Abstract
Cancer is a very complex disorder, and it is urgent to find new ways to treat it. This study aims to evaluate the inhibition mechanism of ALK-5 (target related to breast cancer) from an electronic point of view. Computational simulations (QM/MM, NBO, QTAIM) were performed, and the ONIOM method (B3LYP/cc-pVDZ:UFF) was used to obtain the optimized geometries of the studied systems. The NBO analyses indicated that the most important electron transfer occurs between LP N (inhibitor 1) and BD* O–H (Tyr249) orbitals (ΔE2 = 11.89 kcal/mol). The weakest interaction occurs between the LP N (inhibitor 4) and BD* N–H (His283) orbitals (ΔE2 = 0.81 kcal/mol). The QTAIM analyses suggested that the most active inhibitors perform a greater number of hydrogen bonds with the major residues. Therefore, quantum mechanics methods proved to be important to better understand the inhibition of ALK-5, as well as helping the design of new inhibitors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Cancer is characterized by uncontrolled proliferation and growth of transformed cells. In the world, according to data from GLOBOCAN 2018, an estimated 18.1 million new cases of cancer and 9.6 million cancer-related deaths happened in that year [1]. Several studies show the importance of studying biological targets related to this chronic disease [2, 3]. One of the biological targets involved in the cancer cases is known as TGF-β receptor type I (transforming growth factor β), also called ALK-5 (activin receptor-like kinase 5), which is a biological receptor of the TGF-β superfamily [4]. This growth factor is responsible for several physiological processes, such as wound healing, immune system control, homeostasis and tumor suppression, and is also related to the recruitment of immune cells, apoptosis, among other biological events [5]. Thus, biological signaling of TGF-β occurs by type 1 receptors (TGF-β RI or ALK-5) and type 2 receptor (TGF-β RII). If this signaling is unregulated, TGF-β transforms into a tumor promoter, causing the spread of cancer cells and metastasis [6].
Several studies have shown that the ALK-5 inhibition is related to various types of tumors, such as pancreatic, breast and colon, and these studies have been done to induce the inhibition of TGF-β signaling [7,8,9]. In this paper, six molecules ((1 (4-([1, 2, 4] triazole [1,5-a] pyridin-6-yl)-N-(4-methoxyphenyl)-3-(6-methylpyridin-2-yl)-1H-pyrazole-1 carbothioamide, two molecules of 1-substituted-3-(6-methylpyridin-2-yl)-4-([1, 2, 4] triazole [1,5-a]pyridin-6-yl) pyrazole group (the most active inhibitors) and 3-((3-(6-methylpyridin-2-yl)-4-(1,5-naphthyridin-2-yl)-1H-pyrazol-1-yl) methyl) benzonitrile), N-(3-cyanophenyl)-3-(3-(6-methylpyridin-2-yl)-4-(quinolin-6-yl)-1H-pyrazol-1-yl) propanethioamide and 3-((3-(6-methylpyridin-2-yl)-4-(quinolin-6-yl)-1H-pyrazole-1-yl) methyl) benzonitrile (the least active compounds)) [10,11,12] capable of inhibiting ALK-5 were studied. In addition, the main interactions of these molecules with the major amino acid residues of the target binding site (structural water molecule, Lys232, Glu245, Tyr249, His283 and Asp351) [13] were also analyzed. Currently, there are several ways to study the ALK-5 inhibition, and one of them is the use of molecular modeling techniques and theoretical chemistry.
Of the diverse computational methodologies that can be employed, the present work uses a quantum mechanical approach to perform an electronic analysis of the system such as analyses on the electron transfers involved in the main interactions between inhibitors and the biological target. The first employed method was ONIOM (own n-layered integrated molecular orbital and molecular mechanics) [14,15,16] to generate the conformations related to the electronic structure of the six inhibitors, the structural water molecule and the main residues of ALK-5. After obtaining these conformations, NBO (natural bond orbital) [17,18,19,20,21] technique was used to verify the intensity of the hydrogen bonds. Moreover, QTAIM (quantum theory of atoms in molecules) [22,23,24] method was used to obtain the topology of the electron density of the molecular systems under study. Finally, calculations of interaction energy (ΔE) [25] were performed to analyze the electronic levels related to the stability of the hydrogen bonds being studied. In this way, studying the electronic interactions of a protein–inhibitor system is very important in the development of new possible drug candidates.
2 Computational details
2.1 Dataset
The tridimensional structure of ALK-5 used in all analyses was selected from PDB (Protein Data Bank [26]) with the code 3HBM [13] (Fig. S1). A set of six ALK-5 inhibitors has been chosen from the literature [10,11,12], which have IC50 values that were converted to pIC50 (−log IC50). Table 1 shows the structures of the selected compounds and their respective IC50 values.
2.2 Calculations of bond lengths and angles
To validate the basis set to be used in the ONIOM, NBO and QTAIM analyses, the bond lengths and angle values of 2-(6-methylpyridin-2-yl)-N-pyridin-4-ylquinazoline-4-amine (crystallographic inhibitor, PDB: 3HMM) [13] were analyzed by using DFT (density functional theory), with the B3LYP functional [27,28,29,30] and eight basis sets (Pople functions and a Dunning double—ζ base—cc-pVDZ, correlation consistent basis set with double ζ) [31, 32]. After determining the RMSD (root-mean-square deviation) between the theoretical and experimental values, the ONIOM method was used to obtain the conformation of the ligand–receptor complexes.
2.3 ONIOM analyses
The initial conformation of each ligand–receptor complex, used in the QM/MM (quantum mechanics/molecular mechanics, using the ONIOM method) calculations, was previously obtained [8], and, for these calculations, we employed Gaussian09 [33]. Initially, the binding site of ALK-5 was delimited at a radius of 5 Å, and this cut was done by using Pymol [34]. After selecting the binding site of the biological target, GaussView was used to select the high and low regions (quantum mechanics and classical mechanics, respectively) [14, 35]. The high layer of ONIOM has the following residues: Lys232, Glu245, Tyr249, His283 and Asp351, the inhibitors and the structural water molecule. For the top layer of the ONIOM calculations, B3LYP/cc-pVDZ (basis set that generated the best correlation with the experimental values of bond lengths and angles for the reference compound) was employed; the layer that compasses the protein environment was analyzed with the universal force field (UFF) [31, 36]. After obtaining the conformations of the ligand–receptor complexes from ONIOM, the next stage was the analysis of the NBO orbitals, whose main purpose is to evaluate the electron transfer levels that occur in the interactions between inhibitors and the main residues at the binding site of ALK-5.
2.4 NBO calculations
From the optimized geometries obtained from B3LYP/cc-pVDZ (high layer of ONIOM), natural bond orbitals (NBO) were determined with this same calculation level [20, 37]. The computational code used was NBO 3.1. The NBO model describes the electron density in terms of hybridization and covalent effects for polyatomic wave functions. This model can also be used in the study of electronic interactions that occur from a donor–acceptor point of view. For the study of ALK-5 inhibitors, NBO analysis was used to assess the formation of hydrogen bonds and electronic transfers related to the following interactions: inhibitors + Lys232, Glu245, Tyr249, His283, Asp351 and structural H2O. This type of intermolecular interaction occurs between electron-rich regions, in the case of lone pair (LP) and anti-ligand orbitals, forming the hydrogen bonds. About the second-order stabilization energy (ΔE2) of the NBO orbitals, the values for the interactions considered were higher than 0.7 kcal/mol. Equation 1 presents the calculation of (ΔE2) [20, 38]
From Eq. 1, it is possible to verify that the Fock operator is the term F and \( \varepsilon_{\sigma } {\text{ and }}\varepsilon_{{\sigma^{* } }} \) are the NBO orbital energies [20]. The NBO model was also used in the study of the electronic transfers that occur in the phosphorylation mechanism of ALK-5. The methodology used in the NBO calculations for this step was the same as previously used (B3LYP/cc-pVDZ). Afterward, the next step of this study was to analyze the electronic topology of the ligand–receptor interactions from the QTAIM technique [22, 39].
2.5 QTAIM analyses
The same structures of the complexes obtained from ONIOM (B3LYP/cc-PVDZ:UFF) were used to carry out the QTAIM analyses. This approach is based on the topological variables of electronic density (ρ) and Laplacian of electronic density \( (\nabla^{2} \rho ) \) and performs the mapping of the location and behavior of atoms in a chemical bond. Bond critical points (BCPs) were analyzed, which can be classified according to their classification (\( \omega \)) and signature (\( \sigma \)), being that \( \omega \) is related to the number of nonzero curvatures of \( \rho \) at a critical point and \( \sigma \) is the algebraic sum of the signs of the curvatures, that is, each curvature contributes ± 1 depending on whether it is positive or negative. Therefore, there are four types of stable and nonzero critical points: [40].
- (a)
(3, − 3) three negative curvatures: ρ is a local maximum;
- (b)
(3, − 1) two negative curvatures: ρ is a maximum in the plane defined by the corresponding eigenvectors, but is a minimum along the third axis that is perpendicular to that plane;
- (c)
(3, + 1) two positive curvatures: ρ is a minimum in the plane defined by the corresponding eigenvectors, but is a maximum along the third axis that is perpendicular to that plane;
- (d)
(3, + 3) three positive curvatures: ρ is a local minimum.
In addition to the definition of the different types of critical point, the Laplacian of electronic density is defined according to the QTAIM theory as an energy density, as is presented in Eq. 2 [23]:
where \( T(\overrightarrow {r)} \) and \( V (\overrightarrow {r)} \) are the kinetic and potential energy of the electron density \( \rho \left( {\vec{r}} \right) \), respectively. Equation 2 clearly shows the relation of Laplacian (\( \nabla^{2} \rho \left( {\vec{r}} \right) \)) with the atomic virial theorem, satisfying Eq. 3.
In other words, when \( \nabla^{2} \rho \left( {\vec{r}} \right) \) is negative, the potential energy will prevail, i.e., the electronic density is compressed in BCP. When \( \nabla^{2} \rho \left( {\vec{r}} \right) \) is positive, the kinetic energy in BCP is higher, indicating that the electronic charge is expanded with the charge density values of less than 0.1 u.a. [23].
Thus, single-point calculations (B3LYP/cc-pVDZ) were initially performed to obtain the wfx (AIM Extended Wavefunction Files) file, which is a newer version of the wfn file (AIM Traditional Wavefunction Files), and the data are presented cleanly and allow the writing of the data related to the wave function of the system [41]. All these single-point calculations were performed with Gaussian09. Hereafter, this file was used to obtain the topological variables ρ and \( \nabla^{2} \rho \) for the BCPs of the interactions between the six inhibitors, the main amino acid residues at the binding site active of ALK-5 and the structural water molecule. For this, we employed the AIMQB program (AIMALL subprogram) [41]. Several output files are obtained, and from the sumviz extension files, it is possible to obtain the molecular graphs, the gradient maps and the values obtained for ρ and \( \nabla^{2} \rho \) of the BCPs of the interactions of the studied systems (using the AIMStudio subprogram).
2.6 Stabilization of hydrogen bonds
From the QTAIM topological analysis, it was possible to verify the formation of intermolecular closed-layer interactions (hydrogen bonds) that are important in the study of the ALK-5 inhibition. To evaluate the level of stabilization of these closed-loop interactions, we calculated the interaction energy values, and the first step of this technique was the single-point calculation by using Gaussian09 (B3LYP/cc-pVDZ). The interaction energy calculation considers the energy values of the total system between inhibitor (or structural water molecule) + residues (or water molecule) as EAB and EA and EB for the energy values for the isolated molecules, as presented in Eq. 4. Therefore, from this approach, it is possible to evaluate the hydrogen bonding interactions that present greater stability (lower values of ΔE) [25]:
3 Results and discussion
3.1 Calculations of bond lengths and angles
The geometry optimization of 2-(6-methylpyridin-2-yl)-N-pyridin-4-ylquinazoline-4 (Fig. 1) was performed to choose a reliable basis set to be used in the ONIOM, NBO and QTAIM calculations. These calculations were performed to evaluate the basis set that presents the highest correlation with the experimental data (bond lengths and angles). This analysis was performed by varying only the basis set, while the functional was kept fixed (B3LYP). Supplementary Tables 1 and 2 (S1 and S2) display the experimental and calculated values of bond lengths and angles for the reference compound.
From Tables S1 and S2, it is possible to analyze the values of the experimental and theoretical bond lengths. The RMSD calculation was used to verify the highest correlation with the experimental data, as presented in Eq. 5 [42]:
Regarding the terms of Eq. 5, \( \hat{y}_{i} \) = experimental values and yi = values obtained computationally. Table 2 shows the RMSD values for the bond lengths.
The data presented in Table 2 show that the B3LYP density functional shows a higher correlation with the experimental data when it is calculated using the basis set cc-pVDZ (RMSE = 0.020 Å). For the analysis of the atomic angles, the obtained results are given in Tables S3 and S4, and these results show the values of experimental and theoretical bond angles. To evaluate the correlation between these data, RMSE errors were also calculated, which are given in Table 3.
From the results presented in Table 3, it is possible to note that the highest correlation between theoretical and experimental data is obtained from cc-pVDZ (RMSE = 3.11°). In addition to the comparison between the calculated and experimental values for the bond lengths and angles, the total energy values (E) were evaluated and the results are displayed in Fig. 2.
Values of total energy for the reference compound (2-(6-methylpyridin-2-yl)-N-pyridin-4-ylquinazoline-4-amine) obtained from different basis set and B3LYP. The blue baseline refers to the average of all calculated energies. The orange line corresponds to the individual energy values obtained for each basis set
From Fig. 2, it is possible to observe that the individual energies obtained from different basis sets are below the average (blue line), except for the smaller basis set (3-21G), which ended up raising the average. It is important to observe that the energy of the molecule tends to decrease with the increase in the size of the basis set. In the case of 6-31G, the addition of diffuse and polarization functions decreased the energy value. Since the energy values obtained with 6-311++G (d, p) are equivalent, we decided to use the Dunning basis due to its lower computational cost. Thus, from these analyses, we can conclude that the most appropriate basis set for the ONIOM calculations is cc-pVDZ.
3.2 ONIOM analyses
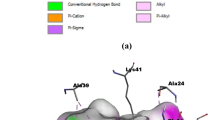
The ONIOM calculations were performed for the six ligand–receptor complexes using B3LYP/cc-pVDZ (for the region containing the inhibitors, the structural water molecule and the main residues—Lys232, Glu245, Tyr249, His283 and Asp351) and the universal force field (UFF) for the remaining environment. Figures 2 and 3 show the conformations obtained from the ONIOM calculations for the inhibitors 1 and 4 (the most and least active compounds, respectively). Figures S2 to S5 (Supplementary Material) display the conformations obtained for the other inhibitors at the binding site of ALK-5.
Figures 3 and 4 show the conformations obtained for the most and the least active inhibitors at the binding site of ALK-5 (high layer = B3LYP/cc-pVDZ; low layer = UFF). Thus, the ONIOM method was employed to generate the conformations based on the electronic structure of the six inhibitors, the major residues of the ALK-5 binding site and the structural water molecule. In this way, it is possible to analyze the hydrogen bonds that occur at the ALK-5 binding site by using the NBO method.
3.3 NBO calculations
The first step in the NBO calculations was to isolate the QM (quantum mechanics) part (the inhibitors, the structural water molecule and the residues from the conformations obtained by ONIOM (B3LYP/cc-pVDZ:UFF)) of the high region (B3LYP/cc-pVDZ). From the isolated structures, the single-point calculation (B3LYP/cc-pVDZ) was also performed from the NBO technique. From this model, hundreds of intramolecular and intermolecular interactions between the inhibitors, the amino acid residues and the structural water were obtained. However, to perform the analysis of the donor or acceptor natural bond orbitals, GaussView 4.1.2 was used to generate the figures containing the electronic clouds of the NBO orbitals. (Only the hydrogen bonds were considered.) The orbitals of the inhibitors 1 and 4 are displayed in Figs. 5 and 6. (The orbitals for the other four inhibitors are shown in Figs. S6–S9.)
Taking into account the donor and acceptor NBOs, it is possible to verify from Fig. 5 the electron density involved in the formation of six intermolecular hydrogen bonds between the inhibitor 1 and the binding site of ALK-5. Regarding the NBO orbitals related to the hydrogen bonds between the inhibitor 4 and the binding site of ALK-5, Fig. 6 shows that three intermolecular hydrogen bonds were formed. Thus, from the NBO analysis related to hydrogen bonds, it is possible to suggest that there is a greater inhibition of ALK-5 from the intermolecular interactions (hydrogen bonds) made with the inhibitor 1. For the analysis of the energy intensities of these hydrogen bonds formed between ALK-5 and the inhibitors 1 and 4, Table 4 presents the values of ΔE2 and the distances among the atoms that are part of the interactions. (For the other four inhibitors, these values are displayed in Tables S5–S9.)
From Table 4, it is possible to verify the intensities of the six intermolecular interactions that occur between the inhibitor 1, the structural water and the main residues at the binding site of ALK-5 from a NBO donor–acceptor point of view. The most intense interactions occur between the isolated pair of electrons of the nitrogen atom (LP N) present in the inhibitor 1 and the antibonding orbital located at the OH bond of Tyr249 (11.89 kcal/mol) and between the isolated pair of electrons of the oxygen atom (LP O) of the structural water and the antibonding π orbital located at the NH bond of Glu245 (4.78 kcal/mol). For the inhibitor 4, the most intense interactions occur between the isolated electron pair of the oxygen atom (LP O) of the structural water and the antibonding π orbital at the NH bond of Asp351; another interaction is also formed between the isolated pair of the oxygen atom (LP O) of the structural water and the OH antibonding π orbital located at Tyr249 (2.12 kcal/mol). From the NBO analysis for this complex, it was also possible to verify that the hydrogen bond between the inhibitor 4 and His283 presents a low value of ΔE2 (0.81 kcal/mol), suggesting that ALK-5 presents lower stability when interacting with this inhibitor.
Therefore, the NBO analyses indicated that the most active inhibitors (1, 2 and 3) present a greater number of hydrogen bonds from the highest values of ΔE2 (greater intensity of the NBO interaction). Also, the values of ΔE2 suggest that the electron transfer levels are higher when ALK-5 performs hydrogen bonds with the most active inhibitors. It is important to highlight that these results contribute to understand the electronic interactions that occur between the set of selected molecules and the binding site of ALK-5. Afterward, the QTAIM technique was employed to analyze the topology of intermolecular interactions.
3.4 QTAIM analyses
For the interactions of ALK-5 and the selected compounds, the same interactions obtained from NBO were analyzed with QTAIM. The BCP search for the six complexes was performed from the electronic density gradient \( \nabla \rho (\overrightarrow {r)} \). The gradient vector lines for the inhibitors 1 and 4 and their respective BCPs are displayed in Figs. 7, 8, 9 and 10. (For the other four inhibitors, these results are presented in Figs. S10–15.)
Figures 7, 8, 9 and 10 show lines around the atomic nuclei that refer to gradients of the electronic density \( \nabla \rho \) (r). At points where \( \nabla \rho \) (r) = 0, singularities are called critical points (CP). The Hessian trace of the electronic density characterizes these critical points. If the sum is equal to − 3, the critical point is an atomic nucleus (nuclear attractor critical point—NACP) characterized by the symbology (3, − 3). If the sum of the Hessian trace is equal to − 1, the critical point is called bond critical point (BCP), indicating that the electronic density is maximum in the plane formed by two atomic nuclei, remaining in the middle of the connection path between two atoms. Analyzing Figs. 7, 8, 9 and 10, we observed that the gradient of electronic density starts at infinity and ends at NACPs. BCPs are displayed between the red dot binding pathways, and they are related to the intra- and intermolecular interactions that occur between the six inhibitors and ALK-5. From Table 5, it is possible to identify the covalent bonds (\( \nabla^{2} \rho \) negative) and the hydrogen bonds (\( \nabla^{2} \rho \) positive) carried out in the complexes 1 and 4. (All BCPs obtained for the other complexes are shown in Supplementary Material.)
Table 5 shows the QTAIM results that characterize the chemical bonds and hydrogen bonds between the inhibitors and ALK-5. Taking into account the covalent bonds, the values of \( \nabla^{2} \rho \) are negative; for the unsaturated bonds, these values are positive (greater than 0.20 a.u.) and, for the intermolecular hydrogen bonds (BCPs in bold in Table 5), the signal of the Laplacian is positive and the values of \( \nabla^{2} \rho \) are smaller than 0.1 a.u. Thus, from the QTAIM analysis on the chemical bonds and intermolecular interactions, the next step involved the calculation of interaction energy (ΔE), which allows verifying the stability of the hydrogen bonds.
3.5 Stabilization of hydrogen bonds
To carry out a more detailed analysis of the hydrogen bonds, the interaction energy values were also calculated. For this, the isolated energies of the complexes (taking into account the studied intermolecular interactions, Ecomplex), the inhibitors, the structural water molecule and the main residues were calculated to obtain the ΔE values. The results obtained for the inhibitors 1 and 4 are presented in Table 6. (The results obtained for the other compounds are displayed in Supplementary Material.)
About the hydrogen bonds that occur between the most active inhibitors, the structural water and the binding site of ALK-5, we can see from the BCPs analyzed by the QTAIM method that the unbound electronic pair at nitrogen is the main responsible for the following intermolecular interactions:
- (a)
inhibitor 1 + H2O
- (b)
inhibitor 1 + Tyr249
- (c)
H2O + Lys232 (at the binding site with the inhibitor 1)
- (d)
H2O + Glu245 (at the binding site with the inhibitor 1)
- (e)
inhibitor 4 + His283
- (f)
H2O + Tyr249 (at the binding site with the inhibitor 4)
Analyzing the hydrogen bonds formed between the structural water and the main residues of ALK-5, we can observe that the electronic pair on the oxygen atoms is the main responsible for the following intermolecular interaction:
- (a)
H2O + Asp351 (at the binging site with the inhibitor 1)
- (b)
H2O + Asp351 (at the binding site with the inhibitor 4)
From Table 6, it is possible to verify that the inhibitor 1 makes the most stable interactions with ALK-5. One clue to this statement is the interaction that occurs between the inhibitors and His283. For the inhibitor 1, ΔE is equal to − 0.91 a.u., and for the inhibitor 4, ΔE is equal to +10.99 a.u. This result can be discussed using the QTAIM parameters, since there is a BCP between O2 and C17 (ρ = 0.0036 a.u. and \( \nabla^{2} \rho \) = 0.01 a.u., see Supplementary Material). The formation of an intermolecular BCP between these atoms can be explained by the stereochemical factor of the conformation of the inhibitor 4 at the binding site of ALK-5. In addition, these analyses also indicate hydrogen bonds with lower ΔE values for the inhibitor 1, suggesting its greater stability at the ALK-5 binding site. From these results, it is also possible to verify that the least active inhibitors make less stable hydrogen bonds and, in the case of the least active inhibitor (4), there is an unstable interaction. Thus, from the topological QTAIM analysis and the mapping of the closed-shell interactions, it was possible to analyze the importance of deep electronic analysis on the role of hydrogen bonds to understand the biological activity against ALK-5 presented by the selected inhibitors.
4 Conclusions
The computational study of ALK-5 and its inhibition was carried out to understand the electronic factors involved in this process. Initially, the validation of the theoretical methodology was performed by calculating the values of bond lengths and angles and compared them to experimental data. The results obtained in this step showed that the combination of B3LYP-cc-pVDZ provided better results when compared to experimental data obtained from the ligand structure found in the PDB structure (PDB code: 3HMM). From the choice of the theoretical methodology, QM/MM simulations were performed using the ONIOM technique. In this way, the conformations obtained from ONIOM (B3LYP/cc-pVDZ:UFF) were used in the NBO analyses, where information was obtained on the electron transfer levels of the hydrogen bonds that occur between ALK-5 and the six studied inhibitors. From this analysis, it was possible to verify a greater number of hydrogen bonds to the complex 1 (containing the most active inhibitor). Values of second-order perturbation energy (ΔE2) showed that the most intense interaction occurs between the isolated pair of electrons at the nitrogen atom (LP N) present in the inhibitor 1 and the antibonding orbital located at the OH group of Tyr249 (11.89 kcal/mol). The least intense interaction occurs between the isolated pair of electrons at the nitrogen atom (LP N) of the inhibitor 4 and the antibonding orbital NH at His283 (0.81 kcal/mol). Also, the QTAIM methodology indicated the formation of chemical interactions, such as covalent bonds and closed-shell interactions (intermolecular hydrogen bonds) for the major amino acid residues of ALK-5 and the respective inhibitors. For the most active inhibitor of the series (compound 1), it was possible to verify hydrogen bonds with the following species: structural water (ρ = 0.01 au and \( \nabla^{2} \rho \) = 0.04 au), Tyr249 (ρ = 0.02 au and \( \nabla^{2} \rho \) = 0.07 au) and His283 (ρ = 0.01 au and \( \nabla^{2} \rho \) = 0.04 au). The inhibitor 4 (the least active) makes only one hydrogen bond with His283 (ρ = 0.01 a.u. and \( \nabla^{2} \rho \) = 0.03 a.u.). To verify the stability of hydrogen bonds analyzed from the NBO and QTAIM techniques, interaction energy calculations were also performed, and the results indicated that the inhibitor 1 achieves the most stable hydrogen bonding (His283, ΔE = − 0.91 a.u.) and the least active inhibitor (4) makes a hydrogen bond with His283, but the value is 10.99 a.u., suggesting instability. Therefore, from the results obtained in this study, a contribution to the electronic features related to the main interactions of some inhibitors and ALK-5 was made, in particular to understand the mechanisms of action of ALK-5 and to design new drug candidates to treat illnesses like cancer.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Cancer J Clin 68(6):394–424. https://doi.org/10.3322/caac.21492
Hantschel O, Grebien F, Superti-Furga G (2012) The growing arsenal of ATP-competitive and allosteric inhibitors of BCR–ABL. Can Res 72:4890–4895
Duffy MJ (2013) The war on cancer: are we winning? Tumor Biol 34:1275–1284. https://doi.org/10.1007/s13277-013-0759-2
Huse M, Muir TW, Xu L, Chen Y-G, Kuriyan J, Massagué J (2001) The TGFβ receptor activation process. Mol Cell 8:671–682. https://doi.org/10.1016/S1097-2765(01)00332-X
Arjaans M, Oude Munnink T, Timmer-Bosscha H, Reiss M, Walenkamp AME, Lub-de Hooge MN, Au-de Vries EGE, Au-Schröder CP (2012) Transforming growth factor (TGF)-β expression and activation mechanisms as potential targets for anti-tumor therapy and tumor imaging. Pharmacol Therap 135
Massagué J (1998) TGF-β signal transduction. Annu Rev Biochem 67:753–791. https://doi.org/10.1146/annurev.biochem.67.1.753
de Caestecker MP, Piek E, Roberts AB (2000) Role of transforming growth factor-β signaling in cancer. J Natl Cancer Inst 92:1388–1402
Almeida MO, Costa CHS, Gomes GC, Lameira J, Alves CN, Honorio KM (2018) Computational analyses of interactions between ALK-5 and bioactive ligands: insights for the design of potential anticancer agents. J Biomol Struct Dyn 36(15):4010–4022. https://doi.org/10.1080/07391102.2017.1404938
Araujo SC, Maltarollo VG, Honorio KM (2013) Computational studies of TGF-βRI (ALK-5) inhibitors: analysis of the binding interactions between ligand–receptor using 2D and 3D techniques. Eur J Pharm Sci 49:542–549
Jin CH, Sreenu D, Krishnaiah M, Subrahmanyam VB, Rao KS, Nagendra Mohan AV, Park C-Y, Son J-Y, Son D-H, Park H-J, Sheen YY, Kim D-K (2011) Synthesis and biological evaluation of 1-substituted-3(5)-(6-methylpyridin-2-yl)-4-(quinoxalin-6-yl)pyrazoles as transforming growth factor-β type 1 receptor kinase inhibitors. Eur J Med Chem 46:3917–3925. https://doi.org/10.1016/j.ejmech.2011.05.063
Jin CH, Krishnaiah M, Sreenu D, Subrahmanyam VB, Rao KS, Mohan AVN, Park C-Y, Son J-Y, Sheen YY, Kim D-K (2011) Synthesis and biological evaluation of 1-substituted-3-(6-methylpyridin-2-yl)-4-([1,2,4]triazolo[1,5-a]pyridin-6-yl)pyrazoles as transforming growth factor-β type 1 receptor kinase inhibitors. Bioorg Med Chem Lett 21:6049–6053. https://doi.org/10.1016/j.bmcl.2011.08.064
Kim D-K, Lee Y-I, Lee YW, Dewang PM, Sheen YY, Kim YW, Park H-J, Yoo J, Lee HS, Kim Y-K (2010) Synthesis and biological evaluation of 4(5)-(6-methylpyridin-2-yl)imidazoles and -pyrazoles as transforming growth factor-β type 1 receptor kinase inhibitors. Bioorg Med Chem 18:4459–4467. https://doi.org/10.1016/j.bmc.2010.04.071
Gellibert F, Fouchet MH, Nguyen VL, Wang R, Krysa G, de Gouville AC, Huet S, Dodic N (2009) Design of novel quinazoline derivatives and related analogues as potent and selective ALK5 inhibitors. Bioorg Med Chem Lett 19:2277–2281. https://doi.org/10.1016/j.bmcl.2009.02.087
Dapprich S, Komáromi I, Byun KS, Morokuma K, Frisch MJ (1999) A new ONIOM implementation in Gaussian98. Part I. The calculation of energies, gradients, vibrational frequencies and electric field derivatives. J Mol Struct (Thoechem) 461–462:1–21. https://doi.org/10.1016/S0166-1280(98)00475-8
Vreven T, Morokuma K, Farkas Ö, Schlegel HB, Frisch MJ (2003) Geometry optimization with QM/MM, ONIOM, and other combined methods. I. Microiterations and constraints. J Comput Chem 24:760–769. https://doi.org/10.1002/jcc.10156
Vreven T, Byun KS, Komáromi I, Dapprich S, Montgomery JA, Morokuma K, Frisch MJ (2006) Combining quantum mechanics methods with molecular mechanics methods in ONIOM. J Chem Theory Comput 2:815–826. https://doi.org/10.1021/ct050289g
Reed AE, Weinhold F (1983) Natural bond orbital analysis of near-Hartree–Fock water dimer. J Chem Phys 78:4066–4073
Reed AE, Weinstock RB, Weinhold F (1985) Natural population analysis. J Chem Phys 83:735–746
Reed AE, Weinhold F, Curtiss LA, Pochatko DJ (1986) Natural bond orbital analysis of molecular interactions: theoretical studies of binary complexes of HF, H2O, NH3, N2, O2, F2, CO, and CO2 with HF, H2O, and NH3. J Chem Phys 84(10):5687–5705. https://doi.org/10.1063/1.449928
Reed AE, Curtiss LA, Weinhold F (1988) Intermolecular interactions from a natural bond orbital, donor–acceptor viewpoint. Chem Rev 88:899–926. https://doi.org/10.1021/cr00088a005
Glendening ED, Landis CR, Weinhold F (2012) Natural bond orbital methods. Wiley Interdisciplinary Reviews. Comput Mol Sci 2:1–42. https://doi.org/10.1002/wcms.51
Bader RFW (1985) Atoms in molecules. Acc Chem Res 18:9–15. https://doi.org/10.1021/ar00109a003
Bader RFW (1990) Atoms in molecules: a quantum theory. Oxford University Press, Oxford
Bader RFW (2002) Atoms in molecules. In: Encyclopedia of computational chemistry. Wiley. https://doi.org/10.1002/0470845015.caa012
Kirschner KN, Sorensen JB, Bowen JP (2007) Calculating interaction energies using first principle theories: consideration of basis set superposition error and fragment relaxation. J Chem Educ 84(7):1225. https://doi.org/10.1021/ed084p1225
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28(1):235–242
Cramer CJ (2004) Essentials of computational chemistry: theories and models, 2nd edn. Wiley, New York
Becke AD (1988) Density-functional exchange-energy approximation with correct asymptotic behavior. Phys Rev A 38:3098–3100
Lee C, Yang W, Parr RG (1988) Development of the Colle–Salvetti correlation-energy formula into a functional of the electron density. Phys Rev B 37(2):785–789. https://doi.org/10.1103/PhysRevB.37.785
Parr RG (1983) Density functional theory. Annu Rev Phys Chem 34:631–656. https://doi.org/10.1146/annurev.pc.34.100183.003215
Dunning TH (1989) Gaussian basis sets for use in correlated molecular calculations. I. The atoms boron through neon and hydrogen. J Chem Phys 90(2):1007–1023. https://doi.org/10.1063/1.456153
Ditchfield R, Hehre WJ, Pople JA (1971) Self-consistent molecular-orbital methods. IX. An extended Gaussian-type basis for molecular-orbital studies of organic molecules. J Chem Phys 54:724–728
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA (2009) Gaussian 09, Revision A. 1. Gaussian, Wallingford CT
Delano WL (2002) The PyMOL molecular graphics system
Gaussview (2007) Gaussian Inc. 4.1.2. edn
Rappe AK, Casewit CJ, Colwell KS, Goddard WA, Skiff WM (1992) UFF, a full periodic table force field for molecular mechanics and molecular dynamics simulations. J Am Chem Soc 114:10024–10035. https://doi.org/10.1021/ja00051a040
Almeida MO, Barros DAS, Araujo SC, Faria SHDM, Maltarollo VG, Honorio KM (2017) Study on molecular structure, spectroscopic properties (FTIR and UV–Vis), NBO, QTAIM, HOMO-LUMO energies and docking studies of 5-fluorouracil, a substance used to treat cancer. Spectrochim Acta Part A Mol Biomol Spectrosc 184(Supplement C):169–176. https://doi.org/10.1016/j.saa.2017.04.070
Foster JP, Weinhold F (1980) Natural hybrid orbitals. J Am Chem Soc 102:7211–7218. https://doi.org/10.1021/ja00544a007
Bader RFW, Larouche A, Gatti C, Carroll MT, MacDougall PJ, Wiberg KB (1987) Properties of atoms in molecules: dipole moments and transferability of properties. J Chem Phys 87:1142–1152
Matta CF, Boyd RJ (2007) The quantum theory of atoms in molecules: from solid state to DNA and drug design. Wiley-VCH, New York
Keith TA (2017) AIMAll (Version 17.11.14). TK Gristmill Software
Byrski A, Oplatkov Z, Carvalho M, Kisiel-Dorohinicki M (2014) Advances in intelligent modelling and simulation: simulation tools and applications. Springer, Berlin
Acknowledgements
The authors dedicated this work to Professor Fernando Rei Ornellas for his great work in computational chemistry in Brazil. The authors would like to thank FAPESP (2014/27189-9, 2016/24524-7, 2017/10118-0), CAPES and CNPq for funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
“Festschrift in honor of Prof. Fernando R. Ornellas” Guest Edited by Adélia Justino Aguiar Aquino, Antonio Gustavo Sampaio de Oliveira Filho & Francisco Bolivar Correto Machado.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Almeida, M.O., Faria, S.H.D.M. & Honorio, K.M. An electronic point of view on the inhibition of ALK-5 by bioactive candidates related to cancer. Theor Chem Acc 139, 91 (2020). https://doi.org/10.1007/s00214-020-02602-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00214-020-02602-2