Abstract
AC microsatellites have proved particularly useful as genetic markers. For some purposes, such as in population biology, the inferences drawn depend on the quantitative values of their mutation rates. This, together with intrinsic biological interest, has led to widespread study of microsatellite mutational mechanisms. Now, however, inconsistencies are appearing in the results of marker-based versus non-marker-based studies of mutational mechanisms. The reasons for this have not been investigated, but one possibility, pursued here, is that the differences result from structural differences between markers and genomic microsatellites. Here we report a comparison between the CEPH AC marker microsatellites and the global population of AC microsatellites in the human genome. AC marker microsatellites are longer than the global average. Controlling for length, marker microsatellites contain on average fewer interruptions, and have longer segments, than their genomic counterparts. Related to this, marker microsatellites show a greater tendency to concentrate the majority of their repeats into one segment. These differences plausibly result from scientists selecting markers for their high polymorphism. In addition to the structural differences, there are differences in the base composition of flanking sequences, marker flanking regions being richer in C and G and poorer in A and T. Our results indicate that there are profound differences between marker and genomic microsatellites that almost certainly affect their mutation rates. There is a need for a unified model of mutational mechanisms that accounts for both marker-derived and genomic observations. A suggestion is made as to how this might be done.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There has been considerable recent interest in the mutational processes leading to length polymorphisms within microsatellite loci, with several studies using microsatellite marker loci. A mutation model is sometimes assessed on the basis of its agreement with observed allele frequencies of microsatellite markers, although it is typical to use relatively small numbers of loci and modestly sized populations (e.g., Chakraborty et al. 1997; Di Rienzo et al. 1994; Estoup and Cornuet 1999; Shriver et al. 1993; Valdes et al. 1993). Mutation rates may also be more directly estimated from the observation of marker allele transmissions between generations using parent–child pedigree data (Brinkmann et al. 1998; Brohede et al. 2002; Ellegren 2000; Huang et al. 2002; Whittaker et al. 2003; Xu et al. 2000) or by genotyping sperm from a few individuals of known genotype (Leeflang et al. 1995). In addition, phylogenetic approaches have been used using microsatellite markers (Harr et al. 2000). The alternative strategy adopted by many workers is to extract many microsatellite loci from across the genome, a system known as genomic data mining (e.g., Bell and Jurka 1997; Kruglyak et al. 1998; Rose and Falush 1998; Sibly et al. 2003). This approach, which has become possible thanks to the availability of genomic sequences, involves recording relevant features of all the microsatellites of a given type across the entire genome. The observed data for a selected motif type are then assessed for goodness of fit against a mutation model for microsatellite slippage mutation.
There are currently inconsistencies between the results of marker-based and non-marker-based studies. For instance, the former suggest that the expansion rate exceeds the contraction rate for short microsatellites (e.g., Whittaker et al. 2003; Xu et al. 2000), although this thesis appears incompatible with observations made from genomic mining (e.g., Kruglyak et al. 1998; Sibly et al. 2003). The reasons for this discrepancy are unresolved. One possibility, pursued here, is that the differences arise from the way that microsatellite markers have been selected for their utility as genetic markers, which involves a complex ascertainment process. For example, for the AC microsatellites in the CEPH database (http://www.cephb.fr/cephdb/), which we concentrate on here, potential markers had to satisfy the requirements that (i) the sequence including flanking sequences had to reliably identify a long AC microsatellite; (ii) the microsatellite and its flanking region had to allow reliable amplification by PCR, and (iii) the microsatellite had to show high allelic polymorphism in the CEPH reference families (Weissenbach et al. 1992).
It is therefore possible that the inconsistencies between the results of marker-based and non-marker-based studies may result from structural differences between markers and genomic microsatellites. Thus far, however, no quantitative comparison of the two classes of loci has been performed. Accordingly, we here compare the structure of marker and genomic microsatellites in terms of total length, base composition of the regions flanking the microsatellite, interruptions that interrupt the microsatellite into segments of perfect repeats, and the numbers and lengths of perfect segments. It turns out the structures of marker and genomic AC microsatellites differ in several important respects. This is perhaps not surprising, but it is important: proper modeling of the effects of these differences on mutation rates is essential if we are to resolve the current inconsistencies in our understanding of microsatellite evolution.
Methods
The Fondation Jean Dausset CEPH database contains genotypes for all the genetic markers that have been used in the human linkage mapping project (Dib et al. 1996). On 9/4/2003, 5282 AC microsatellite sequences were retrieved automatically using the 5761 (AC) n microsatellite records in CEPH database version 9.0. Some records lacked or had invalid accession numbers, and so the corresponding sequence was not retrieved. For the purpose of analysis, an operational definition of the boundary between a microsatellite and its flanking sequence is needed, although it is not clear how this should be formulated (Santibanez-Koref et al. 2001). The retrieved sequences were tabulated according to the following criteria: (1) the microsatellite consists of a series of uninterrupted (AC)n sequences, which we call segments, in which two consecutive segments are separated by an interruption, here meaning a dinucleotide different from AC; (2) at least one of the segments comprises five or more repeats; and (3) the 10-base flanking sequences do not contain either an AC or a CA dinucleotide. These criteria are a minor variation of those of Sibly et al. (2003). Of the 5282 available sequences evaluated in this way, 940 were accepted. For each accepted microsatellite, we recorded its total length, number of segments, length of each segment, base sequence of the 10-base flanking regions, and interrupting dinucleotide motifs.
The complete sequence of the human genome was downloaded from ftp.ncbi.nih.gov/genomes/H_sapiens/April_14_2003/ on 16/04/2003 and the AC microsatellites within it were tabulated using the criteria above. This resulted in the extraction of the features of 18,308 AC microsatellites. In the following these are called “genomic microsatellites,” as opposed to the “marker microsatellites” obtained from the CEPH database.
Results
As expected marker microsatellites show a distinct tendency to be longer than genomic microsatellites, with the majority of markers containing between 11 and 25 repeats compared with less than 10 repeats in most genomic microsatellites (Fig. 1). The changed length distribution of marker microsatellites is an inevitable result of the way the markers were chosen [see requirement (i) in the Introduction]. The length distribution of genomic microsatellites agrees with previously published results (Sibly et al. 2003).
Microsatellites are sometimes broken into segments by interruptions. Attention in this paper is restricted to the case that the interruptions are dinucleotides. The frequencies of the 15 possible interruptions did not differ between marker microsatellites and the length-corrected genomic microsatellites (χ 214 = 16.7; n.s.). It follows that there has been neither conscious nor unconscious bias for (or against) a particular interruption motif during the process of designating microsatellite marker loci. We now consider analysis of the number and length of segments.
Number and Length of Segment
The length differences between genomic and marker microsatellites could confound further analyses. To circumvent this problem we controlled for length by creating a “length-corrected genomic” dataset, in which genomic microsatellites were matched by length to marker microsatellites. This was achieved by randomly selecting, for each marker microsatellite locus, two genomic microsatellite loci of the same length. Two was the largest number of genomic loci available for sampling since there are only six genomic microsatellites of length 33 but there are three markers of that length. Sampling was carried out without replacement.
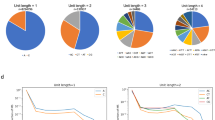
Intriguingly, there are dramatic differences in the number of segments in marker and genomic microsatellites after controlling for total microsatellite length (Fig. 2) (χ 24 = 48.4, p ≈ 10−9). Marker microsatellites possess fewer segments than genomic microsatellites; for instance; there are six times as many four-segment microsatellites among the genomic set as among the markers. In each case, frequency declines approximately geometrically with number of segments, as demonstrated previously by Sibly et al. (2003) for genomic microsatellites, although it should be noted that estimates for five-segment microsatellites have a large standard error because there are few such loci. It is also noteworthy that the fraction of microsatellites lacking interruptions is higher among marker microsatellites than it is in genomic microsatellites. These results suggest that there was bias against the presence of multiple interruptions when selecting the microsatellite loci to be used as markers.
Further evidence of bias comes from comparing the segment lengths of the two types of microsatellite for each possible total length (Fig. 3). The results for the marker data set are most reliable for microsatellites containing between 11 and 24 repeats. As expected from Fig. 2, marker microsatellites within this range possess significantly fewer and longer segments than genomic microsatellites. Given that microsatellite markers were selected partly on the basis of exhibiting high levels of polymorphism between individuals, it is difficult to establish whether these differences reflect increased slippage rates, relative to point mutation rates, among the marker microsatellites or if the difference is entirely attributable to bias against the selection of “imperfect” microsatellite arrays for marker purposes. Note that the left-hand side in Fig. 3 is difficult to interpret because our definition of microsatellite requires that one segment be at least five repeats long. Hence microsatellites of length 5 have exactly one segment by definition, those of length 6 have at most two, length 7 at most three, and so on.
Lastly, we show that the longest segment in marker microsatellites is disproportionately long. We saw in Fig. 3 that for a given length, marker microsatellites have longer segments than genomic microsatellites. For both microsatellite types, much of the length of the microsatellite is contained within the longest segment (Fig. 4). The proportion of total length accounted for by the longest segment naturally declines as the number of segments increases but surprisingly remains above 50% of the length on average in all cases. This pattern is noticeably more pronounced among the marker microsatellites, where the longest segment comprises a significantly greater proportion of the total sequence for all numbers of segments. This difference may result from direct (probably unconscious) bias for asymmetry among imperfect microsatellites for use as markers so that they contain at least one long segment. Alternatively, it may simply be a correlative association caused by experimenter selection for increased polymorphism, if long uninterrupted segments show a greater tendency for slippage.
Composition of Flanking Sequences
Table 1 shows the base composition of the flanking regions for the marker microsatellites (M) and length-corrected genomic microsatellites (CG). Intriguingly, there are significant differences in the base composition of these flanking regions (p < 0.001), as shown by a randomization test controlling for length. Overall, marker microsatellite flanking regions are far richer in C and G content, and so poorer in A and T. It is difficult to see how unconscious bias during the designation of loci for use as markers could account for this discrepancy. It seems rather more plausible that the base composition of the flanking region in some way influences the likelihood of slippage and/or interruptions such that the loci selected on the basis of being polymorphic tend also to have a shared base composition in the flanking regions. A very similar result, albeit on disease loci, has been obtained by (Brock et al. 1999). Specifically, they found an association between the GC content of flanking regions and the “relative expandability” of the locus.
Discussion
We have identified three major differences between markers and genomic microsatellites: markers are longer (Fig. 1); controlling for length, markers contain fewer interruptions (Fig. 2); and segment lengths are longer (Fig. 3).
The reasons for these differences are readily understood in terms of the way markers were designed. They probably all result from scientists selecting for high polymorphism. Thus, length increases mutation rate (Brinkmann et al. 1998; Ellegren 2000; Kruglyak et al. 1998; Sibly et al. 2001; Whittaker et al. 2003; Wierdl et al. 1997). Controlling for overall length, markers have fewer interruptions, and this increases slippage rates (Petes et al. 1997; Rolfsmeier and Lahue 2000). And the longest segment of markers is disproportionately long (Fig. 4). If slippage rate increases with segment length faster than linearly as argued (Brinkmann et al. 1998; Lai and Sun 2004; Rose and Falush 1998; Sibly et al. 2001), then the longest segment will have an overriding effect on the overall slippage rate of the microsatellite. Unconscious preference for long perfect arrays at the marker selection stage could also account for this difference. To sum up, the internal characteristics of marker microsatellite loci produce higher mutation rates in marker than in genomic microsatellites and, hence, are likely to produce higher rates of polymorphism. These differences are therefore plausibly the result of the marker design process.
While the internal structure of markers seems to be associated with higher mutation rates, this is not true of their flanking sequences. The flanking sequences of marker microsatellites contain more C and G, and fewer A and T, than do genomic microsatellites. C–G pairings contain an extra hydrogen bond and so should be more stable than A–T pairings and so less prone to slippage. A possible reason is that added stability in the flanking region stabilize the microsatellite during PCR amplification and so renders it less prone to forming artifact “stutter” bands. Certainly, loci generating such bands are less likely to be selected as markers because of difficulty in allele designation after fractionation by electrophoresis.
In conclusion, our results indicate that marker and genomic microsatellite loci differ profoundly in terms of their length and composition and in the sequence of their flanking regions. Future analyses should take account of these differences. A unified theory is needed that can account both for the genomic distribution of interrupted microsatellites as shown in Figs. 1–4 for experimental observations of mutation frequencies and step sizes in relation to microsatellite structure. A unified model might perhaps write mutation rate for a given step size = f(l1, m1, l2, m2, 13, m3, ⋖) where l i is the length of the ith segment, and m i is the identity of the ith interruption. Models of this type would ideally be fitted in all types of analysis, to provide a unified account of the effects of structural characteristics on the mutational mechanism.
References
GI Bell J Jurka (1997) ArticleTitleThe length distribution of perfect dimer repetitive DNA is consistent with its evolution by an unbiased single-step mutation process J Mol Evol 44 414–421 Occurrence Handle1:CAS:528:DyaK2sXis12msLs%3D Occurrence Handle9089081
B Brinkmann M Klintschar F Neuhuber J Huhne B Rolf (1998) ArticleTitleMutation rate in human microsatellites: influence of the structure and length of the tandem repeat Am J Hum Genet 62 1408–1415 Occurrence Handle1:CAS:528:DyaK1cXlslCiu70%3D Occurrence Handle9585597
GJR Brock NH Anderson DG Monckton (1999) ArticleTitleCis-acting modifiers of expanded CAG/CTG triplet repeat expandability: associations with flanking GC content and proximity to CpG islands Hum Mol Genet 8 1067–1067
J Brohede CR Primmer AP Moller H Ellegren (2002) ArticleTitleHeterogeneity in the rate and pattern of germline mutation at individual microsatellite loci Nucleic Acids Res 30 1997–2003 Occurrence Handle1:CAS:528:DC%2BD38XktFGhtbc%3D Occurrence Handle11972338
RM Chakraborty DN Kimmel J Strivers InstitutionalAuthorNameDavison R Deka (1997) ArticleTitleRelative mutation rates at di-, tri-, and tetranucleotide microsatellite loci Proc Natl Acad Sci USA 94 1041–1046 Occurrence Handle10.1073/pnas.94.3.1041 Occurrence Handle1:CAS:528:DyaK2sXhtVyht70%3D Occurrence Handle9023379
A Di Rienzo AC Petersom JC Garza AM Valdes M Slatkin NB Freimer (1994) ArticleTitleMutational processes of simple-sequence repeat loci in human populations Proc Natl Acad Sci USA 91 3166–3170 Occurrence Handle1:CAS:528:DyaK2cXktVantrk%3D Occurrence Handle8159720
C Dib C Faure C Fizames et al. (1996) ArticleTitleA comprehensive genetic map of the human genome based on 5,264 microsatellites Nature 380 152–154 Occurrence Handle1:CAS:528:DyaK28Xhs1Gns7c%3D Occurrence Handle8600387
H Ellegren (2000) ArticleTitleHeterogeneous mutation processes in human microsatellite DNA sequences Nat Genet 24 400–402 Occurrence Handle1:CAS:528:DC%2BD3cXisVCjtr4%3D Occurrence Handle10742106
A Estoup J-M Cornuet (1999) Microsatellite evolution: inferences from population data DB Goldstein C Schlötterer (Eds) Microsatellites: Evolution and applications Oxford University Press Oxford 49–65
B Harr B Zanger C Schlotterer (2000) ArticleTitleRemoval of microsatellite interruptions by DNA replication slippage: phylogenetic evidence from Drosophila Mol Biol Evol 17 1001–1009 Occurrence Handle1:CAS:528:DC%2BD3cXksVOmu70%3D Occurrence Handle10889213
QY Huang FH Xu H Shen et al. (2002) ArticleTitleMutation patterns at dinucleotide microsatellite loci in humans Am J Hum Genet 70 625–634 Occurrence Handle10.1086/338997 Occurrence Handle1:CAS:528:DC%2BD38XislChsbs%3D Occurrence Handle11793300
S Kruglyak RT Durrett MD Schug CF Aquadro (1998) ArticleTitleEquilibrium distributions of microsatellite repeat length resulting from a balance between slippage events and point mutations Proc Natl Acad Sci USA 95 10774–10778 Occurrence Handle10.1073/pnas.95.18.10774 Occurrence Handle1:CAS:528:DyaK1cXlvFWgtbg%3D Occurrence Handle9724780
Y Lai F Sun (2003) ArticleTitleThe relationship between microsatellite slippage mutation rate and the number of repeat units Mol Biol Evol 20 2123–2131 Occurrence Handle1:CAS:528:DC%2BD2cXnt12nsA%3D%3D Occurrence Handle12949124
EP Leflang L Zhang S Tavare et al. (1995) ArticleTitleSingle sperm analysis of the trinucleotide repeats in the Huntington’s disease gene: quantification of the mutation frequency spectrum Hum Mol Genet 4 1519–1526 Occurrence Handle8541834
TD Petes PW Greenwell M Dominska (1997) ArticleTitleStabilization of microsatellite sequences by variant repeats in the yeast Saccharomyces cerevisiae Genetics 146 491–498 Occurrence Handle1:CAS:528:DyaK2sXmsVaku7c%3D Occurrence Handle9178000
ML Rorfsmeier RS Lahue (2000) ArticleTitleStabilizing effects of interruptions on trinucleotide repeat expansions in Saccharomyces cerevisiae Mol Cell Biol 20 173–180 Occurrence Handle10594019
O Rose D Falush (1998) ArticleTitleA threshold size for microsatellite expansion Mol Biol Evol 15 613–615 Occurrence Handle1:CAS:528:DyaK1cXislensLk%3D Occurrence Handle9580993
MF Santibanez-Koref R Gangeswaran JM Hancock (2001) ArticleTitleA relationship between lengths of microsatellites and nearby substitution rates in mammalian genomes Mol Biol Evol 18 2119–2123 Occurrence Handle1:CAS:528:DC%2BD3MXotVSltr8%3D Occurrence Handle11606708
MD Shriver L Jim R Chakraborty LE Boerwinkle (1993) ArticleTitleVNTR allele frequency distribution under the stepwise mutation model Genetics 134 983–993 Occurrence Handle1:CAS:528:DyaK3sXms1emt7o%3D Occurrence Handle8349120
RM Sibly A Meade N Boxall MJ Wilkinson DW Corne JC Whittaker (2003) ArticleTitleThe structure of interrupted human AC microsatellites Mol Biol Evol 20 453–459 Occurrence Handle1:CAS:528:DC%2BD3sXisFaqtrg%3D Occurrence Handle12644566
RM Sibly JC Whittaker M Talbot (2001) ArticleTitleA maximum-likelihood approach to fitting equilibrium models of microsatellite evolution Mol Biol Evol 18 413–417 Occurrence Handle1:CAS:528:DC%2BD3MXhvVKrt74%3D Occurrence Handle11230542
AM Valdes M Slaktkin NB Frejner (1993) ArticleTitleAllele frequencies at microsatellite loci: the stepwise mutation model revised Genetics 133 737–749 Occurrence Handle1:STN:280:ByyB3c3kvVA%3D Occurrence Handle8454213
J Weisssenbach G Gyapay A Vignbal et al. (1992) ArticleTitleA second-generation linkage map of the human genome Nature 359 794–801
JC Whittaker RM Hardoard N Boxall I Mackay G Dawson RM Sibly (2003) ArticleTitleLikelihood based estimation of microsatellite mutation rates Genetics 164 781–787 Occurrence Handle12807796
M Wierdl M Dominska TD Petes (1997) ArticleTitleMicrosatellite instability in yeast: dependence on the length of the microsatellite Genetics 146 769–779 Occurrence Handle1:CAS:528:DyaK2sXmsVartLw%3D Occurrence Handle9215886
X Xu M Peng Z Fang X Xu (2000) ArticleTitleThe direction of microsatellite mutations is dependent upon allele length Nat Genet 24 396–399 Occurrence Handle1:CAS:528:DC%2BD3cXisVCjsbc%3D Occurrence Handle10742105
Author information
Authors and Affiliations
Corresponding author
Additional information
Reviewing Editor: Dr. Magnto Nordborg
Rights and permissions
About this article
Cite this article
Pardi, F., Sibly, R.M., Wilkinson, M. et al. On the Structural Differences Between Markers and Genomic AC Microsatellites. J Mol Evol 60, 688–693 (2005). https://doi.org/10.1007/s00239-004-0274-6
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00239-004-0274-6