Abstract
With the maturation of microfluidic technologies, microchip electrophoresis has been widely employed for amino acid analysis owing to its advantages of low sample consumption, reduced analysis time, high throughput, and potential for integration and automation. In this article, we review the recent progress in amino acid analysis using microchip electrophoresis during the period from 2007 to 2012. Innovations in microchip materials, surface modification, sample introduction, microchip electrophoresis, and detection methods are documented, as well as nascent applications of amino acid analysis in single-cell analysis, microdialysis sampling, food analysis, and extraterrestrial exploration. Without doubt, more applications of microchip electrophoresis in amino acid analysis may be expected soon.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amino acids are a group of organic molecules that play essential roles in various biological functions. They are not only the structural units that make up proteins, but some of them have also been recognized as neurotransmitters regulating important physiological activities, such as synaptic transmission, aging, learning, and memory [1–3]. Amino acids are widespread in the human body. Variations in the amino acid levels reflect changes in the biological system, which are of great interest for researchers to understand many fundamental biological processes, entailing accurate analysis of amino acids. However, analysis of amino acids is a challenging task because most amino acids are small aliphatic molecules without fluorescence or strong UV absorption characteristics. The optical properties of amino acids often necessitate the partitioning of l/d isomers. In addition, the complex biological system may induce a large amount of interference during the sampling processes, demanding separation of amino acids before quantitative analysis.
Capillary electrophoresis (CE) has been one of the dominant separation methods for analyzing amino acids. Several reviews have recently been published on amino acid analysis by CE [4–6]. With rapid advances in microfluidic technologies or “lab-on-a-chip” [7–14], microchip electrophoresis (MCE) has drawn increasing attention of researchers since its first introduction by Manz and coworkers [15, 16]. MCE allows amino acid analysis on a small scale using microfluidic devices. The small volume and short separation channel significantly reduce sample consumption and the analysis time. The potential of MCE for integration and miniaturization makes possible the development of portable devices that can operate on-site, which is ideal for point-of-care applications and even extraterrestrial explorations.
Pumera [17] has summarized the developments of MCE-based methods for analyzing amino acids before 2007. Since then, microfluidic technologies have been growing fast, and many new approaches have been developed for amino acid analysis, necessitating an update of this topic. Here, we document the recent advances in amino acid analysis using MCE-based methods during the period from 2007 to 2012, covering innovations in microchip materials, surface modification, sample introduction, MCE and detection methods. Selected applications of MCE will be discussed, including single-cell analysis, microdialysis sampling, food analysis, and extraterrestrial exploration. Analyses of both proteinogenic and nonproteinogenic amino acids will be discussed, with emphasis on proteinogenic amino acids.
Microchip materials
Design and fabrication of the microchips are the first step in developing microfluidic-based methods for amino acid analysis. Cross-shaped microchannels [18–20] are still the commonest chip design owing to their convenience for sample introduction and ease of fabrication. Slight tweaking of the cross-shaped microchannels has been reported to improve analytical performance, such as double-T-shaped microchannels for sample introduction [21–23], addition of a side channel for online reaction [24–26], elongation of the separation channel for enhanced resolution [27, 28], and multiplexing of microchannels for higher throughput [29, 30]. Complex microchip designs with multiple layers have also been used for integration and automation [31].
Microchip fabrications mostly follow standard lithography or soft lithography methods [32, 33]. A variety of materials have been used to construct microfluidic chips, including silicon [34, 35], glass [36–39], polydimethylsiloxane (PDMS) [40–42], poly(methyl methacrylate) (PMMA) [43–46], polycarbonate [47], cyclic olefin copolymer [48–50], thermoset polyester [51], and poly(glycidyl methacrylate)-co-(methyl methacrylate) [52]. Glass and PDMS have been the dominant choice in the past few years. Glass has good optical properties and chemical stability. The fabrication of glass microchips is relatively complex. For sealing of glass microchips, many researchers chose a flat layer of PDMS to irreversibly bond with the glass substrate with patterned microchannels [53, 54]. PDMS also has good chemical stability and optical transparency. The rapid prototyping method makes possible mass production of PDMS microchips with high reproducibility. However, a thin layer of PDMS may lack rigidity, resulting in unwanted distortion of microchannels. Thus, a glass substrate is sometimes used to irreversibly bond to the PDMS structure to seal open channels and to improve the structural rigidity [55, 56]. In some cases, the flexibility of PDMS may also be exploited for fabricating hybrid PDMS/glass microchips, such as the construction of diaphragm-based micropumps [57].
In addition to the materials mentioned above, a few new materials have been reported recently for fabricating microfluidic chips, such as poly(ethylene glycol) diacrylate (PEGDA) [58, 59] and OrmoComp [60]. PEGDA is a poly(ethylene glycol) (PEG)-functionalized polymer that is inherently resistant to protein adsorption. Fabrication of PEGDA microchips follows the photopolymerization-based method [61], which is similar to the soft lithography method, and both patterning and bonding can be completed within 10 min. OrmoComp is a new commercial hybrid ceramic polymer [60]. Both UV lithography and UV embossing can be used to fabricate OrmoComp microfluidic separation chips. OrmoComp chips exhibits stable cathodic electroosmotic flow (EOF) that can be used for MCE.
Surface modification
The properties of the microchip materials, including chemical stability, surface chemistry, and optical properties, are essential for amino acid analysis. In most cases, it is the hydrophobicity of microfluidic chips that complicates their use in separating amino acids. This is because high hydrophobicity often means undesired nonspecific adsorption of samples onto the microchannel surface. Thus, surface modification is sometimes necessary to enhance the resistance of microchannel walls to amino acids.
Glass-based microfluidic chips have been extensively used in amino acid analysis owing to their good chemical stability and transparency. However, it is sometimes necessary to suppress the strong EOF of glass microchips for better separation performance. Schulze and Belder [62] recently reported the use of a hydrophilic-coating material, PEG-1 M-100, to modify the channel surface in a glass microchip. The modified device exhibited a suppressed EOF and reduced analyte adsorption, and was successfully applied in MCE of fluorescein isothiocyanate (FITC)-labeled amino acids.
PDMS is the most frequently used polymer material for fabricating microfluidic chips. Its high hydrophobicity often induces nonspecific adsorption of amino acids and proteins. Many efforts have been dedicated to the surface modification of PDMS to improve its hydrophilicity and separation performance. Dynamic adsorption [20, 42, 63–65] and chemical grafting [18, 19, 66, 67] are two common methods for PDMS modifications. For example, Qiu et al. [42] used a dynamic coating method to layer-by-layer immobilize poly(diallyldimethylammonium chloride) and TiO2 nanoparticles on a PDMS surface. The modified PDMS microchip exhibited a decreased and stable EOF, which was favorable for separating biomolecules with similar migration times. Efficient separation of arginine, phenylalanine, serine, and threonine was demonstrated within 100 s in a 3.7-cm-long separation channel. Liang et al. [20] used the reaction between dopamine and HAuCl4 to trigger dopamine polymerization and generation of metallic nanoparticles in the microchannel, resulting in an in situ well-distributed and robust polydopamine/gold nanoparticle coating on the microchannel walls (Fig. 1). Compared with the native PDMS microchannel, the modified surfaces exhibited much better wettability, high stability, and suppressed electroosmotic mobility, and less nonspecific adsorption towards biomolecules. Recently, Zhang et al. [18] developed an environmentally friendly chemical grafting strategy in which PEG-NH2 was covalently attached to a silanized PDMS surface. EOF measurements and protein adsorption studies revealed noticeable EOF suppression and resistance to nonspecific adsorption for more than 30 days. Separation of four FITC-labeled amino acids was demonstrated with high repeatability and reproducibility. A “click” chemistry based modification strategy was further proposed by the same research group [19], where alkyne-PEG was “click” grafted to azido-PDMS for enhanced EOF and resistance to nonspecific adsorption. In addition to dynamic coating and chemical grafting, it is also possible to modify PDMS microchips by bulk modification. Xiao et al. [68] reported a simple and rapid bulk-modification method by adding an amphiphilic copolymer, poly(lactic acid)–PEG, during the fabrication of PDMS microchips. The bulk-modified PDMS microchips exhibited reproducible and stable EOF behavior, and were successfully used for separating mixtures of amino acids.
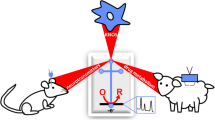
Coating of a polydimethylsiloxane (PDMS) channel with polydopamine (PDA)/gold nanoparticles (Au NP) for separation of amino acids. The construction of the PDA/Au NP modified PDMS microfluidic devices (A). Separation mechanism of the electrophoretic microchip system for analysis of five amino acids integrated with end-channel electrochemical detection (B). DA dopamine, PBS phosphate-buffered saline. (Adapted from [20])
Sample introduction
For microfluidic separation of amino acids, sample introduction is a key step to achieve efficient analysis. Karlinsey [69] lately reviewed sample introduction techniques for MCE. Here, we will only give an overview of the recent development of sample introduction methods in amino acid analysis.
In the past 5 years, electrokinetic injection has still been the commonest technique for mobilizing samples on microchips using EOF. Various modes have been reported for electrokinetic injection on microfluidic systems, including floating, pinched, gated, and dynamic loading modes. Recently, Blas et al. [70] conducted a comparative study of floating and dynamic injection modes in an electrokinetic microfluidic separation system by separating a mixture of fluorescently labeled arginine, glycine, glutamic acid, aspartic acid, and γ-aminobutyric acid (GABA), a nonproteinogenic amino acid. The dynamic loading mode was superior to the floating mode in terms of efficiency and reproducibility, because the sample plug was less dispersed. A monolithic sampling probe system was fabricated by He et al. [71] using simple tools, including a glass cutter and a bench drill. In combination with a slotted-vial array sample/reagent presentation system, a train of different samples could be continuously introduced by electrokinetic injection without interruption. Separation of FITC-labeled amino acids showed relative standard deviations (RSD) of 3.6 %, 3.3 %, and 3.5 % in peak height for arginine, FITC, and phenylalanine, respectively (n = 11).
As an alternative to mobilizing samples by EOF, new sample loading methods have been reported for electrokinetic injection. Zhang and Yin [72] developed a microvacuum-pump-based negative pressure sampling device for parallel separation of multiple samples on a microfluidic chip array. The negative pressure sampling device consists of a microvacuum air pump, a buffer vessel, a three-way electromagnet valve, and a vacuum gauge. Six samples were simultaneously loaded by negative pressure to form pinched sample plugs at the channel crossings, which could then be injected into the separation channels by EOF after the release of the negative pressure. A hybrid microfluidic system was reported by Abdelgawad et al. [73] to interface discrete droplet samples with MCE separations (Fig. 2). Discrete droplets were manipulated on an array of electrodes by dielectrophoresis for on-chip labeling of amino acids, which was then moved to the inlet of the separation channel, and were injected by electrokinetic means for MCE analysis.
A hybrid microfluidic system interfacing discrete droplet samples with microchip electrophoresis (MCE) separations. A The hybrid device comprises an electrode array for sample preparation by digital microfluidics and a network of microchannels for chemical separations. The inset is a schematic of the interface (not to scale). B Frames from a movie (from left to right) depicting droplets containing colored dyes being moved, merged, and mixed by digital microfluidics and then delivered to the interface. C Frames from a movie (from left to right) demonstrating electrokinetic loading of the contents of a droplet into a microchannel. The arrow in panel 2 indicates the front of the reagent (purple dye) being loaded into the channel from the droplet. D MCE separation of naphthalene-2,3-dicarboxyaldehyde (NDA)-labeled glycine, alanine and valine by micellar electrokinetic chromatography (MEKC). E MCE separation of NDA-labeled HeLa cell lysate by MEKC. The inset was generated using an identical protocol without the cell lysate. The y-axis of the inset was scaled identically to that of the main panel. Peaks were assigned by spiking with NDA-labeled standards. (Adapted from [73])
Aside from its widespread use, electrokinetic injection suffers from biased sample introduction owing to the different electrophoretic mobility of molecules. In contrast, hydrodynamic injection allows unbiased sample introduction. PDMS-based pneumatic valves have been developed for reproducible MCE of amino acids. For example, Li and Martin [74] reported integration of PDMS-based pneumatic valves for the rapid injection of analytes from a continuously flowing stream into a channel network for MCE. Continuous injections of a 0.39-nL fluorescein plug into the separation channels showed that the injection process was highly reproducible (RSD of 0.7 %, n = 10). A similar strategy was employed by Sun et al. [75], where a rapidly actuated PDMS pneumatic valve made possible injection of discrete sample plugs as small as 100 pL for MCE separation. PDMS-based micropumps can also be used for sample introduction on microfluidic chips. For example, Li et al. [76] integrated a PDMS-based peristaltic pump on a cross-shaped MCE system to generate a well-defined sample plug. Sequential injections of 1 × 106 M sodium fluorescein resulted in RSD of 2.17 % and 4.96 % (n = 25) for peak height and peak area. Price and Culbertson [77] reported a PDMS microchip that incorporates miniaturized and fully integrated dielectric elastomer actuators in order to perform sample injection for MCE. Separations of FITC-labeled amino acids revealed that injections by integrated dielectric elastomer actuators resulted in a stabler chemical composition than electrokinetic injections. The RSD for peak areas were less than 1.1 % over 30 injections at six different volumes.
Microchip electrophoresis
Achiral separation
Various separation methods have been reported in the past few years for amino acid analysis, including microchip capillary zone electrophoresis (μCZE), microchip micellar electrokinetic chromatography (μMEKC), microchip capillary electrochromatography (μCEC), and multiplexed and multidimensional separation methods.
Owing to its simplicity, μCZE is the commonest separation mode. For example, Noblitt et al. [78] integrated a track-etched polycarbonate membrane into the reservoirs of a PDMS microchip for selective filtering of insoluble particles based on the membrane pore diameter. μCZE of amino acids was conducted without being hindered by the addition of membranes. Zhang et al. [19] used μCZE to evaluate the performance of a PEG-functionalized PDMS microchip for separating five amino acids. Yamamoto et al. [79] fabricated an acidic polyacrylamide gel in a PMMA microchip using in situ photopolymerization. Simultaneous concentration, enrichment, and μCZE separation of FITC-labeled amino acids was demonstrated. In fact, μCZE has been the dominant separation mode for investigating the performance of microchips in separating amino acids after surface modifications [18–21, 64–68]. A few more examples can be found in “Surface modification.”
μMEKC was first introduced by Moore et al. [80], and is based on the partition equilibrium of analytes between the micellar pseudostationary phase and the surrounding medium. Separation of 20 amino acids was attempted using one-dimensional μMEKC by Culbertson et al. [81] on a glass microchip patterned with a 25-cm-long spiral-shaped separation channel (Fig. 3). As a result, 19 amino acids were separated in 165 s with an average plate number of 280,000. However, the 20th amino acid, histidine, could not be resolved. Kitagawa and Otsuka [82] documented the development of μMEKC prior to 2008. Among various available pseudostationary phases, sodium dodecyl sulfate (SDS) is the commonest surfactant for μMEKC. Recently, Sueyoshi et al. [83] combined transient trapping with μMEKC for the separation of amino acids. Transient trapping provided effective preconcentration of samples on the basis of the trap-and-release mechanism. Separation of BODIPY-labeled amino acids showed that a 106-fold to 125-fold increase in detectability was achieved relative to μCZE. Qiao et al. [84] conducted a comparative study of three surfactants, namely, SDS, polyoxyethylene lauryl ether (Brij 35), and ethylene oxide/propylene oxide block copolymer (Pluronic F127), for μMEKC separation of amino acids. The optimized separation of seven amino amides was achieved with FITC as the labeling reagent and Brij 35 as the surfactant in 20.0 mM borate at pH 9.2. As a result, linearity of l-asparagine was obtained in the range from 6.6 ×10-6 to 2.6 × 10-4 M with a detection limit of 0.7 μM. In addition to the common buffers mentioned above, new buffers have also been reported for μMEKC separation of amino acids. For example, Hoeman and Culbertson [85] used a commercially available dishwashing soap from Seventh Generation (Burlington, VT, USA) for amino acid analysis by μMEKC. The optimized buffer contained 5.0 % w/w Seventh Generation Free & ClearTM dishwashing soap and 10 mM sodium borate, which had a different selectivity and provided higher separation efficiencies than SDS-based buffers. Guan et al. [41] used a mixture of ionic and zwitterionic surfactants for μMEKC in a PDMS microchip. The mixed surfactant system allowed tuning of the EOF across a range of pH and concentration conditions.
Microchip MEKC (μMEKC) separation of 20 amino acids in a spiral-shaped separation channel. A Image of a microchip with a spiral separation channel. The separation channel was 24.9 cm long and 40 μm wide (half-depth). The arrows in the inset indicate the vertexes of the polygonally shaped channels. B MEKC separation of 19 tetramethylrhodamine-labeled amino acids in a 10 mM sodium tetraborate/50 mM sodium dodecyl sulfate (SDS) buffer with 10 % (v/v) 2-propanol. The field strength was 770 V/cm, and the detection point was 11.87 cm from the injection cross. The peak locations of the amino acids are indicated by their standard one-letter abbreviations. (Adapted from [81])
μCEC provides an alternative means for separating amino acids which combines the selectivity of liquid chromatography with the efficiency of MCE. Huo and Kok [86] summarized the applications of μCEC in amino acid analysis. Lately, Blas and et al. [87] integrated a fused-silica capillary containing a stationary phase into a PDMS microfluidic chip for μCEC. Separation of a mixture of proteinogenic (arginine, glycine, glutamic acid, and aspartic acid) and nonproteinogenic (GABA) amino acids was demonstrated. Since there are number of capillary modification methods, it is possible to functionalize the assembled device for different applications. Park et al. [48] developed a cyclic olefin copolymer microchip and packed the cross-shaped microchannels with 0.8-μm monodisperse colloidal silica beads by self-assembly. Owing to the large surface-to-volume ratio of the silica packing, it is possible to control the EOF within the channels with high reproducibility (1.3 % RSD in migration rate). As a result, four FITC-labeled amino acids were successfully separated with a 2-mm separation channel length by μCEC.
New separation modes have been developed for achiral separation of amino acids. Xu et al. [88] proposed a microchip free-flow electrophoresis (μFFE) analytical system in which voltages were applied in two dimensions so that both fluid transport and separation were driven electrokinetically. Separation of FITC-labeled amino acids, including l-lysine, l-phenylalanine, and l-aspartic acid, was successfully realized. The research group of Belder [89] developed a μFFE device that allowed high-speed continuous separations of amino acid samples. Free-flow zone electrophoretic separations of labeled amino acids were demonstrated. With a similar design, they fabricated a microfluidic chip using a spacerless approach with the photodefinable polymer PEGDA [90]. Hydrophilic microfluidic channels of only 25 μm in height were generated. Separation of labeled amino acids was realized by μFFE. Shameli et al. [56] reported a hybrid PDMS/glass microfluidic chip integrating a planar heater for generating temperature gradients. A bilinear temperature gradient along the separation channel was realized, which improved both peak capacity and separation resolution simultaneously. Using the developed device, they were able to separate amino acids that have similar electrophoretic mobilities.
In addition to above-mentioned methods, multiplexed approaches have also been reported. An eight-channel multiplexed microfluidic device was fabricated by Shackman et al. [47] which allowed parallel separation of amino acid samples using gradient elution moving boundary electrophoresis. Shadpour et al. [30] developed a 16-channel microfluidic chip integrating a gold electrode array for independent contact conductivity detection in each channel. Parallel analysis of amino acid was performed using μCZE, showing a separation efficiency of 2.0 × 103 plates.
By coupling different separation modes orthogonally, multidimensional MCE is able to separate amino acids with enhanced resolution. Recently, Xu et al. [27] developed a two-dimensional MCE system coupling μMEKC with μCZE for quantitative amino acid analysis. The design of the two-dimensional MCE system is shown in Fig. 4A. Incomplete separation of 20 amino acids was found by using one-dimensional μMEKC (Fig. 4B). By coupling μMEKC with μCZE, they successfully realized complete separation of 20 amino acids with linear dynamic ranges above three orders of magnitude and linear correlation coefficients γ > 0.99 (Fig. 4C). Quantitative analysis of a commercial nutrition supplement liquid was further demonstrated.
Two-dimensional electrophoretic separation coupling μMEKC with microchip capillary zone electrophoresis (CZE) for amino acid analysis. A The microfluidic chip. Arrows indicate the detection points for one-dimensional and two-dimensional separations. The reservoirs are labeled as follows: S sample, B1 buffer 1, SW1 sample waste 1, B2 buffer 2, SW2 sample waste 2, and W waste. B The effect of the borate concentration on the separation of 20 fluorescein isothiocyanate (FITC)-labeled amino acids by the first dimension MEKC. C A gel-like two-dimensional electropherogram of 20 FITC-labeled amino acids. The separation conditions were as follows: 25 mM borate buffer and 100 mM SDS (pH 11.0) in the first dimension; 100 mM borate buffer (pH 9.5) in the second dimension. The electric field strength was 300 V/cm in the first dimension and 1,000 V/cm in the second dimension. (Adapted from [27])
Chiral separation
Chiral separation of amino acid enantiomers is an important issue, since enantiomers often exhibit different pharmaceutical or bioactive effects. MCE-based methods have been widely employed for fast amino acid enantioseparations owing to advantages such as low sample consumption and high separation efficiency. However, it is difficult to separate a pair of amino acid enantiomers using simple separation modes. Generally, chiral selectors are added in the mobile phase (μMEKC) or immobilized as a stationary phase (μCEC) to achieve effective separation. Nagl et al. [91, 92] reviewed the advances in microchip enantioseparations. Here, we summarize the recent progress in chiral separation of amino acid enantiomers using MCE-based approaches.
Huang et al. [93] performed chiral separation of amino acids on a PDMS/glass hybrid microfluidic chip using γ-cyclodextrin as a chiral selector in the running buffer. Using this method, they determined the concentration of d-aspartic acid and d-glutamate in rat brain and human cerebrospinal fluid. The same research group further developed a double-T-shaped microchannel for the chiral separation of d/l-tyrosine enantiomers with γ-cyclodextrin as the chiral selector [94]. The concentration of d-tyrosine in human plasma was found to differ significantly from normal humans to patients with renal failure. Sun et al. [59] developed a PEG-functionalized polymeric microchip. By introducing β-cyclodextrin into the running buffer as a chiral selector, they separated ten different pairs of d,l-amino acid pairs with high reproducibility. Microfluidic open tubular capillary electrochromatography was reported by Zhang et al. [36] for chiral recognition of dansyl enantiomeric amino acids. Avidin was employed as the chiral selector and was immobilized on the microchannel wall by dynamic adsorption. With addition of methanol in the running buffer, four dansyl racemic amino acids were successfully separated by microfluidic open tubular capillary electrochromatography within 100 s. Qu et al. [95] developed a microfluidic device integrating three polymer retainers and a quartz capillary coated with molecularly imprinted polymer. The molecularly imprinted polymer was in situ chemically polymerized on the microchannel wall using acrylamide as the functional monomer and ethylene glycol dimethacrylate as the cross-linker. Baseline separation of tert-butoxycarbonyl-d-tryptophan and tert-butoxycarbonyl-L-tryptophan was realized within 75 s with detection limits of 20 and 140 μM, respectively. The same research group further developed a multitemplate imprinted microchannel using a one-pot in situ imprinting process [55]. Using l-tyrosine and l-tryptophan as the template molecules, they simultaneously baseline-separated two pairs of enantiomers in a 6-cm separation channel within 120 s under the optimized μCEC conditions.
Multidimensional separation methods have also been reported for amino acid enantioseparations. For example, Kim et al. [43] developed a three-dimensional microfluidic device coupling achiral separation with chiral separation for amino acid analysis (Fig. 5). The chiral separation was implemented using μMEKC with β-cyclodextrin as the chiral selector and sodium taurocholate as the micelle-forming agent. Enantioseparation of FITC–d/l-aspartic acid and FITC–d/l-serine was demonstrated. Ross et al. [38] presented a microfluidic chip for two-dimensional separations coupling gradient elution moving boundary electrophoresis with μCZE. The simplicity of the gradient elution moving boundary electrophoresis allowed its implementation in the injection channel of a conventional MCE microchip, simplifying the design and operation of the device. The enantiomers of aspartic acid, glutamic acid, serine, alanine, and valine were successfully separated using the method developed.
A three-dimensional microfluidic device coupling achiral separation with chiral separation for amino acid analysis. A Structure of the device: a the 1.5-mm-thick polycarbonate cap; b, c, e (patterned, 100 μm × 20 μm), and f 20-μm-thick poly(methyl methacrylate) layers; d a 10-μm-thick nanocapillary array membrane (NCAM). B Layout of the channels for two-dimensional separation of an FITC-labeled amino acid mixture. Asterisks indicate the focused excitation/data collection locations for each separation channel. LIF 1 and LIF 0 monitor the first- and second-dimension separations, respectively. C Two-dimensional separation of FITC-Asp and FITC-Ser, where FITC-Asp is selectively transferred to the second-dimension channel for chiral separation. The peak shoulder of the d/l-Ser peak is part of the d/l-Asp peak, recognized and collected for the second-dimension chiral separation. (Adapted from [43])
Detection methods
Optical detection
Optical detection has been extensively used for detecting amino acids. Two review articles have summarized the developments of optical methods for separation on microfluidic chips prior to 2007 [96, 97]. In the following paragraphs, we will highlight the recent innovations in detecting amino acids by optical means.
Laser-induced fluorescence (LIF) detection remains the dominant optical means for detection of amino acid. Since most amino acids are naturally nonfluorescent, it is necessary to label amino acids with fluorophores so that they can be detected optically. Common fluorescent labeling reagents include FITC [34, 35, 43, 44, 48], naphthalene-2,3-dicarboxyaldehyde [73, 87], 4-chloro-7-nitrobenzofurazan [67], 5-(4,6-dichloro-striazin-2-ylamino) fluorescein [64], and BODIPY fluorescent dyes [83, 85]. New fluorescence labeling reagents have also been reported, such as 4-amino-1,8-naphthalimides (UR-431) [98]. A fluorescence microscope is usually employed as the detection platform. For excitation, the light emitted from a laser or a mercury lamp is filtered by an excitation filter, reflected by a dichroic mirror, and then focused on the separation channel. The fluorescence is collected through the same objective with an emission filter and monitored by a photomultiplier or a CCD camera. Recent innovations in LIF detection of amino acids are highlighted in the following paragraphs.
Liu et al. [99] reported a PDMS microchip integrating a direct-contacting optical fiber for detecting amino acids by LIF. A blue-light-emitting diode was applied as the excitation source. Separation of FITC-labeled amino acids was demonstrated. Yassine et al. [100] described the use of two optical fibers for LIF detection on a microfluidic chip. One optical fiber was connected to a laser and positioned in a dedicated channel in close proximity to the detection point. Another optical fiber was placed below the microchip to collect the fluorescence signal. The integration of optical fibers simplified the detection system and freed the researchers from the alignment procedure each time to perform an experiment. Yang et al. [54] presented a novel detection system based on small-angle optical deflection from the collinear configuration of a microfluidic chip. For excitation, the incident light beam was focused on the separation channel through the edge of a lens, resulting in a small deflection angle that deviated 20° from the collinear configuration. The fluorescence was collected through the center of the same lens by a photomultiplier in the vertical direction. As a result, the background level from the light source and from the reflection of the microchip surface was significantly eliminated. An enhanced signal-to-noise ratio was obtained for separating FITC-labeled amino acids.
Chemiluminescence (CL) detection is another common optical detection method for amino acid analysis. CL detection does not require an external light source. The absence of a strong background level enhances the signal-to-noise ratio, improving the detection sensitivity. Liu et al. [101] developed a PDMS MCE system integrated with CL detection. A detection limit down to submicromolar concentrations was achieved with good reproducibility and symmetric peak shape. Chiral recognition of dansyl phenylalanine enantiomers was further achieved using the developed system. Recently, Ye et al. [53] demonstrated MCE separation with CL detection for the determination of amino acids in single mice cells. The contents of tryptophan, glycine, alanine, glutamic acid, and aspartic acid in single mice fibrosarcoma cells were found to be in the ranges 0.95–2.31, 1.08–6.87, 1.03–4.05, 0.84–2.61, and 0.82–3.68 fmol, respectively. Kamruzzaman et al. [102] demonstrated a microfluidic-chip-based system coupled with CL detection for the determination of l-phenylalanine. The detection was based on the enhancement effect of l-phenylalanine on the CL signals of the luminol–H2O2–Cu2+ system in an alkaline medium. The detection limit was 2.4 × 10-10 M with an RSD of 1.8 %.
Optical means can also be used to detect unlabeled amino acids. For example, Khurana and Santiago [103] reported an indirect detection method for nonfluorescent amino acids using fluorescent mobility markers. A mixture of fluorescent markers and nonfluorescent amino acids was separated by isotachophoresis. Unlabeled amino acids were detected as gaps in the fluorescent signals of mobility markers. Preconcentration, separation, and detection of unlabeled serine, glycine, and phenylalanine were successfully demonstrated.
Electrochemical detection
Electrochemical detection has been widely used for label-free amino acid analysis. It offers advantages such as low cost, high efficiency, and potential for integration and miniaturization. Here, we focus on the recent advances in conductivity and amperometric detection methods.
Conductivity detection is a relatively simple approach, which theoretically can be applied to all charged analytes that have low electrochemical activity for amperometric detection or weak optical absorbance for optical measurements. Either a contact arrangement or a contactless arrangement can be implemented in conductivity detection. However, contact mode is generally less popular than contactless mode owing to the difficulty in decoupling the detection circuit from the high separation voltage and degradation of the electrode surface. A floating resistivity detector was developed by Tay et al. [22] for conductivity detection using the contact mode. The floating resistivity detector measured the signal generated from the separation field, which permitted decoupling of the detection circuit from the high separation voltage without compromising separation efficiency. Chen et al. [23] reported a thin cover glass microchip for MCE with capacitively coupled contactless conductivity detection (C4D). Electrodes were placed in contact with a thin cover glass (100-μm thick) outside the microchannel. MCE separation of amino acids with C4D was demonstrated. Xuan and et al. [104] described the use of a polycarbonate microchip with C4D for the separation of a nonproteinogenic amino acid, nicotianamine, and its copper(II) and iron(III) chelates. Nicotianamine is an unusual amino acid from plants which has received the attention of researchers in the past few years as a metal chelator. Liu et al. [105] reported a dual detection system that allowed LIF detection and C4D simultaneously on a microfluidic chip. Separation of inorganic ions and FITC-labeled amino acids showed that the performance of both LIF detection and C4D was satisfactory for routine analysis of biological specimens. Kuban and Hauser [106] lately reviewed the developments of C4D in both CE and MCE.
Amperometric detection is achieved by measuring current while applying a modest potential to the working electrode, which is an effective means for detecting electroactive analytes. Zhai et al. [107] reported amperometric detection of amino acids on an assembled microfluidic device that integrated three polymer retainers with a quartz capillary and a copper microdisk electrode. An end-column detection mode was used with a potential of 0.8 V on the working electrode relative to the reference electrode. Detection of 12 amino acids was demonstrated with high sensitivity and reproducibility. Chen et al. [40] fabricated nanoband microelectrodes for amperometric detection on an MCE microchip. A thin layer of gold and copper (100 nm thick) was deposited on a PDMS substrate by selective region plasma oxidation through shadow masking. By casting another PDMS layer on top of the metal film, they formed a sandwich structure. Cutting the sandwich structure resulted in nanoband microelectrodes at the cross section which could be used for amperometric detection. MCE separation of amino acids was successfully achieved using the nanoband microelectrodes with an end-column setup. By integrating the gold–copper dual metal detector with a dual-channel MCE system, they further distinguished electroactive amino acids from nonelectroactive amino acids. Recently, Ghanim and Abdullah [108] reviewed amperometric detection methods on MCE microfluidic devices.
Applications
Owing to the importance of amino acids to living organisms, analyses of amino acids have shown a wide range of applications. In the following paragraphs, we summarize the recent applications of amino acid analysis in single-cell analysis, microdialysis, food analysis, and extraterrestrial exploration.
Single-cell analysis is an important issue in biology, because seemingly identical cells are often quite heterogeneous in their chemical composition and biological activity. MCE separation of amino acids has proven to be an effective approach for single-cell analysis. Shi et al. [109] applied MCE separation to determine the contents of amino acids in human vascular endothelial cells. The average amounts of amino acids in single human vascular endothelial cells were estimated to be 5.84, 1.15, 3.10, and 1.30 fmol for alanine, glycine, glutamic acid, and aspartic acid, respectively. MCE separation with CL detection was reported by Zhao et al. [24] to quantify the amount of amino acids, including tryptophan, glycine, and alanine, present in single rat hepatocytes. The same research group further used the MCE system developed with a sensitive optical detection scheme based on CL resonance energy transfer to determine the amount of amino acids in individual human red blood cells [25]. Nine amino acids were detected with an amount ranging from 3 to 31 amol.
Combination of microdialysis sampling with a microfluidic analytical system provides near real-time information on changes of analyte levels in the extracellular space or other aqueous environments [110–113]. For example, the research group of Kennedy [110] developed a capillary–PDMS hybrid chip that combined low-flow push–pull perfusion sampling, online derivatization, and flow-gated injection onto an embedded fused-silica capillary for high-speed separation of amine neurotransmitters, including both proteinogenic and nonproteinogenic amino acids, from the brain of living animals. The same research group further demonstrated a dual-chip system for monitoring in vivo chemical changes by analyzing segmented flows from a microdialysis probe [111] (Fig. 6). In this system, a microdialysis probe was integrated with a PDMS microchip that merged dialysate with fluorogenic reagent and segmented the flow into 8–10-nL plugs in perfluorodecalin. A glass microchip was then used to extract the fluid from the sample plugs for MCE analysis with LIF detection. With this approach, they were able to monitor rapid concentration changes of both proteinogenic amino acids, including serine, glycine, glutamate, and aspartate, and nonproteinogenic amino acids, such as GABA, evoked by infusing l-trans-pyrrolidine-2,4-dicarboxylic acid, a glutamate uptake inhibitor, into the striatum of anesthetized rats. Lately, two review articles documented the advances in microfluidic-based microdialysis sampling [114, 115].
A dual-chip system for monitoring in vivo chemical changes by analyzing segmented flows from a microdialysis probe. A The dual-chip system. B Typical electropherogram obtained in vivo from rat striatum with online derivatization using the setup. Peaks for serine, glycine, glutamate, and aspartate are labeled. The electric field was 647 V/cm. The injection width was 80 ms. C Effect of l-trans-pyrrolidine-2,4-dicarboxylic acid (PDC) infusion on glutamate and aspartate basal levels. The black bar denotes infusion of PDC corrected for dead time (approximately 6.5 min). Each point represents the peak height of glutamate or aspartate from one electropherogram collected every 25 s. aCSF artifical cerebrospinal fluid, CE capillary electrophoresis, HPFA high-performance frontal analysis, LIF laser-induced fluorescence, RFU relative fluorescence units. (Adapted from [111])
Since amino acids are employed as ingredients in a large number of food products, such as sports beverages, amino acid analysis is often applied to characterize the properties of foods and to determine their quality and safety. Escarpa et al. [116] reviewed the applications of MCE in food analysis. Recently, Ueno et al. [117] reported the use of a PMMA microchip for μMEKC separation and quantitative analysis of amino acids in three different kinds of functional foods, sports beverages, jelly-form beverages, and tablet-form functional foods. Ohla et al. [118] reported an MCE-based method coupling deep UV fluorescence detection for the determination of biologically active compounds in bananas, including dopamine, serotonin, and their precursors tryptophan and tyrosine. Zhai et al. [119] developed a UV detection method for separation of amino acids on a disposable MCE device. On-column conjugation of amino acids with cupric cation led to UV absorption at 232 nm that could be used for detection of amino acids. Analysis of amino acids in beverage samples was demonstrated with recovery of more than 85.0 %.
Amino acids are excellent biomarkers for extraterrestrial exploration owing to their universal presence in terrestrial biology and the fact that they have been detected in meteorites and comet particles. The Mars Organic Analyzer is a portable MCE instrument for amino acid analysis in extraterrestrial samples. Chiesl et al. [120] recently demonstrated the use of a fluorescent amine reactive probe, Pacific Blue succinimidyl ester (PB), for the detection of trace amounts of amino acids by μMEKC with the Mars Organic Analyzer. Using PB-labeled standards, they achieved a limit of detection of 75 pM. Samples from the Murchison meteorite were successfully analyzed, showing the presence of abiotic amino acids such as β-alanine and ε-aminocaprioc acid as well as several neutral and acidic amino acids. The Multichannel Mars Organic Analyzer was developed by Benhabib et al. [29]. It employs a four-layer microchip, containing eight MCE analysis systems integrated with a microfluidic network for autonomous fluidic processing. A limit of detection of 6 pM was achieved for glycine. Separation of amino acids was demonstrated with minimal sample-to-sample carryover, indicating good reusability of the MCE systems. Mora et al. [31] presented a fully integrated microfluidic device capable of performing automated end-to-end analysis of amino acids by MCE with LIF detection (Fig. 7). The device was validated by electrophoretic analysis of PB-labeled amino acids, and could be used for future in situ extraterrestrial exploration. Mora et al. [121] further gave an overview of MCE instrumentation for in situ analysis in the search for extraterrestrial life.
Automated analysis of amino acids by MCE with LIF detection. A Assembly of the four-layer device and the features of each layer. B Results from a completely automated on-chip analysis of amino acid mixtures. a blank run (buffer only); b standard solution containing 100 nM Pacific Blue succinimidyl ester labeled citrulline (1), valine (2), serine (3), and glycine (5); (c) approximately 160 nM sample of citrulline (1), valine (2), and alanine (4) diluted from a 10 μM sample labeled on-chip; d mixture of b and c. The conditions were as follows: 25 mM tetraborate buffer, pH 9.2, separation voltage 5 kV. (Adapted from [31])
Conclusions and future perspectives
In this review, we have documented the recent progress in MCE-based amino acid analysis. As noted, most articles published in the past 5 years have focused on fundamental research and proof of concept, such as novel surface modification methods, new sampling introduction approaches, and innovation in separation and detection. However, it is interesting to find that a growing number of articles illustrated the use of MCE-based amino acid analysis for a variety of applications, such as food analysis and extraterrestrial exploration. Integration of MCE systems for automated amino acid analysis has also been demonstrated [31] for the first time, which shed light on the future instrumentation of such systems for more practical applications, such as point-of-care testing (POCT). POCT is still a nascent field. There have been few publications on MCE-based amino acid analysis in POCT. But we believe more applications in this field will be published soon.
In addition, a new trend has emerged in MCE-based amino acid analysis, in which the miniaturization of MCE starts to shift towards the nanofluidic domain owing to the maturation of nanofabrication technologies. Janssen et al. [122] lately investigated the scalability of conventional isotachophoresis to the nanofluidic domain. Separation of fluorescently labeled amino acids in biomatrix was realized in a 330-nm-deep channel with a sample injection volume of 0.4 pL. However, separation could not be achieved in a 50-nm-deep channel owing to the restriction of the applicable voltage by bubble formation. This work demonstrated the feasibility of MCE in nanofluidic channels. More efforts are required to further explore MCE in nanofluidic domains.
References
Perry M, Li Q, Kennedy RT (2009) Anal Chim Acta 653:1–22
Lapainis T, Sweedler JV (2008) J Chromatogr A 1184:144–158
Jahr CE, Lester RAJ (1992) Curr Opin Neurobiol 2:270–274
Poinsot V, Carpene MA, Bouajila J, Gavard P, Feurer B, Couderc F (2012) Electrophoresis 33:14–35
Poinsot V, Gavard P, Feurer B, Couderc F (2010) Electrophoresis 31:105–121
Poinsot V, Rodat A, Gavard P, Feurer B, Couderc F (2008) Electrophoresis 29:207–223
DeMello AJ (2006) Nature 442:394–402
Dittrich PS, Tachikawa K, Manz A (2006) Anal Chem 78:3887–3907
Janasek D, Franzke J, Manz A (2006) Nature 442:374–380
Vilkner T, Janasek D, Manz A (2004) Anal Chem 76:3373–3385
West J, Becker M, Tombrink S, Manz A (2008) Anal Chem 80:4403–4419
Thorsen T, Maerkl SJ, Quake SR (2002) Science 298:580–584
Feng XJ, Du W, Luo QM, Liu BF (2009) Anal Chim Acta 650:83–97
Kovarik ML, Gach PC, Ornoff DM, Wang YL, Balowski J, Farrag L, Allbritton NL (2012) Anal Chem 84:516–540
Harrison DJ, Fluri K, Seiler K, Fan Z, Effenhauser CS, Manz A (1993) Science 261:895–897
Manz A, Graber N, Widmer HM (1990) Sens Actuators B 1:244–248
Pumera M (2007) Electrophoresis 28:2113–2124
Zhang ZW, Feng XJ, Luo QM, Liu BF (2009) Electrophoresis 30:3174–3180
Zhang ZW, Feng XJ, Xu F, Liu X, Liu BF (2010) Electrophoresis 31:3129–3136
Liang RP, Meng XY, Liu CM, Qiu JD (2011) Electrophoresis 32:3331–3340
Yoon TH, Li M, Hong LY, Lee J, Kim DP (2011) Anal Chem 83:1901–1907
Tay ETT, Law WS, Sim SPC, Feng H, Zhao JH, Li SFY (2007) Electrophoresis 28:4620–4628
Chen ZG, Li QW, Li OL, Zhou X, Lan Y, Wei YF, Mo JY (2007) Talanta 71:1944–1950
Zhao SL, Huang Y, Liu YM (2009) J Chromatogr A 1216:6746–6751
Zhao SL, Huang Y, Shi M, Liu RJ, Liu YM (2010) Anal Chem 82:2036–2041
Zhao SL, Huang Y, Ye FG, Shi M, Liu YM (2010) J Chromatogr A 1217:5732–5736
Xu B, Feng XJ, Xu YZ, Du W, Luo QM, Liu BF (2009) Anal Bioanal Chem 394:1911–1917
Nandi P, Desai DP, Lunte SM (2010) Electrophoresis 31:1414–1422
Benhabib M, Chiesl TN, Stockton AM, Scherer JR, Mathies RA (2010) Anal Chem 82:2372–2379
Shadpour H, Hupert ML, Patterson D, Liu CG, Galloway M, Stryjewski W, Goettert J, Soper SA (2007) Anal Chem 79:870–878
Mora MF, Greer F, Stockton AM, Bryant S, Willis PA (2011) Anal Chem 83:8636–8641
Duffy DC, McDonald JC, Schueller OJA, Whitesides GM (1998) Anal Chem 70:4974–4984
Xia YN, Whitesides GM (1998) Annu Rev Mater Sci 28:153–184
Xu Y, Hu XG, Liang J, Sun JX, Gu WW, Zhao TM, Wen ZY (2009) Anal Bioanal Chem 394:1947–1953
Xu Y, Liang J, Liu HT, Hu XG, Wen ZY, Wu YJ, Cao MX (2010) Anal Bioanal Chem 397:1583–1593
Zhang GS, Qian CE, Xu YZ, Feng XJ, Du W, Liu BF (2009) J Sep Sci 32:374–380
Baker CA, Roper MG (2010) J Chromatogr A 1217:4743–4748
Ross D, Shackman JG, Kralj JG, Atencia J (2010) Lab Chip 10:3139–3148
Mu XY, Qi L, Qiao J, Zhang HZ, Ma HM (2012) Anal Biochem 421:499–505
Chen SP, Wu J, Yu XD, Xu JJ, Chen HY (2010) Anal Chim Acta 665:152–159
Guan Q, Noblitt SD, Henry CS (2012) Electrophoresis 33:379–387
Qiu JD, Wang L, Liang RP, Wang JW (2009) Electrophoresis 30:3472–3479
Kim BY, Yang J, Gong MJ, Flachsbart BR, Shannon MA, Bohn PW, Sweedler JV (2009) Anal Chem 81:2715–2722
Fuentes HV, Woolley AT (2008) Anal Chem 80:333–339
Sun XH, Peeni BA, Yang W, Becerril HA, Woolley AT (2007) J Chromatogr A 1162:162–166
Yang WC, Sun XH, Pan T, Woolley AT (2008) Electrophoresis 29:3429–3435
Shackman JG, Munson MS, Ross D (2007) Anal Chem 79:565–571
Park J, Lee D, Kim W, Horiike S, Nishimoto T, Lee SH, Ahn CH (2007) Anal Chem 79:3214–3219
Ladner Y, Cretier G, Faure K (2010) J Chromatogr A 1217:8001–8008
Wang Q, Zhang Y, Ding H, Wu J, Wang LL, Zhou L, Pu QS (2011) J Chromatogr A 1218:9422–9427
Pan T, Fiorin GS, Chiu DT, Woolley AT (2007) Electrophoresis 28:2904–2911
Sun XF, Liu JK, Lee ML (2008) Electrophoresis 29:2760–2767
Ye FG, Huang Y, Xu Q, Shi M, Zhao SL (2010) Electrophoresis 31:1630–1636
Yang L, Li XT, Li J, Yuan HY, Zhao SL, Xiao D (2012) Electrophoresis 33:1996–2004
Qu P, Lei JP, Sheng J, Zhang L, Ju HX (2011) Analyst 136:920–926
Shameli SM, Glawdel T, Liu Z, Ren CL (2012) Anal Chem 84:2968–2973
Guillo C, Karlinsey JM, Landers JP (2007) Lab Chip 7:112–118
Rogers CI, Pagaduan JV, Nordin GP, Woolley AT (2011) Anal Chem 83:6418–6425
Sun XF, Li D, Lee ML (2009) Anal Chem 81:6278–6284
Sikanen T, Aura S, Heikkila L, Kotiaho T, Franssila S, Kostiainen R (2010) Anal Chem 82:3874–3882
Liu J, Sun X, Lee ML (2007) Anal Chem 79:1926–1931
Schulze M, Belder D (2012) Electrophoresis 33:370–378
Liang RP, Wang L, Meng XY, Wang JW, Qiu JD (2011) Microfluid Nanofluid 11:227–233
Xiao Y, Yu XD, Wang K, Xu JJ, Huang J, Chen HY (2007) Talanta 71:2048–2055
Wang AJ, Xu JJ, Chen HY (2007) J Chromatogr A 1147:120–126
Wang AJ, Feng JJ, Fan J (2008) J Chromatogr A 1192:173–179
Miyaki K, Zeng HL, Nakagama T, Uchiyama K (2007) J Chromatogr A 1166:201–206
Xiao Y, Yu XD, Xu JJ, Chen HY (2007) Electrophoresis 28:3302–3307
Karlinsey JM (2012) Anal Chim Acta 725:1–13
Blas M, Delaunay N, Rocca JL (2007) Electrophoresis 28:4629–4637
He QH, Fang Q, Du WB, Fang ZL (2007) Electrophoresis 28:2912–2919
Zhang L, Yin XF (2007) Electrophoresis 28:1281–1288
Abdelgawad M, Watson MWL, Wheeler AR (2009) Lab Chip 9:1046–1051
Li MW, Martin RS (2007) Electrophoresis 28:2478–2488
Sun XF, Kelly RT, Danielson WF, Agrawal N, Tang KQ, Smith RD (2011) Electrophoresis 32:1610–1618
Li BW, Jiang L, Wang Q, Oin JH, Lin BC (2008) Electrophoresis 29:4906–4913
Price AK, Culbertson CT (2009) Anal Chem 81:8942–8948
Noblitt SD, Kraly JR, VanBuren JM, Hering SV, Collett JL, Henry CS (2007) Anal Chem 79:6249–6254
Yamamoto S, Watanabe Y, Nishida N, Suzuki S (2011) J Sep Sci 34:2879–2884
Moore AW, Jacobson SC, Ramsey JM (1995) Anal Chem 67:4184–4189
Culbertson CT, Jacobson SC, Ramsey JM (2000) Anal Chem 72:5814–5819
Kitagawa F, Otsuka K (2008) J Sep Sci 31:794–802
Sueyoshi K, Hashiba K, Kawai T, Kitagawa F, Otsuka K (2011) Electrophoresis 32:1233–1240
Qiao J, Qi L, Ma HM, Chen Y, Wang MX, Wang DX (2010) Electrophoresis 31:1565–1571
Hoeman KW, Culbertson CT (2008) Electrophoresis 29:4900–4905
Huo Y, Kok WT (2008) Electrophoresis 29:80–93
Blas M, Delaunay N, Rocca JL (2007) J Sep Sci 30:3043–3049
Xu Y, Cao MX, Gan J, Zeng X, Wen ZY (2010) Anal Sci 26:1139–1143
Kohler S, Weilbeer C, Howitz S, Becker H, Beushausen V, Belder D (2011) Lab Chip 11:309–314
Jezierski S, Gitlin L, Nagl S, Belder D (2011) Anal Bioanal Chem 401:2651–2656
Nagl S, Schulze P, Ludwig M, Belder D (2009) Electrophoresis 30:2765–2772
Nagl S, Schulze P, Ohla S, Beyreiss R, Gitlin L, Belder D (2011) Anal Chem 83:3232–3238
Huang Y, Shi M, Zhao SL (2009) J Sep Sci 32:3001–3006
Huang Y, Shi M, Zhao SL, Liang H (2011) J Chromatogr B 879:3203–3207
Qu P, Lei JP, Ouyang RZ, Ju HX (2009) Anal Chem 81:9651–9656
Gotz S, Karst U (2007) Anal Bioanal Chem 387:183–192
Kuswandi B (2007) Nuriman, Huskens J, Verboom W. Anal Chim Acta 601:141–155
Schulze P, Link M, Schulze M, Thuermann S, Wolfbeis OS, Belder D (2010) Electrophoresis 31:2749–2753
Liu CC, Cui DF, Chen X (2007) J Chromatogr A 1170:101–106
Yassine O, Morin P, Dispagne O, Renaud L, Denoroy L, Kleimann P, Faure K, Rocca JL, Ouaini N, Ferrigno R (2008) Anal Chim Acta 609:215–222
Liu BF, Ozaki M, Utsumi Y, Hattori T, Terabe S (2003) Anal Chem 75:36–41
Kamruzzaman M, Alam AM, Kim KM, Lee SH, Kim YH, Kim GM, Dang TD (2012) Food Chem 135:57–62
Khurana TK, Santiago JG (2008) Anal Chem 80:279–286
Xuan Y, Scheuermann EB, Meda AR, Jacob P, von Wiren N, Weber G (2007) Electrophoresis 28:3507–3519
Liu C, Mo YY, Chen ZG, Li X, Li OL, Zhou X (2008) Anal Chim Acta 621:171–177
Kuban P, Hauser PC (2011) Electrophoresis 32:30–42
Zhai C, Qiang W, Lei JP, Ju HX (2009) Electrophoresis 30:1490–1496
Ghanim MH, Abdullah MZ (2011) Talanta 85:28–34
Shi BX, Huang WH, Cheng JK (2008) J Sep Sci 31:1144–1150
Cellar NA, Kennedy RT (2006) Lab Chip 6:1205–1212
Wang M, Roman GT, Perry ML, Kennedy RT (2009) Anal Chem 81:9072–9078
Wang M, Hershey ND, Mabrouk OS, Kennedy RT (2011) Anal Bioanal Chem 400:2013–2023
Mecker LC, Martin RS (2007) J Assoc Lab Autom 12:296–302
Nandi P, Lunte SM (2009) Anal Chim Acta 651:1–14
Guilhen E, O'Connor WT (2010) Electrophoresis 31:55–64
Escarpa A, Gonzalez MC, Crevillen AG, Blasco AJ (2007) Electrophoresis 28:1002–1011
Ueno H, Wang J, Kaji N, Tokeshi M, Baba Y (2008) J Sep Sci 31:898–903
Ohla S, Schulze P, Fritzsche S, Belder D (2011) Anal Bioanal Chem 399:1853–1857
Zhai C, Sheng J, Qiang W, Lei JP, Ju HX (2010) Talanta 82:67–71
Chiesl TN, Chu WK, Stockton AM, Amashukeli X, Grunthaner F, Mathies RA (2009) Anal Chem 81:2537–2544
Mora MF, Stockton AM, Willis PA (2012) Electrophoresis 33:2624–2638
Janssen KGH, Li JJ, Hoang HT, Vulto P, van den Berg R, Overkleeft HS, Eijkel JCT, Tas NR, van der Linden HJ, Hankemeier T (2012) Lab Chip 12:2888–2893
Acknowledgements
The authors gratefully acknowledge financial support from the National Basic Research Program of China (2011CB910403) and the National Natural Science Foundation of China (30970692, 21075045).
Author information
Authors and Affiliations
Corresponding author
Additional information
Published in the topical collection Amino Acid Analysis with guest editor Toshimasa Toyo'oka.
Rights and permissions
About this article
Cite this article
Ou, G., Feng, X., Du, W. et al. Recent advances in microchip electrophoresis for amino acid analysis. Anal Bioanal Chem 405, 7907–7918 (2013). https://doi.org/10.1007/s00216-013-6830-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-013-6830-4