Abstract
The evolutionary properties of a metabolic network may be determined by the topology of the network. One attribute of pathways that make up the network is the number of enzymatic steps between initial substrates and final products. To determine the effect of pathway length on evolutionary lability of pathway structure, we examined amino acid biosynthetic pathways across 48 sequenced organisms. We demonstrate that longer pathways exhibit lower rates of change in pathway structure than shorter pathways. This finding suggests that increasing complexity may increase constraint on evolutionary change.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Selection can both foster and inhibit evolutionary changes in metabolism (Feder 1996; Whitt et al. 2002). Changes in the metabolic properties of a species can be crucial in adaptive responses to a changed environment or in exploiting a new ecological niche (Tatusov et al. 1996; Huynen et al. 1999). However, changes in metabolic processes that are fundamental to the function of an organism may be severely constrained (Li et al. 2001; Chalker et al. 2001). Certain topological and genetic features of the metabolic network may help to predict which elements of the network are most easily changed and which are likely to remain static (Wagner 2001; Cornish-Bowden and Cardenas 2002; Gu et al. 2003). For example, the highly connected nature of certain gene products within the genome has been linked to evidence of purifying selection on those genes (Fraser et al. 2002). Furthermore, in yeast, knockouts of highly connected genes have been shown to lead to severe fitness effects relative to knockouts of less connected genes (Wagner 2001).
One characteristic of metabolic networks that has not previously been examined and that may influence the action of selection upon the network is the length of individual biochemical pathways within the network (Rison et al. 2002). We define longer pathways as those that require more enzymes or reactions to convert an initial substrate into the required product (and thus more genes coding for those enzymes) than “shorter” pathways. This characteristic has also been termed pathway distance (Rison et al. 2002) or metabolic distance (Kolesov et al. 2001). Although many pathways are branching, certain substrates are sufficiently common and certain products sufficiently important that the reaction flux is likely to frequently flow from the initial substrate to the final product. Consequently, the number of steps involved in this process could have a direct impact on evolutionary changes in the system.
For example, if each step in a pathway evolves independently of the other steps, there will be more opportunity for evolutionary change in longer pathways than in shorter pathways simply as a result of the larger number of steps. Each enzyme involved in a longer pathway could potentially be specialized for the organisms’ particular environment. Regions of the metabolic network that involve relatively longer pathways might diverge more rapidly than areas involving shorter pathways because of the vast number of combinations of character states possible if every step varies independently. In this case, changes at any particular gene within the network would be independent of pathway length, but longer pathways would be more likely to show more change overall. Alternatively, longer pathways might require precise interactions between enzymes that would select against changes in those pathways. In this case, the maintenance of epistatic interactions by selection would be expected to be less important in shorter pathways with fewer potential interactions, and the lack of constraint would be manifest in greater evolutionary change in shorter pathways.
The recent accumulation of complete genome sequences has created the opportunity to test these hypotheses about the relationship between pathway length and evolutionary lability. BLAST searches and other methods of identifying genes that are likely to share function across organisms allow the construction of putative metabolic networks in organisms for which every metabolic step has not been tested with knockout mutants or biochemical methods (found in databases such as KEGG [Kanehisa et al. 2002], ERGO [Overbeek et al. 2003], and MetaCyc [Karp et al. 2002]). These putative metabolic networks can then be compared across organisms to assess the ways in which the network has changed over time (Forst and Schluten 2001; Alves et al. 2002).
We investigated the relationship between pathway length and evolutionary lability by examining the biosynthesis of the coded amino acids across 48 sequenced organisms. Amino acid biosynthesis pathways are highly informative because these pathways vary in length, are well characterized both biochemically and genetically, and are utilized for similar functions across organisms (Herrmann and Somerville 1983). We used the pattern of evolutionary change in amino acid biosynthesis to derive an empirical estimate of the relationship between pathway length and evolutionary lability, and tested the above hypotheses about the effect of pathway length on evolutionary change.
Methods
We examined biosynthetic pathways of 19 of the coded amino acids across a wide range of sequenced organisms (the biosynthesis of isoleucine and valine are enzymatically identical). Organisms were chosen to represent the widest diversity of taxa from all completely sequenced and annotated genomes at the time of analysis. We included a single representative of each of 48 species, including 5 Archaea, 41 Eubacteria, and 2 Eukaryotes. Organisms whose phylogenetic positions were not clearly resolved or which did not have complete pathway data at the start of the study were excluded from the analysis.
Determining Pathway Character States
We used ERGO (Overbeek et al. 2003), a database of metabolic pathways curated by Integrated Genomics, to determine the structure of all biosynthetic pathways used to produce the common amino acids in each species. In this database, organisms are scored for the presence or absence of each gene that could be involved in a previously characterized amino acid biosynthetic pathway. Based on these data, and on confirmation of the function of those genes by expert annotators, Integrated Genomics asserts whether or not an organism uses a particular biosynthetic pathway. Our analyses were conducted only on these asserted pathways. While there is a degree of arbitrariness in defining the linear structure of any metabolic pathway, all pathways were defined based on biochemical convention (Herrmann and Sommerville 1983; Bender 1985).These may be simplistic representations of the true network of pathways, but there is no reason to suspect that using biochemical convention to define pathways will systematically bias the results. Furthermore, we chose pathways such that there were no overlapping pathways across amino acids, and thus the pathway length within each synthesis pathway was independent of all other synthesis pathways. Aminotransferase steps were not included in the analysis because these are highly variable and frequently shared across pathways, thus it is difficult to accurately assign a particular aminotransferase to a specific pathway (Herrmann and Somerville 1983).
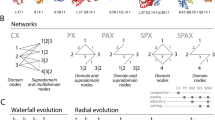
Pathways were considered different if their enzymatic composition differed at any step. The pathway structure character state for each species is the pathway (or set of different pathways) used by that species to synthesize a given amino acid. Because some organisms have multiple functional pathways to produce a single amino acid, each combination of pathways used by an organism was defined as a unique character state (Fig. 1A). We assumed that each change between character states involves either the loss or the gain of at least a single enzyme and that losses and gains are equally likely to occur. We use this simple model because we have no prior expectation as to the relative frequency of losses and gains (see Discussion for further elaboration). We then defined the distance between two pathway character states as the number of changes, i.e., enzyme additions or deletions, that would be necessary to convert one pathway, or set of pathways, into another (Fig. 1B).
Some of the possible ways that organisms make methionine. The number above each arrow corresponds to the EC number of the enzyme which catalyzes the reaction. Some organisms (type X) have two alternative pathways for producing methionine. Other organisms have just the first (type Y) or the second (type Z) of the two pathways found in the X organisms. B If these sets of methionine biosynthesis pathways are considered different character states, this matrix defines the number of steps necessary to change from one character state to another when gain or loss of an enzyme is counted as one step. C The total change in an amino acid biosynthetic pathway is computed from the total number of changes across the phylogeny using a parsimony criterion (see Methods). Each character state is represented by a color: white = X, hatched = Y, and black = Z.
Determining Evolutionary Lability
A phylogeny of the study organisms was produced based on maximum-likelihood analysis of 1415 bp from 16S rRNA (Fig. 2). Sequences were obtained from the ssu rRNA database (Wuyts et al. 2002). Sequences were aligned in ClustalX (Thompson et al. 1997) and phylogenetic analyses were performed in PAUP* 4.0 (Swofford 1998). A maximum likelihood tree was obtained based on a general time reversible model plus among site variation plus gamma (this was determined to be the best model based on hierarchical likelihood ratio tests in Modeltest version 3.0 [Posada and Crandall 1998]). The tree is shown rooted with the eukaryotic lineage; the results are not dependent on choice of rooting.
For each amino acid, character state information from the set of organisms capable of synthesizing that amino acid was used to estimate the evolutionary lability of its synthesis pathway. Lability was determined by quantifying the total change in that pathway that occurred across the study organisms. The total amount of change in the biosynthetic pathways for each amino acid was determined by mapping the pathways onto our reconstructed phylogeny. The distance, or number of changes, between character states was simply the number of enzymes that were lost or gained between those states. Given the character state of each organism and distance between each state, we used MacClade (Maddison and Maddison 2000) to calculate the total number of changes that occurred across all organisms for each amino acid (Fig. 1C). This total number of changes across the phylogeny is our measure of lability.
Pathway Length
We also determined a phylogenetic average of pathway length for each amino acid using MacClade (Maddison and Maddison 2000); in general, the lengths of the multiple pathways involved in synthesis of a particular amino acid were similar (average variance in length within an amino acid = 0.85; variance across all amino acids = 5.76; mean pathway length = 3.18 steps).
The Relationship of Pathway Length and Evolutionary Lability
To determine the relationship between the length of a pathway and the amount of evolutionary change in that pathway, we performed a linear regression analysis of pathway lability on pathway length (general linear models procedure in SAS (SAS Institute 1990). To meet the assumption of normality of the residuals, the lability measure was transformed with the Box–Cox (1964) procedure. The Box–Cox procedure estimates the best transformation to normality within the family of power transformations. A λ value of −0.25 was determined to be optimal (maximizing the log-likelihood function) through an iterative procedure and was used in transformation. Lability was transformed as (lability + 1) to account for pathways that did not change. Thus, under the Box–Cox transformation, lability was transformed to [(lability + 1)−0.25 − 1]/(−0.25). This transformation resulted in a normal distribution of residuals in the relationship between the transformed lability value and the measure of pathway length. After this transformation, linear regression is appropriate to determine if pathway length influences evolutionary change in amino acid biosynthesis pathways. We also conducted an additional nonparametric analysis, using Kendall’s rank correlation test on the untransformed measures of lability and pathway length, to test if lability is a monotonically increasing or decreasing function of pathway length. For either of these analyses, a positive relationship between lability and length would be consistent with the hypothesis that longer pathways change faster because of their greater number of parts or neutrality of change. A negative relationship between lability and length would conform to the prediction that epistatic interactions lead to greater constraint in longer pathways.
Results
We found a negative linear relationship between pathway length and lability across taxa and pathways (Fig. 3) (r 2 = 0.332, p < 0.01). This finding indicates that longer biosynthetic pathways are less likely to have changed in any steps than shorter pathways. An analogous but less powerful nonparametric test, Kendall’s rank correlation, also found that lability declined with pathway length (p = 0.05).
The relationship between pathway length, calculated as the phylogenetic average of pathway lengths for those amino acids with multiple pathways that differ in length, and evolutionary lability, calculated as the total change in a pathway as in fig. 1C. The “essential” amino acids, which humans cannot synthesize, are shown as open circles. The linear regression on transformed data (not shown) detected a significant negative effect of pathway length on change in the pathway (F = 6.41, p = 0.0215).
The number of character states, i.e., pathways or combinations of pathways, ranged from 1 to 26. Glutamate had 26 character states, glycine 22, methionine 9, and the remaining amino acids had 5 or fewer character states. Seven of the 19 amino acid pathways did not vary across the studied organisms, i.e., all organisms that produce these amino acids use the same biosynthetic pathway (Fig. 3). All amino acids have identifiable pathways in more than 60% of the study organisms except for asparagine, methionine, and lysine, which are absent in 28, 28, and 31 organisms, respectively. Removing these amino acids from the analyses does not qualitatively change the results. In addition, the results do not seem to be driven by any particular lineage, i.e., no one lineage is unusually variable or invariant. For example, removing the eukaryotes or archaea or both does not change the result.
A complicating factor in the interpretation of these results stems from the fact that several of these pathways are interconnected, i.e., they share enzymatic steps. Although our pathways were independent in that they were chosen to be nonoverlapping, if selection particularly constrains pathway steps that are necessary for the synthesis of multiple amino acids, this could obscure any direct relationship between pathway length and evolutionary lability. In order to determine whether sharing enzymes across multiple pathways can explain the observed variation, we performed an analysis of variance including the number of shared steps in a pathway as a covariate with pathway length. Specifically, for a given pathway the covariate was the number of synthesis steps shared with the synthesis pathway of any other amino acid. This analysis reveals that the number of shared steps is not a significant factor (F = 0.47, p = 0.50), while pathway length remained significant (F = 7.66, p = 0.014). In addition, in order to determine whether the observed trend might be due to specific features of the amino acids, we performed multiple ANOVAs with each of a number of covariates: hydrophobicity (following Barrett and Elmore 1998), pK a of the protonated α-amino group, pK a of the carboxylic acid group on the α-carbon, pK a of the side chain (for the appropriate subset of amino acids), pI, and molecular weight of the amino acids. None of these factors were able to explain a significant amount of variation in pathway lability (p ≥ 0.31 for the covariates), while the effect of pathway length remained significant in every analysis.
The trend we observe, greater change in shorter pathways, could be produced by short pathways having a larger number of character states or a higher frequency of alternating between a small number of character states. For example an amino acid pathway structure with only two character states but four independent originations of one pathway structure from another could be measured as having the same amount of evolutionary lability as a case in which four different character states were present in the organisms studied, each of which emerged only once. These two scenarios represent very different evolutionary phenomena. To test which of these has occurred, we calculated, for each amino acid, a character switching index, where

Thus, high lability with a low number of character states will have a high degree of switching, indicating that there is frequent change between character states. An analysis of the relationship between the pathway length and the character switching index reveals no significant relation between the two (p = 0.31). In addition, there is a significant positive correlation between the number of pathway character states available for a given amino acid and the total amount of change observed for that amino acid (Pearson’s correlation coefficient, 0.619; p < 0.005). This indicates that short pathways have a greater diversity of possible pathways and/or combinations of pathways, as opposed to simply switching more frequently between a small number of pathways. Thus, there is not significant convergence of pathways, but rather when an evolutionary change in amino acid biosynthesis occurs (that does not involve loss of the entire pathway), it usually involves the production of an entirely novel pathway to produce that amino acid.
While the observed patterns appear to be robust, there are some amino acids that do not fit the observed trends. For example, glutamine is produced via a very short pathway, one step, and is invariant across the study organisms (Herrmann and Somerville 1983). Arginine has a large total amount of change, but among only a few pathways, indicating a high degree of switching.
Discussion
To our knowledge, this study is the first empirical evidence that the length of a biochemical pathway has the potential to constrain the rate of evolution in that system. In the set of 48 sequenced organisms studied here, there was a strong tendency for longer amino acid biosynthesis pathways to remain evolutionarily static in their structure. Shorter pathways demonstrated greater evolutionary lability, and this occurred through pathway diversification instead of alternation between set pathway character states. This finding suggests that the connection of steps in long pathways may deter evolutionary change. Radical change in upstream genes may require simultaneous compensatory change in so many downstream genes as to render any such change extremely deleterious (Rausher et al. 1999). In addition, if multiple enzyme-catalyzed reactions are crucial to producing the final product, it may be very difficult for alternative pathways to develop rapidly enough to compete with the original pathway. Wholesale substitution of genes at different pathway steps, which would be common if each step evolved independently of the others, appears not to occur in the longest pathways.
These results are consistent with theoretical studies that suggest that more complex systems may show slower rates of evolutionary adaptation. Pathways with a high number of distinct steps can be considered more complex than pathways with fewer steps (McShea 1996). Fisher (1930) first used a simple geometric model to suggest that overall rates of adaptive change might be lower in more complex systems. In his model, the chance that a mutation of a given size will be favorable decreases as complexity increases. Orr (2000) described a refinement of this model and found that organisms pay a “cost of complexity” even larger than Fisher’s analysis suggested. Although this theoretical background is derived from simplified descriptions of evolutionary pathways (Barton and Partridge 2000), our data support its prediction that changes in complex biosynthetic pathways are less likely than changes in simple pathways due to increased constraint on complex pathways. While we have shown that one aspect of complexity, i.e., number of parts, negatively affects rates of change, there are also other factors that influence lability in metabolic systems. For example, Fraser et al. (2002) found that the number of interactions in a protein interaction network is negatively correlated with the substitution rate in those proteins. Their finding, together with the results presented here, suggests that in pathway evolution multiple aspects of complexity, in both number of components and connectivity among those components, play a role in determining the rate of evolution.
Our approach is simplifying in several respects. Our model of distance between character states assumes that the loss of enzymes from pathways is equally probable to the gain of enzymes in a pathway, but the validity of this assumption is difficult to assess in the macroevolutionary context of our study. While loss-of-function mutations are more common than gain-of-function mutations, it is less clear how often a loss of a step in a pathway occurs without loss of function of the entire pathway. Thus, we have no prior expectation that either loss or gain of nodes in the metabolic network is more probable across our studied phylogeny. In addition, our analysis does not incorporate information about genetic distances between species, and addition of these data in future analyses may provide a more nuanced description of the evolutionary lability of these pathways. Finally, because we examined changes in existing pathways, we may be neglecting important evolutionary trends in the loss or gain of entire biosynthetic pathways. For example, most of the “essential” ammo acids (those not synthesized by humans) are produced via highly complex pathways with little variability in structure across the organisms that produce them (Fig. 3). This suggests that pathways that are most constrained in their structure are also most likely to be lost completely. Furthermore, many pathway genes are organized as operons, and thus entire pathways can be transferred across organisms (Goldman and Kranz 1998; Wilding et al. 2000; Zuniga et al. 2002). However, we did not detect evidence in most pathways for a high character switching index, which would be consistent with sporadic horizontal transfer of entire pathways. Finally, it will be of particular interest to determine if the pattern of molecular evolution within the enzymes of these pathways corresponds to change in overall structure, i.e., if rates of evolutionary change of genes from longer pathways are slower than those of genes from shorter pathways. These further analyses could determine whether the increased rate of change in shorter pathways is adaptive or neutral.
References
R Alves RAG Chaleil MJE Sternberg (2002) ArticleTitleEvolution of enzymes in metabolism: a network perspective. J Mol Biol 320 751–770 Occurrence Handle10.1016/S0022-2836(02)00546-6 Occurrence Handle1:CAS:528:DC%2BD38XltVShtLk%3D Occurrence Handle12095253
GC Barrett DT Elmore (1998) Amino acids and peptides. Cambridge University Press New York
N Barton L Partridge (2000) ArticleTitleLimits to natural selection. BioEssays 22 1075–1084 Occurrence Handle10.1002/1521-1878(200012)22:12<1075::AID-BIES5>3.0.CO;2-M Occurrence Handle1:CAS:528:DC%2BD3MXmslWmtbs%3D Occurrence Handle11084623
DA Bender (1985) Amino acid metabolism. John Wiley & Sons New York
GEP Box DR Cox (1964) ArticleTitleAn analysis of transformations. J Roy Stat Soc Ser B 26 211–243 Occurrence Handle1:STN:280:BiqC283ptFU%3D Occurrence Handle3982271
AF Chalker HW Minehart NJ Hughes KK Koretke MA Lonetto KK Brinkman PV Warren A Lupas JR Stanhope, Brown PS Hoffman (2001) ArticleTitleSystematic identification of selective essential genes in Helicobacter pylori by genome prioritization and allelic replacement mutagenesis. J Bacteriol 183 1259–1268 Occurrence Handle10.1128/JB.183.4.1259-1268.2001 Occurrence Handle1:CAS:528:DC%2BD3MXhtVGmsbw%3D Occurrence Handle11157938
A Cornish-Bowden ML Cardenas (2002) ArticleTitleSystems biology: Metabolic balance sheets. Nature 420 129–130 Occurrence Handle10.1038/420129a Occurrence Handle1:CAS:528:DC%2BD38Xos1Ggu7c%3D Occurrence Handle12432369
ME Feder NV Cartano L Milos RA Krebs SL Lindquist (1996) ArticleTitleEffect of engineering Hsp 70 copy number on Hsp 70 expression and tolerance of ecologically relevant heat shock in larvae and pupae of Drosophila melanogaster. J Exp Biol 199 1837–1844 Occurrence Handle1:CAS:528:DyaK28XlsVCktL8%3D Occurrence Handle8708583
RA Fisher (1930) The genetical theory of natural selection. Oxford University Press Oxford
HB Fraser AE Hirch LM Steinmetz C Scharfe MW Feldman (2002) ArticleTitleEvolutionary rate in the protein interaction network. Science 296 750–752 Occurrence Handle10.1126/science.1068696 Occurrence Handle1:CAS:528:DC%2BD38XjtlOntrc%3D Occurrence Handle11976460
CV Forst K Schulten (2001) ArticleTitlePhylogenetic analysis of metabolic pathways. J Mol Evol 52 471–489 Occurrence Handle1:CAS:528:DC%2BD3MXksFOitbc%3D Occurrence Handle11443351
BS Goldman RG Kranz (1998) ArticleTitleEvolution and horizontal transfer of an entire biosynthetic pathway for cytochrome c biogenesis: Helicobacter, Deinococcus, Archae and more. Mol Microbiol 27 871–873 Occurrence Handle10.1046/j.1365-2958.1998.00708.x Occurrence Handle1:CAS:528:DyaK1cXhsVGit7k%3D Occurrence Handle9515711
ZL Gu LM Steinmetz X Gu C Scharfe RW Davis WH Li (2003) ArticleTitleRole of duplicate genes in genetic robustness against null mutations. Nature 421 63–66 Occurrence Handle10.1038/nature01198 Occurrence Handle1:CAS:528:DC%2BD3sXos1M%3D Occurrence Handle12511954
KM Herrmann RL Somerville (1983) Amino acids: Biosynthesis and genetic regulation. Addison–Wesley Reading, MA
MA Huynen T Dandekar P Bork (1999) ArticleTitleVariation and evolution of the citric acid cycle: a genomic perspective. Trends Microbiol 7 281–291 Occurrence Handle10.1016/S0966-842X(99)01539-5 Occurrence Handle1:STN:280:DyaK1Mzit1eisg%3D%3D Occurrence Handle10390638
M Kanehisa S Goto S Kawashima A Nakaya (2002) ArticleTitleThe KEGG databases at GenomeNet. Nucleic Acids Res 30 42–46 Occurrence Handle10.1093/nar/30.1.42 Occurrence Handle1:CAS:528:DC%2BD38Xht12ksbc%3D Occurrence Handle11752249
PD Karp M Riley S Paley A Pellegrini-Toole (2002) ArticleTitleThe MetaCyc Database. Nucleic Acids Res 30 IssueID1 59 Occurrence Handle1:CAS:528:DC%2BD38Xht12ktro%3D Occurrence Handle11752254
G Kolesov HW Mewes D Frishman (2001) ArticleTitleSNAPping up functionally related genes based on context information: a colinearity-free approach. J Mol Biol 311 639–656 Occurrence Handle10.1006/jmbi.2001.4701 Occurrence Handle1:CAS:528:DC%2BD3MXmtFGgtrk%3D Occurrence Handle11518521
J Li S Saxena D Pain A Dancis (2001) ArticleTitleAdrenodoxin reductase homolog (Arhlp) of yeast mitochondria required for iron homeostasis. J Biol Chem 276 1503–1509 Occurrence Handle1:CAS:528:DC%2BD3MXmtV2gtw%3D%3D Occurrence Handle11035018
WP Maddison DR Maddison (2000) MacClade: Analysis of phylogeny and character evolution, version 4.0. Sinauer Associates Sunderland, MA
DW McShea (1996) ArticleTitleMetazoan complexity: Is there a trend? Perspect Evol 50 477–492
HA Orr (2000) ArticleTitleAdaptation and the cost of complexity. Evolution 54 13–20 Occurrence Handle1:STN:280:DC%2BD3cvjsFOmuw%3D%3D Occurrence Handle10937178
R Overbeek N Larsen T Walunas M D’Souza G Pusch E Selkov Jr. K Liolios V Joukov D Kaznadzey I Anderson A Bhattacharyya H Burd W Gardner P Hanke V Kapatral N Mikhailova O Vasieva A Osterman V Vonstein M Fonstein N Ivanova N Kyrpides (2003) ArticleTitleThe ERGO(TM) genome analysis and discovery system. Nucleic Acids Res 31 164–171 Occurrence Handle10.1093/nar/gkg148 Occurrence Handle1:CAS:528:DC%2BD3sXhvFSntrk%3D Occurrence Handle12519973
D Posada KA Crandall (1998) ArticleTitleMODELTEST: Testing the model of DNA substitution. Bioinformatics 14 817–818 Occurrence Handle10.1093/bioinformatics/14.9.817 Occurrence Handle1:CAS:528:DyaK1MXktlCltw%3D%3D Occurrence Handle9918953
MD Rausher RE Miller P Tiffin (1999) ArticleTitlePatterns of evolutionary rate variation among genes of the anthocyanin biosynthesis pathway. Mol Biol Evol 16 266–274 Occurrence Handle1:CAS:528:DyaK1MXht1elt7o%3D Occurrence Handle10028292
SCG Rison SA Teichmann JM Thornton (2002) ArticleTitleHomology, pathway distance, and chromosomal localization of the small molecule metabolism enzymes in Escherichia coli. J Mol Biol 318 911–932 Occurrence Handle10.1016/S0022-2836(02)00140-7 Occurrence Handle1:CAS:528:DC%2BD38Xks1Krsb4%3D Occurrence Handle12054833
InstitutionalAuthorNameSAS Institute (1990) SAS/STAT user’s guide, version, 6. SAS Institute Cary, NC
DL Swofford (1998) Paup* 4.0: Phylogentic analysis using parsimony (and other methods). Sinauer Associates Sunderland, MA
RL Tatusov AR Mushegian P Bork NP Brown WS Hayes M Borodovsky KE Rudd EV Koonin (1996) ArticleTitleMetabolism and evolution of Haemophilus influenzae deduced from a whole-genome comparison with Escherichia coli. Curr Biol 6 279–291 Occurrence Handle1:CAS:528:DyaK28XhvFymurs%3D Occurrence Handle8805245
JD Thompson TJ Gibson F Plewniak F Jeanmougin DG. Higgins (1997) ArticleTitleThe ClustalX windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24 4876–4882 Occurrence Handle10.1093/nar/25.24.4876
A Wagner (2001) ArticleTitleEstimating coarse gene network structure from large-scale gene perturbation data. Genome Res 12 309–315 Occurrence Handle10.1101/gr.193902
SR Whitt LM Wilson MI Tenaillon BS Gaut ES Buckler (2002) ArticleTitleGenetic diversity and selection in the maize starch pathway. Proc Natl Acad Sci USA 99 12959–12962 Occurrence Handle10.1073/pnas.202476999 Occurrence Handle1:CAS:528:DC%2BD38XnvFGht7Y%3D Occurrence Handle12244216
EI Wilding JR Brown AP Bryant AF Chalker DJ Holmes KA Ingraham S Iordanescu YS Chi M Rosenberg MN Gwynn (2000) ArticleTitleIdentification, evolution, and essentiality of the mevalonate pathway for isopentenyl diphosphate biosynthesis in gram-positive cocci. J Bacteriol 182 4319–4327 Occurrence Handle1:CAS:528:DC%2BD3cXlt1Sgs7c%3D Occurrence Handle10894743
J Wuyts Y Van de Peer T Winkelmans R De Wachter (2002) ArticleTitleThe European database on small subunit ribosomal RNA. Nucleic Acids Res 30 183–185 Occurrence Handle10.1093/nar/30.1.183 Occurrence Handle1:CAS:528:DC%2BD38Xht12ktbY%3D Occurrence Handle11752288
M Zuniga G Perez F Gonzalez-Candelas (2002) ArticleTitleEvolution of arginine deiminase (ADI) pathway genes. Mol Phylogenet Evol 25 429–444 Occurrence Handle10.1016/S1055-7903(02)00277-4 Occurrence Handle1:CAS:528:DC%2BD38Xosl2gtL0%3D Occurrence Handle12450748
Acknowledgements
We thank T. Oakley for invaluable advice and discussion and C. Fenster, A. Gibson, M, Hahn, D. McShea, C. Murren, M. Rausher, and the Duke University Population Biology group for technical assistance, helpful discussion, and comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rutter, M.T., Zufall, R.A. Pathway Length and Evolutionary Constraint in Amino Acid Biosynthesis . J Mol Evol 58, 218–224 (2004). https://doi.org/10.1007/s00239-003-2546-y
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00239-003-2546-y