Abstract
HIV, the cause of AIDS in humans, is characterized by great genetic heterogeneity. In particular, HIV-1 group M subtypes are responsible for most of the infections worldwide. We investigate the demographic history of HIV-1B and HIV-1C subtypes in South Africa and Brazil using both a parametric and a nonparametric approach based on coalescent theory. Our results show that although both subtypes are spreading exponentially in Brazil, the HIV-1C growth rate is about twice that of Brazilian HIV-1B or South African HIV-1C, providing evidence, for the first time, of a different epidemic potential between two HIV-1 subtypes. The present study not only may have important consequences for devising future vaccination and therapeutic strategies, but also offers additional evidence that skyline plots are indeed a simple and powerful tool for monitoring and predicting the behavior of viral epidemics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
HIV, the etiologic agent of AIDS, is classified into two distinct but clearly related types of viruses, HIV-1 and HIV-2, characterized by an extraordinary genetic variability (Hahn et al. 1984; Clavel et al. 1986). Three major groups can be distinguished within HIV-1: group M (for main), O (for outlier), and N (for neither, non-M–non-O, or new) (Robertson et al. 2000; Simon et al. 1998). Moreover, 9 phylogenetically distinct subtypes, 2 two subsubtypes, and at least 14 intersubtype HIV-1 recombinants, known as circulating recombinant forms (CRFs), have been repeatedly identified within group M so far (Robertson et al. 2000; McCutchan 2000; Salminen et al. 1995). Indeed, recombination coupled with the elevated error rate of the reverse transcriptase and the rapid turnover of HIV-1 in infected individuals are at the origin of the high genetic variability of the virus (Peeters and Sharp 2000).
Most subtypes, as well as CRFs, are present in Africa, reflecting the African origin of the epidemic (Gao et al. 1999). HIV-1B is the subtype responsible for most of the infections in Europe, United States, and Australia, whereas HIV-1C is the most prevalent worldwide, accounting for more than 56% of all infections (Esparza and Bhamarapravati 2000). It has been shown that the HIV-1 group M epidemic in humans originated from a zoonotic transmission (Gao et al. 1999) and that the cenancestor (most recent common ancestor) of group M probably dates back to the 1930s (Korber et al. 2000; Salemi et al. 2001). The existence of genetic subtypes could be the result of certain viral strains being involved in extensive transmission chains in a given geographic area, the so-called “founder effect.” In this view, in spite of their genetic differences, HIV-1 subtypes would be biologically equivalent. It is also hypothesized that characteristics such as a higher or lower transmissibility and/or fitness could explain the success or failure of different subtypes in different regions (Bjorndal et al. 1999). No definitive answer has been reached so far. In addition, the ability of a virus to spread in a population could be related to specific transmission routes like sexual contact and injecting drug use (Salemi et al. 1999; Pybus et al. 2001).
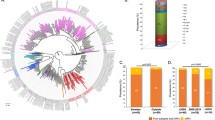
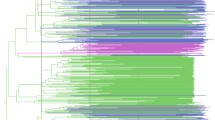
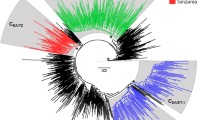
In the present study we investigate the epidemic behavior of HIV-1B in Brazil and HIV-1C in Brazil and South Africa. These two countries represent an interesting model to explore the potential differences between the two subtypes. The epidemic of HIV-1C is relatively old in South Africa, where it is the most prevalent subtype (Fig. 1), whereas HIV-1B is dominant in Brazil. However, HIV-1C has been introduced recently in Brazil, and within the last few years it has become the most common subtype in recently diagnosed individuals in the southern states (Fig. 1) (Soares et al. 2003).
Seroprevalence of different HIV-1 subtypes in South Africa and Brazil. The data for South Africa are based on a study of the South African National Health Department (HIV prevalence: HIV Antenatal Prevalence of the National Health Department 2001). The data for Brazil are based on a study by the Brazilian Institute of Health (HIV prevalence: Brazilian Antenatal HIV Prevalence in Sentinel Sites in 2000).
Materials and Methods
Data Sets
Six data sets have been compiled using B and C sequences that were nonrecombinant in the RT and protease genes: HIV-1B and HIV-1C Brazilian strains collected between 1992 and 2001 and HIV-1C South African strains collected between 1990 and 2002 (Cornelissen et al. 1997; Dumans et al. 2002; Gordon et al. 2003; Soares et al. 2003). To reduce nonrandom sampling, sequences were excluded if they came from the same patient or from individuals known to be related by a direct transmission. The data sets are summarized in Table 1. Sequences were aligned with the Clustal algorithm (Thompson et al. 1994) followed by manual editing. Positions containing gaps were removed from the final alignment. The alignments are available from the authors upon request.
Phylogeny and Evolutionary Rate Estimates
Maximum likelihood (ML) phylogenies were estimated for each data set. The best-fitting nucleotide substitution model was tested with a hierarchical likelihood ratio test following the strategy described by Swofford and Sullivan (2003), using a neighbor-joining tree with HKY85 estimated distances. ML phylogenies were then reestimated with the selected model, using a neighbor-joining tree as starting tree, and the TBR algorithm for branch swapping. Calculations were performed with PAUP* 4.0b10 (D.L. Swofford, Sinauer Associates, Sunderland, MA).
Because the sequences in our data set have been collected over several years, the evolutionary rate μ can be estimated via ML directly from the phylogenetic tree assuming a molecular clock with noncontemporaneous tips (Rambaut 2000). The molecular clock hypothesis can be tested with the likelihood ratio test with n − 3 degrees of freedom, where n is the number of taxa (Rambaut 2000).
Coalescent Models
By using coalescent theory, we can study how the ancestral relationships of individuals sampled from a population are influenced by its demographic history (Griffiths and Tavarè 1994; Kingman 1982).
A genealogy reconstructed from randomly sampled HIV sequences contains information about population-level processes such as change in population size and growth rate (Pybus et al. 2000). Given a viral phylogeny P and a vector φ representing the parameters of the model N(t), it is possible to calculate the log of the conditional probability ln[φ|P] (Pybus et al. 2000). ML estimates of φ can be found by numerical optimization of ln[(φ|P] and 95% CIs for the estimates obtained with the likelihood ratio statistic (Pybus et al. 2000). The estimated parameters are, in fact, N(0)μ. and r|μ, where μ is the evolutionary rate in nucleotide substitutions per site per year (the parameter c in the logistic model is unaffected by linear scaling of time). Notice that time runs backward into the past so that N(0) is the effective number of the infections at the present, and N(t) represents the effective number of infections at time t.
We consider two demographic models: exponential growth
and logistic growth
where r is the growth rate achieved in a wholly susceptible population and c (Eq. [2]) is a logistic shape parameter related to the population carrying capacity (Pybus et al. 2001). Since the two models are nested they can be compared with the likelihood ratio test. The logistic model reduces to the exponential one when c is infinite (degrees of freedom [df] = 1). In practice, c must be evaluated within a finite range and can be considered infinite when the upper limit of the 95% CI is 100,000 or more.
We also obtained nonparametric estimates of demographic history through the skyline plots (Pybus et al. 2000). However, the phylogenetic trees obtained for the HIV data sets show several zero or near-zero internal branch lengths, which make the skyline plots very noisy and more difficult to interpret. Therefore, we estimated the generalized skyline plots (Strimmer and Pybus 2001) for clock-like phylogenetic trees with dated tips. In such plots adjacent intervals smaller than a threshold of size ε in a tree are grouped together before obtaining the nonparametric estimates of the population size at any given time. For each data set, the optimal ε value used was the one maximizing the AICC (corrected Akaike information criterion) of the plot. All calculations were performed with GENIE version 3.0 (Pybus and Rambaut 2002).
Results
Evolutionary Rate Estimates
The estimated HIV-1 evolutionary rates for the different data sets were 1.40 ± 0.28 × 10−3 (HIV-B protease; Brazil), 1.54 ± 0.38 × 10−3 (HIV-B RT; Brazil), 3.38 ± 0.95 × 10−3 (HIV-C protease; Brazil), 3.55 ± 0.64 × 10−3 (HIV-C RT; Brazil), 1.71 ± 0.27 × 10−3 (HIV-C protease; South Africa), 1.47 ± 0.21 × 10−3 (HIV-C RT; South Africa). The molecular clock hypothesis was rejected by the likelihood ratio test for each data set. However, simulation studies have shown that if there is only a small amount of rate variation among lineages, then the 95% confidence limits of the rate estimate still contain the true mean rate about 95% of the time, even if the clock is rejected (Jenkins et al. 2002). In other words, the clock is very easily rejected, but the rate inferred enforcing the clock is still a good estimate of the mean rate and can still be used as a useful time scale.
Parametric Estimates of HIV-1B and 1CDemographic History in South Africa and Brazil
Using clock-like phylogenetic trees with noncontemporaneous tips and the estimated evolutionary rates, we tested the demographic hypotheses described by Eqs. (1) and (2) for each of the data sets listed in Table 1. For none of them could the exponential model be rejected (see Table 1). Evidently, the number of infected individuals cannot grow forever and should eventually plateau according to the logistic model. At present, however, B and C subtypes in South Africa and Brazil appear to be spreading exponentially among the susceptible population, indicating that the population carrying capacity has not yet been reached. Table 2 shows the parametric estimates of N(0) and r using the exponential model. For each HIV-1 data set similar results were obtained analyzing the protease or the RT gene. The confidence intervals, CIs, of N(0) are quite large compared to those of r, but the estimates are in good agreement with the epidemiological data on the seroprevalence of subtypes B and C in Brazil and subtype C in South Africa (Department of Health, South Africa 2001; UN AIDS/WHO Working Group on Global HIV/AIDS and STD Surveillance 2000; see also Fig. 1). Note that N(0) gives an estimate of the effective number of infections, i.e., those that are effectively transmitted in a population mating at random, not the absolute number of the infected people, which is usually larger.
The robustness of r estimates to change in N(0) was tested by reestimating r while constraining N(0) to vary over a range of values including its lower and upper 95% confidence limit given in Table 2 for each data set. In every case, the new estimates of r fell within the CIs of r reported in Table 2 (data not shown).
The r estimates also allow the estimation of the epidemiological quantity R0, the basic reproductive number (infectivity) of a pathogen, with the equation R0 = rD+1, where D is the average duration of infectiousness (Pybus et al. 2001). In Table 2 we use a putative but plausible range for D. For D = 10 years, on average eight secondary infections are generated by each primary HIV-1C infection in Brazil, versus about four secondary infections generated by HIV-1B in South Africa and Brazil or by the South African HIV-1C.
Generalized Skyline Plots of HIV-1B and 1Cin South Africa and Brazil
Figure 2 shows the generalized skyline plots (Strimmer and Pybus 2001) with the nonparametric estimates of N(t) for the different data sets. ML and nonparametric estimates of N(0) appear to be in good agreement (see Table 2 and Fig. 2). According to the analysis of both protease and RT, the HIV-IB Brazilian epidemic appears to date back to between the middle of the 1950s and the middle of the 1960s, and probably originated, based on the phylogenetic analysis of subtype B and C Brazilian isolates (Soares et al. 2003), by one single introduction caused by one to three effective infections (see Fig. 2). The HIV-1C epidemics in South Africa and Brazil show a quite different story. The HIV-1-C South African epidemic originated sometime in the 1980s, with about three effective infections at its origin (see Fig. 2). The HIV-1C Brazilian epidemic is more recent (beginning of the 1990s), with its origin estimated to be a single effective infection (see Fig. 2). The data in Brazil are in accordance with a single recent introduction of the new subtype in the country (Soares et al. 2003). On the other hand, the skyline plot for HIV-1C in South Africa is consistent with phylogenetic analyses showing that the South African HIV-1C strains are not monophyletic and implying multiple separate introductions of the subtype around the beginning of the 1980s (Gordon et al. 2003). The plot also matches a report showing that the adult population infected in the country changed from 1% in 1990 to 10% in 1995 and to 24% in 2000 (Department of Health, South Africa 2001). By superimposing the protease skyline plots of HIV-1C in South Africa and Brazil (Fig. 3), it is clear that the Brazilian epidemic is spreading even faster and that the number of effective infections in Brazil might be expected to increase greatly within the next few years. RT skyline plots give similar results (data not shown).
Generalized skyline plots representing nonparametric estimates of N(t) (Strimmer and Pybus 2001) using the protease (black line) and RT (gray or blue line) genes. The topmost step in the plot is the nonparametric estimate of N(0). Genetic distances were transformed into a time scale of years using the evolutionary rates μ inferred for each data set. Since these plots are point estimates of N(t), they do not contain information about uncertainty N(0) and r (Table 2), or μ.
Generalized skyline plots representing nonparametric estimates of N(t) (Strimmer and Pybus 2001) using the protease gene for the South African (black line) and Brazilian (gray line) HIV-1C. The figure was obtained by superimposing the relative skyline plots reported in Fig. 1. Dashed lines represent the parametric estimates of N(t) according to the exponential model (Table 2).
Discussion
South Africa is one of the epicenters of the epidemic in the world, with about 22% of the adult population infected with HIV-1 (Department of Health, South Africa 2001), mostly with subtype C infections. Brazil, on other hand, is considered to be one of the best examples where therapeutic implementation has slowed down the course of the epidemic. Today less than 1% of the adult population is infected with HIV, about 600,000 people, mostly by subtype B and, more recently, by subtype C in the southern region of the country (Dumans et al. 2002). HIV-1 growth rates are similar, except for HIV-1C in Brazil, which is spreading about two times faster than either HIV-1B or the South African HIV-1C. In particular, R0 estimates in South Africa are in good agreement with current epidemiological data showing that each infected person has transmitted HIV to at least three new persons within 5 years (Department of Health, South Africa 2001). No evidence has been reported so far indicating that the average duration of infectiousness may vary among HIV-1 subtypes. Therefore, the twofold increase in infectivity of the Brazilian HIV- 1C compared to the Brazilian HIV-1B and the South African HIV-IC may reflect a difference in the efficiency of different transmission routes in different geographic areas.
The above results depend on the assumptions of the coalescent model used: the evolutionary rate constancy, the absence of positive selection, recombination, and migration. We analyzed the protease and RT of naïve patients to reduce the effect of positive selection, and excluded recombinant strains, but it is difficult to assess the importance of migration among subpopulations of infected HIV-1 patients in the countries studied. Also, the uncertainty in the evolutionary rate estimates may confound the interpretation of the analysis. However, the consistency of the results with current epidemiological data stren-gthens our confidence. Moreover, since our estimates of demographic history are consistent among genes, it appears that the level of rate heterogeneity among HIV sequences is not large enough to systematically bias demographic inferences.
Overall we have shown that HIV-1C in Brazil is spreading at an increased rate. Following this trend the subtype may eventually become prevalent in the entire country, as has happened in the southern Brazilian states. Phylogenetic inference is too indirect to establish firmly whether the Brazilian HIV-1C is a new, more infectious strain or whether the virus is spreading faster because of a more favorable transmission route as suggested above. Yet the recently introduced HIV-1C is outcompeting HIV-1B in a country where the latter subtype was virtually the only one present until a few years ago, and a rapid escalation of HIV-1C infections has been occurring throughout sub-Saharan Africa, in India, and in China (Esparza and Bhamarapravati 2000; UNAIDS/WHO 2000). A similar scenario may be possible for other Western countries and deserves to be taken into account for future planning of vaccination and therapeutic campaigns around the world. These results also underline the need for refocusing prevention strategies in Brazil to stop the spreading of this viral variant. In this light, the use of viral gene sequences coupled with the results of coalescent theory appears to be a promising and important tool for monitoring and predicting the epidemic behavior of HIV subtypes and of other pathogens as well (Pybus et al. 2001; Robbins et al. 2003; Tanaka et al. 2002).
References
A Bjorndal A Sonnerborg J Tscherning C Albert EM Fenyo (1999) ArticleTitlePhenotypic characteristics of human immunodeficiency virus type 1 subtype C isolates of Ethiopian AIDS patients AIDS Res Hum Retrovir 15 647–653 Occurrence Handle1:CAS:528:DyaK1MXjtFCrsbs%3D Occurrence Handle10331443
F Clavel M Guyader D Guetard M Salle L Montagnier M Alizon (1986) ArticleTitleMolecular cloning and polymorphism of the human immune deficiency virus type 2 Nature 324 691–695 Occurrence Handle1:CAS:528:DyaL2sXnvFCmsA%3D%3D Occurrence Handle3025743
M Cornelissen R Burg Particlevan den F Zorgdrager V Lukashov J Goudsmit (1997) ArticleTitlePol gene diversity of five human immunodeficiency virus type 1 subtypes: evidence for naturally occurring mutations that contribute to drug resistance, limited recombination patterns, and common ancestry for subtypes B and D J Virol 71 6348–6358 Occurrence Handle1:CAS:528:DyaK2sXlsVKqtLg%3D Occurrence Handle9261352
Department of Health/Directorate Health Systems Research (2001) Seventh national HIV survey of woman attending antenatal clinics of the public health service of South Africa. October/November 2000. Directorate Health Systems Research. Department of Health, Pretoria, South Africa
AT Dumans MA Soares D Pieniazek ML Kalish V Vroey ParticleDe K Hertogs A Tanuri (2002) ArticleTitlePrevalence of protease and reverse transcriptase drug resistance mutations over time in drug-naive human immunodeficiency virus type 1-positive individuals in Rio de Janeiro, Brazil Antimicrob Agents Chemother 46 3075–3079 Occurrence Handle1:CAS:528:DC%2BD38Xmtl2kurk%3D Occurrence Handle12183276
J Esparza N Bhamarapravati (2000) ArticleTitleAccelerating the development and future availability of HIV-1 vaccines: why, when, where, and how? Lancet 355 2061–2066 Occurrence Handle1:STN:280:DC%2BD3czjvFyktQ%3D%3D Occurrence Handle10885368
F Gao E Bailes DL Robertson Y Chen CM Rodenburg SF Michael LB Cummins LO Arthur M Peeters GM Shaw PM Sharp BH Hahn (1999) ArticleTitleOrigin of HIV-1 in the chimpanzee Pan troglodytes troglodytes Nature 397 436–441 Occurrence Handle1:CAS:528:DyaK1MXhtFOht7g%3D Occurrence Handle9989410
M Gordon T Oliveira ParticleDe K Bishop HM Coovadia L Madurai S Engelbrecht E Janse Rensburg Particlevan A Mosam A Smith S Cassol (2003) ArticleTitleMolecular characteristics of human immunodeficiency virus type 1 subtype C viruses from KwaZulu-Natal, South Africa: implications for vaccine and antiretroviral control strategies J Virol 77 2587–2599 Occurrence Handle1:CAS:528:DC%2BD3sXpvVGmug%3D%3D Occurrence Handle12551997
RC Griffiths S Tavarè (1994) ArticleTitleSampling theory for neutral alleles in a varying environment Philos Trans. R Soc London Ser B 344 403–410 Occurrence Handle1:STN:280:ByqD1MfovFA%3D
BH Hahn GM Shaw SK Arya M Popovic RC Gallo F Wong-Staal (1984) ArticleTitleMolecular cloning and characterization of the HTLV-III virus associated with AIDS Nature 312 166–169 Occurrence Handle1:CAS:528:DyaL2MXhtlOg Occurrence Handle6095086
GM Jenkins A Rambaut OG Pybus EC Holmes (2002) ArticleTitleRates of molecular evolution in RNA viruses: a quantitative phylogenetic analysis J Mol Evol 54 156–165 Occurrence Handle1:CAS:528:DC%2BD38XhtVCgurs%3D Occurrence Handle11821909
JFC Kingman (1982) ArticleTitleThe coalescent Stoch Proc Appl 13 235–248
B Korber M Muldoon J Theiler F Gao R Gupta A Lapedes BH Hahn S Wolinsky T Bhattacharya (2000) ArticleTitleTiming the ancestor of the HIV-1 pandemic strains Science 288 1789–1796 Occurrence Handle1:CAS:528:DC%2BD3cXjvFCisro%3D Occurrence Handle10846155
FE McCutchan (2000) ArticleTitleUnderstanding the genetic diversity of HIV-1 AIDS 14 S31–S44 Occurrence Handle1:CAS:528:DC%2BD3cXosFymsL8%3D Occurrence Handle11086847
M Peeters PM Sharp (2000) ArticleTitleGenetic diversity of HTV-1: the moving target AIDS 14 IssueIDSuppl 13 S129–S140 Occurrence Handle1:CAS:528:DC%2BD3cXosFymsLw%3D Occurrence Handle11086856
OG Pybus A Rambaut (2002) ArticleTitleGENIE: estimating demographic history from molecular phylogenies Bioinformatics 18 1404–1405 Occurrence Handle1:CAS:528:DC%2BD38XnvF2ru7o%3D Occurrence Handle12376389
OG Pybus A Rambaut PH Harvey (2000) ArticleTitleAn integrated framework for the inference of viral population history from reconstructed genealogies Genetics 155 1429–1437 Occurrence Handle1:STN:280:DC%2BD3cvgsVSnuw%3D%3D Occurrence Handle10880500
OG Pybus MA Charleston S Gupta A Rambaut EC Holmes PH Harvey (2001) ArticleTitleThe epidemic behavior of the hepatitis C virus Science 292 2323–2325 Occurrence Handle1:CAS:528:DC%2BD3MXkslSht78%3D Occurrence Handle11423661
A Rambaut (2000) ArticleTitleEstimating the rate of molecular evolution: incorporating non-contemporaneous sequences into maximum likelihood phylogenies Bioinformatics 16 395–399 Occurrence Handle1:CAS:528:DC%2BD3cXltFOlu7c%3D Occurrence Handle10869038
KE Robbins P Lemey OG Pybus HW Jaffe AS Youngpairoj TM Brown M Salemi A-M Vandamme ML Kalish (2003) ArticleTitleU.S. human immunodeficiency virus type 1 epidemic:date of origin, population history, and characterization of early strains J Virol 77 6359–6366 Occurrence Handle1:CAS:528:DC%2BD3sXktV2nsrc%3D Occurrence Handle12743293
DL Robertson JP Anderson JA Bradac et al. (2000) ArticleTitleHIV-1 nomenclature proposal Science 288 55–56 Occurrence Handle1:CAS:528:DC%2BD3cXisVCqsLo%3D Occurrence Handle10766634
M Salemi M Lewis JF Egan WW Hall J Desmyter A-M Vandamme (1999) ArticleTitleDifferent population dynamics of human T cell lymphotropic virus type II in intravenous drug users compared with endemically infected tribes Proc Natl Acad Sci USA 96 13253–13258 Occurrence Handle1:CAS:528:DyaK1MXns1Gmtrg%3D Occurrence Handle10557307
M Salemi K Strimmer WW Hall M Duffy E Delaporte S Mboup M Peeters A-M Vandamme (2001) ArticleTitleDating the common ancestor of SIVcpz and HIV-1 group M and the origin of HIV-1 subtypes using a new method to uncover clock-like molecular evolution FASEB J 15 276–278 Occurrence Handle1:CAS:528:DC%2BD3MXhsFentbc%3D Occurrence Handle11156935
MO Salminen JK Carr DS Burke FE McCutchan (1995) ArticleTitleIdentification of breakpoints in intergenotypic recombinants of HIV type 1 by bootscanning AIDS Res Hum Retrovir 11 1423– 1425 Occurrence Handle1:CAS:528:DyaK28XitlKlsg%3D%3D Occurrence Handle8573403
Simon F, Mauclere P, Roques P, Loussert-Ajaka I, Muller-Trutwin MC, Sargosti S, Georges-Courbot MC, Barre-Sinoussi F, Brun-Vezinet F (1998) Indentification of a new human immunodeficiency virus type 1 distinct from group M and group O. Nat Med 4:1032–1037
MA Soares T Oliveira ParticleDe RM Brindeiro et al. (2003) ArticleTitleA specific subtype C of human immunodeficiency virus type 1 circulates in Brazil AIDS 17 11–21 Occurrence Handle1:CAS:528:DC%2BD38XpsVSrsrc%3D Occurrence Handle12478065
K Strimmer OG Pybus (2001) ArticleTitleExploring the demographic history of DNA sequences using the generalized skyline plot Mol Biol Evol 18 2298–2305 Occurrence Handle1:CAS:528:DC%2BD3MXptFCrsLY%3D Occurrence Handle11719579
DL Swofford J Sullivan (2003) Phylogeny inference based on parsimony and other methods with PAUP* A-M Salemi M Vandame (Eds) The phylogenetic handbook—A practical approach to DNA and protein phylogeny Cambridge University Press New York
Y Tanaka K Hanada M Mizokami AET Yeo J Wai-Kuo Shih T Gojobori HJ Alter (2002) ArticleTitleA comparison of the molecular clock of hepatitis C virus in the United States and Japan predicts that hepatocellular carcinoma incidence in the United States will increase over the next two decades Proc Natl Acad Sci USA 99 15584–15589 Occurrence Handle1:CAS:528:DC%2BD3sXjvVyl Occurrence Handle12438687
JD Thompson DG Higgins TJ Gibson (1994) ArticleTitleCLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties and weight matrix choice Nucleic Acids Res 22 4673– 4680 Occurrence Handle1:CAS:528:DyaK2MXitlSgu74%3D Occurrence Handle7984417
InstitutionalAuthorNameUN AIDS/WHO Working Group on Global HIV/AIDS and STD Surveillance (2000) Report on the global HIV/AIDS epidemic UN AIDS Geneva
Acknowledgment
This work was supported by the Flemish Funds voor Wetenschappelijk Onderzoek (FWO Grants G.0288.01 and KAN2002 1.5.193.02, Postdoctoral Onderzoeker Contract 530).
Author information
Authors and Affiliations
Corresponding author
Additional information
Reviewing Editor: Dr. Yves Van de Peer
Rights and permissions
About this article
Cite this article
Salemi, M., de Oliveira, T., Soares, M.A. et al. Different Epidemic Potentials of the HIV-1B and C Subtypes. J Mol Evol 60, 598–605 (2005). https://doi.org/10.1007/s00239-004-0206-5
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00239-004-0206-5