Abstract
The emergence of a crystal nucleus from disordered states is a critical and challenging aspect of the crystallization process, primarily due to the extremely short length and timescales involved. Methods such as liquid-cell or low-dose focal-series transmission electron microscopy (TEM) are often employed to probe these events. In this study, we demonstrate that ion mobility spectrometry-mass spectrometry (IMS-MS) offers a complementary and insightful perspective on the nucleation process by examining the sizes and shapes of small clusters, specifically those ranging from n = 2 to 40. Our findings reveal the significant role of sulfate ions in the growth of adeninediium sulfate clusters, which are the precursors to the formation of single crystals. Specifically, sulfate ions stabilize adenine clusters at the 1:1 ratio. In contrast, guanine sulfate forms smaller clusters with varied ratios, which become stable as they approach the 1:2 ratio. The nucleation size is predicted to be between n = 8 and 14, correlating well with the unit cell dimensions of adenine crystals. This correlation suggests that IMS-MS can identify critical nucleation sizes and provide valuable structural information consistent with established crystallographic data. We also discuss the strengths and limitations of IMS-MS in this context. IMS-MS offers rapid and robust experimental protocols, making it a valuable tool for studying the effects of various additives on the assembly of small molecules. Additionally, it aids in elucidating nucleation processes and the growth of different crystal polymorphs.
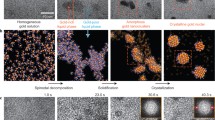
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Considerable effort has been dedicated to investigating the self-assemblies of small molecules using mass spectrometry (MS) [1,2,3,4], initially spurred by the discovery of amyloid-like fibrils made of metabolites [3, 5,6,7,8,9,10], amino acids [4, 11,12,13,14,15,16,17], and lipids [18]. This endeavor has deepened our understanding of the amyloid hypothesis and forged connections between metabolic disorders and protein-aggregation diseases [5, 6, 19,20,21]. While intriguing, the differentiation between amyloid-like fibril formation and metabolite crystallization presents a significant challenge due to their concurrent occurrence under similar concentrations and conditions. In a recent investigation involving cysteine and cystine, we observed a temporal sequence wherein the fibril formation of cysteine preceded the emergence of hexagonal cystine crystals [4]. Such findings suggest analogous behaviors in other metabolites, prompting a shift in focus towards elucidating the self-assembly, nucleation, and growth mechanisms of small-molecule crystals using MS.
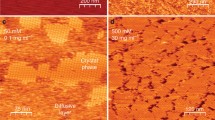
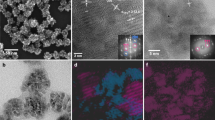
Gaining a thorough understanding of the evolution of molecular or atomic order during the initial stages of ordered structure formation, such as crystals or fibrils, has remained a persistent challenge. At the same time, probing the transformation from amorphous aggregates into crystalline matter can potentially rationalize structural polymorphism and non-covalent interactions [22], identify novel materials and catalysis [23,24,25,26,27,28,29], and provide mechanistic details to modify existing crystallization theories [30,31,32,33,34,35,36]. At present, the state-of-the-art techniques for investigating the crystallization mechanisms of small molecules are transmission electron microscopy (TEM) [29, 31,32,33,34,35,36,37] and molecular dynamics (MD) [38,39,40]. Although TEM operates in the condensed phase and offers real-time visualization of the self-assembly process, it may overlook crucial aspects such as initial molecular interactions or the influence of solvent additives. Furthermore, in classic nucleation theory, a metastable disordered dense liquid or amorphous solid cluster spontaneously and irreversibly transforms into a crystalline nucleus. However, recent electron microscopy studies have suggested more dynamic processes with reversible fluctuations between disordered and ordered states and between different polymorphs [34, 35, 37]. For small molecules, these processes occur at sizes of a dozen or fewer molecules and cluster sizes of a few nanometers [35,36,37,38]. Observing a few molecules associate and dissociate while searching for metastable states during crystallization remains challenging and time-consuming. Considering this, the scope of analytes that have been and can be studied by TEM and MD is constrained. This includes small organic molecules and model systems like ionic salts [38,39,40].
Why is mass spectrometry useful for understanding assemblies and nucleation?
As both crystallization and fibrillization gradually eliminate water from the assemblies, a comparable process, such as dehydration in the gas phase, could offer an alternative approach to examining the initial assemblies and the roles of additives in the assembly of small molecules like adenine. The gas phase provides an extreme environment where undesired interferences from the surface and solvent can be eliminated. The mass selection capability of MS allows for a robust characterization of individual species that are otherwise heterogeneous in the condensed phase. Gas-phase techniques such as ion mobility spectrometry-mass spectrometry (IMS-MS) have emerged as novel tools at the forefront of structural biology and amyloid studies [41,42,43,44,45]. In early studies, experimental IMS-MS data (e.g., collisional cross sections (CCS) are often directly compared with theoretical values derived from X-ray crystal structures [45]. As the MS conditions can be fine-tuned to kinetically trap condensed-phase structures, IMS-MS has provided a unique perspective into the early stages of the bimolecular self-assembly process. With IMS-MS, the very early stages of assembly, involving only a few molecules, can be observed. One can distinguish between stable and unstable arrangements and model the structures using CCS data [1,2,3, 45].
While MS does offer significant advantages, it also comes with limitations. One primary constraint is its ability to detect only clusters with non-zero net charges, whereas most clusters in the condensed phase maintain charge neutrality. Additionally, interferences from charge carriers or protonation (deprotonation) introduced during desolvation but before structural measurements cannot be overlooked. Consequently, MS has not emerged as a competitive tool for studying nucleation and crystallization of small molecules. However, it presents an excellent opportunity to investigate systems where the charge carriers are the crystallization additives. In such cases, the presence of a chargeable species is essential for the growth of crystals in the condensed phase and the detection of clusters in the gas phase. Although these systems are limited in scope, they serve as valuable platforms for proof-of-concept experiments.
Nucleobase crystallization is biologically relevant
Nucleobases are an integral part of the genetic blueprint of all living organisms and carry important hereditary information [46]. Adenine, thymine, guanine, and cytosine make up the DNA double-helix strand, while uracil replaces thymine in the case of RNA [47]. Nucleobase-containing molecules are indispensable in biology due to the critical roles of nucleic acids, particularly G-quadruplex DNA and RNA. For example, adenine is an essential component of high-energy molecules such as adenosine triphosphate (ATP), nicotinamide adenine dinucleotide (NAD), and flavin adenine dinucleotide (FAD) [48], all are needed for cellular metabolism [49]. In addition, adenine’s diverse binding to proteins as a ligand has been exploited for designing different inhibitors by characterizing potential molecules based on similar active site classification that mimics adenine [50].
Gazit and co-workers proposed an intriguing hypothesis linking metabolostasis to proteostasis [19, 51], suggesting that disruptions in metabolostasis, as in inborn errors of metabolism (IEM) or severe combined immunodeficiency (SCID) [52], increases the risk of protein amyloidosis. This is supported by compelling evidence; for example, the fibril formation of homocysteine precedes and colocalizes with the aggregation of amyloid-β [1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42] in Alzheimer’s disease mouse brains [3]. Specific to adenine, defects in adenine phosphoribosyltransferase and adenosine deaminase result in the overaccumulation of adenine and adenosine in SCID [53]. In yeast models, increasing levels of adenine have been shown to lead to fibril-like formation and decreased cell growth [7]. In vivo, xanthine dehydrogenase oxidizes excess adenine to 2,8-dihydroxyadenine, of which the low solubility leads to crystal precipitation and chronic kidney disease [54, 55]. Clearly, both the fibrillization of adenine and the crystallization of its oxidized derivatives are harmful.
Interestingly, adenine, characterized by its simple structure, versatile coordination abilities, and hydrogen bonding capacity, presents a novel avenue for various ligands to engage in assemblies, generating diverse polymorphs. The reported crystal structures of adenine include adenine hydrochloride [56], adenine dihydrochloride (or adeninediium dichloride) [57, 58], adenine sulfate and hemisulfate [59], adenine-water-halide [60], adeninediium dinitrate [61], adenine with metal salts [62, 63], adenine with organic acids [64], adenine trihydrate [65], and anhydrous adenine [66, 67]. The small ligands increase adenine’s solubility and drive forward the formation of crystals [68]. Based on these findings, we posit that metabolites like adenine have their small-molecule chaperones [69]. These chaperones include but are not limited to sulfate and chloride ions, which are ubiquitous but essential macronutrients. Crystallization, or the formation of “solid solutions,” in which the molecule host harbors metabolically available molecules into its crystals, could provide a strategy to protect overproduced metabolites from aggregation and formation of cytotoxic derivatives. Unlike irreversible aggregation or fibrillization, in vivo biogenic crystals serve as an economical and physiological reservoir to maintain a balance between the need for high concentrations and retaining functional forms, allowing the molecules to be reversibly transformed from crystalline to soluble forms when needed.
While there has not been enough study on the biological activities of adenine-salt crystals, biogenic, functional crystals of guanine are found in living organisms [70,71,72,73,74]. Due to their high reflective index, these crystals perform functions such as coloration. Analysis of these crystals shows that they contain not only guanine but also hypoxanthine and xanthine [70].
In this work, our main goal is to assess the stability of adenine clusters in the gas phase and investigate whether gas-phase structures can provide insights into the oligomer growth and crystallization of adenine and the influence of ligands. For this purpose, we opt for hemisulfate/sulfate as the ligand, with adenine hemisulfate chosen due to its ability to accelerate crystallization and produce high-quality crystals [75]. This choice underscores the significant role of sulfate ions in stabilizing the structures. Of note, anhydrous adenine is more sensitive to crystallization conditions [66]. Since the crystal structure is typically obtained in a hydrated form [23, 56, 59, 65], we will also examine the impact of dehydration on the system. Moreover, we will compare the cluster formation of adenine in the presence of sulfates with the clusters of guanines. Strategies and interpretation of the structural data regarding crystal growth and formation will also be discussed.
Materials and methods
Ion mobility spectrometry-mass spectrometry (IMS-MS)
Adenine, guanine sulfate dihydrate (both from VWR), and adenine hemisulfate (Sigma-Aldrich) at > 99% purity were purchased and used without further purification. Each adenine sample was reconstituted in LC–MS grade water to a stock concentration of 10 mM. The stock solution was diluted using 20 mM ammonium acetate pH 7 to the desired sample concentration (1 mM) for the IMS-MS experiments. To prepare the working concentration of guanine sulfate dihydrate, the stock solution was first reconstituted in LC–MS H2O to a concentration of 10 mM with about 25% HCl (1N, Acros Organics) introduced into the solvent to aid sample dissolution for MS experiments. The stock solution was diluted to 1 mM using 20 mM ammonium acetate pH 7. All multi-field ion-mobility spectrometry experiments were performed using an Agilent 6560 IMS-Q-TOF instrument (Agilent Technologies, Santa Clara, CA). The mass calibration and CCS were validated using the Agilent ESI-L tuning mix (diluted in 95:5 v/v ACN: H2O). The ions were generated by electrospray ionization (ESI) by a dual ESI/Agilent Jet stream (JS) source and a syringe pump at the rate of 30 µL/min. Instrument parameters were tuned based on the early work by Gabelica and co-workers [76]. The ions were stored in a source funnel and then pulsed into a 78.1-cm drift cell filled under a drift-gas pressure of 3.94 Torr. The ions then traversed through the drift tube under the influence of a weak electric field and simultaneously collided with the stationary buffer gas (nitrogen). The size, shape, and net charge determined the velocity at which the ions drifted through the cell. Data were obtained in positive polarity over the course of 5.2 min with drift cell voltages ΔV = 1490, 1390, 1290, 1190, 1090 V in nitrogen. For each species, the drift velocity is related to the reduced ion mobility \({K}_{0}\) and used to calculate the momentum-transfer collision integral through the Mason-Schamp equation, which is reported as the experimental CCS [77].
where e is the elementary charge, N is the gas number density of the drift-gas, µ is the reduced mass of the ion and drift gas, kB is the Boltzmann constant, and T is the drift-gas temperature. The mass spectra were analyzed on IM-MS browser B.08.00 software (Agilent, Santa Clara). The arrival time distributions (ATDs) and the mass spectra graphs were made using OriginPro. The instrument method parameters can be found in the Supporting Information.
Crystal growth
Adenine hemisulfate (≥ 99% purity, Sigma-Aldrich) stock was prepared in LC–MS grade water to a final concentration of 38 mM, from which two crystal growth trials were set up. The first trial was prepared by introducing 500 µL of 38 mM stock each into an NMR tube. Then, 500 µL of ethanol was added to the NMR tube, resulting in a final concentration of 19 mM adenine hemisulfate in the NMR tube. The remaining set was prepared using the same protocol except that the stock concentration of adenine hemisulfate used here was 19 mM (the 38 mM stock was diluted), yielding a final concentration of 9.5 mM in the NMR tube.
X-ray crystallographic analysis
The single-crystal X-ray diffraction data of adenine hemisulfate were measured on a Bruker D8 Venture diffractometer equipped with a Photon 100 detector (Mo Kα radiation, λ = 0.71 Å). The cylindrical-like shaped crystal was coated in Paratone oil and mounted on a MiTeGen loop. Data were collected at 100 K after placing the crystal in a cold stream of nitrogen generated by an Oxford Cryostream low-temperature apparatus. Data were reduced with Bruker SAINT and corrected for absorption using SADABS. The structure was solved and refined using SHELX and OLEX2, respectively. The other parameters for structural refinement are listed in Table S2 in the Supporting Information.
Results and discussion
Crystal structure of adenine: structural determination using single crystal X-ray diffraction
The structural determination of adenine hemisulfate at 100 K reveals that the compound crystallizes into a triclinic space group crystal (P1) with the lattice constants of a = 7.7601(3)Å; b = 9.4163(3)Å; c = 11.6085(4)Å; α = 83.908(1)°; β = 76.379(1)°; γ = 81.742(1)°. The X-ray structure is shown in Fig. 1, consisting of two adeninium cations, a sulfate anion, and two water. Also, it is evident that the sulfate ion is situated in between the two adeninium cations and connected via hydrogen bonds. Additionally, each water molecule interacts with the adeninium cation and sulfate ion through a hydrogen bond connection. These hydrogen bond lengths lie within the range of 1.9–2.0 Å. The crystal structure with the atoms labeled and the ORTEP structure are shown in Figure S1. The same crystal structure was previously reported in the CCDC repository as adeninium hemisulfate monohydrate [78, 79].
The growth mechanism of adeninediium sulfate clusters probed by IMS-MS
The mass spectrum of pure adenine does not show any adenine cluster (Figure S2). Of note, the X-ray crystal structure of anhydrous adenine showed two adenine molecules held together via N–H···N hydrogen bonds in the asymmetric unit. Because no other molecule is participating in the stabilization, protonation (or deprotonation) during ESI may dramatically destabilize the complexes. The low signal-to-noise ratios in the high mass-to-charge (m/z) region indicate the formation of high molecular weight aggregates that cannot be resolved, suggesting that adenine quickly aggregates at this concentration (1 mM).
On the other hand, the mass spectrum of adenine hemisulfate (Fig. 2A, B) shows many peaks that could be assigned to clusters of adenine sulfate. No hydrated clusters were detected. The species have varied compositions of adenine, sulfate, and sodium ions (Na+ is ubiquitous in MS experiments). Each set of mass spectral peaks is assigned based on monoisotopic m/z and isotopic spacing, which subsequently yields information regarding the cluster size. The ADE to sulfate ratio in these complexes is often close to 1:1, and thus are derived from adeninediium sulfate; [ADE + 2H+] ∙ SO42−.
A ESI-mass spectrum of 1 mM adenine hemisulfate. Most mass spectral peaks can be assigned to three types of adeninediium sulfate clusters. The notation (n, k, x, y) is used where n is the number of adenines, k is the number of sulfuric acids, x is the number of additional protons (H+), and y is the number of sodium cations (Na+). B A partial mass spectrum showing the relative intensities of three types of clusters. Types I and II are much more abundant than type III. Within type II, n > k is much more intense than n \(\le\) k. C A representative 2D plot of m/z vs. drift time (ms) shows two overlapping isotope distributions of two distinct species. The first species is type II, i.e., (21, 19, 2, 0) = 19 ADE** + 19 SO42− + 2 ADE* (or ADE** + ADE), and the second species is type I, i.e., (10, 10, 0, 1) = 10 ADE** + 10 SO42− + Na+
We abbreviate [ADE + 2H+] hereafter as ADE** (doubly protonated adenine or adeninediium). The initial complexes could have the general formula of nADE** + kSO42− + xH+ + yNa+. Thus, we expect our MS data to resemble more adeninediium sulfate (1:1 ratio) than adenium hemisulfate (2:1 ratio). Furthermore, for the adeninediium sulfate clusters to be detected in positive ESI polarity, the complexes need to either acquire a cation (e.g., Na+ or K+; referred to as type I) or lose an SO42− or possess an unpaired ADE* (adeninium) (types II and III).
We will discuss the different types of clusters observed by IMS-MS. Figure 1B shows an example of the three types of clusters. Type I clusters have the overall nADE** + nSO42− + Na+ compositions. The arrival time distributions (ATDs) corresponding to these m/z values exhibit multiple features, discernible in two distinct sections. Specifically, only the rightmost segment of the ATDs, often characterized by three closely spaced yet well-resolved features, pertains to type I adenine clusters (more discussion later). When aligning the drift times of the features, we can assign them to clusters and post-IM dissociated clusters. Post-IM dissociation happens when a cluster moves through the drift cell as a cohesive, non-covalent complex. However, before reaching the detector, it dissociates into product ions with reduced masses. This dissociation process can result in either neutral or charged losses, affecting the appearance of their IM features in the ATDs [80, 81]. Neutral loss causes the species to travel slower than expected with the detected m/z [80, 81]. Conversely, charged loss causes the species to appear to travel faster than anticipated. Therefore, because post-IM dissociation changes the mass (and charge) of the dissociated species, it is possible for different adenine clusters, which have different compositions, to be detected at the same m/z values as if they were isobaric species. For example, 3ADE** + 3SO42− + Na+ (i.e., a type I adenine trimer), which has an m/z of 722, could dissociate post-IM measurements into a sodiated dimer 2ADE** + 2SO42− + Na+ and a neutral monomer ADE** + SO42− (undetectable). In other words, a species can travel with the drift time of a trimer or a tetramer but is then detected as a dimer. This dissociated trimer species will have a longer drift time than the intact dimer but identical to the intact trimer. While at first glance, these events cause the ATDs to become more complicated, the procedure to analyze data is straightforward once we align the ATDs and look for those with consistent arrival times (Fig. 3). The CCS values of the type I adenine clusters are reported in Table S4. Because Na+ or K+ adduction occurs during the ESI process, the metal cation does not participate in the growth of the clusters in solution. The metal ion adduction in type I clusters is expected to have a smaller effect on the structural integrity of the adeninediium sulfate clusters as compared to the addition of charged adenines or sulfates (protonation).
Because several combinations of adenine and sulfate give rise to overlapping isotope distributions, there are species with different monoisotopic m/z but greatly overlapping when shown in one-dimensional ATD. However, these species can be unambiguously resolved and assigned in a 2D plot of m/z versus drift time, such as Fig. 2C. Their monoisotopic masses differ, but the isotope distributions overlap with type I clusters. In type II clusters where n and k may not always be the same, the clusters can contain either ADE** and ADE* or sulfate and sulfuric acid. These unique features form the basis for further classifying type II clusters into two subtypes.
In type II-a clusters, where n > k, there are more adenines than sulfates. These complexes contain mostly adeninediium (z = + 2) and a few neutral adenines. For example, we observed 13ADE** + 12SO42− + 2ADE; (n, k) = (15, 12) at m/z 1601.5, in which there are 13 adeninediium, 2 adenine molecules, and 12 sulfate ions. Similarly, we also observed 14ADE** + 13 SO42− + 1ADE; (n, k) = (15, 13) at m/z 1650, in which there are 14 adeninediium, one adenine, and 13 sulfate ions.
In type II-b clusters, n \(\le\) k. Type II-b clusters are much less abundant than type II-a. For example, we observed 15ADE** + 14SO42− + 1H2SO4; (n, k) = (15, 15) in which there are 15 adeninediium, 14 sulfate ions, and one sulfuric acid. Similarly, at (n, k) = (15, 16), we observed 15ADE** + 14 SO42− + 2H2SO4, i.e., 15 adeninediium, 14 sulfate ions, and two sulfuric acid molecules. The type II-a clusters are observed as early as n = 2 (dimer), whereas the type II-b clusters are only seen at n = 6 or above. This indicates that adding adenine is preferable over adding sulfate (or sulfuric acid) as the clusters grow.
Type III clusters share common properties with types I and II. Like type II, type III clusters contain mostly large m with a net charge of z = + 2. Like type I, these clusters also contain a sodium cation as a charge carrier. In these clusters, at least one adenine molecule is an adeninium (singly protonated adenine); as such, to maintain the interaction with sulfate, a sodium cation is needed. For example, the molecular composition of type III’s (n, k) = (12, 11) is 11ADE** + 11SO42− + ADE* + Na+ at m/z 1361, and that of (n, k) = (12, 12) is 11ADE** + 11SO42− + 1H2SO4 + ADE* + Na.
Together, we drew the following conclusions. Firstly, due to desolvation preceding MS measurements, our observation of the ESI of adenine hemisulfate samples primarily revealed clusters of adeninediium sulfate. The doubly protonated adenine establishes robust hydrogen bonding with the sulfate anion, producing a predominantly 1:1 ratio. Secondly, the observed clusters shed light on potential growth mechanisms. Neutral adenine can incrementally join adeninediium sulfate clusters. However, because neutral adenine molecules do not form stable coordination with sulfates, only two to three adenine molecules can be added sequentially. The close interactions between neutral adenine and H2SO4 help protonate adenine into adeninium and adeninediium. This growth mechanism suggested that until the cluster reaches a balanced state with an equal number of ADE** and SO42−, the growth end of the cluster would be relatively disordered. When we compare clusters with the same n and k of the ordered and disordered states, we consistently observe that the CCS values of type I clusters are smaller than type II, which supports our claim (see Fig. 4).
Comparisons of types I and II adenine clusters with the same adenine and sulfate compositions. A Representative ATDs at m/z 1421 and m/z 1399. These monoisotopic m/z values correspond to (n, k, x, y) = (6, 6, 0, 1) and (6, 6, 1, 0). Because of post-IM dissociations, the IM features corresponding to (n, k) = (7, 7) and (8, 8) can also be labeled. The type I clusters with larger MWs have smaller CCSs than type II clusters. B A partial plot of n vs. CCS showing that the differences get smaller as n increases
Next, the CCS comparison between types II and III clusters gives us a glimpse into the structural transition when the clusters accommodate an additional adenine. For the same (n, k) where n = k, type II clusters contain an “extra” ADE** while type III clusters contain an ADE* (and a Na+). The observed trend remains consistent: despite its greater mass, the type III clusters have about the same CCS as the type II clusters. Collectively, although the growth motif is simple and predictable, this is the first time the stepwise process could be followed and demonstrated unambiguously.
IMS-MS can distinguish the roles of sulfates in the growth of adenine and guanine clusters
Adenine and guanine, both purine bases, share similar shapes and structures, yet they exhibit distinct patterns of hydrogen donors and acceptors. Consequently, the self-assembly of guanine sulfate into small clusters and the involvement of sulfate ions in this process are anticipated to differ significantly from those of adenine sulfate. Notably, there have not been any reports on the crystal structure of guanine sulfate. However, substituting guanine with isoguanine yields a crystal structure similar to adenine sulfate [82]. Thus, guanine sulfate serves as an excellent model system for negative-control experiments, illustrating the distinct roles of sulfate in adenine sulfate crystallization compared to guanine.
As shown in Fig. 5, the mass spectrum of guanine sulfate exhibits numerous peaks corresponding to (n, k, x), where n represents the guanine cluster size, k is the number of sulfate groups, and x denotes the number of protons (H+). Notably, no significant sodiated or potassiated adducts are observed. Successive additions of sulfate groups to the same guanine clusters are evident. For instance, within the inset of Fig. 5, transitions from (6, 3, 1) to (6, 4, 1) and from (14, 3, 2) to (14, 4, 2) and (14, 5, 2), as well as (13, 5, 2) to (13, 6, 2) and (13, 7, 2) are observed. The ATDs become more intense and well-resolved as the guanine to sulfate ratio approaches 2:1 (see Figure S3).
A Representative ESI-mass spectrum of guanine sulfate. Most mass spectral peaks can be assigned to protonated clusters of (n, k) where n is the number of guanine and k is the number of H2SO4. The inset shows the compositions of some complexes where sulfates are sequentially added to the clusters of guanines without following a specific ratio. B Representative 2D plot of m/z vs. drift time (ms) showing species with overlapping isotopic distributions
Figure 6 compares the clusters of adenine and guanine observed in IMS-MS experiments. Large clusters of adenines are detected, reaching sizes of up to n = 42, likely constrained only by the mass range limitations of our MS method. In contrast, smaller clusters of guanines are detected, suggesting that the self-assembly of guanine (in the presence of sulfate) is slower and may not progress to the nucleation stage for crystal formation. Therefore, Figs. 5 and 6 collectively illustrate a clear distinction in adenine assemblies and potential crystal growth behaviors compared to guanine under the influence of sulfate ligands.
Cluster compositions of A guanine sulfate and B adenine sulfate. The clusters are classified based on the charge states and types. The ratio of ADE to sulfate for adenine is close to 1:1, whereas that of GUA to sulfate is broadly distributed. This indicates that sulfate is essential to the stabilities of the adenine clusters. In contrast, sulfate is more important as a charge carrier for detecting GUA clusters than for their structures and stabilities
Finally, Fig. 7 presents the plot of the experimental CCS of adenine clusters plotted against cluster size (n). This offers an indirect insight into the assembly and growth of adenine clusters into single crystals at an early stage. Initially, the CCS values exhibit isotropic growth [45], evidenced by a rapid increase from n = 2 to n = 8. Subsequently, the rate of change slows down, and a new trend emerges around n = 10. Starting at n = 10, the CCS trend begins unidirectional growth, allowing linear fitting. This linear fit yields an R2 value of 0.9828 and a y-intercept of 325.1 Å2. Notably, this value intriguingly aligns with the experimental CCS of a hexamer to an octamer (n = 6 to 8, CCS = 320.0 to 353.5 Å2). Furthermore, it is noteworthy that the unit cell of adeninediium sulfate comprises precisely eight copies of ADE** + SO42−. Collectively, the CCS data from IMS-MS measurements suggest that (a) the nucleation for the crystal formation of adenine sulfate occurs at n = 8 to 14, where there are reversible changes from disordered (isotropic) to ordered (linear) growth, (b) linear, unidirectional growth of adenine sulfate clusters is observed for n > 10.
Conclusion
Capturing the emergence of a crystal nucleus from disordered states is the most challenging aspect of the crystallization process due to the short length and time scales involved. These events are often probed by liquid-cell or low-dose focal-series TEM. Here, we demonstrate that IMS-MS can provide a complementary view of this process by probing the sizes and shapes of small clusters (n = 2 to 40). Interestingly, this method illustrates the crucial role of sulfate in the growth of adeninediium sulfate clusters leading up to single crystals. The nucleation size is predicted to be between n = 8 and 12, aligning well with the unit cells of the adenine crystal. We also highlight the strengths and weaknesses of the method, including data analysis and interpretation. Due to the speed and robustness of IMS-MS experiments, this technique can be used to study the effects of different additives on the assembly of small molecules, elucidate nucleation processes, and investigate the growth of various crystal polymorphs.
It is important to address some methodological weaknesses in this study. Firstly, while we anticipate that all three types of clusters will grow to similar sizes, only a fraction of the mass spectral peaks from each type can be resolved and unambiguously analyzed. For example, the type I clusters depicted in Fig. 7 abruptly end at ~ n = 11. Although evidence for significantly larger clusters of this type can be clear, they are not well-resolved, leading to their exclusion from the final analysis. Figure S4 illustrates this issue. Another factor limiting the detection of larger clusters is the mass range, which is capped at m/z 3200. This restriction biases the results by potentially excluding large, low-charge clusters outside this range.
Secondly, our measurements are conducted in the gas phase, which may not fully preserve the molecular interactions present in the condensed phase. Consequently, the CCS values obtained here cannot be directly compared to those derived from X-ray crystallography. Nonetheless, IMS-MS presumably ionizes clusters formed in solution, and the relative intensities of the mass spectral peaks reflect the abundance of the clusters in solution. Even when a subset of these clusters is promoted by desolvation, the interactions (i.e., sizes and shapes of these clusters) still contain relevant structural information about their packing in a solventless environment, resembling that of crystalline materials. Since similar interactions stabilize all clusters, they should undergo comparable structural contraction (compactness) after desolvation. This suggests that despite the gas-phase limitations, the data still offer valuable insights into the relative stabilities and abundances of the clusters in the condensed phase.
References
Do TD, Kincannon WM, Bowers MT. Phenylalanine oligomers and fibrils: the mechanism of assembly and the importance of tetramers and counter ions. J Am Chem Soc. 2015;137(32):10080–3.
Do TD, de Almeida NE, LaPointe NE, Chamas A, Feinstein SC, Bowers MT. Amino acid metaclusters: implications of growth trends on peptide self-assembly and structure. Anal Chem. 2016;88(1):868–76.
Sade Yazdi D, Laor Bar-Yosef D, Adsi H, Kreiser T, Sigal S, Bera S, et al. Homocysteine fibrillar assemblies display cross-talk with Alzheimer’s disease beta-amyloid polypeptide. Proc Natl Acad Sci U S A. 2021;118(24). https://doi.org/10.1073/pnas.2017575118.
Oluwatoba DS, Chakraborty P, Bar-Yosef DL, Limbach MN, Gazit E, Do TD. Self-assembly of cysteine into nanofibrils precedes cystine crystal formation: implications for aggregation inhibition. ACS Appl Mater Interfaces. 2023;15(27):32177–87.
Gazit E. Metabolite amyloids: a new paradigm for inborn error of metabolism disorders. J Inherit Metab Dis. 2016;39:483–8.
Shaham-Niv S, Adler-Abramovich L, Schnaider L, Gazit E. Extension of the generic amyloid hypothesis to nonproteinaceous metabolite assemblies. Science Adv. 2015;1(7):e1500137.
Laor D, Sade D, Shaham-Niv S, Zaguri D, Gartner M, Basavalingappa V, et al. Fibril formation and therapeutic targeting of amyloid-like structures in a yeast model of adenine accumulation. Nat Commun. 2019;10(1):62.
Tavassoly O, Sade D, Bera S, Shaham-Niv S, Vocadlo DJ, Gazit E. Quinolinic acid amyloid-like fibrillar assemblies seed alpha-synuclein aggregation. J Mol Biol. 2018;430(20):3847–62.
Zaguri D, Shaham-Niv S, Naaman E, Mimouni M, Magen D, Pollack S, et al. Induction of retinopathy by fibrillar oxalate assemblies. Commun Chem. 2020;3(1):2.
Anand BG, Prajapati KP, Dubey K, Ahamad N, Shekhawat DS, Rath PC, et al. Self-assembly of artificial sweetener aspartame yields amyloid-like cytotoxic nanostructures. ACS Nano. 2019;13(5):6033–49.
Adler-Abramovich L, Vaks L, Carny O, Trudler D, Magno A, Caflisch A, et al. Phenylalanine assembly into toxic fibrils suggests amyloid etiology in phenylketonuria. Nat Chem Biol. 2012;8(8):701–6.
Nandi S, Mukhopadhyay A, Nandi PK, Bera N, Hazra R, Chatterjee J, et al. Amyloids formed by nonaromatic amino acid methionine and its cross with phenylalanine significantly affects phospholipid vesicle membrane: an insight into hypermethioninemia disorder. Langmuir. 2022;38(27):8252–65.
Shaham-Niv S, Rehak P, Vuković L, Adler-Abramovich L, Král P, Gazit E. Formation of apoptosis-inducing amyloid fibrils by tryptophan. Isr J Chem. 2016;57(7–8):729–37.
Zaguri D, Kreiser T, Shaham-Niv S, Gazit E. Antibodies towards tyrosine amyloid-like fibrils allow toxicity modulation and cellular imaging of the assemblies. Molecules. 2018;23(6):1273.
Ménard-Moyon C, Venkatesh V, Krishna KV, Bonachera F, Verma S, Bianco A. Self-assembly of tyrosine into controlled supramolecular nanostructures. Chem Eur J. 2015;21(33):11681–6.
Kreiser T, Sogolovsky-Bard I, Zaguri D, Shaham-Niv S, Bar-Yosef DL, Gazit E. Branched-chain amino acid assembly into amyloid-like fibrils provides a new paradigm for maple syrup urine disease pathology. Int J Mol Sci. 2023;24(21):15999.
Tiwari OS, Rencus-Lazar S, Gazit E. Advances in self-assembly of metabolite nanostructures: physiology, pathology and nanotechnology. ChemNanoMat. 2022;8(7):e202200055.
Paul A, Jacoby G, Laor Bar-Yosef D, Beck R, Gazit E, Segal D. Glucosylceramide associated with Gaucher disease forms amyloid-like twisted ribbon fibrils that induce alpha-synuclein aggregation. ACS Nano. 2021;15(7):11854–68.
Levkovich SA, Gazit E, Laor B-Y. The metabolostasis network and the cellular depository of aggregation-prone metabolites. Angew Chem Int Ed Engl. 2023;62(38):e202217622.
Procaccini C, Santopaolo M, Faicchia D, Colamatteo A, Formisano L, de Candia P, et al. Role of metabolism in neurodegenerative disorders. Metabolism. 2016;65(9):1376–90.
Shen C, Liu C, Qiu A. Metabolism-related brain morphology accelerates aging and predicts neurodegenerative diseases and stroke: a UK Biobank study. Transl Psychiatry. 2023;13(1):233.
Mishra MK, Kelley SP, Smetana V, Dixon DA, McNeill AS, Mudring AV, et al. Crystallographic evidence of Watson-Crick connectivity in the base pair of anionic adenine with thymine. Proc Natl Acad Sci U S A. 2020;117(31):18224–30.
Verma S, Mishra AK, Kumar J. The many facets of adenine: coordination, crystal patterns, and catalysis. Acc Chem Res. 2010;43(1):79–91.
Vijayakanth T, Dasgupta S, Ganatra P, Rencus-Lazar S, Desai AV, Nandi S, et al. Peptide hydrogen-bonded organic frameworks. Chem Soc Rev. 2024;53(8):3640–55.
Wang Y, Rencus-Lazar S, Zhou H, Yin Y, Jiang X, Cai K, et al. Bioinspired amino acid based materials in bionanotechnology: from minimalistic building blocks and assembly mechanism to applications. ACS Nano. 2024;18(2):1257–88.
Wang Y, Liu S, Li L, Li H, Yin Y, Rencus-Lazar S, et al. Manipulating the piezoelectric response of amino acid-based assemblies by supramolecular engineering. J Am Chem Soc. 2023;145(28):15331–42.
Tiwari OS, Aizen R, Meli M, Colombo G, Shimon LJW, Tal N, et al. Entropically-driven co-assembly of l-histidine and l-phenylalanine to form supramolecular materials. ACS Nano. 2023;17(4):3506–17.
Shipps C, Kelly HR, Dahl PJ, Yi SM, Vu D, Boyer D, et al. Intrinsic electronic conductivity of individual atomically resolved amyloid crystals reveals micrometer-long hole hopping via tyrosines. Proc Natl Acad Sci U S A. 2021;118(2). https://doi.org/10.1073/pnas.2014139118.
Ogata AF, Rakowski AM, Carpenter BP, Fishman DA, Merham JG, Hurst PJ, et al. Direct observation of amorphous precursor phases in the nucleation of protein-metal-organic frameworks. J Am Chem Soc. 2020;142(3):1433–42.
Biran I, Rosenne S, Weissman H, Tsarfati Y, Houben L, Rybtchinski B. Organic crystal growth: hierarchical self-assembly involving nonclassical and classical steps. Cryst Growth Des. 2022;22(11):6647–55.
Biran I, Houben L, Weissman H, Hildebrand M, Kronik L, Rybtchinski B. Real-space crystal structure analysis by low-dose focal-series TEM imaging of organic materials with near-atomic resolution. Adv Mater. 2022;34(26):e2202088.
Tsarfati Y, Biran I, Wiedenbeck E, Houben L, Colfen H, Rybtchinski B. Continuum crystallization model derived from pharmaceutical crystallization mechanisms. ACS Cent Sci. 2021;7(5):900–8.
Houben L, Weissman H, Wolf SG, Rybtchinski B. A mechanism of ferritin crystallization revealed by cryo-STEM tomography. Nature. 2020;579(7800):540–3.
Mashiach R, Weissman H, Avram L, Houben L, Brontvein O, Lavie A, et al. In situ NMR reveals real-time nanocrystal growth evolution via monomer-attachment or particle-coalescence. Nat Commun. 2021;12(1):229.
Jeon S, Heo T, Hwang SY, Ciston J, Bustillo KC, Reed BW, et al. Reversible disorder-order transitions in atomic crystal nucleation. Science. 2021;371(6528):498–503.
Nakamuro T, Sakakibara M, Nada H, Harano K, Nakamura E. Capturing the moment of emergence of crystal nucleus from disorder. J Am Chem Soc. 2021;143(4):1763–7.
Nielsen MH, Aloni S, De Yoreo JJ. In situ TEM imaging of CaCO3 nucleation reveals coexistence of direct and indirect pathways. Science. 2014;345(6201):1158–62.
Zimmermann NE, Vorselaars B, Quigley D, Peters B. Nucleation of NaCl from aqueous solution: critical sizes, ion-attachment kinetics, and rates. J Am Chem Soc. 2015;137(41):13352–61.
Chakraborty D, Patey GN. How crystals nucleate and grow in aqueous NaCl solution. J Phys Chem Lett. 2013;4(4):573–8.
Finney AR, Salvalaglio M. Multiple pathways in NaCl homogeneous crystal nucleation. Faraday Discuss. 2022;235:56–80.
Wyttenbach T, Bowers MT. Structural stability from solution to the gas phase: native solution structure of ubiquitin survives analysis in a solvent-free ion mobility–mass spectrometry environment. J Phys Chem B. 2011;115(42):12266–75.
Morris CB, Poland JC, May JC, McLean JA. Fundamentals of ion mobility-mass spectrometry for the analysis of biomolecules. Ion Mobility-Mass Spectrometry: Methods and Protocols. Methods Mol Biol. 2020;2084:1–31.
Dodds JN, Baker ES. Ion mobility spectrometry: fundamental concepts, instrumentation, applications, and the road ahead. J Am Soc Mass Spectrom. 2019;30(11):2185–95.
Lanucara F, Holman SW, Gray CJ, Eyers CE. The power of ion mobility-mass spectrometry for structural characterization and the study of conformational dynamics. Nat Chem. 2014;6(4):281–94.
Bleiholder C, Dupuis NF, Wyttenbach T, Bowers MT. Ion mobility-mass spectrometry reveals a conformational conversion from random assembly to beta-sheet in amyloid fibril formation. Nat Chem. 2011;3(2):172–7.
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P. The universal features of cells on Earth. Molecular Biology of the Cell. 4th ed. New York: Garland Science; 2002.
Minchin S, Lodge J. Understanding biochemistry: structure and function of nucleic acids. Essays Biochem. 2019;63(4):433–56.
Matsumoto S, Raivio K, Willis R, Seegmiller J. Interactions between energy metabolism and adenine nucleotide metabolism in human lymphoblasts. Purine Metabolism in Man—III: Biochemical, Immunological, and Cancer Research. 1980:277–82. https://doi.org/10.1007/978-1-4684-8559-2_45.
Denessiouk KA, Rantanen VV, Johnson MS. Adenine recognition: a motif present in ATP-, CoA-, NAD-, NADP-, and FAD-dependent proteins. Proteins. 2001;44(3):282–91.
Cappello V, Tramontano A, Koch U. Classification of proteins based on the properties of the ligand-binding site: the case of adenine-binding proteins. Proteins. 2002;47(2):106–15.
Gazit E. Metabolite amyloids: a new paradigm for inborn error of metabolism disorders. J Inherit Metab Dis. 2016;39(4):483–8.
Fischer A, Notarangelo LD, Neven B, Cavazzana M, Puck JM. Severe combined immunodeficiencies and related disorders. Nat Rev Dis Primers. 2015;1:15061.
Van Acker KJ, Simmonds HA, Potter C, Cameron JS. Complete deficiency of adenine phosphoribosyltransferase. Report of a family. N Engl J Med. 1977;297(3):127–32.
Runolfsdottir HL, Palsson R, Agustsdottir IM, Indridason OS, Edvardsson VO. Long-term renal outcomes of APRT deficiency presenting in childhood. Pediatr Nephrol. 2019;34(3):435–42.
Khan MA, Nag P, Grivei A, Giuliani KTK, Wang X, Diwan V, et al. Adenine overload induces ferroptosis in human primary proximal tubular epithelial cells. Cell Death Dis. 2022;13(2):104.
Kistenmacher TJ, Shigematsu T. Adenine hydrochloride hemihydrate: three-dimensional data and refinement. Acta Cryst. 1974;B30:166–8.
Kistenmacher TJ, Shigematsu T. The crystal structure of adenine dihydrochloride. Acta Cryst. 1974;B30:1528–33.
Schmidt A, Lindner A, Nieger M, Ruiz-Delgado Mdel C, Ramirez FJ. Syntheses, pi-stacking interactions and base-pairings of uracil pyridinium salts and uracilyl betaines with nucleobases. Org Biomol Chem. 2006;4(16):3056–66.
Moravcova H. Crystal Data for Some Salts of Adenine. Z Kristallogr. 1975;141(1–2):156–7.
Cheng YJ, Wang ZM, Liao CS, Yan CH. Novel sheet-like supramolecular architectures constructed from infinite hydrogen-bonded, protonated adenine water halide and polyiodide ribbons. New J Chem. 2002;26(10):1360–4.
Mineki H, Hanasaki T, Matsumoto A, Kawasaki T, Soai K. Asymmetric autocatalysis initiated by achiral nucleic acid base adenine: implications on the origin of homochirality of biomolecules. Chem Commun (Camb). 2012;48(85):10538–40.
Armentano D, Barquero MA, Rojas-Dotti C, Moliner N, De Munno G, Brechin EK, et al. Enhancement of intermolecular magnetic exchange through halogen...halogen interactions in bisadeninium rhenium(IV) salts. Cryst Growth Des. 2017;17(10):5342–8.
Savchenkov A, Demina L, Safonov A, Grigoriev M, Solovov R, Abkhalimov E. Syntheses and crystal structures of new aurate salts of adenine or guanine nucleobases. Acta Crystallogr C Struct Chem. 2020;76(Pt 2):139–47.
Byres M, Cox PJ, Kay G, Nixon E. Supramolecular structures of six adenine-carboxylic acid complexes. CrystEngComm. 2009;11(1):135–42.
Tret’yak SM, Mitkevich VV, Sukhodub LF. Adenine trihydrate: its structure at 295 and 105 K. Kristallografiya. 1987;32:1268–71.
Mahapatra S, Nayak SK, Prathapa S, Guru Row TN. Anhydrous adenine: crystallization, structure, and correlation with other nucleobases. Cryst Growth Des. 2008;8(4):1223–5.
Stolar T, Lukin S, Pozar J, Rubcic M, Day GM, Biljan I, et al. Solid-state chemistry and polymorphism of the nucleobase adenine. Cryst Growth Des. 2016;16(6):3262–70.
Chen AX, Zito SW, Nash RA. Solubility enhancement of nucleosides and structurally related compounds by complex formation. Pharm Res. 1994;11:398–401.
Adsi H, Levkovich SA, Haimov E, Kreiser T, Meli M, Engel H, et al. Chemical chaperones modulate the formation of metabolite assemblies. Int J Mol Sci. 2021;22(17):9172.
Pinsk N, Wagner A, Cohen L, Smalley CJH, Hughes CE, Zhang G, et al. Biogenic guanine crystals are solid solutions of guanine and other purine metabolites. J Am Chem Soc. 2022;144(11):5180–9.
Wagner A, Upcher A, Maria R, Magnesen T, Zelinger E, Raposo G, et al. Macromolecular sheets direct the morphology and orientation of plate-like biogenic guanine crystals. Nat Commun. 2023;14(1):589.
Hu H, Xue R, Chen F. Biomineralization and properties of guanine crystals. Molecules. 2023;28(16):6138.
Oaki Y, Kaneko S, Imai H. Morphology and orientation control of guanine crystals: a biogenic architecture and its structure mimetics. J Mat Chem. 2012;22(42):22686–91.
Pavan ME, Movilla F, Pavan EE, Di Salvo F, Lopez NI, Pettinari MJ. Guanine crystal formation by bacteria. BMC Biol. 2023;21(1):66.
Zachová J. Growing and morphology of crystals of adenine sulphate. Czechoslovak J Phys B. 1973;23(10):1038–44.
Gabelica V, Livet S, Rosu F. Optimizing native ion mobility Q-TOF in helium and nitrogen for very fragile noncovalent structures. J Am Soc Mass Spectrom. 2018;29(11):2189–98.
Oluwatoba DS, Islam MF, Som B, Sindt AJ, Smith MD, Shimizu LS, et al. Evaluating the effects of metal adduction and charge isomerism on ion-mobility measurements using m-xylene macrocycles as models. J Am Soc Mass Spectrom. 2022;33(5):840–50.
Langer V, Huml K, Lessinger L. The crystal and molecular structure of adeninium hemisulfate hydrate, C5H5N5. 0.5 H2SO4. H2O. Acta Crystallogr B Struct Crystallogr Cryst Chem. 1978;34(7):2229–34.
Hassen CB, Boujelbene M, Mhiri T. Structural determination; vibration study and thermal decomposition of [C5H6N5] 2SeO4⋅ 2H2O. J Mol Struct. 2015;1079:147–54.
Gray AL, Antevska A, Oluwatoba DS, Schonfeld GE, Lazar Cantrell KL, Do TD. Cytotoxicity of α-helical, Staphylococcus aureus PSMα3 investigated by post-ion-mobility dissociation mass spectrometry. Anal Chem. 2020;92(17):11802–8.
Link BA, Sindt AJ, Shimizu LS, Do TD. Selective host-guest chemistry, self-assembly and conformational preferences of m-xylene macrocycles probed by ion-mobility spectrometry mass spectrometry. Phys Chem Chem Phys. 2020;22(17):9290–300.
Subramanian E, Marsh RE. The crystal structure of isoguanine sulfate monohydrate. Acta Cryst. 1971;B27:753–9.
Funding
Eli Lilly and Co. Young Investigator Award in Analytical Chemistry.
Author information
Authors and Affiliations
Contributions
Conceptualization: Thanh D. Do. Methodology: Damilola Oluwatoba, Thanh D. Do. Formal analysis and investigation: Damilola Oluwatoba. Writing—original draft preparation: Damilola Oluwatoba. Writing—review and editing: Thanh D. Do, Happy A. Safoah, Damilola Oluwatoba. Funding acquisition: Thanh D. Do. Resources: Thanh D. Do. Supervision: Thanh D. Do.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
ABC Highlights: authored by Rising Stars and Top Experts.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Oluwatoba, D.S., Safoah, H.A. & Do, T.D. The rise and fall of adenine clusters in the gas phase: a glimpse into crystal growth and nucleation. Anal Bioanal Chem 416, 5037–5048 (2024). https://doi.org/10.1007/s00216-024-05442-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-024-05442-2