Abstract
The genetic code is not random but instead is organized in such a way that single nucleotide substitutions are more likely to result in changes between similar amino acids. This fidelity, or error minimization, has been proposed to be an adaptation within the genetic code. Many models have been proposed to measure this adaptation within the genetic code. However, we find that none of these consider codon usage differences between species. Furthermore, use of different indices of amino acid physicochemical characteristics leads to different estimations of this adaptation within the code. In this study, we try to establish a more accurate model to address this problem. In our model, a weighting scheme is established for mistranslation biases of the three different codon positions, transition/transversion biases, and codon usage. Different indices of amino acids’ physicochemical characteristics are also considered. In contrast to pervious work, our results show that the natural genetic code is not fully optimized for error minimization. The genetic code, therefore, is not the most optimized one for error minimization, but one that balances between flexibility and fidelity for different species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Since the characterization of the entire genetic code in the 1960s, the internal order of the code, in which codons for biochemically similar amino acids are grouped together, has been noted (e.g., Woese 1965a). The genetic code appears to be organized in such a way that when single nucleotide changes result in amino acid substitutions, the new amino acids are likely to be similar to the old ones (Woese 1965a). This apparent organization, now called “translation error load minimization” in most of literature cited below, was proposed to be an adaptation via natural selection (Woese 1965b). It is now accepted that the organization of genetic code results in translation error minimization (Di Giulio 1989; Haig and Hurst 1991).
Researchers have measured the effect of the genetic code in error minimization by using the mean square method (Alff-Steinberger 1969). Different physicochemical characteristics of individual amino acids can be used in this method. In 1998, Freehand and Hurst (1998) developed a modified mean square measurement (WMS measurement), which added the weighting of mistranslation and of the transition/transversion bias across the three codon positions to the mean square measurement. Their work, however, did not consider the effect of codon usage. Different species have different patterns of codon usage, and thus the same genetic code may be more or less optimized in different species. Furthermore, their methodology ignores difference of base composition (Lehninger 1982) across species in their coding sequences. Here, we introduce a method for measuring error minimization within the genetic code that incorporates differences in codon usage across taxa. We call it “usage weighted mean square measurement (UWMS). The UWMS is a mean square measure which incorporates transition/transversion bias, mistranslation, and codon usage weight. UWMS0 corresponds to all possible single base changes in all codon positions and is the sum of UWMS1 (measures the first position of a triple), UWMS2 (the second), and UWMS3 (the third). Refer to Appendix A for the mathematic expression of UWMS.
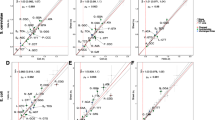
Using our model and codon usage of E. coli, we get the frequency distribution of the UWMS0 of randomly generated genetic codes shown in Fig. 1 and Tables 1 and 2 refer to Appendix B for the generation method and algorithm. Data for UWMS1, UWMS2, and UWMS3 not shown.
Frequency distribution for the UWMS0 values obtained from 4,000,000 randomly generated codes. The X-axis shows the range of UWMS0 values, and the Y-axis shows the possibilities of certain codes corresponding to a certain UWMS0 range. Most of the codes have medium adaptation. Codes that are poorly or highly optimized are both very scarce. Most of the codes are less conservative than the natural one. There are only a few codes more optimized for error minimization than the canonical code. In these calculations Woese’s (1966) “polar requirement” index is used, and the codon usage of E. coli is weighted.
We find that with inclusion of the codon usage of E. coli, the genetic code no longer shows as high an error minimization effect as reported earlier (Freehand and Hurst 1998). This means that codon usage actually decreases the error minimization of the genetic code. Different species have different codon usage, so the same genetic code is differently optimized for fidelity in different species (Table 3).
Further study shows that due to the similar code usage, the adaptation measurements are similar in closely related species (data not shown). It seems that the optimization measurement of the genetic code can reflect the genetic distance between different species on the level of the whole coding sequence. However, because different indices of amino acids will lead to different measurements, it is hard to interpret this correlation between level of optimization and genetic relatedness.
Originally, Woese (1965b) used “polar requirement” to measure the “distance” between amino acids. In Haig and Hurst’s work (1991), they also tested hydrophobicity, molecular volume, and isoelectric point. They found that the apparent error minimization in the genetic code is different when measured by different characteristics of amino acids. Miller et al. (1987) suggested that the hydrophobicity of amino acids is important in determining the protein three-dimensional structures. Following this idea, we introduce a new index of amino acid hydrophobicity based on Miller’s statistical data. The effect of different amino acid residues can be measured by the quotient of f(interior) to f(external), where f(interior) and f(external) represent the frequencies of a certain amino acid located inside and on the surface of globular proteins, respectively. In order to apply such an index to our UWMS measure, we modified it to ln[f(interior)/f(external)] to make the variation comparable to other indices.
We calculated several other frequently used indices of amino acid hydrophobicity (Table 4), finding that using different hydrophobicity indices can cause the measurements of error minimization to differ (Table 5).
Table 4 lists a set of most popular indices of amino acid hydrophobicity from the many such indices suggested previously. It is unclear which of these many indices is the best for measuring translation error minimization. Freeland and Hurst (1998) explained that they chose the polar requirement because it gives the most significant evidence of minimization of error. That is, using this hydrophobicity index, they found fewer codes “better” than the natural code than when using other hydrophobicity indices. However, this does not necessarily mean that this index is more accurate than others are. The polar requirement index results in a greater apparent level of optimization than using other indices at least partially because the polar requirement index underestimates the distance between Ile, Leu, Met, Val, Cys, Phe, Trp, and Tyr compared to other indices listed in Table 4. Kyte and Doolittle’s (1982) index is often used and is efficient in predicting transmembrane domains. However, some of the key values of the Kyte–Doolittle index were adjusted arbitrarily to meet prior expectations, limiting the application of this index to other types of protein domains.
Recent research shows that the genetic code is a product of selection for error minimization (Di Giulio 2000; Freeland 1998, 2000a, b; Knight 1999), and it is generally accepted that error minimization is adaptive. It is also accepted that the code is “frozen” or “fixed” for being adaptive. The strongest evidence of this selection and fixation event would be to show that genetic code is highly adaptive or even the “best” possible one. Many publications have tried to show this, however, these previous studies have ignored two important points.
First, studies have underestimated the number of codes “better” at error minimization than the natural genetic code. Freeland et al. (2000a) argued that overestimation of the number of “better” codes is due to the inaccuracy of amino acid polarity/hydrophobicity indices and overestimation of the total number of possible codes. When a correct amino acid index is chosen and the total number of possible codes is carefully estimated, other work shows that the canonical genetic code may be the “best” one (Freeland et al. 2000b). We agree with the argument that the total number of possible codes should be carefully estimated. However, as discussed above for Freeland and Hurst (1998), the effort to choose a correct amino acids index can be slightly subjective. Facing this problem, our suggestion is to using different indices to measure error minimization and compare the results, as done in this work.
Second, since codon usage can at least alter the error minimization effect, in order to know whether, and to what extent, the genetic code is optimized for error minimization, it is necessary to investigate the codon usage of individual species. In fact, in order to know whether the code is the “best” one and fixed for being the “best” in error minimization, it would be necessary to know the codon usage of the common cell ancestor 4 billion years ago. This is, of course, not possible. Therefore, it is difficult to demonstrate that the natural code is the “best” possible for error minimization, and consequently, it becomes hard to tell whether the code is “frozen” or “fixed” for fidelity.
Our results (Table 3) also indicate that because different species have different codon usages, the canonical genetic code is differently optimized for error minimization in different species. In fact, our result shows that when codon usage is considered, the code appears to be less optimized for our sampling species. This suggests that codon usage acts to increase the flexibility of the translational control beyond the genetic code. The same result also shows that there is a tendency for the genetic code to be more flexible in developmentally simple creatures than in complex creatures. All life forms today are believed to be descended from a single pool of primitive cells in which that genetic code was frozen (Crick 1968) from which the codon usage biases have diverged. With the evolution of more complex structures and ontogenesis processes, the usage may have been selected to increase the fidelity of the genetic code, in order to stabilize the protein system of species. However, if the complexity of species did not increase during the course of evolution, the usage may have been selected to increase the flexibility of the genetic code, leading to a higher evolution rate. These opposing forces drive the deviation of codon usage. Our work suggests that codon usage is a product of selection for flexibility in translation. The canonical genetic code is not the most optimized one for minimizing error but the one that reaches a balance between fidelity and flexibility. So, adaptation is a balance, rather than optimized for error minimization.
There are still some problems in the methodology. For example, when codon usages of different species are applied to the measurement, a major flaw is that transition/transversion biases also actually differ. Our measurement, like Freeland and Hurst’s (1998) work, uses a single proposed transition/transversion bias, since not all the biases were known, in the landscape of the genome. Further work should take the different transition/transversion biases, both within genome and among genomes, into account.
References
C Alff-Steinberger (1969) ArticleTitleThe genetic code and error transmission. Proc Natl Acad Sci USA 64 203–207
FHC Crick (1968) ArticleTitleThe origin of genetic code. J Mol Biol 38 367–293 Occurrence Handle1:CAS:528:DyaF1MXksVSiug%3D%3D Occurrence Handle4887876
M Di Giulio (1989) ArticleTitleThe extension reached by minimization of the polarity distances during the evolution of the genetic code. J Mol Evol 29 288–293 Occurrence Handle1:CAS:528:DyaL1MXmt1Ogurc%3D Occurrence Handle2514270
M Di Giulio (2000) ArticleTitleGenetic code origin and the strength of natural selection. J Theoret Biol 205 659–661 Occurrence Handle10.1006/jtbi.2000.2115 Occurrence Handle1:CAS:528:DC%2BD3cXlsVSrsrw%3D
DM Engelman TA Steitz A Goldman (1986) ArticleTitleIdentifying nonpolar transbilayer helices in amino acid sequences of membrane proteins. Annu Rev Biophys Biophys Chem 15 321–353
SJ Freeland LD Hurst (1998) ArticleTitleThe genetic code is one in a million. J Mol Evol 47 238–248 Occurrence Handle1:CAS:528:DyaK1cXmt1ansrs%3D Occurrence Handle9732450
SJ Freeland RD Knight LF Landweber (2000a) ArticleTitleMeasuring the adaptation within the genetic code. Trends Biochem Sci 25 44–45 Occurrence Handle1:CAS:528:DC%2BD3cXhsFejtr4%3D
SJ Freeland et al. (2000b) ArticleTitleEarly fixation of an optimal genetic code. Mol Biol Evol 17 511–518 Occurrence Handle1:CAS:528:DC%2BD3cXisVSgt70%3D
D Haig LD Hurst (1991) ArticleTitleA quantitative measure of error minimization in the genetic code. J Mol Evol 33 412–417 Occurrence Handle1:CAS:528:DyaK3MXmsleqtrs%3D Occurrence Handle1960738
RD Knight SJ Freeland LF Landweber (1999) ArticleTitleSelection, history and chemistry: The three faces of the genetic code. Trends Biochem Sci 24 241–247 Occurrence Handle10.1016/S0968-0004(99)01392-4 Occurrence Handle1:CAS:528:DyaK1MXks1ansLg%3D Occurrence Handle10366854
J Kyte RF Doolittle (1982) ArticleTitleA simple method for displaying the hydropathic character of a protein. J Mol Biol 157 105–132 Occurrence Handle1:CAS:528:DyaL38Xks1yjtro%3D Occurrence Handle7108955
AL Lehninger (1982) Principles of biochemistry. Worth New York
S Miller J Janin AM Lesk C Chothia (1987) ArticleTitleInterior and surface of monomeric proteins. J Mol Biol 196 641–656 Occurrence Handle1:CAS:528:DyaL1cXitlWh Occurrence Handle3681970
Y Nakamura T Gojobori T Ikemura (1999) ArticleTitleCodon usage tabulated from the international DNA sequence databases; Its status 1999. Nucleic Acids Res 27 292–292 Occurrence Handle1:CAS:528:DyaK1MXpsVKgtw%3D%3D Occurrence Handle9847205
Y Nozaki C Tanford (1971) ArticleTitleThe solubility of amino acids and two glycine peptides in aqueous ethanol and dioxane solutions. Establishments of a hydrophobicity scale. J Mol Chem 246 2111
CR Woese (1965a) ArticleTitleOrder in the genetic code. Proc Natl Acad Sci USA 54 71–75 Occurrence Handle1:CAS:528:DyaF2MXks1Cmsrk%3D
CR Woese (1965b) ArticleTitleOn the evolution of the genetic code. Proc Natl Acad Sci USA 54 1546–1552 Occurrence Handle1:CAS:528:DyaF28Xltlagsg%3D%3D
CR Woese DH Dugre SA Dugre M Kondo WC Saxinger (1966) ArticleTitleOn the fundamental nature and evolution of the genetic code. Cold Spring Habor Symp Quant Biol 31 723–736 Occurrence Handle1:CAS:528:DyaF2sXks1Srtrw%3D
Acknowledgements
The authors sincerely appreciate Zhen Yao and Zhi-Hong Zhang for instructive advice on programming. In addition, we thank Prof. Yang Zhong, Dr. Li-Ying Cui, and Dr. Thomas Merritt for helpful discussions. We are also very gratitude to anonymous referees for very informative comments on the manuscript. C.-T. Zhu (Zhu Lei) is supported by a Chun-Tsung fellowship at Fudan University endowed by Tsung-Dao Lee, 1957 Nobel Prize laureate in physics.
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix A
Let us, for convenience in describing the sums performed, refer to four bases by number; U = 1, C = 2, A = 3, and G = 4. Let the characteristic Q be specified by a corresponding base triple (L, M, N) of the amino acid. T is the quantification of combined weighting of mistranslation, transition, and transversion. U (L, M, N) is the codon usage of the triple (L, M, N). UWMS measurement can be defined as follows (stop codons are excluded).

Codon usage of a certain species is calculated from all the coding sequences of that species reported to GenBank. The codon usage of each codon is measured by the relative frequency of the count of that codon versus the count of all codons. Codon usage of a certain species is consequently a vector of 61 variables.
Codon usage data were obtained from the Codon Usage Database (ftp://www.kazusa.or.jp/ ) in August 1999 (Nakamura 1999).
Appendix B
A Delphi 3.0 program was written for calculation. The random codes have the same degenerate pattern as the natural code, only the positions of amino acids were changed. Four million codes are generated using a cycle function; in one cycle, 4000 codes are generated and calculated. In every cycle, the 4000 codes are all different. Although a code may be generated and calculated more than once in 1000 cycles, in such a big population, sampling with or without replacement has the same meaning.
The program and source code are available on request to the authors.
Rights and permissions
About this article
Cite this article
Zhu, CT., Zeng, XB. & Huang, WD. Codon Usage Decreases the Error Minimization Within the Genetic Code . J Mol Evol 57, 533–537 (2003). https://doi.org/10.1007/s00239-003-2505-7
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00239-003-2505-7