Abstract
Urine is an attractive and non-invasive alternative source to tissue, blood or other biofluids for biomarker screening in clinical research. In normal human adult urine, 48% of the total urinary protein is in the sediment, 49% is soluble and the remaining 3% is contained in urinary extracellular vesicles (EVs). The soluble proteins and EV proteins in urine have attracted particular attention in recent years as cancer diagnostics. Furthermore, considering the important role of N-glycoproteins in practically all physiological processes, including regulating receptor-ligand binding, cell-cell interactions, inflammatory response and tumour progression, N-glycoproteome in human urine is an invaluable target for monitoring the physiological status and pathological changes of the kidney and urinary tract. Given the different origins of the soluble proteins and EV proteins in the urine, different N-glycoproteome patterns exist. Therefore, isolating the soluble N-glycoproteins and EV N-glycoproteins for separate analysis will provide a more specific and comprehensive view and provide a deeper understanding of human urinary N-glycoproteome. In this work, we developed a sequential separation method that isolates urinary soluble proteins and EV proteins via stepwise ultrafiltration based on their obvious size difference. A facile and reproducible protein isolation was achieved using this strategy. Subsequent N-glycoproteome enrichment and identification revealed distinct patterns in the two sub-proteomes of urine with more than 60% differential N-glycopeptides. A more comprehensive picture of the urinary N-glycoproteome with close to 1800 identified N-glycopeptides was obtained by this new analysis strategy, therefore making it advantageous for urinary biomarker screening.

A sequential separation method that isolates urinary soluble proteins and EV proteins via stepwise ultrafiltration was developed in this work. Subsequent N-glycopeptides enrichment and mass spectrometry analysis reveals distinct N-glycoproteome patterns in the two sub-proteomes of urine and a deep mapping of close to 1800 N-glycopeptides.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Urine is a combined product of glomerular filtration of plasma and excretion from epithelial cells of the urinary system [1]. In addition to proteins and metabolites, extracellular vesicles are also important contents in the urine. In normal human urine, 48% of the total urinary protein is in the sediment, 49% is soluble and the remaining 3% is in the urinary extracellular vesicles [2]. Since the soluble proteins in the urine are mainly derived from glomerular filtration, alteration of the proteome profile of the soluble proteins reflects changes in the blood and the function variation of the glomerular filter system [3, 4]. Therefore, these proteins have been widely studied in physiological and pathological status monitoring, as biomarkers for screening for an early diagnosis of cancer, cancer progression and therapeutic response assessment [5, 6].
Urinary extracellular vesicles (EVs) were first reported in 2004 [7] and have been extensively studied for clinical applications in recent years. EVs are cell-secreted nanometre to micrometre-sized lipid-bilayer-enclosed vesicles that incorporate various components (proteins, lipids and nucleic acids) [8]. Their composition varies depending on their cell type and origin, making them a potential source of urinary biomarkers and therapeutic reagents for many diseases, such as renal diseases, diabetes and carcinomas [9,10,11]. The important role of EVs in cell-cell communication has also gained increasing attention, which may be attributed to the various glycoproteins on their surface [12, 13].
Glycosylation, especially N-glycosylation, is one of the most important and prevalent protein post-translational modifications (PTMs) [14,15,16] and is vital for a number of physiological processes, such as cell adhesion, intercellular communication and immune response [17,18,19]. In clinical studies, there is increasing evidence demonstrating the role of aberrant glycosylation in inflammation, tumour formation and metastasis [20, 21]. Therefore, studies on the glycoproteome of the soluble proteins and the EV proteins in human urine may provide a facile way to monitor the physiological and pathological changes of the urinary system [22, 23]. Although the glycoprotein profiles of the soluble proteins and EV proteins in urine have been studied before, no direct comparison was reported [24,25,26,27]. Considering the different origins of the two sub-proteomes in urine [28] and the recent discovery of distinct protein profiles of the cell type-resolved and region-specific proteomes [29, 30], we expect a diverse N-glycoproteome composition may exist in the two sub-proteomes. Therefore, methods that are capable of facilely isolating both urinary soluble protein and EV proteins are urgently needed to gain a more comprehensive view of disease-related N-glycoproteins in urine.
Due to the dilute concentration of soluble protein and low abundance of EV proteins in human urine (3%), various isolation/enrichment strategies have been reported based on the physicochemical properties of the urinary protein and EVs, such as acetone precipitation [31] for soluble protein extraction, ultracentrifugation [32, 33], and ultrafiltration [34, 35] for EVs isolation. Although acetone precipitation and ultracentrifugation are commonly adopted, these techniques are labour-intensive with low throughput. Twelve hours are needed when combining the 2 strategies for isolation of both soluble proteins and EV proteins due to the prolonged centrifugation and cold precipitation steps, which is not suitable for large-scale processing of clinical samples in precision medicine research.
A promising alternative for fast EVs isolation is ultrafiltration. Cheruvanky et al. reported the successful purification of EVs using nanomembrane concentrators within 10–30 min with high isolation efficiency [36], although the valuable soluble proteins in the supernatant were discarded in this strategy.
To obtain a more comprehensive view of the N-glycoproteome profile in human urine, we developed a sequential isolation strategy to separately extract urinary EV proteins and soluble proteins by stepwise ultrafiltration using 300 kDa (MWCO) and 10 kDa (MWCO) centrifugal filter devices, respectively. In this strategy, urine EVs are retained and recovered by the 300 kDa (MWCO) centrifugal filter device, due to their large size. The soluble proteins in the flow-through solution are retained by the 10 kDa (MWCO) centrifugal filter with similar recovery rates compared to the use of acetone precipitation. In this way, both the soluble proteins and EV proteins in the urine can be obtained easily within 60 min, which is more than ten times faster than using conventional methods. The obtained proteins are digested and then enriched by hydrophilic interaction liquid chromatography (HILIC) [37] for N-glycopeptides enrichment and identification by mass spectrometry. Highly reproducible N-glycoprotein identification (> 80%) can be achieved for both the soluble proteins and the EV proteins. However, distinct N-glycoproteome patterns are found in the two sub-proteomes with only ~ 50%/40% overlap in N-glycoprotein/N-glycopeptide identification. Therefore, an obviously increased N-glycoproteome identification scale was achieved with close to 1800 N-glycopeptides by combined analysis of the 2 sub-proteomes. The presented results have demonstrated the feasibility and advantages of using our sequential isolation strategy for comprehensive N-glycoproteome analysis of human urine.
Materials and methods
Vivaspin 20,300 kDa and Amicon® Ultra 10 kDa devices were purchased from Sartorius Stedim Biotech GmbH (Goettingen, Germany) and Merck (Darmstadt, Germany). Anti-TSG101 antibody and Anti-AQP2 antibody were obtained from Abcam (Cambridge, MA, USA). Total Exosome Isolation kit (from urine), Pierce™ BCA Protein Assay Kit and Pierce™ ECL Western Blotting Substrate were purchased from ThermoFisher scientific (Shanghai, China). Sequencing grade trypsin was obtained from Promega (Madison, WI, USA). H218O (97% 18O) was purchased from Chemical Industry Research Institute (Shanghai, China). Peptide-N-glycosidase F (PNGase F), acetone and protease inhibitor cocktail were obtained from Sigma-Aldrich (St. Louis, MO, USA). All of the solutions were prepared with ultrapure water from an ELGA DV25 purification system (UK).
Urine sample collection
First-morning void urine (midstream) samples from six healthy volunteers (three men and three women) with no known medical conditions were collected under the approval of the Ethics Committee of Beijing Proteome Research Centre.
Isolation of urinary EVs and soluble proteins by ultrafiltration
After adding a protease inhibitor cocktail, ten millilitres of fresh urine samples was centrifuged at 12,000 g under 4 °C for 20 min to remove large cellular debris and polymers of the abundant Tamm–Horsfall glycoprotein (THP). After that, the supernatant of the urine sample was transferred to a 300 kDa (MWCO) centrifugal ultrafiltration device and was centrifuged at 8000 g for 10 min to collect EVs. The flow-through solution was subsequently transferred to a 10 kDa (MWCO) ultrafiltration device and was centrifuged at 5000 g for 10 min to obtain the soluble proteins. Next, the 300 kDa ultrafiltration device was washed twice with 1 × PBS to remove residual proteins that had adsorbed on the EVs and the filter membrane. The washing buffer was also transferred to the 10 kDa ultrafiltration device for soluble protein collection. Finally, 200 μL lysis buffer (8 M urea, 100 mM Tris-HCl pH = 8.0) was added to the 300 kDa filter devices. After shaking, the solution was transferred to a 1.5 mL centrifuge tube and subjected to ultrasound for 10 min in an ice-bath for EVs lysis. An enzyme-friendly surfactant (0.1% w/v RapiGest) was used to assist in-solution protein digestion. The soluble protein was directly recovered by adding lysis buffer to the 10 kDa ultrafiltration device.
Isolation of EVs with commercial precipitation kits
Briefly, a 10-mL urine sample was mixed with the same volume of the Total Exosome Isolation reagent and incubated at room temperature for 1 h. Next, the mixture was centrifuged at 10,000×g for 1 h at 4 °C. The exosome pellets on the bottom of the tubes were collected after removing the supernatant.
Characterisation of the EVs isolated by ultrafiltration
EVs suspended in PBS were deposited on formvar/carbon-coated copper/palladium grids for whole-mount analysis. A mixture of methylcellulose and uranyl acetate was embedded. Transmission electron microscopy (TEM) characterisation was carried out using a JEM-2100F (Akishima-shi, Japan) operating at 200 kV. Vesicle size was assessed by the iTEM analysis software (Soft Imaging Systems, Japan). Next, the EVs were diluted in 1 × PBS and loaded into the cell of a nanoparticle tracking analysis (NTA) device. The sizes and concentrations of the sample were measured by an NTA ZetaView® (Meerbusch, Germany). A known sample containing nanoparticles with an average size of 100 nm was used to calibrate the instrument prior to sample reading. The obtained data were analysed using ZetaView 8.02.28 as indicated by the manufacturer.
Western blotting
Equal amounts of proteins (20 μg) measured using a Pierce™ BCA Protein Assay Kit were separated by SDS-PAGE before being transferred to a PVDF membrane (Millipore, USA). The membrane was blocked in a dilute solution of protein-5% w/v bovine serum albumin (BSA) in TBST (0.02% w/v Tween-20, 150 mM NaCl, 10 mM Tris pH 8.0) for 1 h at room temperature (RT). The membrane was then incubated with the anti-TSG101 antibody or anti-AQP2 antibody at a 1:1000 dilution in the blocking buffer (TBST, 0.5% w/v BSA) at 4 °C overnight. After washing with TBST three times, the membrane was incubated with horseradish peroxidase (HRP)-conjugated secondary antibody at a 1:2000 dilution for 1 h in the blocking buffer at RT. After washing three times with TBST, the membrane was treated with ECL reagents and imaged by a LAS-500 instrument (GE Healthcare Sciences, USA).
Acetone precipitation of urine proteins
Fresh urine samples were centrifuged to remove any large cellular debris after adding a protease inhibitor cocktail as described in the previous part. The supernatants were collected and three volumes of ice-cold acetone were added. After incubation at − 20 °C overnight, the samples were centrifuged at 14,000 g for 15 min at 4 °C. The supernatants were removed and the obtained pellets were allowed to air dry. Finally, the protein pellets were re-suspended in 200 μL lysis buffer (8 M urea, 100 mM Tris-HCl pH = 8.0).
Protein digestion and N-glycopeptide enrichment
The soluble and EV proteins were digested by trypsin using the FASP protocol [38] after protein reduction and alkylation with dithiothreitol and iodoacetamide. The digested peptides were lyophilized under vacuum and stored at − 80 °C. Triblock hydrophilic copolymer-grafted silica microparticles prepared using our previously reported method were used for HILIC-based N-glycopeptides enrichment [37]. In brief, 2 mg of HILIC materials were sequentially washed with 100 μL of 0.5% v/v formic acid (FA) for 10 min and equilibrated with 100 μL of binding buffer (acetonitrile (ACN):water:FA = 60:35:5, v/v/v 10 mM Ca2+) for 1 h. The HILIC materials were mixed with the peptide samples reconstituted in the binding buffer with further gentle rotation mixing for 2 h. After washing with 80 μL × 5 binding buffer, the enriched glycopeptides were eluted with 0.5% v/v FA followed by 5% ACN containing 0.5% v/v FA. Finally, the combined elution was dried using an Eppendorf concentrator. The enriched N-glycopeptides were re-suspended in 20 μL of 50 mM ammonium bicarbonate in H218O (pH 8.0) containing 1 μL (100 U) of PNGase F and incubated at 37 °C for 16 h for de-N-glycosylation and 18O labelling. Finally, the sample was dried and re-suspended in 18 μL of 0.1% v/v FA for LC-MS/MS analysis.
Mass spectrometry analysis and database searching
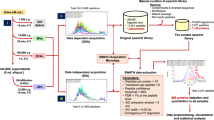
The resulting peptide mixture was analysed by a Q-Exactive™ HF mass spectrometer (ThermoFisher Scientific, USA) coupled with an EASY-nLC 1000 system through a nanoelectrospray ion source. Separation was accomplished by a 120-mm-long column (100 μm internal diameter) packed with C18 packing materials (1.9 μm diameter with a pore size of 100 Å). A linear gradient of A and B buffers (buffer A = 0.1% v/v FA; buffer B = 95% v/v ACN, 0.1% v/v FA) from 6 to 32% buffer B for 78 min at a flow rate of 0.6 μL/min was used to elute the peptides into the mass spectrometer. All of the MS and MS2 spectra were acquired using data-dependent acquisition (DDA) mode and the mass spectrometer was set to a scan range of 300–1400 m/z followed by a top 20 data-dependent MS/MS scans. Raw files were searched using MaxQuant version 1.5.2.8 against the Uniprot Human Protein Database (release 20150729, 2015; 20,203 entries) with a tolerance level of 6 ppm for MS and 20 ppm for MS/MS. Enzyme specificity was set as trypsin, a minimum of seven amino acids per identified peptide was required and two missed cleavages were allowed. Methionine oxidation (+ 15.994 Da), deamidation (+ 2.988 Da) (for N-glycopeptide identification) and acetylation (+ 42.010 Da) of protein N-terminals were set as variable modifications and carbamidomethylation of cysteine (+ 57.021 Da) was set as a fixed modification. A 1% false discovery rate (FDR) was used at the peptide-to-spectrum match (PSM) and protein level. The minimum and delta scores were set as 40 and 8, respectively. N-glycopeptides containing the motif (N-X-T/S/C; X ≠ P) and 18O labelling were screened out.
Results and discussion
Sequential ultrafiltration for facile isolation of urinary EVs and soluble protein for comprehensive N-glycoproteome analysis
To study the diversity of N-glycoproteome of the urinary EVs and soluble proteins, we developed a sequential isolation strategy using stepwise ultrafiltration based on the obvious size difference between the EVs and soluble proteins (Fig. 1). According to the working principle, after removing the cell debris and polymer proteins, the urine sample was processed by 300 kDa MWCO (pore size < 15 nm) filter devices, which retain the urinary EVs and allow the soluble urine proteins to pass through. Next, the soluble proteins in the flow-through are retained by the 10 kDa MWCO filter devices. In this way, the EV (proteins) and soluble proteins in the urine can be easily separated and recovered for subsequent N-glycoproteome analysis.
Characterisation of the EVs obtained by ultrafiltration
Morphology and the size of the EVs obtained by ultrafiltration were characterised by transmission electron microscopy (TEM) and nanoparticle tracking analysis (NTA). TEM images showed that the EVs (boxed in the figure) were roughly spherical with a diameter of approximately 100 nm and presented with a cup-shaped morphology (Fig. 2a), indicating that the structure of the EVs were intact after isolation by ultrafiltration. NTA analysis showed that > 70% of the obtained EVs were within a size range of 30–150 nm (Fig. 2c), which is consistent with the reported size of the exosomes [39], while the rest of the EVs with a size > 150 nm are microvesicles. Protein markers, such as AQP2 and TSG101 in the EVs, were obtained from three technical replicates of ultrafiltration and were analysed by Western blotting (Fig. 2d). TSG101 and AQP2 were observed at the 50 and 30 kDa molecular weight band. The band shape and grey-scale value were very similar among the three tests, indicating that the ultrafiltration-based EVs isolation is highly reproducible.
To further evaluate the efficiency and purity of the EVs isolated by ultrafiltration, we compared the proteins obtained from the EVs isolated by ultrafiltration to the commercial precipitation kits. The same amount of urine and protein lysates was used. As shown in the results of Western blotting in Fig. S1 (see Electronic Supplementary Material (ESM)), no obvious differences were found between the bands of TSG101 and AQP2 obtained by ultrafiltration and the precipitation kit. Next, proteome analysis was conducted to characterise the marker proteins of the EVs obtained by ultrafiltration in a large-scale experiment. The experiments were repeated three times and the identified proteins were compared with the 100 most common EV proteins (Top100 EV proteins) reported in EVpedia and Vesiclepedia [40]. As shown in Table 1, at least 93 proteins of the Top100 EV proteins were identified in the fraction from the 300 kDa filter device. In contrast, only 11 proteins were discovered in the fraction from the 10 kDa filter device, suggesting that EVs can be successfully retained and recovered by the 300 kDa filter device. Further comparison with the commercial precipitation kit in Fig. S2a (see ESM) shows that 75.2% (583) of the 775 protein groups identified by the precipitation kit are covered by our ultrafiltration method (the EV fraction obtained by the 300 kDa filter device). GO analysis of the co-identified protein groups by the two methods (583) and the EV protein groups identified only by our ultrafiltration method (513) shows similarly enriched GO terms that are closely associated with vesicle, extracellular regions and membrane (Fig. S2b, c, see ESM). The above results indicate that our method not only covers the EV protein groups obtained by the precipitation kit but also provides deeper proteome coverage.
Recovery of the soluble proteins in urine
Currently, acetone precipitation is the most commonly adopted technique for urinary soluble protein extraction, although the whole process takes 6 h to overnight. Our sequential ultrafiltration-based strategy, which takes only 1 h to complete, has obvious advantages in sample process throughput. To evaluate the protein isolation efficiency, we added a fluorescent protein (BSA-FRITC) into urine samples and divided them into two for a comparison between the ultrafiltration and acetone precipitation strategies. The molecular mass of BSA-FRITC is 67 kDa, and therefore, it flows through the 300 kDa filter and is retained by the 10 kDa filter. The fluorescence intensity of BSA-FRITC measured by fluorescence spectrometer shows that the recovery of the ultrafiltration strategy is 76.8% and is a little higher than that of the acetone precipitation strategy (71.4%) as shown in Fig. S3 (see ESM), demonstrating the feasibility of using this strategy for fast urinary soluble protein extraction.
Application of the sequential ultrafiltration strategy for study of the urinary N-glycoproteome
Reproducibility of the sequential ultrafiltration strategy was evaluated at the N-glycoproteome level by HILIC enrichment of the tryptic digest of the urinary EVs and soluble proteins. Figure 3a, b and Table S2 (see ESM) show good repeatability with 88.3 and 82.5% overlaps for EV N-glycoprotein groups and N-glycopeptide identification in at least two technical replicates. Similar results were obtained for the soluble proteins since a 93.2 and 91.5% overlap were achieved for the N-glycoprotein groups and N-glycopeptide identification in at least two technical replicates (Fig. 3c, d, ESM Table S3). Good quantification reproducibility was also obtained in three tests with a Pearson correlation coefficient ranging from 0.80 to 0.94 (ESM, Fig. S4 and Table S4).
The reproducibility of N-glycoproteome identification of the urinary EVs and soluble proteins. a, b The Venn diagrams showing the overlap of the EV N-glycoproteins groups/N-glycopeptides obtained in three technical replicates. c, d Exhibit overlap of the identified N-glycoproteins groups/N-glycopeptides from the soluble proteins obtained in three technical replicates
Diversity study of the N-glycoproteome in urinary EVs and soluble proteins
As we hypothesised, diverse N-glycoproteome profiles exist in the EVs and soluble proteins in human urine due to their different origins. Separating the low abundant EV proteins (which account for only 3% of the total protein mass in the urine) from the highly abundant soluble proteins for individual N-glycoproteome analysis may lead to enhanced identification sensitivity and a more comprehensive view of the human urinary N-glycoproteome. Table S1 (see ESM) shows the summary of the identified N-glycopeptides in the EVs and the soluble proteins from six healthy volunteers. In total, we identified 1210 and 1256 N-glycopeptides assigned to 586 and 619 N-glycoprotein groups in EVs and soluble proteins, respectively. As expected, approximately 50 and 60% (the calculating equation was listed on Page S4, ESM) of the N-glycoprotein groups and N-glycopeptides were only identified in either EVs or soluble proteins (Fig. 4a, b, ESM Table S5). Since around 80% reproducibility can be achieved using this sequential ultrafiltration strategy with tandem HILIC enrichment, we attribute the distinct N-glycoproteome profiles between the EVs and soluble proteins to their differences in N-glycoprotein group composition, which may be potentially important for future application in biomarker studies. To provide an overall picture of the functional role of the N-glycoproteins identified in urine, gene ontology (GO) annotation analysis was conducted using the bioinformatics tool DAVID [41]. All of the identified N-glycoproteins were mainly located in extracellular exosomes, the extracellular space and the plasma membrane, which is consistent with the reported cellular component of N-glycoproteome [42]. The top enriched molecular functions include calcium ion binding, heparin binding and receptor activity. In the category of biological processes, cell adhesion, extracellular matrix organisation and haemophilic cell adhesion constitute a large proportion of the identified N-glycoproteins. For N-glycoproteins identified both in the EVs and in the soluble fraction, though the majority of the enriched GO terms are similar to those obtained using all of the identified N-glycoproteins, differences are found. Lysosomal lumen, collagen binding, receptor binding, negative regulation of endopeptidase activity and leukocyte migration are only enriched in the N-glycoproteins identified both in the EVs and the soluble fraction (Fig. 5).
Pie chart showing the overlap of the N-glycoproteome of the urinary EV proteins and soluble proteins. a Overlap of the identified N-glycoprotein groups obtained from the urinary EV proteins and soluble proteins. b Overlap of the identified N-glycopeptides obtained from the urinary EV proteins and soluble proteins. (green circle: the identified N-glycoprotein groups or N-glycopeptides of the urinary EV proteins, black circle: the identified N-glycoprotein groups or N-glycopeptides of the soluble proteins)
Gene ontology classification of the identified glycoproteins. Lists of the top five enriched terms associated with cellular component, molecular function, biological process. a Gene ontology classification of the identified N-glycoproteins in urine. b Gene ontology categories of the glycoproteins identified in both EVs and in soluble urine proteins
Conclusion
In this work, we developed a sequential ultrafiltration strategy for facile isolation of urinary EVs (proteins) and soluble proteins. A greater than tenfold shortened sample processing time was achieved while maintaining a similar EV and soluble protein extraction efficiency. Isolating the low abundant EV proteins is beneficial for more sensitive N-glycoproteome analysis since interference from the high abundant urinary soluble proteins is avoided. Distinct N-glycoproteome profiles are found in the urinary EV proteins and soluble proteins, indicating the necessity of analysing both sub-proteomes to obtain a more comprehensive view of the human urinary N-glycoproteome.
References
Brown N, Segev G, Francey T, Kass P, Cowgill L. Glomerular filtration rate, urine production, and fractional clearance of electrolytes in acute kidney injury in dogs and their association with survival. J Vet Intern Med. 2015;29(1):28–34.
Moon PG, You S, Lee JE, Hwang D, Baek MC. Urinary exosomes and proteomics. Mass Spectrom Rev. 2011;30(6):1185–202.
Zoja C, Abbate M, Remuzzi G. Progression of renal injury toward interstitial inflammation and glomerular sclerosis is dependent on abnormal protein filtration. Nephro Dial Transpl. 2014;30(5):706–12.
Pontillo C, Jacobs L, Staessen JA, Schanstra JP, Rossing P, Heerspink HJ, et al. A urinary proteome-based classifier for the early detection of decline in glomerular filtration. Nephro Dial Transpl. 2016;32(9):1510–6.
Mok CC, Soliman S, Ho LY, Mohamed FA, Mohamed FI, Mohan C. Urinary angiostatin, CXCL4 and VCAM-1 as biomarkers of lupus nephritis. Arthritis Res Ther. 2018;20(1):6–16.
Leyten GH, Hessels D, Jannink SA, Smit FP, de Jong H, Cornel EB, et al. Prospective multicentre evaluation of PCA3 and TMPRSS2-ERG gene fusions as diagnostic and prognostic urinary biomarkers for prostate cancer. Eur Urol. 2014;65(3):534–42.
Pisitkun T, Shen RF, Knepper MA. Identification and proteomic profiling of exosomes in human urine. Proc Natl Acad Sci U S A. 2004;101(36):13368–73.
Gámez-Valero A, Lozano-Ramos SI, Bancu I, Lauzurica-Valdemoros R, Borràs FE. Urinary extracellular vesicles as source of biomarkers in kidney diseases. Front Immunol. 2015;6(6):1–10.
Dear JW, Street JM, Bailey MA. Urinary exosomes: a reservoir for biomarker discovery and potential mediators of intrarenal signalling. Proteomics. 2013;13(10–11):1572–80.
Zhang Y, Wang X-F. A niche role for cancer exosomes in metastasis. Nat Cell Biol. 2015;17(6):709–11.
Buzas EI, György B, Nagy G, Falus A, Gay S. Emerging role of extracellular vesicles in inflammatory diseases. Nat Rev Rheumatol. 2014;10(6):356–64.
Huang-Doran I, Zhang C-Y, Vidal-Puig A. Extracellular vesicles: novel mediators of cell communication in metabolic disease. Trends Endocrinol Metab. 2017;28(1):3–18.
Chen I-H, Aguilar HA, Paez Paez JS, Wu X, Pan L, Wendt MK, et al. An analytical pipeline for discovery and verification of glycoproteins from plasma-derived extracellular vesicles as breast cancer biomarkers. Anal Chem. 2018;90(10):6307–13.
Yang Q, Wang L-X. Chemoenzymatic glycan remodeling of natural and recombinant glycoproteins. Methods Enzymol. 2017;597(1):265–81.
Chen Y, Xiong Z, Zhang L, Zhao J, Zhang Q, Peng L, et al. Facile synthesis of zwitterionic polymer-coated core–shell magnetic nanoparticles for highly specific capture of N-linked glycopeptides. Nanoscale. 2015;7(7):3100–8.
Jiang B, Liang Y, Wu Q, Jiang H, Yang K, Zhang L, et al. New GO–PEI–Au–L-Cys ZIC-HILIC composites: synthesis and selective enrichment of glycopeptides. Nanoscale. 2014;6(11):5616–9.
Hang Q, Isaji T, Hou S, Wang Y, Fukuda T, Gu J. A key regulator of cell adhesion: identification and characterization of important N-glycosylation sites on integrin α5 for cell migration. Mol Cell Biol. 2017;37(9):1–14.
Zhuo Y, Yang J-Y, Moremen KW, Prestegard JH. Glycosylation alters dimerization properties of a cell-surface signaling protein, carcinoembryonic antigen-related cell adhesion molecule 1 (CEACAM1). J Biol Chem. 2016;291(38):20085–95.
Zhao D, Liang L, Wang S, Nakao T, Li Y, Liu L, et al. Glycosylation of the hemagglutinin protein of H5N1 influenza virus increases its virulence in mice by exacerbating the host immune response. J Virol. 2017;91(7):1–14.
Pinho SS, Reis CA. Glycosylation in cancer: mechanisms and clinical implications. Nat Rev Cancer. 2015;15(9):540–55.
Stowell SR, Ju T, Cummings RD. Protein glycosylation in cancer. Annu Rev Pathol. 2015;10(1):473–510.
Guo Z, Liu X, Li M, Shao C, Tao J, Sun W, et al. Differential urinary glycoproteome analysis of type 2 diabetic nephropathy using 2D-LC–MS/MS and iTRAQ quantification. J Transl Med. 2015;13(1):371–88.
Saraswat M, Joenväära S, Musante L, Peltoniemi H, Holthofer H. Renkonen R. N-linked (N-) glycoproteomics of urimary exosomes. Mol Cell Proteomics. 2015;14(2):263–76.
Pan L, Aguilar HA, Wang L, Iliuk A, Tao WA. Three-dimensionally functionalized reverse phase glycoprotein array for cancer biomarker discovery and validation. J Am Chem Soc. 2016;138(47):15311–4.
Jia X, Chen J, Sun S, Yang W, Yang S, Shah P, et al. Detection of aggressive prostate cancer associated glycoproteins in urine using glycoproteomics and mass spectrometry. Proteomics. 2016;16(23):2989–96.
Wang L, Li F, Sun W, Wu S, Wang X, Zhang L, et al. Concanavalin A-captured glycoproteins in healthy human urine. Mol Cell Proteomics. 2006;5(3):560–2.
Saraswat M, Joenväära S, Musante L, Peltoniemi H, Holthofer H. Renkonen R. N-linked (N-) glycoproteomics of urinary exosomes. Mol Cell Proteomics. 2015;14(2):263–76.
Pisitkun T, Johnstone R, Knepper MA. Discovery of urinary biomarkers. Mol Cell Proteomics. 2006;5(10):1760–71.
Sharma K, Schmitt S, Bergner CG, Tyanova S, Kannaiyan N, Manrique-Hoyos N, et al. Cell type–and brain region–resolved mouse brain proteome. Nat Neurosci. 2015;18(12):1819–31.
Doll S, Dreßen M, Geyer PE, Itzhak DN, Braun C, Doppler SA, et al. Region and cell-type resolved quantitative proteomic map of the human heart. Nat Commun. 2017;8(1):1469–82.
Santa C, Anjo SI, Manadas B. Protein precipitation of diluted samples in SDS-containing buffer with acetone leads to higher protein recovery and reproducibility in comparison with TCA/acetone approach. Proteomics. 2016;16(13):1847–51.
Livshits MA, Khomyakova E, Evtushenko EG, Lazarev VN, Kulemin NA, Semina SE, et al. Isolation of exosomes by differential centrifugation: theoretical analysis of a commonly used protocol. Sci Rep. 2015;5(1):17319–32.
Salih M, Zietse R, Hoorn EJ. Urinary extracellular vesicles and the kidney: biomarkers and beyond. Am J Physiol Renal Physiol. 2014;306(11):F1251–9.
Motamedinia P, Scott AN, Bate KL, Sadeghi N, Salazar G, Shapiro E, et al. Urine exosomes for non-invasive assessment of gene expression and mutations of prostate cancer. PLoS One. 2016;11(5):1–15.
Liang L-G, Kong M-Q, Zhou S, Sheng Y-F, Wang P, Yu T, et al. An integrated double-filtration microfluidic device for isolation, enrichment and quantification of urinary extracellular vesicles for detection of bladder cancer. Sci Rep. 2017;7(1):46224–34.
Cheruvanky A, Zhou H, Pisitkun T, Kopp JB, Knepper MA, Yuen PS, et al. Rapid isolation of urinary exosomal biomarkers using a nanomembrane ultrafiltration concentrator. Am J Physiol Renal Physiol. 2007;292(5):F1657–61.
Pan Y, Ma C, Tong W, Fan C, Zhang Q, Zhang W, et al. Preparation of sequence-controlled triblock copolymer-grafted silica microparticles by sequential-ATRP for highly efficient glycopeptides enrichment. Anal Chem. 2014;87(1):656–62.
Wiśniewski JR, Zougman A, Nagaraj N, Mann M. Universal sample preparation method for proteome analysis. Nat Methods. 2009;6(5):359–62.
Wu M, Ouyang Y, Wang Z, Zhang R, Huang P-H, Chen C, et al. Isolation of exosomes from whole blood by integrating acoustics and microfluidics. Proc Natl Acad Sci U S A. 2017;114(40):10584–9.
Merchant ML, Rood IM, Deegens JK, Klein JB. Isolation and characterization of urinary extracellular vesicles: implications for biomarker discovery. Nat Rev Nephrol. 2017;13(12):731–49.
Huang DW, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2008;37(1):1–13.
Hu S, Musante L, Tataruch D, Xu X, Kretz O, Henry M, et al. Purification and identification of membrane proteins from urinary extracellular vesicles using triton X-114 phase partitioning. J Proteome Res. 2017;17(1):86–96.
Acknowledgments
This study was supported by the National Key Program for Basic Research of China (No. 2017YFC0906703, 2016YFA0501403, 2018YFF0212505, 2017YFA0505002), National Natural Science Foundation of China (No. 21675172), National Key Laboratory of Proteomics Grant (SKLP-K201706), BPRC-Tianjin Baodi Hospital Joint Center Grant (TMRC2014Z03), Innovation Project (16CXZ027) and Beijing Science and Technology Plan Project (No. Z161100002616036).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Live subject statement
The authors state that all experiments were performed in compliance with the relevant laws and institutional guidelines. The institutional committee of the National Centre for Protein Science Beijing has approved the experiments with live subjects. The authors also state that informed consent was obtained for any experimentation with human subjects and National Centre for Protein Science Beijing is committed to the protection and safety of human subjects involved in research.
Rights and permissions
About this article
Cite this article
Huo, B., Chen, M., Chen, J. et al. A sequential separation strategy for facile isolation and comprehensive analysis of human urine N-glycoproteome. Anal Bioanal Chem 410, 7305–7312 (2018). https://doi.org/10.1007/s00216-018-1338-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-018-1338-6