synthesis of ribose and tumor cell proliferation Cancer Res 57 4242–4248 Occurrence Handle1:CAS:528:DyaK2sXmsFChu74%3D Occurrence Handle9331084
R Breathnach C Benoist F Gannon P Chambon (1978) ArticleTitleOvalbumin gene: evidence for a leader sequence in mRNA and DNA sequences at the exon-intron boundaries Proc Natl Acad Sci USA 75 4853–4857 Occurrence Handle1:CAS:528:DyaE1MXhvFCitg%3D%3D Occurrence Handle283395
BR Brooks RE Bruccoleri BD Olafson DJ States S Swaminathan M Karplus (1983) ArticleTitleCHARMM a program for macromolecular energy minimization and dynamics calculations J Comput Chem 4 165–175 Occurrence Handle10.1002/jcc.540040211
Bush GL, Kitto GB (1979) Research on the genetic structure of wild and laboratory strains of the olive fruit fly. FAO Report: Development of pest management systems for olive culture program. Food and Agriculture Organization of the United Nations, Rome, p 27
A Carne B Walker (1983) ArticleTitleAmino acid sequence of ovine 6-phosphogluconate dehydrogenase J Biol Chem 258 12895–12906 Occurrence Handle1:CAS:528:DyaL2cXislal Occurrence Handle6685125
P Caprari MP Caforio P Cianciulli D Maffi MX Pasquino A Tarzia S Amadori AM Salvati (2001) ArticleTitle6-Phosphogluconate dehydrogenase deficiency in an Italian family Ann Hematol 80 41–44 Occurrence Handle10.1007/s002770000233 Occurrence Handle1:CAS:528:DC%2BD3MXhtFemtL8%3D Occurrence Handle11233775
N Cosmides M Loukas E Zouros (1997) ArticleTitleDifferences in fitness components among alcohol dehydrogenase genotypes of the olive fruit fly (Diptera: Tephritidae) under artificial rearing Ann Entomol Soc Am 90 363–371 Occurrence Handle1:CAS:528:DyaK2sXktV2jurY%3D
Economopoulos AP (1980) SIRM against the olive fruit fly: Differences between wild and lab-reared (normal or sterilized) insects. Proc. XVI Int. Congr. Entomol., Symp. Fruit Fly Problems, Kyoto and Naha, Japan, August, pp 17–26
D Fischer C Barret K Bryson A Elofsson A Godzik D Jones KJ Karplus LA Kelley RM Maccallum K Pawowski B Rost L Rychlewski MJ Sternberg (1999) ArticleTitleCAFASP-1: Critical assessment of fully automated structure prediction methods Proteins Suppl 3 209–217 Occurrence Handle10.1002/(SICI)1097-0134(1999)37:3+<209::AID-PROT27>3.0.CO;2-Y
Y Fujita T Fujita Y Miwa J Nihashi Y Aratani (1986) ArticleTitleOrganization and transcription of the gluconate operon, GNT, of Bacillus subtilis J Biol Chem 261 13644–13653
E Gordillo A Machado (1991) ArticleTitleImplication of lysine residues in the loss of 6-phosphogluconate dehydrogenase activity in aging human erythrocytes Mech Age Dev 59 291–297 Occurrence Handle10.1016/0047-6374(91)90139-Q Occurrence Handle1:CAS:528:DyaK3MXlvVShsLw%3D
E Gordillo A Ayala J Bautista A Machado (1989) ArticleTitleImplication of lysine residues in the loss of enzymatic activity in rat liver 6-phosphogluconate dehydrogenase found in aging J Biol Chem 264 17014–17019
S Hanau M Rippa M Bertelli F Dallocchio MP Barrett (1996) ArticleTitle6-Phosphogluconate dehydrogenase from Trypanosoma brucei. Kinetic analysis and inhibition by trypanocidal drugs Eur J Biochem 240 592–599 Occurrence Handle10.1111/j.1432-1033.1996.0592h.x Occurrence Handle1:CAS:528:DyaK28XlvFKlsrw%3D Occurrence Handle8856059
I Hanukoglu T Gutfinger (1989) ArticleTitlecDNA sequence of adrenodoxin reductase, identification of NADP-binding sites in oxidoreductases Eur J Biochem 180 479–484 Occurrence Handle10.1111/j.1432-1033.1989.tb14671.x Occurrence Handle1:CAS:528:DyaL1MXht12ksrg%3D Occurrence Handle2924777
I Harbitz R Chowdhary S Kran E Frengen I Gustavsson W Davies (1990) ArticleTitleIsolation, characterization and chromosomal assignment of a partial cDNA for porcine 6-phosphogluconate dehydrogenase Hereditas 112 83–88 Occurrence Handle1:CAS:528:DyaK3cXkvFant7s%3D Occurrence Handle2361879
DS Holmes J Bonner (1973) ArticleTitlePreparation, molecular weight, base composition and secondary structure of giant nuclear ribonucleic acid Biochemistry 12 2330–2338 Occurrence Handle10.1021/bi00736a023 Occurrence Handle1:CAS:528:DyaE3sXkt1ymt7Y%3D Occurrence Handle4710584
GN Jonges IM Vogels CJ Noorden Particlevan (1995) ArticleTitleEffects of partial hepatectomy, phenobarbital and 3-methylcholanthrene on kinetic patameters of glucose-6-phosphate and phosphogluconate dehydrogenase in situ in periportal, intermediate and pericentral zones of rat liver lobules Biochim Biophys Acta 1243 59–64 Occurrence Handle1:CAS:528:DyaK2MXjt1Shtb4%3D Occurrence Handle7827108
WE Karsten L Chooback PF Cook (1998) ArticleTitleGlutamate 190 is a general acid catalyst in the 6 phosphogluconate dehydrogense-catalyzed reaction Biochemistry 37 15691–15697 Occurrence Handle10.1021/bi9812827 Occurrence Handle1:CAS:528:DyaK1cXmsleitLg%3D Occurrence Handle9843373
AS Kolaskar PC Tongaonkar (1990) ArticleTitleA semi-empirical method for prediction of antigenic determinants on protein antigens FEBS Lett 276 172–174 Occurrence Handle10.1016/0014-5793(90)80535-Q Occurrence Handle1:CAS:528:DyaK3MXktVGntw%3D%3D Occurrence Handle1702393
PJ Kraulis (1991) ArticleTitleMOLSCRIPT: A program to produce both detailed and schematic plots of protein structure J Appl Crystallogr 24 246–250 Occurrence Handle10.1107/S0021889891004399
J Kwiatowski D Skarecky S Hernandez D Pham F Quijas F Ayala (1991) ArticleTitleHigh fidelity of polymerase chain reaction Mol Biol Evol 8 IssueID6 884–887 Occurrence Handle1:CAS:528:DyaK38Xhs1elsg%3D%3D Occurrence Handle1775068
HD Langohr (1980) ArticleTitleBiochemical studies on muscles in neurogenic atrophies and central paralysis. Studies of the atrophic functions of neurons Fortschr Maed 98 1512–1516 Occurrence Handle1:STN:280:Bi6D2crjs1M%3D
M Loukas AP Economopoulos E Zouros Y Vergini (1985) ArticleTitleGenetic changes in artificially reared colonies of the olive fruit fly (Diptera, Tephritidae) Ann Enomol Soc Am 78 159–165 Occurrence Handle1:CAS:528:DyaL2MXkt1Sjurk%3D
RN Martins CG Harper GB Stokes CL Masters (1986) ArticleTitleIncreased cerebral glucose-6-phosphate dehydrogenase activity in Alzheimer’s disease may reflect oxidative stress J Neurochem 46 1042–1045 Occurrence Handle1:CAS:528:DyaL28XhsVKmsb0%3D Occurrence Handle3950618
EA Merrit DJ Bacon (1997) ArticleTitleRaster 3D: Photorealistic molecular graphics Methods Enzymol 277 505–524
MS Nasoff HV Baker RE Wolfe SuffixJr. (1984) ArticleTitleDNA sequence of the Escherichia coli gene, gnd, for 6-phosphogluconate dehydrogenase Gene 27 253–264 Occurrence Handle10.1016/0378-1119(84)90070-2 Occurrence Handle1:CAS:528:DyaL2cXksV2gtLs%3D Occurrence Handle6329905
F Novello P McClean (1968) ArticleTitleThe pentose phosphate pathway of glucose metabolism Biochem 1107 775–791
AM Palmer (1999) ArticleTitleThe activity of the pentose pathway is increased in response to oxidative stress in Alzheimer’s disease J Neural Transm 106 317–328 Occurrence Handle10.1007/s007020050161 Occurrence Handle1:CAS:528:DyaK1MXjvFKqsb0%3D Occurrence Handle10392540
WR Pearson DJ Lipman (1988) ArticleTitleImproved tools for biological sequence comparison Proc Natl Acad Sci USA 85 2444–2448 Occurrence Handle1:CAS:528:DyaL1cXktFyit78%3D Occurrence Handle3162770
MC Peitsch (1995) ArticleTitleProMod: automated knowledge-based protein modelling tool PDB Q Newslett 72 4
Quanta ((1998) A molecular graphics program Molecular Simulations San Diego, CA
P Rice I Longden A Bleasby (2000) ArticleTitleEMBOSS: The European Molecular Biology Open Software Suite Trends Genet 16 276–277 Occurrence Handle10.1016/S0168-9525(00)02024-2 Occurrence Handle1:CAS:528:DC%2BD3cXjvVygsbs%3D Occurrence Handle10827456
MA Rosemeyer (1987) ArticleTitleThe biochemistry of glucose 6-phosphate dehydrogenase, 6-phosphogluconate dehydrogenase and glutathione reductase Cell Biochem Funct 5 79–95 Occurrence Handle1:CAS:528:DyaL2sXktVaqt74%3D Occurrence Handle3581436
MG Rossmann A Liljas C-I Branden LJ Banaszac (1975) Lactate dehydrogenase PD Boyer (Eds) The enzymes, Vol XIA EditionNumber3 Academic Press Orlando, FL 61–102
J Sambrook EF Fritsch T Maniatis (1989) Molecular cloning: A laboratory manual EditionNumber2 Cold Spring Harbor Laboratory Press Cold Spring Harbor
MJ Scott JC Lucchesi (1991) ArticleTitleStructure and expression of the Drosophila melanogaster gene encoding 6-phosphogluconate dehydrogenase Gene 109 177–183 Occurrence Handle10.1016/0378-1119(91)90607-D Occurrence Handle1:CAS:528:DyaK3sXhs1ajs74%3D Occurrence Handle1765265
MJ Scott D Kriticou AS Robinson (1993) ArticleTitleIsolation of cDNAs encoding 6-phosphogluconate and glucose-6-phosphate dehydrogenase from the Mediterranean fruit fly Ceratitis capitata: Correlating genetic and physical maps of chromosome 5 Insect Mol Biol 1 213–222 Occurrence Handle1:CAS:528:DyaK2cXisVGktA%3D%3D Occurrence Handle8269100
NS Scrutton A Berry RN Perham (1990) ArticleTitleRedesign of the coenzyme specificity of a dehydrogenase by protein engineering Nature 343 38–43 Occurrence Handle10.1038/343038a0 Occurrence Handle1:CAS:528:DyaK3cXhslClsrk%3D Occurrence Handle2296288
JC Street AA Alfieri JA Koutcher (1997) ArticleTitleQuantitation of metabolic and radiobiological effects of 6-aminonicotinamide in RIF-1 tumor cells in vitro Cancer Res 57 3956–3962 Occurrence Handle1:CAS:528:DyaK2sXmtFehsL4%3D Occurrence Handle9307279
JD Thompson TJ Gibson F Plewniak F Jeanmougin DG Higgins (1997) ArticleTitleThe Clustal X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools Nucleic Acids Res 24 4876–4882 Occurrence Handle10.1093/nar/25.24.4876
SK Tsui JY Chan MM Waye KP Fung CY Lee (1996) ArticleTitleIdentification of a cDNA encoding 6-phosphogluconate dehydrogenase from a human heart cDNA library Biochem Genet 34 367–373 Occurrence Handle10.1007/BF00554412 Occurrence Handle1:CAS:528:DyaK2sXjtlaj Occurrence Handle8978909
T Wood (1985) The pentose phosphate pathway Academic Press New York
L Zhang L Chooback PF Cook (1999) ArticleTitleLysine 183 is the general base in the 6-phosphogluconate dehydrogenase-catalyzed reaction Biochemistry 38 11231–11238 Occurrence Handle10.1021/bi990433i Occurrence Handle1:CAS:528:DyaK1MXltVGht70%3D Occurrence Handle10471272
Author information
Authors and Affiliations
Corresponding author
Additional information
Abbreviations used: 6Pgd, 6-Phosphogluconate dehydrogenase gene; 6PGD, 6-Phosphogluconate dehydrogenase enzyme; NADPH, nicotinamide adenine dinucleotide phosphate; ADP, adenine dinucleotide phosphate; TNBS, 2,4,6 trinitrobenzensulfonic acid.
Rights and permissions
About this article
Cite this article
Goulielmos, G.N., Eliopoulos, E., Loukas, M. et al. Functional Constraints of 6-Phosphogluconate Dehydrogenase (6-PGD) Based on Sequence and Structural Information. J Mol Evol 59, 358–371 (2004). https://doi.org/10.1007/s00239-004-2630-y
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00239-004-2630-y