Abstract
Rhodopsin is the light receptor in photoreceptor cells of the retina and a prototypical G protein-coupled receptor. Two types of quaternary structures can be adopted by rhodopsin. If rhodopsin folds and attains a proper tertiary structure, it can then form oligomers and nanodomains within the photoreceptor cell membrane. In contrast, if rhodopsin misfolds, it cannot progress through the biosynthetic pathway and instead will form aggregates that can cause retinal degenerative disease. In this review, emerging views are highlighted on the supramolecular organization of rhodopsin within the membrane of photoreceptor cells and the aggregation of rhodopsin that can lead to retinal degeneration.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Vision is initiated in photoreceptor cells, which are located in the outer retina. The initiation of vision occurs when light activates rhodopsin or cone opsins in photoreceptor cells, initiating a prototypical G protein-mediated signaling cascade called phototransduction. Rod photoreceptor cells are responsible for scotopic vision and contain rhodopsin whereas cone photoreceptor cells are responsible for photopic vision and contain cone opsins. Rod photoreceptor cell biology and rhodopsin’s structure and function have been characterized more extensively compared to cone photoreceptor cell biology and cone opsin structure and function, in part, because of the abundance of rod photoreceptor cells in most vertebrate retina. The focus here will be on the quaternary structures formed by rhodopsin.
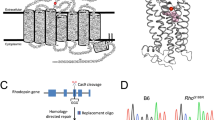
Rhodopsin is a prototypical G protein-coupled receptor (GPCR), exhibiting a characteristic 7 α-helical transmembrane architecture. Rhodopsin is a two-component system that includes the apoprotein opsin and the covalently bound chromophore 11-cis retinal. Light causes the isomerization of 11-cis retinal to all-trans retinal, thereby activating rhodopsin and initiating phototransduction (Park 2014). The biosynthesis and activity of rhodopsin occur in distinct compartments of the rod photoreceptor cell. Photoreceptor cells are compartmentalized into an inner segment and an outer segment (Fig. 1). The biosynthetic machinery is in the rod inner segment. Rhodopsin must be synthesized and adopt a proper three-dimensional structure to bypass quality-control mechanisms in the endoplasmic reticulum (ER). If rhodopsin misfolds, it must be discarded or aggregation can occur leading to dysfunction. Several mutations in rhodopsin cause misfolding and aggregation of the receptor. Emerging views on rhodopsin aggregation will be presented here.

(This figure is reprinted from (Rakshit et al. 2017), with permission from Elsevier) (Color figure online)
Cartoon of a rod photoreceptor cell. Rod photoreceptor cells contain a rod outer segment and rod inner segment. The rod outer segment contains stacks of membranous disks, which are shown in a side view and top view. Rhodopsin (red) is densely packed within the membrane of the disks, forming nanodomains of oligomeric receptor
After passing quality control, rhodopsin is transported to the base of the connecting cilium, which separates the inner and outer segments, and then transported across the connecting cilium into the rod outer segment (ROS) (Goldberg et al. 2016; Insinna and Besharse 2008; Nemet et al. 2015; Sung and Chuang 2010; Wang and Deretic 2014; Wensel et al. 2016). The ROS contains a structured system of membranes containing stacks of disks, which are double lamellar membranes connect by a rim, that are encased by a plasma membrane (Gilliam et al. 2012; Nickell et al. 2007). Rhodopsin is incorporated into disks at the base of the ROS (Ding et al. 2015; Volland et al. 2015). Rhodopsin is densely packed into the membrane of the disks. Emerging views on rhodopsin supramolecular organization within the disk membrane will be presented here.
Rhodopsin Oligomers and Nanodomains
Rhodopsin is unique among GPCRs in that it is densely packed within its native membrane. The average density of rhodopsin is estimated to be about 20,000 molecules/μm2, which is orders of magnitude higher than that of other GPCRs, with estimates ranging from 5 to 30 molecules/μm2 (Hegener et al. 2004; Herrick-Davis et al. 2015). This high density of rhodopsin within the membrane can be both beneficial and detrimental in facilitating the exquisite sensitivity exhibited by rod photoreceptor cells, which can respond to a single photon of light (Baylor et al. 1979). The high density of rhodopsin can maximize the probability of photon capture, such as it occurs in thylakoid membranes of plants, green algae, and cyanobacteria, which are densely packed with photosynthetic membrane proteins to maximize absorption of sunlight (Kirchhoff 2014). While the high density of rhodopsin is advantageous from the perspective of photon capture probability, G protein signaling is a diffusion-mediated process, which would be adversely affected by a crowded membrane environment densely packed with the receptor. This dichotomy raises an important question: how do you achieve efficient and sensitive signaling within a crowded membrane environment? A general solution nature has designed for this problem is to impart order within biological membranes (Bethani et al. 2010; Mugler et al. 2013; Radhakrishnan et al. 2012).
What kind of order is present within ROS disk membranes? Historically, rhodopsin was envisioned to be present within the disk membrane as randomly dispersed monomers (Chabre et al. 2003; Chabre and le Maire 2005). More recent studies by atomic force microscopy (AFM) and cryo-electron microscopy (cryo-EM) paint a different picture involving order within the membrane (Gunkel et al. 2015; Liang et al. 2003; Rakshit et al. 2015; Whited and Park 2015). Order within the membrane is achieved by oligomeric rhodopsin, arranged as rows of dimers, dispersed within the membrane forming nanodomains (Fig. 1). This type of order is predicted to overcome the impedance imparted by densely packed rhodopsin and facilitate efficient and sensitive signaling (Cangiano and Dell’Orco 2013; Dell’Orco 2013; Dell’Orco and Schmidt 2008; Gunkel et al. 2015; Schoneberg et al. 2014).
Investigating the organization of rhodopsin within native photoreceptor cell membranes is a challenge, and there are few available methods amenable for such pursuits. AFM has particularly been useful for visualizing membrane proteins under physiologically relevant conditions (Muller 2008; Whited and Park 2014), and a recently developed method utilizing AFM has provided insights to advance our understanding about the packing of rhodopsin in ROS disk membranes (Senapati and Park 2019). Although there are caveats to observations made by AFM, as there are with any method, a similar arrangement of rhodopsin within ROS disk membranes is observed by both AFM and cryo-EM, indicating that observations of oligomeric rhodopsin forming nanodomains is method independent. Moreover, artifacts related to phase separation of lipids due to low sample preparation temperatures, adsorption of samples on a mica substrate, and lateral forces imparted by the AFM tip have been ruled out (Rakshit and Park 2015; Rakshit et al. 2015). Thus, AFM provides a unique window to observe the native organization of rhodopsin within photoreceptor cell membranes. Some of the insights gained by AFM are discussed here.
Rod photoreceptor cells contain 500–2000 stacked disks within a single ROS, depending on the species (Daemen 1973). Although disks within a ROS are often presumed to be identical to each other when describing their properties, they are in fact quite heterogeneous. Different amounts of rhodopsin can be packed into a disk (Haeri et al. 2013; Hsu et al. 2015; Organisciak and Noell 1977; Penn and Anderson 1987), the lipid composition of membranes can be different among disks (Albert et al. 1998; Andrews and Cohen 1979; Boesze-Battaglia et al. 1990; Caldwell and McLaughlin 1985), and the functional properties of the disks can be heterogeneous (Baylor and Lamb 1982; Makino et al. 1990; Mazzolini et al. 2015; Williams and Penn 1985; Young and Albert 2000). It is not surprising then that properties of the disks visualized by AFM are heterogeneous as well. Heterogeneity has been observed in the size of disks and the size, number, and density of rhodopsin nanodomains (Rakshit and Park 2015; Whited and Park 2015). This heterogeneity points to plasticity in the ROS.
Photoreceptor cells must adapt to the environment of the organism for optimal function and survival of the organism. Adaptations occurring at the level of individual disks have not been studied in much detail. An optimal density of rhodopsin for signaling within ROS disk membranes has been previously proposed (Saxton and Owicki 1989). The packing density of about 20,000 molecules/μm2 for rhodopsin observed by AFM appears to be optimal for rod photoreceptor cells, at least under normal lighting conditions. Photoreceptor cells appear to adapt to different numbers of rhodopsin incorporated into disks by modulating the size of the disks to maintain an average rhodopsin density of 20,000 molecules/μm2 (Whited and Park 2015). This packing density is even maintained under conditions where the amount of rhodopsin expressed in photoreceptor cells is reduced by half (Rakshit and Park 2015). Although this packing density of rhodopsin may be optimal under normal conditions, it can be modulated by changes in the environmental lighting condition. Under conditions where animals are housed under constant light or constant dark conditions, the density of rhodopsin decreases or increases, respectively (Rakshit et al. 2017). These adaptations in photoreceptor cells are dependent on the signal from phototransduction and the changes in rhodopsin density within the disk membrane can impact visual function as assessed by electroretinography (Rakshit et al. 2017; Senapati et al. 2018).
The concentration of rhodopsin within the membrane can impact the complement of oligomeric forms that are present within the membrane. Studies on heterologously expressed rhodopsin in cultured cells have demonstrated the existence of an equilibrium of oligomeric forms of rhodopsin (Fig. 2a). A monomer–dimer equilibrium was detected by pulsed-interleaved excitation fluorescence cross-correlation spectroscopy (PIE-FCCS) and a dimer–tetramer equilibrium was detected using Förster resonance energy transfer (FRET) spectrometry (Comar et al. 2014; Mishra et al. 2016). An equilibrium of oligomeric forms has also been detected for other GPCRs as well (Calebiro et al. 2013; Hern et al. 2010; Kasai et al. 2011; Patowary et al. 2013; Stoneman et al. 2017; Ward et al. 2015, 2017), thereby suggesting that oligomerization of all GPCRs can be described by schemes based on chemical equilibria (Fig. 2a). Within such a scheme, the equilibrium constants and concentration of the receptor will dictate the complement of oligomeric forms of the receptor present in the membrane.
Equilibrium of different oligomeric forms of rhodopsin. a A general schematic showing a chemical equilibrium among differently sized oligomers (denoted by the superscript) of a receptor (R). The equilibrium dissociation constants determined for rhodopsin are denoted (Mishra et al. 2016). b–e A summary of analysis of AFM images of ROS disk membranes is shown. Data from mice housed under cyclic light, 10 days constant dark, or 10 days constant light conditions are shown in b, c, and data from mice housed under 10, 20, or 30 days constant dark conditions are shown in d, e. Histograms of nanodomain sizes (b, d) and rhodopsin density within the membrane (c, e) are shown. The nanodomain size reflects the size of the rhodopsin oligomer. The histograms are reproduced from (Rakshit et al. 2017), with permission from Elsevier. f The predominant oligomeric species of rhodopsin in photoreceptor membranes is a 24-mer. The changes in the complement of rhodopsin nanodomains/oligomers illustrated in b, d can be described in terms of a shift in equilibrium between a 24-mer- and larger-sized oligomers (Color figure online)
The oligomerization of GPCRs is most often investigated in heterologous expression systems, where the receptors are considered to be overexpressed relative to their native expression levels. In the case of rhodopsin, this is not the case. Heterologous expression levels in cells like Chinese hamster ovary (CHO) cells is far below the native expression levels of rhodopsin in photoreceptor cells in the retina (1350 molecules/μm2 versus 20,000 molecules/μm2, on average) (Mishra et al. 2016). The difference in concentration of rhodopsin in heterologous expression systems and native photoreceptor cells can explain, at least in part, the difference in oligomeric forms observed in the two systems. Heterologous expression in CHO cells results in a mixture of dimers and tetramers, with higher order oligomers becoming detectable at concentrations of rhodopsin greater than 1150 molecules/μm2 (Mishra et al. 2016). The size of oligomers in native photoreceptor cell membranes can be estimated from the size of rhodopsin nanodomains detected by AFM (Liang et al. 2003; Senapati and Park 2019). The size of rhodopsin nanodomains/oligomers within ROS disk membranes is heterogeneous, exhibiting a skewed distribution (Fig. 2b, d). The predominant oligomeric species has a size of 335 nm2 (Rakshit and Park 2015), which corresponds to an oligomer with 24 rhodopsin molecules (i.e., 24-mer). These observations by AFM are consistent with observations in heterologous expression systems suggesting that there is an equilibrium of oligomeric forms of rhodopsin.
If an equilibrium of oligomeric forms of rhodopsin were present in photoreceptor cell membranes, then the complement of oligomeric forms present within the membrane should be adjustable by changing the concentration of the receptor within the membrane or any factor that alters the equilibrium constant. Evidence for both concentration-dependent and equilibrium constant-dependent changes in the oligomeric status of rhodopsin have been demonstrated in photoreceptor cells (Rakshit et al. 2017). Mice housed in constant light for 10 days have lower concentrations of rhodopsin in the membrane compared to mice housed in constant darkness for 10 days, and mice housed under normal cyclic lighting conditions have intermediary concentrations of rhodopsin (Fig. 2c). Examining histograms of the size of rhodopsin nanodomains/oligomers present in photoreceptor cell membranes from each of these mice reveals that all exhibit a skewed distribution with the predominant oligomeric species corresponding to a 24-mer (Fig. 2b). There is, however, a difference in the proportion of the 24-mer (main peak in histogram) and larger sized oligomers (shown in inset). A lower concentration of rhodopsin (10 days constant light) results in a higher level of the 24-mer and lower level of larger oligomers compared to that at higher concentrations of rhodopsin (10 days constant dark). Thus, lower concentrations of rhodopsin in the membrane shifts the equilibrium more in favor of the 24-mer and higher concentrations of rhodopsin in the membrane shifts the equilibrium more in favor of the larger oligomers (Fig. 2f).
The equilibrium between the 24-mer and larger oligomers can still be shifted even when the concentration of rhodopsin is similar. Mice housed for 10 days, 20 days, or 30 days in constant darkness all exhibit similarly elevated concentrations of rhodopsin in the membrane compared to mice housed under normal cyclic lighting conditions (Fig. 2e). Yet, the complement of oligomeric forms is different in mice housed for 10 days in constant darkness versus mice housed for longer periods in complete darkness. The equilibrium appears to be shifted more in favor of the 24-mer than the larger oligomers for mice housed for longer periods of constant darkness compared to that for mice housed for 10 days in constant darkness (Fig. 2d, f). Thus, in this instance, it is not the concentration responsible for shifting the equilibrium but, rather, some factor that appears to have changed the equilibrium constant. It is unclear what this factor can be at the moment. An obvious source of change may be the lipids in the membrane.
Lipids in the membrane appear to play a role in driving oligomerization or stabilizing the oligomeric complex (Botelho et al. 2006; Jastrzebska et al. 2006; Periole et al. 2007; Soubias et al. 2015). It is unclear, however, which membrane lipids can alter the equilibrium constants. Docosahexaenoic acid (DHA) is highly abundant in ROS disk membranes and can affect the structure and function of rhodopsin (Boesze-Battaglia and Albert 1989; Brown 1994; Bush et al. 1994; Grossfield et al. 2006; Mitchell et al. 2001; Niu et al. 2001, 2004; Wiedmann et al. 1988). Although modulation of DHA in the membrane of photoreceptor cells of mice impacts function and leads to some adaptations in ROS disk membranes, the complement of oligomers in the membrane is not significantly altered (Senapati et al. 2018). Further studies are required to better understand the interplay of specific lipids and oligomerization, and determining what factors can change the equilibrium constants that underlie rhodopsin oligomerization.
Misfolding and Aggregation of Rhodopsin
GPCRs are highly hydrophobic proteins and must adopt a proper tertiary structure to avoid the fate of misfolding and aggregation. Heritable mutations in GPCRs can cause receptor misfolding and a range of human disease (Tao and Conn 2014). The rhodopsin gene is a hot spot for inherited mutations causing retinal disease (Mendes et al. 2005; Nathans et al. 1992; Stojanovic and Hwa 2002). There are over 100 reported mutations in the rhodopsin gene in patients with inherited retinal disease, most of them causing retinitis pigmentosa (RP)—a progressive retinal degenerative disease. Over half of the mutations in rhodopsin with known biochemical defect result in receptor misfolding and aggregation, which lead to autosomal dominant RP (adRP) (Athanasiou et al. 2018). Rhodopsin mutations are the largest cause of adRP (Dalke and Graw 2005; Hartong et al. 2006).
Rhodopsin synthesis requires a network of coordinated processes in the ER that maintains protein homeostasis or proteostasis (Athanasiou et al. 2013; Gorbatyuk and Gorbatyuk 2013; Griciuc et al. 2011; Kroeger et al. 2012). These processes include a chaperone system that aids in the proper folding of rhodopsin and a quality-control system that eliminates improperly folded proteins via the ubiquitin–proteasome system or autophagy. Stresses on the ER perturbing proteostasis activate the unfolded protein response, which restores proteostasis in the short term but can lead to photoreceptor cell death in the long term if chronically activated (Walter and Ron 2011). The precise mechanisms that lead to photoreceptor cell death in retinal disease are still being uncovered (Adekeye et al. 2014; Arango-Gonzalez et al. 2014; Chiang et al. 2014; Sizova et al. 2014). Misfolded rhodopsin forms nonnative oligomers (i.e., aggregates), which can disrupt proteostasis and cause photoreceptor cell death (Bence et al. 2001; Illing et al. 2002). Inhibiting rhodopsin aggregation appears to reduce retinal degeneration (Athanasiou et al. 2012; Gorbatyuk et al. 2010; Parfitt et al. 2014; Vasireddy et al. 2011), establishing a direct link between aggregation and retinal degeneration. The mechanism by which aggregates cause cell toxicity is still unclear. Little is known about the process of rhodopsin misfolding and the nature of aggregates formed.
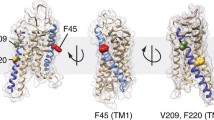
Mutations in rhodopsin that cause misfolding and aggregation are present within the transmembrane helices and extracellular surface of the receptor (Fig. 3). Since the discovery of the first mutation in rhodopsin causing adRP, the P23H mutation (Dryja et al. 1990), misfolding mutants of rhodopsin have been characterized biochemically and by microscopy (Garriga et al. 1996; Hwa et al. 1997; Kaushal and Khorana 1994; Sung et al. 1991a, b). Based on these characterizations, misfolding mutants of rhodopsin have been classified as either complete or partial misfolding mutants (Kaushal and Khorana 1994; Krebs et al. 2010; Sung et al. 1991a; 1993). Complete misfolding mutants cannot bind or be rescued by 11-cis retinal and are predominantly retained in the ER rather than trafficking correctly to the plasma membrane. Partial misfolding mutants can variably bind and be rescued by 11-cis retinal and are variably retained in the ER. The complete misfolding mutations tend to be located in close proximity to the binding pocket for 11-cis retinal whereas partial misfolding mutations are more distal (Fig. 3b).

(The figure is reprinted from (Gragg and Park 2019), with permission from Elsevier) (Color figure online)
Mutations in rhodopsin that cause misfolding and aggregation. Partial (blue) and complete (yellow) misfolding mutations in rhodopsin are illustrated on the secondary structure (a) and tertiary structure (b) of rhodopsin. Mutation of proline at position 267 (green) can be either partial or complete, depending on the specific mutation. The chromophore 11-cis retinal is shown as pink spheres
The biochemical and microscopy-based methods used to classify rhodopsin mutants as complete and partial misfolding mutants do not directly assess the aggregation of the misfolded rhodopsin mutants. Aggregation of misfolding rhodopsin mutants have, in large part, been assessed by light microscopy [e.g., (Illing et al. 2002; Saliba et al. 2002)]. The types of aggregates examined by light microscopy are inclusion bodies or aggresomes (Kopito 2000), which are relatively large structures identifiable by light microscopy. These types of structures, however, are not detected in knockin mouse models of adRP expressing the P23H mutant of rhodopsin (Price et al. 2011; Sakami et al. 2011). A recently developed method utilizing FRET has detected aggregates of misfolded rhodopsin that appear diffusely dispersed in the ER and are indistinguishable from the wild-type receptor (Gragg and Park 2019; Miller et al. 2015). Thus, the pathogenic aggregates in adRP may be smaller aggregates not discernible by light microscopy rather than larger inclusion bodies or aggresomes. The discussion here will focus on these smaller aggregates.
Misfolded rhodopsin mutants are retained in the ER; however, retention of rhodopsin in the ER itself does not cause aggregation (Miller et al. 2015). The receptor must be misfolded to induce aggregation. The biosynthesis of wild-type rhodopsin results in mostly properly folded rhodopsin with little or no misfolded protein because of proteasomal degradation. When the proteasome is inhibited, the misfolded wild-type rhodopsin that would have been degraded persists and then forms aggregates (Gragg et al. 2016). Secondary structure changes are associated with misfolded rhodopsin mutants. Rhodopsin has a high α-helical content because of its 7 α-helical transmembrane architecture. In misfolding mutants of rhodopsin, this α-helical content is not completely abolished. Only a partial reduction of α-helical structure occurs that is accompanied by an increase in β-sheet structure (Liu et al. 1996; Miller et al. 2015). Interestingly, increased β-sheet structure is a hallmark of amyloid aggregates, and this secondary structure mediates the protein–protein interactions within the aggregates (Chiti and Dobson 2006; Knowles et al. 2014). Although further studies are required to determine the role that β-sheets have in the aggregation of misfolded rhodopsin mutants, the formation of this secondary structure raises the possibility that they may contribute to the formation of aggregates and that there is specificity in the aggregation process.
A couple observations suggest that the aggregation of rhodopsin does not occur randomly in a nonspecific manner, but rather, is mediated by specific protein–protein interactions. Misfolded rhodopsin does not aggregate with other proteins known to aggregate, including an unrelated misfolded membrane protein (Rajan et al. 2001). Moreover, misfolded rhodopsin mutants do not aggregate with properly folded wild-type rhodopsin (Gragg et al. 2016; Gragg and Park 2018). Thus, aggregation appears to be mediated by specific interactions, likely requiring the β-sheet structures that form in misfolded rhodopsin. The nature of the protein–protein interactions that underlie the formation of aggregates differs from those of oligomers formed by properly folded rhodopsin. The aggregates formed by rhodopsin are resistant to the mild detergent n-dodecyl-β-d-maltoside (DM) but can be disrupted by the harsher detergent sodium dodecyl sulfate (SDS) (Gragg and Park 2019). This behavior contrasts to those exhibited by wild-type rhodopsin that normally forms oligomers, which can be disrupted by DM (Jastrzebska et al. 2004; Miller et al. 2015). Thus, the protein–protein interfaces within aggregates and oligomers are different in nature.
Pharmacological chaperones have been proposed as a therapeutic approach to combat against the detrimental effects of misfolded GPCR mutants (Beerepoot et al. 2017; Tao and Conn 2014), including misfolded rhodopsin mutants. Retinoid-based pharmacological chaperones, such as the endogenous chromophore 11-cis retinal, have been shown to aid in the folding and proper cellular trafficking of misfolded rhodopsin mutants (Chen et al. 2015; Krebs et al. 2010; Mendes and Cheetham 2008; Noorwez et al. 2004). The therapeutic effectiveness of retinoid-based chaperones, however, is questionable. Retinoid-based chaperones are only effective for partial misfolding rhodopsin mutants and will have no effect on complete misfolding mutants. Moreover, it is unclear how therapeutically beneficial these chaperoning effects will be since the chaperoned misfolded mutants are unstable (Chen et al. 2014; Opefi et al. 2013). Since misfolding mutants of rhodopsin cause adRP, most patients will express both mutant and wild-type rhodopsin. Partial misfolding mutants do not aggregate with wild-type rhodopsin when coexpressed in the absence of a retinoid chaperone; however, in the presence of a retinoid chaperone, aggregation between the mutant and wild-type receptor is surprisingly observed (Gragg and Park 2018). Thus, retinoid-based chaperones are not predicted to be beneficial in these instances, but rather, are predicted to be detrimental, making the condition worse. It is unclear whether or not other types of chaperones will exhibit similar negative effects. Further studies will be required to better understand how to prevent or disrupt aggregates of rhodopsin to combat disease.
Concluding Remarks
Two types of quaternary structures formed by rhodopsin have been discussed here, oligomers and aggregates. Oligomers form under native conditions and are involved in the normal function of photoreceptor cells, whereas aggregates form under pathological conditions and are detrimental to the normal function of photoreceptor cells. The hydrophobic nature of rhodopsin makes it difficult to study oligomers and aggregates under physiologically relevant conditions. Methods that can overcome this barrier are beginning to reveal important insights into the nature of these quaternary structures. Further advances are required to better understand the fundamental properties of these quaternary structures, which will reveal further details about the mechanism of action of rhodopsin in phototransduction and the mechanism by which misfolded rhodopsin mutants cause retinal degeneration.
References
Adekeye A, Haeri M, Solessio E, Knox BE (2014) Ablation of the proapoptotic genes CHOP or Ask1 does not prevent or delay loss of visual function in a P23H transgenic mouse model of retinitis pigmentosa. PLoS ONE 9:e83871. https://doi.org/10.1371/journal.pone.0083871
Albert AD, Young JE, Paw Z (1998) Phospholipid fatty acyl spatial distribution in bovine rod outer segment disk membranes. Biochim Biophys Acta 1368:52–60
Andrews LD, Cohen AI (1979) Freeze-fracture evidence for the presence of cholesterol in particle-free patches of basal disks and the plasma membrane of retinal rod outer segments of mice and frogs. J Cell Biol 81:215–228
Arango-Gonzalez B et al (2014) Identification of a common non-apoptotic cell death mechanism in hereditary retinal degeneration. PLoS ONE 9:e112142. https://doi.org/10.1371/journal.pone.0112142
Athanasiou D et al (2012) BiP prevents rod opsin aggregation. Mol Biol Cell 23:3522–3531. https://doi.org/10.1091/mbc.e12-02-0168
Athanasiou D, Aguila M, Bevilacqua D, Novoselov SS, Parfitt DA, Cheetham ME (2013) The cell stress machinery and retinal degeneration. FEBS Lett 587:2008–2017. https://doi.org/10.1016/j.febslet.2013.05.020
Athanasiou D, Aguila M, Bellingham J, Li W, McCulley C, Reeves PJ, Cheetham ME (2018) The molecular and cellular basis of rhodopsin retinitis pigmentosa reveals potential strategies for therapy. Prog Retin Eye Res 62:1–23. https://doi.org/10.1016/j.preteyeres.2017.10.002
Baylor DA, Lamb TD (1982) Local effects of bleaching in retinal rods of the toad. J Physiol 328:49–71
Baylor DA, Lamb TD, Yau KW (1979) Responses of retinal rods to single photons. J Physiol 288:613–634
Beerepoot P, Nazari R, Salahpour A (2017) Pharmacological chaperone approaches for rescuing GPCR mutants: current state, challenges, and screening strategies. Pharmacol Res 117:242–251. https://doi.org/10.1016/j.phrs.2016.12.036
Bence NF, Sampat RM, Kopito RR (2001) Impairment of the ubiquitin-proteasome system by protein aggregation. Science 292:1552–1555. https://doi.org/10.1126/science.292.5521.1552
Bethani I, Skanland SS, Dikic I, Acker-Palmer A (2010) Spatial organization of transmembrane receptor signalling. EMBO J 29:2677–2688. https://doi.org/10.1038/emboj.2010.175
Boesze-Battaglia K, Albert AD (1989) Fatty acid composition of bovine rod outer segment plasma membrane. Exp Eye Res 49:699–701
Boesze-Battaglia K, Fliesler SJ, Albert AD (1990) Relationship of cholesterol content to spatial distribution and age of disc membranes in retinal rod outer segments. J Biol Chem 265:18867–18870
Botelho AV, Huber T, Sakmar TP, Brown MF (2006) Curvature and hydrophobic forces drive oligomerization and modulate activity of rhodopsin in membranes. Biophys J 91:4464–4477. https://doi.org/10.1529/biophysj.106.082776
Brown MF (1994) Modulation of rhodopsin function by properties of the membrane bilayer. Chem Phys Lipids 73:159–180
Bush RA, Malnoe A, Reme CE, Williams TP (1994) Dietary deficiency of N-3 fatty acids alters rhodopsin content and function in the rat retina. Invest Ophthalmol Vis Sci 35:91–100
Caldwell RB, McLaughlin BJ (1985) Freeze-fracture study of filipin binding in photoreceptor outer segments and pigment epithelium of dystrophic and normal retinas. J Comp Neurol 236:523–537. https://doi.org/10.1002/cne.902360408
Calebiro D et al (2013) Single-molecule analysis of fluorescently labeled G-protein-coupled receptors reveals complexes with distinct dynamics and organization. Proc Natl Acad Sci USA 110:743–748. https://doi.org/10.1073/pnas.1205798110
Cangiano L, Dell’Orco D (2013) Detecting single photons: a supramolecular matter? FEBS Lett 587:1–4. https://doi.org/10.1016/j.febslet.2012.11.015
Chabre M, le Maire M (2005) Monomeric G-protein-coupled receptor as a functional unit. Biochemistry 44:9395–9403
Chabre M, Cone R, Saibil H (2003) Biophysics: is rhodopsin dimeric in native retinal rods? Nature 426:30–31
Chen Y et al (2014) Inherent instability of the retinitis pigmentosa P23H mutant opsin. J Biol Chem 289:9288–9303. https://doi.org/10.1074/jbc.m114.551713
Chen Y, Tang H, Seibel W, Papoian R, Li X, Lambert NA, Palczewski K (2015) A high-throughput drug screening strategy for detecting rhodopsin P23H mutant rescue and degradation. Invest Ophthalmol Vis Sci 56:2553–2567. https://doi.org/10.1167/iovs.14-16298
Chiang WC et al (2014) Robust endoplasmic reticulum-associated degradation of rhodopsin precedes retinal degeneration. Mol Neurobiol. https://doi.org/10.1007/s12035-014-8881-8
Chiti F, Dobson CM (2006) Protein misfolding, functional amyloid, and human disease. Annu Rev Biochem 75:333–366. https://doi.org/10.1146/annurev.biochem.75.101304.123901
Comar WD, Schubert SM, Jastrzebska B, Palczewski K, Smith AW (2014) Time-resolved fluorescence spectroscopy measures clustering and mobility of a G protein-coupled receptor opsin in live cell membranes. J Am Chem Soc 136:8342–8349. https://doi.org/10.1021/ja501948w
Daemen FJ (1973) Vertebrate rod outer segment membranes. Biochim Biophys Acta 300:255–288
Dalke C, Graw J (2005) Mouse mutants as models for congenital retinal disorders. Exp Eye Res 81:503–512
Dell’Orco D (2013) A physiological role for the supramolecular organization of rhodopsin and transducin in rod photoreceptors. FEBS Lett 587:2060–2066. https://doi.org/10.1016/j.febslet.2013.05.017
Dell’Orco D, Schmidt H (2008) Mesoscopic Monte Carlo simulations of stochastic encounters between photoactivated rhodopsin and transducin in disc membranes. J Phys Chem B 112:4419–4426
Ding JD, Salinas RY, Arshavsky VY (2015) Discs of mammalian rod photoreceptors form through the membrane evagination mechanism. J Cell Biol 211:495–502. https://doi.org/10.1083/jcb.201508093
Dryja TP et al (1990) A point mutation of the rhodopsin gene in one form of retinitis pigmentosa. Nature 343:364–366. https://doi.org/10.1038/343364a0
Garriga P, Liu X, Khorana HG (1996) Structure and function in rhodopsin: correct folding and misfolding in point mutants at and in proximity to the site of the retinitis pigmentosa mutation Leu-125→Arg in the transmembrane helix C. Proc Natl Acad Sci USA 93:4560–4564
Gilliam JC et al (2012) Three-dimensional architecture of the rod sensory cilium and its disruption in retinal neurodegeneration. Cell 151:1029–1041. https://doi.org/10.1016/j.cell.2012.10.038
Goldberg AF, Moritz OL, Williams DS (2016) Molecular basis for photoreceptor outer segment architecture. Prog Retin Eye Res 55:52–81. https://doi.org/10.1016/j.preteyeres.2016.05.003
Gorbatyuk M, Gorbatyuk O (2013) Review: retinal degeneration: focus on the unfolded protein response. Mol Vis 19:1985–1998
Gorbatyuk MS et al (2010) Restoration of visual function in P23H rhodopsin transgenic rats by gene delivery of BiP/Grp78. Proc Natl Acad Sci USA 107:5961–5966. https://doi.org/10.1073/pnas.0911991107
Gragg M, Park PS (2018) Misfolded rhodopsin mutants display variable aggregation properties. Biochim Biophys Acta 1864:2938–2948. https://doi.org/10.1016/j.bbadis.2018.06.004
Gragg M, Park PS (2019) Detection of misfolded rhodopsin aggregates in cells by Forster resonance energy transfer. Methods Cell Biol 149:87–105. https://doi.org/10.1016/bs.mcb.2018.08.007
Gragg M, Kim TG, Howell S, Park PS (2016) Wild-type opsin does not aggregate with a misfolded opsin mutant. Biochim Biophys Acta 1858:1850–1859. https://doi.org/10.1016/j.bbamem.2016.04.013
Griciuc A, Aron L, Ueffing M (2011) ER stress in retinal degeneration: a target for rational therapy? Trends Mol Med 17:442–451. https://doi.org/10.1016/j.molmed.2011.04.002
Grossfield A, Feller SE, Pitman MC (2006) A role for direct interactions in the modulation of rhodopsin by omega-3 polyunsaturated lipids. Proc Natl Acad Sci USA 103:4888–4893. https://doi.org/10.1073/pnas.0508352103
Gunkel M, Schoneberg J, Alkhaldi W, Irsen S, Noe F, Kaupp UB, Al-Amoudi A (2015) Higher-order architecture of rhodopsin in intact photoreceptors and its implication for phototransduction kinetics. Structure 23:628–638. https://doi.org/10.1016/j.str.2015.01.015
Haeri M, Calvert PD, Solessio E, Pugh EN Jr, Knox BE (2013) Regulation of rhodopsin-eGFP distribution in transgenic xenopus rod outer segments by light. PLoS ONE 8:e80059. https://doi.org/10.1371/journal.pone.0080059
Hartong DT, Berson EL, Dryja TP (2006) Retinitis pigmentosa. Lancet 368:1795–1809. https://doi.org/10.1016/S0140-6736(06)69740-7
Hegener O, Prenner L, Runkel F, Baader SL, Kappler J, Haberlein H (2004) Dynamics of beta2-adrenergic receptor-ligand complexes on living cells. Biochemistry 43:6190–6199. https://doi.org/10.1021/bi035928t
Hern JA et al (2010) Formation and dissociation of M1 muscarinic receptor dimers seen by total internal reflection fluorescence imaging of single molecules. Proc Natl Acad Sci USA 107:2693–2698. https://doi.org/10.1073/pnas.0907915107
Herrick-Davis K, Grinde E, Lindsley T, Teitler M, Mancia F, Cowan A, Mazurkiewicz JE (2015) Native serotonin 5-HT2C receptors are expressed as homodimers on the apical surface of choroid plexus epithelial cells. Mol Pharmacol 87:660–673. https://doi.org/10.1124/mol.114.096636
Hsu YC, Chuang JZ, Sung CH (2015) Light regulates the ciliary protein transport and outer segment disc renewal of Mammalian photoreceptors. Dev Cell 32:731–742. https://doi.org/10.1016/j.devcel.2015.01.027
Hwa J, Garriga P, Liu X, Khorana HG (1997) Structure and function in rhodopsin: packing of the helices in the transmembrane domain and folding to a tertiary structure in the intradiscal domain are coupled. Proc Natl Acad Sci USA 94:10571–10576
Illing ME, Rajan RS, Bence NF, Kopito RR (2002) A rhodopsin mutant linked to autosomal dominant retinitis pigmentosa is prone to aggregate and interacts with the ubiquitin proteasome system. J Biol Chem 277:34150–34160
Insinna C, Besharse JC (2008) Intraflagellar transport and the sensory outer segment of vertebrate photoreceptors. Dev Dyn 237:1982–1992. https://doi.org/10.1002/dvdy.21554
Jastrzebska B et al (2004) Functional characterization of rhodopsin monomers and dimers in detergents. J Biol Chem 279:54663–54675
Jastrzebska B, Fotiadis D, Jang GF, Stenkamp RE, Engel A, Palczewski K (2006) Functional and structural characterization of rhodopsin oligomers. J Biol Chem 281:11917–11922
Kasai RS, Suzuki KG, Prossnitz ER, Koyama-Honda I, Nakada C, Fujiwara TK, Kusumi A (2011) Full characterization of GPCR monomer-dimer dynamic equilibrium by single molecule imaging. J Cell Biol 192:463–480. https://doi.org/10.1083/jcb.201009128
Kaushal S, Khorana HG (1994) Structure and function in rhodopsin. 7. Point mutations associated with autosomal dominant retinitis pigmentosa. Biochemistry 33:6121–6128
Kirchhoff H (2014) Diffusion of molecules and macromolecules in thylakoid membranes. Biochim Biophys Acta 1837:495–502. https://doi.org/10.1016/j.bbabio.2013.11.003
Knowles TP, Vendruscolo M, Dobson CM (2014) The amyloid state and its association with protein misfolding diseases. Nat Rev Mol Cell Biol 15:384–396. https://doi.org/10.1038/nrm3810
Kopito RR (2000) Aggresomes, inclusion bodies and protein aggregation. Trends Cell Biol 10:524–530
Krebs MP, Holden DC, Joshi P, Clark CL 3rd, Lee AH, Kaushal S (2010) Molecular mechanisms of rhodopsin retinitis pigmentosa and the efficacy of pharmacological rescue. J Mol Biol 395:1063–1078. https://doi.org/10.1016/j.jmb.2009.11.015
Kroeger H, Chiang WC, Lin JH (2012) Endoplasmic reticulum-associated degradation (ERAD) of misfolded glycoproteins and mutant P23H rhodopsin in photoreceptor cells. Adv Exp Med Biol 723:559–565. https://doi.org/10.1007/978-1-4614-0631-0_71
Liang Y, Fotiadis D, Filipek S, Saperstein DA, Palczewski K, Engel A (2003) Organization of the G protein-coupled receptors rhodopsin and opsin in native membranes. J Biol Chem 278:21655–21662
Liu X, Garriga P, Khorana HG (1996) Structure and function in rhodopsin: correct folding and misfolding in two point mutants in the intradiscal domain of rhodopsin identified in retinitis pigmentosa. Proc Natl Acad Sci USA 93:4554–4559
Makino CL, Howard LN, Williams TP (1990) Axial gradients of rhodopsin in light-exposed retinal rods of the toad. J Gen Physiol 96:1199–1220
Mazzolini M et al (2015) The phototransduction machinery in the rod outer segment has a strong efficacy gradient. Proc Natl Acad Sci USA 112:E2715–E2724. https://doi.org/10.1073/pnas.1423162112
Mendes HF, Cheetham ME (2008) Pharmacological manipulation of gain-of-function and dominant-negative mechanisms in rhodopsin retinitis pigmentosa. Hum Mol Genet 17:3043–3054. https://doi.org/10.1093/hmg/ddn202
Mendes HF, van der Spuy J, Chapple JP, Cheetham ME (2005) Mechanisms of cell death in rhodopsin retinitis pigmentosa: implications for therapy. Trends Mol Med 11:177–185. https://doi.org/10.1016/j.molmed.2005.02.007
Miller LM, Gragg M, Kim TG, Park PS (2015) Misfolded opsin mutants display elevated beta-sheet structure. FEBS Lett 589:3119–3125. https://doi.org/10.1016/j.febslet.2015.08.042
Mishra AK et al (2016) Quaternary structures of opsin in live cells revealed by FRET spectrometry. Biochem J 473:3819–3836. https://doi.org/10.1042/bcj20160422
Mitchell DC, Niu SL, Litman BJ (2001) Optimization of receptor-G protein coupling by bilayer lipid composition I: kinetics of rhodopsin-transducin binding. J Biol Chem 276:42801–42806. https://doi.org/10.1074/jbc.m105772200
Mugler A, Tostevin F, ten Wolde PR (2013) Spatial partitioning improves the reliability of biochemical signaling. Proc Natl Acad Sci USA 110:5927–5932. https://doi.org/10.1073/pnas.1218301110
Muller DJ (2008) AFM: a nanotool in membrane biology. Biochemistry 47:7986–7998. https://doi.org/10.1021/bi800753x
Nathans J, Merbs SL, Sung CH, Weitz CJ, Wang Y (1992) Molecular genetics of human visual pigments. Annu Rev Genet 26:403–424. https://doi.org/10.1146/annurev.ge.26.120192.002155
Nemet I, Ropelewski P, Imanishi Y (2015) Rhodopsin trafficking and mistrafficking: signals, molecular components, and mechanisms. Prog Mol Biol Transl Sci 132:39–71. https://doi.org/10.1016/bs.pmbts.2015.02.007
Nickell S, Park PS, Baumeister W, Palczewski K (2007) Three-dimensional architecture of murine rod outer segments determined by cryoelectron tomography. J Cell Biol 177:917–925
Niu SL, Mitchell DC, Litman BJ (2001) Optimization of receptor-G protein coupling by bilayer lipid composition II: formation of metarhodopsin II-transducin complex. J Biol Chem 276:42807–42811. https://doi.org/10.1074/jbc.m105778200
Niu SL, Mitchell DC, Lim SY, Wen ZM, Kim HY, Salem N Jr, Litman BJ (2004) Reduced G protein-coupled signaling efficiency in retinal rod outer segments in response to n-3 fatty acid deficiency. J Biol Chem 279:31098–31104. https://doi.org/10.1074/jbc.m404376200
Noorwez SM, Malhotra R, McDowell JH, Smith KA, Krebs MP, Kaushal S (2004) Retinoids assist the cellular folding of the autosomal dominant retinitis pigmentosa opsin mutant P23H. J Biol Chem 279:16278–16284
Opefi CA, South K, Reynolds CA, Smith SO, Reeves PJ (2013) Retinitis pigmentosa mutants provide insight into the role of the N-terminal cap in rhodopsin folding, structure, and function. J Biol Chem 288:33912–33926. https://doi.org/10.1074/jbc.m113.483032
Organisciak DT, Noell WK (1977) The rod outer segment phospholipid/opsin ratio of rats maintained in darkness or cyclic light. Invest Ophthalmol Vis Sci 16:188–190
Parfitt DA et al (2014) The heat-shock response co-inducer arimoclomol protects against retinal degeneration in rhodopsin retinitis pigmentosa. Cell Death Dis 5:e1236. https://doi.org/10.1038/cddis.2014.214
Park PS (2014) Constitutively active rhodopsin and retinal disease. Adv Pharmacol 70:1–36. https://doi.org/10.1016/b978-0-12-417197-8.00001-8
Patowary S, Alvarez-Curto E, Xu TR, Holz JD, Oliver JA, Milligan G, Raicu V (2013) The muscarinic M3 acetylcholine receptor exists as two differently sized complexes at the plasma membrane. Biochem J 452:303–312. https://doi.org/10.1042/bj20121902
Penn JS, Anderson RE (1987) Effect of light history on rod outer-segment membrane composition in the rat. Exp Eye Res 44:767–778
Periole X, Huber T, Marrink SJ, Sakmar TP (2007) G protein-coupled receptors self-assemble in dynamics simulations of model bilayers. J Am Chem Soc 129:10126–10132. https://doi.org/10.1021/ja0706246
Price BA, Sandoval IM, Chan F, Simons DL, Wu SM, Wensel TG, Wilson JH (2011) Mislocalization and degradation of human P23H-rhodopsin-GFP in a knockin mouse model of retinitis pigmentosa. Invest Ophthalmol Vis Sci 52:9728–9736. https://doi.org/10.1167/iovs.11-8654
Radhakrishnan K, Halasz A, McCabe MM, Edwards JS, Wilson BS (2012) Mathematical simulation of membrane protein clustering for efficient signal transduction. Ann Biomed Eng 40:2307–2318. https://doi.org/10.1007/s10439-012-0599-z
Rajan RS, Illing ME, Bence NF, Kopito RR (2001) Specificity in intracellular protein aggregation and inclusion body formation. Proc Natl Acad Sci USA 98:13060–13065. https://doi.org/10.1073/pnas.181479798
Rakshit T, Park PS (2015) Impact of reduced rhodopsin expression on the structure of rod outer segment disc membranes. Biochemistry 54:2885–2894. https://doi.org/10.1021/acs.biochem.5b00003
Rakshit T, Senapati S, Sinha S, Whited AM, Park PS-H (2015) Rhodopsin forms nanodomains in rod outer segment disc membranes of the cold-blooded Xenopus laevis. PLoS ONE 10:e0141114
Rakshit T, Senapati S, Parmar VM, Sahu B, Maeda A, Park PS (2017) Adaptations in rod outer segment disc membranes in response to environmental lighting conditions. Biochim Biophys Acta 1864:1691–1702. https://doi.org/10.1016/j.bbamcr.2017.06.013
Sakami S et al (2011) Probing mechanisms of photoreceptor degeneration in a new mouse model of the common form of autosomal dominant retinitis pigmentosa due to P23H opsin mutations. J Biol Chem 286:10551–10567. https://doi.org/10.1074/jbc.m1
Saliba RS, Munro PM, Luthert PJ, Cheetham ME (2002) The cellular fate of mutant rhodopsin: quality control, degradation and aggresome formation. J Cell Sci 115:2907–2918
Saxton MJ, Owicki JC (1989) Concentration effects on reactions in membranes: rhodopsin and transducin. Biochim Biophys Acta 979:27–34
Schoneberg J, Heck M, Hofmann KP, Noe F (2014) Explicit spatiotemporal simulation of receptor-g protein coupling in rod cell disk membranes. Biophys J 107:1042–1053. https://doi.org/10.1016/j.bpj.2014.05.050
Senapati S, Park PS (2019) Investigating the nanodomain organization of rhodopsin in native membranes by atomic force microscopy. Methods Mol Biol 1886:61–74. https://doi.org/10.1007/978-1-4939-8894-5_4
Senapati S, Gragg M, Samuels IS, Parmar VM, Maeda A, Park PS (2018) Effect of dietary docosahexaenoic acid on rhodopsin content and packing in photoreceptor cell membranes. Biochim Biophys Acta 1860:1403–1413. https://doi.org/10.1016/j.bbamem.2018.03.030
Sizova OS, Shinde VM, Lenox AR, Gorbatyuk MS (2014) Modulation of cellular signaling pathways in P23H rhodopsin photoreceptors. Cell Signal 26:665–672. https://doi.org/10.1016/j.cellsig.2013.12.008
Soubias O, Teague WE Jr, Hines KG, Gawrisch K (2015) Rhodopsin/lipid hydrophobic matching-rhodopsin oligomerization and function. Biophys J 108:1125–1132. https://doi.org/10.1016/j.bpj.2015.01.006
Stojanovic A, Hwa J (2002) Rhodopsin and retinitis pigmentosa: shedding light on structure and function. Receptors Channels 8:33–50
Stoneman MR, Paprocki JD, Biener G, Yokoi K, Shevade A, Kuchin S, Raicu V (2017) Quaternary structure of the yeast pheromone receptor Ste2 in living cells. Biochim Biophys Acta Biomembr 1859:1456–1464. https://doi.org/10.1016/j.bbamem.2016.12.008
Sung CH, Chuang JZ (2010) The cell biology of vision. J Cell Biol 190:953–963. https://doi.org/10.1083/jcb.201006020
Sung CH et al (1991a) Rhodopsin mutations in autosomal dominant retinitis pigmentosa. Proc Natl Acad Sci USA 88:6481–6485
Sung CH, Schneider BG, Agarwal N, Papermaster DS, Nathans J (1991b) Functional heterogeneity of mutant rhodopsins responsible for autosomal dominant retinitis pigmentosa. Proc Natl Acad Sci USA 88:8840–8844
Sung CH, Davenport CM, Nathans J (1993) Rhodopsin mutations responsible for autosomal dominant retinitis pigmentosa. Clustering of functional classes along the polypeptide chain. J Biol Chem 268:26645–26649
Tao YX, Conn PM (2014) Chaperoning G protein-coupled receptors: from cell biology to therapeutics. Endocr Rev 35:602–647. https://doi.org/10.1210/er.2013-1121
Vasireddy V et al (2011) Rescue of photoreceptor degeneration by curcumin in transgenic rats with P23H rhodopsin mutation. PLoS ONE 6:e21193. https://doi.org/10.1371/journal.pone.0021193
Volland S et al (2015) Three-dimensional organization of nascent rod outer segment disk membranes. Proc Natl Acad Sci USA 112:14870–14875. https://doi.org/10.1073/pnas.1516309112
Walter P, Ron D (2011) The unfolded protein response: from stress pathway to homeostatic regulation. Science 334:1081–1086. https://doi.org/10.1126/science.1209038
Wang J, Deretic D (2014) Molecular complexes that direct rhodopsin transport to primary cilia. Prog Retin Eye Res 38:1–19. https://doi.org/10.1016/j.preteyeres.2013.08.004
Ward RJ, Pediani JD, Godin AG, Milligan G (2015) Regulation of oligomeric organization of the serotonin 5-hydroxytryptamine 2C (5-HT2C) receptor observed by spatial intensity distribution analysis. J Biol Chem 290:12844–12857. https://doi.org/10.1074/jbc.m115.644724
Ward RJ, Pediani JD, Harikumar KG, Miller LJ, Milligan G (2017) Spatial intensity distribution analysis quantifies the extent and regulation of homodimerization of the secretin receptor. Biochem J 474:1879–1895. https://doi.org/10.1042/bcj20170184
Wensel TG, Zhang Z, Anastassov IA, Gilliam JC, He F, Schmid MF, Robichaux MA (2016) Structural and molecular bases of rod photoreceptor morphogenesis and disease. Prog Retin Eye Res 55:32–51. https://doi.org/10.1016/j.preteyeres.2016.06.002
Whited AM, Park PS (2014) Atomic force microscopy: a multifaceted tool to study membrane proteins and their interactions with ligands. Biochim Biophys Acta 1838:56–68. https://doi.org/10.1016/j.bbamem.2013.04.011
Whited AM, Park PSH (2015) Nanodomain organization of rhodopsin in native human and murine rod outer segment disc membranes. Bba-Biomembranes 1848:26–34. https://doi.org/10.1016/j.bbamem.2014.10.007
Wiedmann TS, Pates RD, Beach JM, Salmon A, Brown MF (1988) Lipid-protein interactions mediate the photochemical function of rhodopsin. Biochemistry 27:6469–6474
Williams TP, Penn JS (1985) Intracellular topography of rhodopsin regeneration in vertebrate rods. J Gen Physiol 86:413–422
Young JE, Albert AD (2000) Transducin binding in bovine rod outer segment disk membranes of different age/spatial location. Exp Eye Res 70:809–812. https://doi.org/10.1006/exer.2000.0821
Funding
This work was funded by a Grant from the National Institutes of Health (R01EY021731).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares that he has no conflicts of interest to disclose.
Ethical Approval
This article does not contain any studies with human participants or animals performed by the author.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Park, P.SH. Rhodopsin Oligomerization and Aggregation. J Membrane Biol 252, 413–423 (2019). https://doi.org/10.1007/s00232-019-00078-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-019-00078-1