Abstract
Mitochondria often use genetic codes different from the standard genetic code. Now that many mitochondrial genomes have been sequenced, these variant codes provide the first opportunity to examine empirically the processes that produce new genetic codes. The key question is: Are codon reassignments the sole result of mutation and genetic drift? Or are they the result of natural selection? Here we present an analysis of 24 phylogenetically independent codon reassignments in mitochondria. Although the mutation-drift hypothesis can explain reassignments from stop to an amino acid, we found that it cannot explain reassignments from one amino acid to another. In particular—and contrary to the predictions of the mutation-drift hypothesis—the codon involved in such a reassignment was not rare in the ancestral genome. Instead, such reassignments appear to take place while the codon is in use at an appreciable frequency. Moreover, the comparison of inferred amino acid usage in the ancestral genome with the neutral expectation shows that the amino acid gaining the codon was selectively favored over the amino acid losing the codon. These results are consistent with a simple model of weak selection on the amino acid composition of proteins in which codon reassignments are selected because they compensate for multiple slightly deleterious mutations throughout the mitochondrial genome. We propose that the selection pressure is for reduced protein synthesis cost: most reassignments give amino acids that are less expensive to synthesize. Taken together, our results strongly suggest that mitochondrial genetic codes evolve to match the amino acid requirements of proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A number of alternative genetic codes are now known, most of them from mitochondria. The leading hypothesis for the evolution of such codes is the “codon capture” model of Osawa and Jukes (Osawa 1995; Osawa et al. 1992). According to this model, codon reassignment is a neutral process that avoids the presumably deleterious consequences of reassigning codons that are in use (Crick 1968). The first step in codon capture is for a codon to disappear from the genome through genetic drift. This can happen if, say, nucleotide bias leads to a shift in codon usage for a given amino acid so that one codon disappears while a cognate codon becomes more frequent. Once a codon has been lost, the selection pressure on the translation machinery to recognize that codon disappears. Then the codon can be reassigned through either of two principal routes. In the first, the tRNA for the lost codon mutates to acquire a new aminoacyl-tRNA synthetase, thus becoming charged with a different amino acid. In the second, the tRNA (or the release factor) for the lost codon mutates so that it can no longer read that codon. At the same time, a different tRNA mutates to recognize the lost codon in addition to its usual codons. (For some tRNAs, one point mutation will be sufficient to effect this (Inagaki et al. 1998; Saks et al. 1998).) But whichever route is taken, when mutation causes the lost codon to reappear in the genome it will be translated as a different amino acid.
The observation that most codon reassignments occur in mitochondria is consistent with the codon capture model. The coding regions of mitochondrial genomes, which encode between 2 and 67 polypeptides (Gray et al. 1998; Palmer 1997), are minute compared with those of eukaryotic nuclei, bacteria, or even chloroplasts. In addition, with the exception of those of higher plants (Wolfe et al. 1987), mitochondrial genomes have extraordinary nucleotide bias: some fungal mitochondria—Pichia canadensis, for example—have adenosine or thymine at more than 97% of fourfold degenerate sites. This combination of small genomes and extreme nucleotide bias makes it likely that codons will be rare or absent by chance: indeed, of 111 complete mitochondrial genomes in public databases at the time of this study, 76 lack one or more codons (mean, 1.6), and 101 have at least one codon (mean, 4.3) that occurs fewer than three times.
The codon capture hypothesis predicts that rarely used codons are more likely than common ones to be reassigned, as rarer codons are more likely to be lost through drift (Osawa et al. 1992). This means that the codon that will be reassigned—call it the focal codon—should be rarer than expected by chance: before the reassignment, it should occur in the genome at a frequency of less than 1 in 64. A codon may be rare for two reasons: it may be rare because the entity—by which we mean either an amino acid or stop—to which it is assigned is rare, or it may be rare because, although the entity to which it is assigned is common, other codons are used for that entity. This gives two further predictions. First, the focal codon should, on average, be rarer than other codons for the same entity. Second, the focal entity should be underused in the genome relative to its representation in the genetic code.
We tested these three predictions by compiling information on mitochondrial codon reassignments and amino acid and codon usage. All mitochondrial codon reassignments in the literature were assembled and all complete mitochondrial genomes downloaded, with the addition of partial genomes where necessary to provide independent contrasts. Codon reassignments were mapped onto an 18S rRNA phylogeny using parsimony. The predictions being tested involve usage patterns at the time of the codon reassignment. However, we cannot know directly what those usage patterns were—the organisms are long extinct—and thus must infer them. To do this, for each reassignment the most closely related sister taxon that did not share the reassignment was identified. Usage patterns in these control taxa provide proxies for usage patterns in the ancestor at the time of the reassignment.
Testing the Codon Capture Model
Methods
The literature was searched for all mitochondria with nonstandard genetic codes (as established through a combination of protein sequencing, comparisons of conserved sites, and sequencing of tRNA anticodon loops; for nonstandard codes and sources thereof, see Table 1) and a phylogeny constructed using 18S rRNA data from the Ribosomal Data Project version 7.1 (http://www.cme.msu.edu/) (Maidak et al. 1999), with additional sources to resolve polytomies in the protostomes (Aguinaldo et al. 1997), fungi (Paquin et al. 1997), chlorophytes (Hayashi-Ishimaru et al. 1996), and protistans (Maddison and Maddison 1998).
The codon reassignments were mapped on to the phylogeny by maximum parsimony, with the solution with fewest reversions being chosen where there were ties. Additional information allowed resolution of the polarity of AUA transitions in the lophotrochozoa (Telford et al. 2000).
All mitochondrial genomes that were complete at the start of this project were downloaded from GenBank, Gobase (http://alice.bch.umontreal.ca/genera/gobase/protein.html), and the FMGP (http://megasun.bch.umontreal.ca/People/lang/FMGP/FMGP.html). For clades lacking a complete genome, all genomes possessing the maximum number of sequenced protein genes were downloaded. Genomes with discrepancies between amino acid and nucleotide sequences were rejected, leaving 111 complete and 17 partial genomes (see Supplementary Information Table 10).
For each genome we recorded the observed amino acid and codon frequencies and estimated the expected frequencies based on codon representation in the ancestral genetic code. The comparison of observed and expected allows us to test whether the focal codon is rare. For example, given the standard code, the stop codon UGA is considered rare if its frequency is less than 1 in 64 of all codons, and the tryptophan codon UGG is considered rare if its frequency is less than 1 in 61 of sense codons—the three stop codons are subtracted from the denominator so as to derive the expected frequency of tryptophan as a function of that of the other amino acids.
Noncrossing phylogenetic independent contrasts (Burt 1989) were selected over minimized evolutionary distances, treating each entity as a separate character with three states: codon gained, codon lost, and unchanged. The control taxa in these contrasts were selected so that (i) they had not undergone any reassignment affecting either of the entities in the reassignment of interest and (ii) they were as closely related as possible to the taxon exhibiting the reassignment of interest while maintaining phylogenetic independence within the set of contrasts as a whole.
Each contrast was then scored as to whether the simple majority of genomes in the prereassignment control taxon underused the entity or codon—within-taxon phylogeny was ignored because of the known problems in reconstructing ancestral base composition through parsimony (Eyre-Walker 1998; Galtier and Gouy 1995).
Results and Discussion
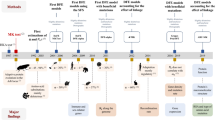
In total, evidence was compiled for 24 mitochondrial codon reassignments, 10 from one amino acid to another (henceforth, sense:sense), 13 from stop to an amino acid (stop:sense), and 1 from an amino acid to stop (Fig. 1).
Mapping of mitochondrial codon reassignments onto eukaryote phylogeny. key: ileAUAmet: AUA is reassigned from isoleucine to methionine. N: any nucleotide. R: adenosine and guanine.*** potentially unassigned. The analysis treats only fully reassigned codons: the inference that a codon absent from the genome is unassigned (not recognized by any tRNA) is unreliable without the characterization of all tRNAs encoded in both the mitochondrion and the nucleus because mitochondrial tRNAs are sometimes imported from the nucleus. The prymnesiophytes (for which only a single gene was obtained) are not used in the analysis, as they could not be contrasted separately from the alveolates (complete genomes with 32 and 44 proteins). The genomes in each clade are listed in Supplementary Information Table 10, and the references for the codon reassignments are given in Table 1. The Metazoa account for 7 of 10 sense:sense codon reassignments. But it is unclear whether this preponderance reflects some special feature of metazoan mitochondria or is merely a study-effort effect, as 88 of 111 completely sequenced mitochondrial genomes at the time of this study are metazoan.
Global analysis of the entire dataset fails to support all three predictions of the codon capture model (Table 2a). Prior to reassignment, focal codons were not used significantly less than random (p = 0.34, one-tailed binomial test, as are all statistics unless otherwise noted; for further details of the statistical analyses and calculations of the binomial coefficients, see Supplementary Information section 3); focal codons were not used significantly less than their cognates (p = 0.14); and focal entities were not significantly underused (p = 0.21). However, if we consider only stop:sense reassignments, the results match the predictions: focal stop codons were used less than random (p = 0.0036), focal stop codons were used less than their cognates (p = 0.017), and stop was underused compared to amino acids (p = 0.0021). The first and third results can be dismissed as trivial, as stop is bound to be underused (if codon usage was random, a genetic code with just one stop codon would produce an average protein length of only 63 amino acids, whereas real mitochondrial proteins average 300 amino acids). However, the comparison with cognate codons cannot be so dismissed and supports the codon capture model. The preponderance of reassignments from stop (13 of 24) is also consistent with the codon capture model, which predicts that stop codons should be more vulnerable to capture than amino acid codons, as stop codons are rarer and thus more likely to disappear from a genome by genetic drift. This difference is clear in Table 2b: note in particular the high a priori probability of stop codons drifting to zero as compared with the tiny probability of the codon(s) involved in sense:sense reassignments doing so. Note, too, that of the 180 codons missing from the 111 complete mitochondrial genomes in our data set, 76 are stop codons.
By contrast, the data for sense:sense reassignments fail to support the codon capture hypothesis (Table 2a). Focal codons were not used less than random (p = 0.99—despite this high p-value, focal codons were not used significantly more than random (p = 0.053)), nor less than their cognates (p = 0.90); and focal amino acids were not underused (p = 0.96). Overall, the focal codons or codon groups are implausible candidates for loss through drift: they are heavily used (median occurrence, 114; quartiles, 77 and 183), and in all cases the majority—usually the great majority—of other codons are more vulnerable to being lost through drift because they are used less (Table 2b). Indeed, in several cases the argument that the codons could vanish through drift is literally incredible (Table 2b “prior prob” column). What might explain this failure of the codon capture hypothesis? One possibility is that our control taxa are not good proxies for the ancestral taxa at the time of reassignment. To test this, for each of the 24 reassignments every control genome was compared with every postreassignment genome, with the number of concordant amino acids for each pair of genomes being recorded (to be concordant an amino acid must be either overused in both genomes or underused in both genomes), and the median calculated. The median of these 24 medians gives 17.5 amino acids of 20 that are concordant in control and postreassignment taxa (Supplementary Information Table 9). However, the evolutionary span between the control taxon and the ancestral taxon will be roughly half that between the control taxon and the postreassignment taxon. Therefore, assuming that the probability of becoming discordant is constant per unit time, to estimate the concordance between the ancestor and the control, the square root of (17.5/20) is taken, giving 19 of 20 amino acids concordant in the control and ancestral taxa. For codon usage, the median of medians is 46 of 64, suggesting that 54 of 64 codons were concordant in control and ancestral taxa. Thus control taxa appear to be good proxies for ancestral taxa. (Strictly speaking, one takes the median of the square roots rather than the square roots of the medians; however, in both cases shown here, the results are mathematically identical.)
Moreover, there are two general reasons for doubting that codon capture can explain sense:sense reassignments in mitochondria: the focal codons are either multiple (CUN, AGR) or composed 100% of AT (AAA, AUA). That multiple codons are less likely to be lost than single ones is obvious. Perhaps less obviously, AT-rich codons are also unlikely to be lost in mitochondrial genomes. This is because the genomes are themselves AT-rich: the genomes available at the time of this study have a median fourfold degenerate GC% of only 0.30 (10th and 90th percentiles, 0.44 and 0.06), with only a single genome (Paramecium aurelia, with a GC% of 0.50) not being AT-rich. The closest relations of mitochondria, the rickettsial group of α-proteobacteria (Andersson et al. 1998), are also AT-rich: Rickettsia prowazekii has a fourfold degenerate GC% of 0.16. We may assume, therefore, that mitochondrial genomes exhibit a general mutational bias toward AT—making AT-rich codons unlikely to be lost through drift (Knight et al. 2001b).
A Weak Selection Alternative
Perhaps, then, sense:sense reassignments occur by some process other than codon capture. One possibility is that the codon is still used at the time of reassignment and that natural selection drives the reassignment (Schultz and Yarus 1994). With selection there is the possibility of various, idiosyncratic reasons for each reassignment, but there is also the possibility of a more general cause. A number of general selective processes based on reductions in mutational load or translational error have been hypothesized to be important in the evolution of the standard genetic code (Knight et al. 1999, 2001a), including error minimization (Alff-Steinberger 1969; Freeland and Hurst 1998; Sella and Ardell 2002) and genetic flexibility (Judson and Haydon 1999; Maeshiro and Kimura 1998). Such forces may well have operated during the early evolution of the standard code, if translation was initially highly ambiguous (i.e., one codon being translated as several amino acids) (Ardell and Sella 2001; Sella and Ardell 2002). However, this condition of ambiguity does not apply to the late derived mitochondrial codes that are the subject of this paper. Thus, in the context of mitochondrial code evolution, mutational load reduction—that is, selection for codes that generate fewer or less deleterious nonsynonymous mutations—and translational load reduction seem unlikely to be important as they give only weak benefits—which in the case of mutational load are not felt upon reassignment, but only when a mutation to the focal codon subsequently occurs—and should be swamped by the immediate consequences of altering the translation of 100-plus codons. Similarly implausible when the focal codon is widely used at the time of reassignment is Andersson and Kurland’s (1991) idea that pressure to shrink the mitochondrial genome selects for codon reassignments that reduce the number of different tRNAs. Our logic is that if the focal codon is widely used, then genome economization under selection requires implausible simultaneous mutations to expand the scope of one tRNA and to delete another: for if the expansion in scope occurs without the deletion, then there is no selection pressure; and if the deletion occurs without the expansion in scope, then the codon will not be translated. In any case, the tRNA reduction hypothesis is not supported by the data (Knight et al. 2001b) (see Supplementary Information section 2).
How, then, can natural selection drive the reassignment of codons that are, at the time of reassignment, in use at an appreciable frequency? One possible scenario is as follows. Suppose that a genome has many sites that are fixed (or polymorphic) for amino acid B. And suppose that, at a substantial number of these sites, amino acid A would actually be favored over amino acid B. (This could happen if, say, amino acid A were cheaper—for example, more readily available or more easily synthesized.) Suppose further that, at any given site, the selection pressure is weak relative to the effects of mutation and drift and is insufficient to cause the replacements. Under these conditions, a compensatory mutation to a tRNA that caused one of the codons for the disfavored amino acid B to be translated as A might be selected for, because the selection pressure would be summed over many sites.
Under this scenario, the focal codon would go through an ambiguous intermediate phase when, following a mutation to a second tRNA that enables this second tRNA to recognize the focal codon, the codon would be translated as two different amino acids (Schultz and Yarus 1994, 1996). (Alternatively, a mutation might allow the original tRNA to be charged by two aminoacyl-tRNA synthetases.) During this intermediate phase, the mitochondrial genome, although haploid, would effectively be heterozygous at sites with the focal codon. Accordingly, new translations at some sites could be deleterious without adverse selective consequences, provided the deleterious translations were not “dominant.” If the selection coefficients summed over all affected sites favored the new translation (perhaps, as suggested earlier, because it is cheaper), then selection would complete the codon reassignment by (i) favoring mutations increasing the ability of the new tRNA—and mutations decreasing the ability of the old tRNA—to recognize the focal codon and (ii) favoring mutations restoring the original amino acid at sites where the new translation is deleterious.
Is this scheme plausible? The notion of the ambiguous intermediate has been controversial, and a number of authors have argued that it cannot work (see, e.g., Osawa and Jukes 1992). However, it is the only mechanism so far proposed that could reassign codons that are in use at an appreciable frequency. Moreover, experimental evidence suggests that ambiguous intermediates can evolve at least in principle: when ambiguous intermediate codon reassignments were artificially engineered into the genomes of Escherichia coli (Doring and Marliere 1998) and Saccharomyces cerevisiae (Santos et al. 1999) by introducing a plasmid-borne nonstandard tRNA, both organisms survived in the laboratory. Turning to the second part of our model—the presence of slightly deleterious mutations—we may note that these are thought to have been accumulated in great numbers in mitochondria (Lynch 1996; Lynch and Blanchard 1998; Nachman 1998; Rand and Kann 1998). However, the details of how the selection coefficients would need to be distributed in order for a codon reassignment to be favored are still to be worked out.
The weak selection scenario predicts that disfavored amino acids are more likely to lose codons, and favored ones are more likely to gain them. In fact, the critical condition is that the amino acid gaining the codon should be favored relative to the amino acid losing it. Disfavored amino acids are presumably selected against and, therefore, used less than expected based on their representation in the code and the nucleotide bias of the genome; favored amino acids are presumably used more than expected (King and Jukes 1969).
Testing the Weak Selection Hypothesis
Methods
The methods are as described for the codon capture model, except that the test for the weak selection hypothesis (Table 3) is whether the focal amino acid is selected against. To establish this, expected amino acid and stop (“entity”) frequencies (under the assumption of neutrality) were estimated from the nucleotide bias at fourfold degenerate sites—i.e., sites where all four nucleotides encode the same amino acid—according to the King and Jukes (1969) method. For example, for the standard code, given nucleotide proportions 0.3 U, 0.2 C, 0.4 A, and 0.1 G, the expected stop codon frequency is (0.3 × 0.4 × 0.1) + (0.3 × 0.4 × 0.4) + (0.3 × 0.1 × 0.4) = 0.072, and the expected frequency of cysteine (codons UGU and UGC) is ((0.3 × 0.1 × 0.3) + (0.3 × 0.1 × 0.2))/(1 − 0.072) = 0.016. (The expected frequency of stop codons, 0.072, is subtracted from the denominator so as to derive the expected frequency of cysteine as a function of that of the other amino acids.)
Results and Discussion
The predictions of the weak selection hypothesis are supported (Table 3, A). Amino acids that lost codons were disfavored prior to the reassignment (p = 0.0026), amino acids that gained codons were favored, albeit not significantly (p = 0.22); and, critically, the amino acid gaining the codon was favored relative to the one losing it (p = 0.0020, one-tailed sign test). When the tests are repeated on the postreassignment taxa—i.e., using the postreassignment genetic code, nucleotide bias, and amino acid composition—these patterns disappear (Supplementary Information Table 7). Amino acids that lost codons are no longer disfavored (p = 0.81), amino acids that gained codons are no longer favored (p = 0.91), and the amino acid that gained the codon is no longer favored relative to the one that lost it (p = 0.91, one-tailed sign test). These negative results show that the positive test results for the prereassignment taxa are not an artifact affecting all mitochondrial genomes equally. Rather, they indicate real selection pressure against the genetic code that is relieved by the codon reassignment.
Two additional observations are consistent with these results. First, nucleotide bias in the control taxa appears to be a reasonable proxy for that in ancestral taxa. A measure of this is the correlation between the fourfold degenerate usage in the control and postreassignment taxa: and the median of medians correlation coefficient is 0.81 (Supplementary Information Table 9). Moreover, the better the proxy, the better the weak selection model fits. The median of medians correlation coefficient for those reassignments where there is any result contrary to the weak selection model in Table 3(A) is only 0.34, whereas that for the perfectly predicted reassignments is 0.96. The difference is significant (p = 0.005, one-tailed Mann–Whitney U-test) and suggests that incorrect predictions may be the result of noise in the data.
The second observation is that conservatively grouping the 10 reassignments into six codon reassignment types still supports the weak selection model (Table 3, B). Amino acids that lost codons were disfavored (p = 0.024), and in all six cases the amino acid that gained the codon was favored relative to the one that lost it (p = 0.016, one-tailed sign test). This suggests that the weak selection model may explain mitochondrial sense-codon reassignments in general, not just the high recurrence of particular types.
The weak selection model generates additional hypotheses. First, under the proposed mechanism, codon reassignment is facilitated if selection can act on a single amino acid property that is the same at all protein sites. A plausible candidate for such a global property is amino acid synthesis cost. And remarkably, in 9 of 10 sense:sense reassignments the amino acid gaining the codon is cheaper than the one losing it (p = 0.0068, Wilcoxon signed-rank test; Table 4). (The codon capture hypothesis does not predict that reassignments will reduce cost, and indeed tryptophan, which gains the codon in 9 of 13 reassignments from stop to an amino acid (Table 2a), is the most expensive amino acid of all.)
Second, the proposed mechanism also depends upon many of the reassigned sites being effectively neutral with respect to function. The number of sites that are effectively neutral for the two amino acids involved in the codon reassignment will be increased if these amino acids are (i) similar to one another physically and chemically and (ii) relatively interchangeable in databases of aligned sequences. Both conditions are met (Table 4): compared with the pairs in all 390 possible sense:sense reassignments, the two amino acids in the observed reassignments are both more similar (p = 0.011, one-tailed sign test) and relatively unconstrained (p = 0.00098). The codon capture hypothesis does not make these predictions, and indeed tryptophan is the least interchangeable amino acid of all.
If many sites are effectively neutral, then a further prediction may be made: codon reassignments will affect amino acid usage, a phenomenon previously reported for three amino acids (Andersson and Kurland 1991; Castresana et al. 1998). The data presented here support this prediction (Table 5). Amino acids that lost codons become less frequent in postreassignment genomes and those that gained them become more frequent (p = 0.00027, one-tailed sign test). Mitochondrial amino acid composition is also known to respond to shifts in nucleotide bias (Jermiin et al. 1994), which provides yet further evidence for weakly selected sites. Note that although codon reassignments clearly alter amino acid usage, there is no evidence in Table 5 that they alter protein length (which they might do for reassignments involving stop): presumably stop codons are rarely neutral.
Conclusions
We found that two different processes seem to drive codon reassignments in mitochondria. Codon reassignments from stop to an amino acid appear to take place via codon capture after a given codon has fallen out of use. The evidence in favor of the codon capture model is the large number of such reassignments and the rarity of the focal stop codon compared with its cognates. The codon capture model fails, however, to explain sense:sense reassignments, because the focal codons appear to be appreciably in use at the time of reassignment. We suggest one scenario—the weak selection hypothesis—for how such reassignments could be favored by natural selection. Apart from the codons being well used at the time of reassignment, we present several results consistent with the model: the amino acid gaining a codon is preferred in the proxy taxa compared to the amino acid losing the codon, it is typically cheaper to synthesize, and the two amino acids are relatively similar and interchangeable. (Interestingly, the same features are also found in the only known nonmitochondrial sense: sense codon reassignment—leuCUGser in the nuclear genome of some Candida yeasts (Kawaguchi et al. 1989) (see Supplementary Information section 1).) However, the evidence cannot be taken as conclusive, and much work needs to be done, for example, to define the distribution of selection coefficients that would allow a weak selection model such as ours to operate and to determine empirically whether such a distribution is plausible.
Regardless of the underlying selective forces, our results indicate that more frequently useful amino acids accrete codons at the expense of less frequently useful amino acids. If there are many such reassignments, then more frequently useful amino acids will come to have more codons, and one would observe a positive correlation between the usage of an amino acid and its representation in the genetic code. Such a correlation has long been known for organisms using the standard genetic code and has been attributed solely to a high frequency of selectively neutral sites (Jukes et al. 1975; King and Jukes 1969; Osawa 1995). Here we see that such appearances of neutrality may also arise through selection.
References
AMA Aguinaldo JM Turbeville LS Linford MC Rivera JR Garey RA Raff JA Lake (1997) ArticleTitleEvidence for a clade of nematodes, arthropods and other moulting animals Nature 387 489–492
C Alff-Steinberger (1969) ArticleTitleThe genetic code and error transmission Genetics 64 584–591
S Anderson AT Bankier BG Barrell MHL Debruijn AR Coulson J Drouin IC Eperon DP Nierlich BA Roe F Sanger PH Schreier AJH Smith R Staden IG Young (1981) ArticleTitleSequence and organization of the human mitochondrial genome Nature 290 457–465 Occurrence Handle1:CAS:528:DyaL3MXlt1OlsL8%3D Occurrence Handle7219534
SGE Andersson CG Kurland (1991) ArticleTitleAn extreme codon preference strategy—Codon reassignment Mol Biol Evol 8 530–544 Occurrence Handle1:CAS:528:DyaK3MXks1Kitb0%3D Occurrence Handle1921708
SGE Andersson A Zomorodipour JO Andersson T Sicheritz-Ponten UCM Alsmark RM Podowski AK Naslund A-S Eriksson HH Winkler CG Kurland (1998) ArticleTitleThe genome sequence of Rickettsia prowazekii and the origin of mitochondria Nature 396 133–140 Occurrence Handle10.1038/24094 Occurrence Handle1:CAS:528:DyaK1cXnslamtL0%3D Occurrence Handle9823893
DH Ardell G Sella (2001) ArticleTitleOn the evolution of redundancy in genetic codes J Mol Evol 53 269–281
BG Barrell ATD Bankier J Drouin (1979) ArticleTitleA different genetic code in human mitochondria Nature 282 189–194
CT Beagley R Okimoto DR Wolstenholme (1998) ArticleTitleThe mitochondrial genome of the sea anemone Metridium senile (Cnidaria): Introns, a paucity of tRNA genes, and a near-standard genetic code Genetics 148 1091–1108
Y Bessho T Ohama S Osawa (1992) ArticleTitlePlanarian mitochondria. II. The unique genetic code as deduced from cytochrome-c-oxidase subunit I gene sequences J Mol Evol 34 331–335
JL Boore LL Daehler WM Brown (1999) ArticleTitleComplete sequence, gene arrangement, and genetic code of mitochondrial DNA of the cephalochordate Branchiostoma floridae (Amphioxus) Mol Biol Evol 16 410–418
C Boyen C Leblanc G Bonnard J-M Grienenberger B Kloareg (1994) ArticleTitleNucleotide sequence of the COX3 gene from Chondrus crispus: Evidence that UGA encodes tryptophan and evolutionary implications Nucleic Acids Res 22 1400–1403
CE Bullerwell J Leigh BF Lang (2003) ArticleTitleA comparison of three fission yeast mitochondrial genomes Nucleic Acids Res 31 759–768 Occurrence Handle10.1093/nar/gkg134 Occurrence Handle1:CAS:528:DC%2BD3sXitlCnsrk%3D Occurrence Handle12527786
G Burger I Plante KM Lonergan MW Gray (1995) ArticleTitleThe mitochondrial DNA of the amoeboid protozoon, Acanthamoeba castellanii: Complete sequence, gene content and genome organization J Mol Biol 245 522–537
A Burt (1989) Comparative methods using phylogenetically independent contrasts PH Harvey L Partridge (Eds) Oxford surveys in evolutionary biology OUP Oxford 33–53
J Castresana G Feldmaier-Fuchs S Paabo (1998) ArticleTitleCodon reassignment and amino acid composition in hemichordate mitochondria Proc Natl Acad Sci USA 95 3703–3707
CL Craig RS Weber (1998) ArticleTitleSelection costs of amino acid substitutions in ColE1 and ColIa gene clusters harbored by Escherichia coli Mol Biol Evol 15 774–776
FHC Crick (1968) ArticleTitleThe origin of the genetic code J Mol Biol 38 367–379
VF de la Cruz N Neckelmann L Simpson (1984) ArticleTitleSequences of six genes and several open reading frames in the kinetoplast maxicircle DNA of Leishmania tarentolae J Biol Chem 259 5136–5147
V Doring P Marliere (1998) ArticleTitleReassigning cysteine in the genetic code of Escherichia coli Genetics 150 543–551
M Ehara Y Hayashi-Ishimaru Y Inagaki T Ohama (1997) ArticleTitleUse of a deviant mitochondrial code in yellow-green algae as a landmark for segregating members within the phylum J Mol Evol 45 119–124
A Eyre-Walker (1998) ArticleTitleProblems with parsimony in sequences of biased base composition J Mol Evol 47 686–690
SJ Freeland LD Hurst (1998) ArticleTitleThe genetic code is one in a million J Mol Evol 47 238–248 Occurrence Handle1:CAS:528:DyaK1cXmt1ansrs%3D Occurrence Handle9732450
N Galtier M Gouy (1995) ArticleTitleInferring phylogenies from DNA sequences of unequal base compositions Proc Natl Acad Sci USA 92 11317–11321
R Grantham (1974) ArticleTitleAmino acid difference formula to help explain protein evolution Science 185 862–864
MW Gray BF Lang R Cedergren GB Golding C Lemieux D Sankoff M Turmel N Brossard E Delage TG Littlejohn I Plante P Rioux D Saint-Louis Y Zhu G Burger (1998) ArticleTitleGenome structure and gene content in protist mitochondrial DNAs Nucleic Acids Res 26 865–878 Occurrence Handle10.1093/nar/26.4.865 Occurrence Handle1:CAS:528:DyaK1cXhvVWnsL0%3D Occurrence Handle9461442
Y Hayashi-Ishimaru T Ohama Y Kawatsu K Nakamura S Osawa (1996) ArticleTitleUAG is a sense codon in several chlorophycean mitochondria Curr Genet 30 29–33
Y Hayashi-Ishimaru M Ehara Y Inagaki T Ohama (1997) ArticleTitleA deviant mitochondrial genetic code in prymnesiophytes (yellow algae): UGA codon for tryptophan Curr Genet 32 296–299
RJ Hoffmann JL Boore WM Brown (1992) ArticleTitleA novel mitochondrial genome organization for the blue mussel, Mytilus edulis Genetics 131 397–412 Occurrence Handle1:CAS:528:DyaK2cXis1anurk%3D Occurrence Handle1386586
MES Hudspeth WM Ainley DS Shumard RA Butow LI Grossman (1982) ArticleTitleLocation and structure of the var1 gene on yeast mitochondrial DNA: nucleotide sequence of the 40.0 allele Cell 30 617–626
Y Inagaki M Ehara KI Watanabe Y Hayashi-Ishimaru T Ohama (1998) ArticleTitleDirectionally evolving genetic code: the UGA codon from stop to tryptophan in mitochondria J Mol Evol 47 378–384
HT Jacobs DJ Elliott VB Math A Farquharson (1988) ArticleTitleNucleotide sequence and gene organization of sea urchin mitochondrial DNA J Mol Biol 202 185–217
LS Jermiin D Graur RM Lowe RH Crozier (1994) ArticleTitleAnalysis of directional mutation pressure and nucleotide content in mitochondrial cytochrome-b genes J Mol Evol 39 160–173
D Jones W Taylor J Thornton (1992) ArticleTitleThe rapid generation of mutation data matrices from protein sequences CABIOS 8 275–282 Occurrence Handle1:CAS:528:DyaK38Xlt1Okt7w%3D Occurrence Handle1633570
OP Judson D Haydon (1999) ArticleTitleThe genetic code: What is it good for? An analysis of the effects of selection pressures on genetic codes J Mol Evol 49 539–550 Occurrence Handle1:CAS:528:DyaK1MXnsVajsbw%3D Occurrence Handle10552035
T Jukes R Holmquist H Moise (1975) ArticleTitleAmino acid composition of proteins: selection against the genetic code Science 189 50–51 Occurrence Handle1:CAS:528:DyaE2MXksleqs7s%3D Occurrence Handle237322
Y Kawaguchi H Honda J Taniguchi-Morimura S Iwasaki (1989) ArticleTitleThe codon CUG is read as serine in an asporogenic yeast Candida cylindracea Nature 341 164–166 Occurrence Handle10.1038/341164a0 Occurrence Handle1:CAS:528:DyaK3cXjtFCgsg%3D%3D Occurrence Handle2506450
JL King TH Jukes (1969) ArticleTitleNon-Darwinian evolution Science 164 788–798
RD Knight SJ Freeland LF Landweber (1999) ArticleTitleSelection, history and chemistry: The three faces of the genetic code TIBS 24 241–247
RD Knight SJ Freeland LF Landweber (2001a) ArticleTitleRewiring the keyboard: evolvability of the genetic code Nature Rev Genet 2 49–58
RD Knight LF Landweber M Yarus (2001b) ArticleTitleHow mitochondria redefine the code J Mol Evol 53 299–313
M-J Laforest I Roewere BF Lang (1997) ArticleTitleMitochondrial tRNAs in the lower fungus Spizellomyces punctatus: tRNA editing and UAG ‘stop’ codons recognized as leucine Nucleic Acids Res 25 626–632
M Lynch (1996) ArticleTitleMutation accumulation in transfer RNAs: Molecular evidence for Muller’s ratchet in mitochondrial genomes Mol Biol Evol 13 209–220 Occurrence Handle1:CAS:528:DyaK28XhtVylu7w%3D Occurrence Handle8583893
M Lynch JL Blanchard (1998) ArticleTitleDeleterious mutation accumulation in organelle genomes Genetica 102 29–39
G Macino G Coruzzi F Nobrega M Li A Tzagoloff (1979) ArticleTitleUse of the UGA terminator as a tryptophan codon in yeast mitochondria Proc Natl Acad Sci USA 76 3784–3785
Maddison DR, Maddison WP (1998) The Tree of Life (a multiauthored, distributed Internet project containing information about phylogeny and biodiversity). http://phylogeny. ;arizona.edu/tree/phylogeny.html/
T Maeshiro M Kimura (1998) ArticleTitleThe role of robustness and changeability on the origin and evolution of genetic codes Proc Natl Acad Sci USA 95 5088–5093
B Maidak J Cole C Parker G Garrity N Larsen B Li T Lilburn M McCaughey G Olsen R Overbeek S Pramanik T Schmidt J Tiedje C Woese (1999) ArticleTitleA new version of the RDP (Ribosomal Database Project) Nucleic Acids Res 27 171–173 Occurrence Handle10.1093/nar/27.1.171 Occurrence Handle1:CAS:528:DyaK1MXpsVKjsw%3D%3D Occurrence Handle9847171
A McLachlan (1971) ArticleTitleTests for comparing related amino-acid sequences cytochrome c and cytochrome c551 J Mol Biol 61 409–424 Occurrence Handle1:CAS:528:DyaE38XhsF2k Occurrence Handle5167087
T Miyata S Miyazawa T Yasunaga (1979) ArticleTitleTwo types of amino acid substitutions in protein evolution J Mol Evol 12 219–236 Occurrence Handle1:CAS:528:DyaE1MXhvVWrtLk%3D Occurrence Handle439147
MW Nachman (1998) ArticleTitleDeleterious mutations in animal mitochondrial DNA Genetica 102 61–69
R Okimoto JL Macfarlane DO Clary DR Wolstenholme (1992) ArticleTitleThe mitochondrial genomes of two nematodes, Caenorhabditis elegans and Ascaris suum Genetics 130 471–498 Occurrence Handle1:CAS:528:DyaK3sXhs1aju7g%3D Occurrence Handle1551572
S Osawa (1995) Evolution of the genetic code OUP Oxford
S Osawa T Jukes T Watanabe A Muto (1992) ArticleTitleRecent evidence for evolution of the genetic code Microbiol Rev 56 229–264
JD Palmer (1997) ArticleTitleThe mitochondrion that time forgot Nature 387 454–455
B Paquin M-J Laforest L Forget I Roewer Z Wang J Longcore BF Lang (1997) ArticleTitleThe fungal mitochondrial genome project: Evolution of fungal mitochondrial genomes and their gene expression Curr Genet 31 380–395
DM Rand LM Kann (1998) ArticleTitleMutation and selection at silent and replacement sites in the evolution of animal mitochondrial DNA Genetica 102 393–407
MS Saks JR Sampson J Abelson (1998) ArticleTitleEvolution of a transfer RNA gene through a point mutation in the anticodon Science 279 1665–1670
MA Santos C Cheesman V Costa P Moradas-Ferreira M Tuite (1999) ArticleTitleSelective advantages created by codon ambiguity allowed for the evolution of an alternative genetic code in Candida spp Mol Microbiol 31 937–947
DW Schultz M Yarus (1994) ArticleTitleTransfer RNA mutation and the malleability of the genetic code J Mol Biol 235 1377–1380
DW Schultz M Yarus (1996) ArticleTitleOn malleability in the genetic code J Mol Evol 42 597–601
G Sella DH Ardell (2002) ArticleTitleThe impact of message mutation on the fitness of a genetic code J Mol Evol 54 638–651
MJ Telford EA Herniou RB Russell DTJ Littlewood (2000) ArticleTitleChanges in mitochondrial genetic codes as phylogenetic characters: Two examples from the flatworms Proc Natl Acad Sci USA 97 11359–11364
M Turmel C Lemieux G Burger BF Lang C Otis I Plante MW Gray (1999) ArticleTitleThe complete mitochondrial DNA sequences of Nephroselmis olivacea and Pedinomonas minor: Two radically different evolutionary patterns within green algae Plant Cell 11 1717–1729 Occurrence Handle10.1105/tpc.11.9.1717 Occurrence Handle1:CAS:528:DyaK1MXms1ahsr4%3D Occurrence Handle10488238
K Wolfe W-H Li P Sharp (1987) ArticleTitleRates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast and nuclear genes Proc Natl Acad Sci USA 84 9054–9058 Occurrence Handle1:CAS:528:DyaL1cXovVyktQ%3D%3D Occurrence Handle3480529
S Yokobori T Ueda K Watanabe (1993) ArticleTitleCodons AGA and AGG are read as glycine in ascidian mitochondria J Mol Evol 36 1–8
Acknowledgments
Many thanks go to Dan Haydon and two anonymous reviewers for extensive comments, criticisms, and suggestions.
Author information
Authors and Affiliations
Corresponding author
Additional information
Reviewing Editor: Dr. Dmitri Petrov
Appendix
Rights and permissions
About this article
Cite this article
Swire, J., Judson, O.P. & Burt, A. Mitochondrial Genetic Codes Evolve to Match Amino Acid Requirements of Proteins. J Mol Evol 60, 128–139 (2005). https://doi.org/10.1007/s00239-004-0077-9
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00239-004-0077-9