Abstract
In recent years, genome-wide association studies (GWAS) have identified more than 300 validated associations between genetic variants and risk of approximately 70 common diseases. A small number of rare variants with a frequency of usually less than 1% are associated with a strongly enhanced risk, such as genetic variants of TP53, RB1, BRCA1, and BRCA2. Only a very small number of SNPs (with a frequency of more that 1% of the rare allele) have effects of a factor of two or higher. Examples include APOE4 in Alzheimer’s disease, LOXL1 in exfoliative glaucoma, and CFH in age-related macular degeneration. However, the majority of all identified SNPs have odds ratios between 1.1 and 1.5. In the case of urinary bladder cancer, all known SNPs that have been validated in sufficiently large populations are associated with odds ratios smaller than 1.5. These SNPs are located next to the following genes: MYC, TP63, PSCA, the TERT-CLPTM1L locus, FGFR3, TACC3, NAT2, CBX6, APOBEC3A, CCNE1, and UGT1A. It is likely that these moderate risk or “wimp SNPs” interact, and because of their high number, collectively have a strong influence on whether an individual will develop cancer or not. It should be considered that variants identified so far explain only approximately 5–10% of the overall inherited risk. Possibly, the remaining variance is due to an even higher number of SNPs with odds ratios smaller than 1.1. Recent studies have provided the following information: (1) The functions of genes identified as relevant for bladder cancer focus on detoxification of carcinogens, control of the cell cycle and apoptosis, as well as maintenance of DNA integrity. (2) Many novel SNPs are far away from the protein coding regions, suggesting that these SNPs are located on distant-acting transcriptional enhancers. (3) The low odds ratio of each individual bladder cancer-associated SNP is too low to justify reasonable preventive measures. However, if the recently identified SNPs interact, they may collectively result in a substantial risk that is of preventive relevance. In addition to the “novel SNPs” identified by the recent GWAS, at least 163 further variants have been reported in relation to bladder cancer, although they have not been consistently validated in independent case–control series. Moreover, given that only 60 of these 163 “old SNPs” are covered by the SNP chips used in the recent GWAS, there are in principle 103 published variants still awaiting validation or disproval. In future, besides identifying novel disease-associated rare variants by deep sequencing, it will also be important to understand how the already identified variants interact.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Frequency and risk factors of urinary bladder cancer
Urinary bladder cancer (UBC) is the 9th most common cancer worldwide, and the 13th most common cause for death from cancer (Parkin 2008). After removal of primary carcinomas, UBC frequently recurs leading to repeated surgery. The strongest known risk factors are cigarette smoking, occupational exposure to bladder carcinogens, particularly to aromatic amines and polycyclic aromatic hydrocarbons, and male gender. Recent substances drawing interest include azo colourants (Golka et al. 2004) and hair dyes (Bolt and Golka 2007).
Single nucleotide polymorphisms and SNP chip analysis
It is well known that single nucleotide polymorphisms contribute to interindividual differences in cancer susceptibility (Kiemeney et al. 2008, 2010; Hengstler et al. 1998; Arand et al. 1996; Saravana Devi et al. 2008; Hewitt et al. 2007; Gehrmann et al. 2008; Carmo et al. 2006; Cadenas et al. 2010; Hellwig et al. 2010). DNA from different individuals is identical for most base positions; however, variants are observed approximately every 500 bases. If a variant occurs in more than 1% of the population, it is defined as a single nucleotide polymorphism (SNP; Fig. 1a). Today, up to 900,000 SNPs and 900,000 copy number variations can be determined in a single analysis on SNP chips. This technique is based on hybridization of DNA from patients to oligonucleotides immobilized on a chip, which contains oligonucleotides with the two alleles on different spots (Fig. 1b). The patients’ DNA (after digestion with endonucleases) is labelled with fluorochromes; therefore, cluster plots can be obtained, which differentiate between the homozygous major allele (green dots in Fig. 1c), the homozygous minor allele (red triangles) and the heterozygous (blue squares) individuals that carry allele A on one and allele B on the other chromosome (Fig. 1c). The automatic clustering of the fluorescence intensities to separate between the three possible genotypes may lead to misclassification of patients in cluster plots—a problem that is frequently underestimated when performing genome-wide association studies (GWAS) with SNP chips. An example is provided in Fig. 1c, where the heterozygous patients have been misclassified, due to the use of a sub-optimal classification algorithm for a specific data set. Although misclassifications can easily be identified by manual inspection, this is not feasible for 900,000 SNPs and large numbers of patients. However, to avoid possible validation of false positive SNPs caused by misclassification in cluster plots, the candidate genes identified from a discovery group (see below) should be manually controlled.
a Exchange of a single base pair present in at least 1% of the population, T:A to C:G in the shown example, is defined as a single nucleotide polymorphism (SNP). The human genome contains approximately 3 million SNPs (picture from: “Dna-SNP.svg”, Wikipedia, author: David Hall, Gringer, licence: CC-by 2.5, http://creativecommons.org/licenses/by/2.5/deed.de). b Principle of a DNA microarray chip. Single-stranded DNA oligonucleotides function as DNA probes by hybridizing DNA fragments from the analysed sample whose nucleotide sequences are homologous. The technique can be applied to differentiate whether an A or a G is present in a certain sequence (from: Carr et al. 2008). c Cluster plot obtained from a DNA microarray. The x-axis (contrast) and the y-axis (strength) indicate transformed values of the two allele intensities SA and SB for the [A] and the [B] allele, respectively. They are defined as follows: Contrast = (SA − SB)/(SA + SB) and Strength = log(SA + SB) (BRLMM whitepaper: BRLMM: an Improved Genotype Calling Method for the GeneChip® Human Mapping 500K Array Set, Revision Date: 2006-04-14; http://media.affymetrix.com/support/technical/whitepapers/brlmm_whitepaper.pdf). Homozygous major alleles are plotted as green dots (right cluster), heterozygous genotypes as blue squares (middle cluster), homozygous minor alleles as red triangles (left cluster), and not determinable genotypes as grey crosses. The example illustrates the problem of misclassification of patients in cluster plots using automated systems. Some heterozygous patients have been misclassified as homozygous (red triangles instead of blue squares)
A second major problem is that false positive results are generally caused by multiple testing. When testing with P < 0.05 without adjustment for multiple testing, approximately five false positive SNPs will occur in 100 tested SNPs, with a potential outcome of approximately 45,000 false positives in 900,000 SNPs. One possibility to avoid false positives is performing an adjustment for multiple testing, for example by the Bonferroni technique. However, this technique is very conservative and rejects a large number of real positives. Therefore, a good strategy is to establish a “discovery group” where the most promising SNPs are identified. Only a small number of hypotheses will then be validated in an independent group, the so-called “follow-up group”. This study design has the additional advantage that analysis in the follow-up groups can be done using much cheaper conventional PCR, whereas the expensive SNP chips are limited to the discovery group.
Combining data from different GWAS to increase the power of the discovery group, it has to be noted that there is little overlap between arrays from different distributors (see Fig. 2a), and also handling data generated with larger arrays of the same distributor results in gaps of thousands of SNPs. To gain information about untyped SNPs, e.g. to investigate candidate regions in more detail or to combine data from different SNP assays, a number of imputation techniques are available (BEAGLE: Browning and Browning 2007, 2009; BIMBAM: Servin and Stephens 2007; fastPHASE: Scheet and Stephens 2006; GenABEL: Aulchenko et al. 2007; IMPUTE/SNPTEST: Marchini et al. 2007; Marchini and Howie 2008; MaCH: Li Y et al. 2010b; Plink: Purcell et al. 2007; TUNA: Nicolae 2006). These approaches are used to infer unknown genotypes using measured loci and linkage disequilibrium information, e.g. from reference panel data usually HapMap (Halperin and Stephan 2009a, b). Comparisons of imputation algorithms can be found in Pei et al. (2008, 2010), Browning (2008), Hao et al. (2009), and Yu and Schaid (2007). Potential difficulties due to ethnic differences between the study group and the reference panel were addressed by Huang et al. (2009).
a The Venn diagram illustrates relatively little overlap between two frequently applied SNP chips, the Affymetrix SNP Array 5.0 and the Illumina Human 610 Quad, and the SNP500Cancer database. b An overview of how many of the previously analysed SNPs (the so-called “old SNPs” in bladder cancer case–control series; Table 2, Supplemental Table 1) are present on currently used SNP chips. Considering that the recent large GWAS (Table 1) have been performed with Illumina chips only 60 of the 163 “old SNPs” could have been discovered by these SNP chip studies. Presence or absence of SNPs if identified by rs numbers was determined using the software R, version 2.12.1 and annotation from the meta-data packages pd.genomewidesnp.5 version 1.1.0 (Affymetrix SNP Chip 5.0) and human610quadv1bCrlmm version 1.0.2 (Illumina Human 610 Quad) and from the SNP500Cancer database (from: ftp://ftp-snp500cancer.nci.nih.gov/snp500Cancer/Genotypes/allgenes.tab, access date 05 Jan 2011)
Strategy for discovery and validation of SNPs
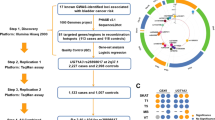
The study design with a discovery group and follow-up groups has recently been applied in order to identify new SNPs that are associated with urinary bladder cancer risk (Kiemeney et al. 2008, 2010; Wu et al. 2009; Rafnar et al. 2009; Rothman et al. 2010). For example, Kiemeney et al. (2010) studied 4,580 bladder cancer cases and 45,269 controls, where the discovery group consisted of 1,889 cases and 39,310 controls. The follow-up groups included 2,691 cases and 5,959 controls. The twenty most significant SNPs from the discovery group, all with P ≤ 2.5 × 10−5, were validated in the follow-up groups. This resulted in the identification of a SNP on chromosome 4p16.3 (rs798766) that was associated with bladder cancer risk in the discovery group (P = 2.4 × 10−5) and in the follow-up group (P = 8.5 × 10−8) (Kiemeney et al. 2010). Odds ratios (OR) for the T allele of rs7998766 were 1.22 (95% confidence interval: 1.11–1.34) in the discovery group and 1.26 (1.16–1.37) in the follow-up group. In the combined group, rs798766[T] was associated with an OR of 1.24 (P = 9.9 × 10−12). This association was significant, even after adjustment to cigarette smoking, age, and gender. No association between rs798766[T] and cigarette smoking or smoking quantity was obtained suggesting that rs798766 does not represent a genetic variation that confers susceptibility to addiction.
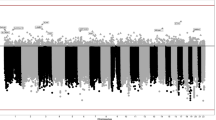
The rs798766 is located on intron 5 of TACC3 (transforming acidic coiled-coil containing protein 3), which is involved in the regulation of microtubule dynamics. TACC3’s relevance for bladder carcinogenesis is currently unknown; however, FGFR3, a neighbouring gene approximately 70 kb away from rs798766, contains activating mutations in about one-third of all bladder carcinomas. The critical role of FGFR3 in urinary bladder carcinogenesis led to the question whether the rs798766 polymorphism correlates with RNA levels of both FGFR3 and TACC3 (Kiemeney et al. 2010). Ideally, the optimal model for this study is cells originating from urinary bladder cancer, namely the epithelial cells of the bladder. Unfortunately, availability of bladder epithelial cells from tissue banks is limited; therefore, the analysis was performed in adipose tissue from 604 individuals. Interestingly, there was a significant correlation between rs798766 and RNA levels of both FGFR3 and TACC3 (Fig. 3). The highest RNA levels were obtained for the homozygous T allele, intermediate expression for the heterozygous C/T genotype, and lowest levels for the homozygous C allele. Currently, it is not known how the polymorphism of rs798766 influences expression of FGFR3 which is 70 kb away. A potential explanation is rs798766[T] leads to a conformational alteration of this region of the chromosome that improves accessibility of the transcriptional machinery. However, this remains speculative.
Correlation between rs798766 and RNA levels of FGFR3 and TACC3 in adipose tissue (from: Kiemeney et al. 2010)
A possible mechanism how rs798766[T] may contribute to bladder cancer risk is via the increased production of the FGFR3 protein as a consequence of enhanced gene expression, leading to an increased rate of proliferation and an increased probability for accumulation of mutations. The example of rs798766 illustrates that GWAS with discovery plus follow-up group design and sufficient case numbers can successfully be applied to identify novel disease-relevant SNPs.
State of the art: overview of recently discovered, validated SNPs
Using the above-described strategy for GWAS, nine SNPs have been identified since 2008 that are associated with bladder cancer risk (Table 1). Importantly, none of these SNPs have been previously described in relation to bladder cancer. In contrast to the newly discovered polymorphisms, the relevance of GSTM1 0/0, a deletion in the glutathione S-transferase M1 gene leading to loss of enzyme activity and the NAT2 polymorphism, was described by our group 15 years ago (Golka et al. 1996; Kempkes et al. 1996; Hengstler et al. 1998) and has been confirmed in the recent studies (Golka et al. 2009; Rothman et al. 2010).
An overview of the closest genes to the newly discovered and validated SNPs is given in Table 1. The well-known proto-oncogene c-Myc encodes a DNA-binding factor that activates or suppresses transcription, thus explaining its regulation of many target genes involved in the proliferation and cell cycle progression (Dominguez-Sola et al. 2007). TP63 shows strong homology to the tumour suppressor P53 and, similar to P53, is also involved in cell cycle and apoptosis control (Sayan et al. 2007; Lefkimmiatis et al. 2009). Prostate stem cell antigen (PSCA) is a cell surface protein associated with prostate and other types of cancer (Watabe et al. 2002). The TERT gene represents the reverse transcriptase component of telomerase that is essential for the maintenance of DNA length and cellular immortality (Cheung and Deng 2008; Florl and Schulz 2008). Little is known about the function of cleft lip and palate transmembrane 1-like gene (CLPTM1L). CLPTM1L is upregulated in cisplatin-resistant cell lines and might be involved in apoptosis control (Liu et al. 2010). Fibroblast growth factor receptor 3 (FGFR3) belongs to a family of polypeptide growth factors containing a cytoplasmic tyrosine kinase domain and extracellular immunoglobulin-like domains and is involved in mitogenesis, angiogenesis, and wound healing (Keegan et al. 1991). TACC3 plays a role in the maintenance of nuclear envelope structure and in cell division control (Gómez-Baldó et al. 2010). NAT2 is a phase II metabolizing enzyme involved in detoxification, but also bioactivation of xenobiotics (Hengstler et al. 1998). Low NAT2 activity is associated with increased bladder cancer risk in Caucasians. Chromobox homolog 7 (CBX7) positively regulates E-cadherin expression by interacting with histone deacetylase 2 (Federico et al. 2009), possibly explaining why loss of CBX7 expression is associated with a highly malignant phenotype of carcinomas. APOBEC3 deaminases cause G to A hypermutation in nascent DNA of hepatitis B viruses which seems to play a role in antiviral defence (Abe et al. 2009). Overexpression of APOBEC3 genes may lead to mutations in the genome and influence the development of tumours (Vartanian et al. 2008). Cyclin E (CCNE1) controls cell cycle progression at the G1/S transition (Koff et al. 1991). UDP-glucuronosyltransferase 1A (UGT1A) is a phase II metabolizing enzyme that catalyses glucuronidation and elimination of numerous lipophilic xenobiotics, thereby acting as a detoxifying enzyme (Hengstler et al. 2000; Strassburg et al. 2008). Glutathione S-transferase M1 is also a phase II metabolizing enzyme (Bolt and Thier 2006) that upon conjugation with glutathione detoxifies numerous xenobiotics including polycyclic aromatic hydrocarbons that are known bladder carcinogens (Golka et al. 2009). In conclusion, the functions of the discussed genes focus on carcinogen detoxification, control of the cell cycle as well as apoptosis, and maintenance of DNA integrity.
More than 75 further studies of SNPs and bladder cancer risk
A literature search on SNPs and bladder cancer identified more than 75 further studies reporting on genetic variants in bladder cancer (Tables 2, 3, Supplemental Table 1). Most of these SNPs (n = 34) affect genes encoding xenobiotic metabolizing enzymes (Table 2). Moreover, variants of genes have been reported that play a role in DNA repair and damage signalling (n = 58), cell cycle control, DNA replication, translesion synthesis and transcription (n = 14), inflammation (n = 13), apoptosis (n = 8), methylation (n = 4), growth factors (n = 5), matrix metalloproteinases (n = 5), mTOR and associated factors (n = 4), and others (n = 18) (Table 3, Supplemental Table 1). However, in contrast to the SNPs summarized in the previous paragraph and in Table 1, the latter variants have not been consistently validated in independent case–control series. Therefore, we analysed the fraction of the 163 SNPs from Table 3, Supplemental Table 1 (“old SNPs”) that are represented on the SNP chips from the previously published GWAS (Table 1), because in this case it is unlikely that the “old SNPs” could be verified in genome-wide studies. Considering that the recent GWAS (Table 1) were performed with Illumina chips, it can be concluded that 60 of the “old SNPs” were re-analysed in the Illumina SNP chip studies but not confirmed. However, 103 of the “old SNPs” are not currently present on the Illumina SNP chip and therefore represent candidates that could be validated (or disproven) in future. Of course, most of the mentioned SNPs are likely to be in linkage disequilibrium with markers on the SNP chips and some of their neighbouring SNPs might indeed show small P values. However, not reaching genome-wide significance, the association with “old SNPs” is unlikely to be recognized. Nevertheless, final answers will most probably be provided when the next generation sequencing studies (see paragraph below) are finished.
Implications for prevention?
A common feature of the novel bladder cancer-associated SNPs is the relatively low odds ratio (<1.5), when compared to heavy cigarette smoking that is associated with an odds ratio of approximately four (e.g. OR = 3.69; 95% CI = 2.97–4.67; P = 8.1 × 10−33 in Lehmann et al. 2010). Generally, an odds ratio of 1.5 is of minor relevance for an individual and would not justify additional medical precautionary measures. However, a relevant question is whether the high-risk alleles of several of the SNPs in Table 1 interact, leading to odds ratios that are similar or even higher than that of cigarette smoking. If so, one should consider that individuals carrying combinations of several high-risk alleles are rare. For example, only approximately eight in 1,000 individuals may carry a combination of four high-risk alleles if the frequency of the individual high-risk alleles is 0.3. Nevertheless, identifying these individuals would be advantageous, especially if adequate measures of precaution such as cystoscopy or specific imaging techniques, in the case of bladder cancer, are too expensive or too invasive to be applied to the general population. The above example illustrates that studies are required to test whether influential SNPs interact and add together to create odds ratios that would justify preventive measures.
Strong impact of “wimp SNPs”
To date, GWAS have identified more than 300 validated associations between genetic variants and approximately 70 common diseases (Hindorff et al. 2009; http://www.genome.gov/gwastudies). Only a small number of rare variants with a frequency of usually much less than 1% is associated with a strongly enhanced risk, such as genetic variants of TP53, RB1, BRCA1, and BRCA2. A very small number of SNPs have effects of a factor of two or higher, for example APOE4 in Alzheimer’s disease, LOXL1 in exfoliative glaucoma, and CFH in age-related macular degeneration (Altshuler et al. 2008). Nevertheless, the majority of all identified SNPs have odds ratios between 1.1 and 1.5. In the case of urinary bladder cancer, all known SNPs increase risk by a factor smaller than 1.5. It is likely that these “wimp SNPs” interact and that complex combinations of several SNPs have a strong influence on whether an individual will develop cancer or not.
However, no comprehensive studies on “wimp SNPs” and “wimp-SNP”-environment interactions are currently available. It should also be noted that variants identified so far, including the novel SNPs identified by GWAS, explain only approximately 5–10% of the overall inherited risk (Altshuler et al. 2008). The remaining variance may be due to an even higher number of SNPs with odds ratios smaller than 1.1; however, the case numbers of most completed GWAS were too small to identify variants associated with such low risk. Nevertheless, it is likely that the “wimpiest-wimp SNPs” with odds ratios smaller than 1.1 are collectively even stronger (Varghese and Easton 2010). Additionally, the locus-attributable risk may have been underestimated, because the marker SNPs identified in GWAS were suboptimal proxies for the causal mutations (Altshuler et al. 2008). The genetic variance explained by the variants identified so far may also have been underestimated, because gene–gene and gene–environment interactions have not yet been adequately considered. Finally, many rare variants may remain undiscovered, because they cannot be identified by SNP chip analysis but require systematic sequencing.
SNPs of distant-acting enhancers
Many of the recently discovered SNPs associated with bladder cancer risk are located in non-coding regions. Examples are the sequence variant rs9642880 on chromosome 8q24, which is 30 kb upstream of Myc (Kiemeney et al. 2008) and rs798766 on 4q16.3, 70 kb from FGFR3 (Kiemeney et al. 2010). Both SNPs are located so far from the exons whose expression they influence that the effect cannot be explained by linkage disequilibrium. In principle, it is not surprising that non-coding sequences can play an important role. It is well known that approximately 5% of the human genome is evolutionary conserved, and only less than one-third of this 5% consists of coding genes (Mouse Genome Sequencing Consortium et al. 2002). One possibility that might therefore explain these hits from GWAS is that the respective non-coding regions contain distant-acting enhancers (Visel et al. 2009). Distant-acting transcriptional enhancers represent sequences that can be located either downstream or upstream of the target gene or even within other genes. They consist of aggregations of transcription factor binding sites. Occupancy of these transcription factor binding sites leads to recruitment of transcriptional co-activators and chromatin remodelling. The protein aggregates at the distant-acting enhancer facilitate DNA looping, whereby the enhancer relocates to physical proximity of the target gene promoter and finally activates transcription by RNA polymerase II (Visel et al. 2009). This mechanism would also explain why some of the novel SNPs act in a tissue specific manner enhancing, for example, the risk of urinary bladder cancer but not that of breast cancer (Kiemeney et al. 2008). In any tissue, only a subset of enhancers is active, because only a specific set of transcription factors is formed. Therefore, it is plausible that rs9642880 and rs798766 are located within urothelium-specific enhancers. In future, it will be interesting to study whether the relatively high number of cancer-associated SNPs in non-coding regions identify distant-acting transcriptional enhancers.
Do SNPs differentiate between bladder cancer with and without exposure to bladder carcinogens?
In most countries, the eligibility criteria for occupational disability compensation are restrictive. Additional criteria that help identify cases where past occupational exposure has contributed to carcinogenesis are welcome. Therefore, it would be relevant to analyse whether specific SNP patterns can differentiate between urinary bladder carcinomas with and without occupational exposure to carcinogens. Recent evidence suggests that such differentiation is possible (Golka et al. 2009). Occupational exposure to aromatic amines and polycyclic aromatic hydrocarbons (PAHs) has been documented in several case–control series (Table 4A). The Wittenberg case–control series is a hospital-based study comprising only a relatively small fraction of individuals with occupational exposure to aromatic amines or PAHs (Table 4A). In contrast, the “Occupational case–control series” comprises individuals that have been evaluated for bladder cancer as an occupational disease, showing a high fraction of individuals exposed to aromatic amines (61%) and PAHs (27%). The Dortmund case–control series comprises former workers from the coal, iron, and steel industries and contains the highest fraction of individuals exposed to PAHs (52%) but not to aromatic amines (0%). Interestingly, GSTM1 0/0 was significantly associated with bladder cancer in the two case–control series with relatively high exposure to PAHs (Table 4B). In contrast, no significant association of GSTM1 0/0 was obtained in the Wittenberg case–control series with a relatively low number of individuals exposed to PAHs. On the other hand, rs9642880[T] was significantly associated with bladder cancer risk in the Wittenberg and not in the occupational as well as the Dortmund case–control series (Table 4B). This result suggests that the quality and quantity of exposure to bladder carcinogens determines which SNPs are relevant. In the case of exposure to certain carcinogens, the influence of SNPs on relevant detoxifying enzymes may increase. However, this concept must be discussed with caution for different reasons: firstly, because of the relatively small case numbers in the current study and secondly because of the lack of a consistent interaction of GSTM1 0/0 and cigarette smoking. In a recently published meta-analysis, GSTM1 0/0 was reported to be associated with a similarly increased risk in smokers and non-smokers (García-Closas et al. 2005), which speaks against an enhanced role of GSTM1 0/0 in the presence of cigarette smoke-associated carcinogens. On the other hand, NAT2 slow acetylators have been reported to be especially susceptible to the adverse effects of cigarette smoking on bladder cancer risk (García-Closas et al. 2005). Further studies are needed to analyse whether bladder carcinomas with and without occupational exposure to carcinogens can be differentiated by SNP patterns. Such gene–environment interactions may be particularly interesting for the recently discovered SNP of the detoxifying enzyme UGT1A (Table 1).
Future perspective: next generation sequencing
Sequencing of the first human genome took approximately 50 years and an investment of more than two billion Euros. With the advent of deep sequencing, the required time has been reduced to weeks. Within the next 10 years, the time required for sequencing of a human genome may be reduced to less than 1 day. Therefore, it can be expected that GWAS with SNP chips will soon be replaced by whole genome sequencing, thus allowing access to critical further information, particularly rare mutations. As a consequence, this will further increase the problem of multiple testing and even larger case–control series will be needed. Nevertheless, deep sequencing will, for the first time, give the opportunity to analyse comprehensively and quantitatively the degree to which interindividual differences within our genome contribute to our overall cancer risk.
References
Abbasi R, Ramroth H, Becher H et al (2009) Laryngeal cancer risk associated with smoking and alcohol consumption is modified by genetic polymorphisms in ERCC5, ERCC6 and RAD23B but not by polymorphisms in five other nucleotide excision repair genes. Int J Cancer 125:1431–1439
Abe H, Ochi H, Maekawa T et al (2009) Effects of structural variations of APOBEC3A and APOBEC3B genes in chronic hepatitis B virus infection. Hepatol Res 39:1159–1168
Ahirwar DK, Mandhani A, Dharaskar A, Kesarwani P, Mittal RD (2009) Association of tumour necrosis factor-alpha gene (T-1031C, C-863A, and C-857T) polymorphisms with bladder cancer susceptibility and outcome after bacille Calmette-Guérin immunotherapy. BJU Int 104:867–873
Ahirwar DK, Mandhani A, Mittal RD (2010) IL-8-251 T> A polymorphism is associated with bladder cancer susceptibility and outcome after BCG immunotherapy in a northern Indian cohort. Arch Med Res 41:97–103
Alakus H, Afriani N, Warnecke-Eberz U et al (2010) Clinical impact of MMP and TIMP gene polymorphisms in gastric cancer. World J Surg 34:2853–2859
Altshuler D, Daly MJ, Lander ES (2008) Genetic mapping in human disease. Science 322(5903):881–888
Andrew AS, Gui J, Sanderson AC et al (2009) Bladder cancer SNP panel predicts susceptibility and survival. Hum Genet 125:527–539
Arand M, Mühlbauer R, Hengstler J et al (1996) A multiplex polymerase chain reaction protocol for the simultaneous analysis of the glutathione S-transferase GSTM1 and GSTT1 polymorphisms. Anal Biochem 236:184–186
Arizono K, Osada Y, Kuroda Y (2008) DNA repair gene hOGG1 codon 326 and XRCC1 codon 399 polymorphisms and bladder cancer risk in a Japanese population. Jpn J Clin Oncol 38:186–191
Aulchenko YS, Ripke S, Isaacs A, van Duijn CM (2007) GenABEL: an R library for genome-wide association analysis. Bioinformatics 23:1294–1296
Bégin P, Tremblay K, Daley D et al (2007) Association of urokinase-type plasminogen activator with asthma and atopy. Am J Respir Crit Care Med 175:1109–1116
Berman DM, Wang Y, Liu Z et al (2004) A functional polymorphism in RGS6 modulates the risk of bladder cancer. Cancer Res 64:6820–6826
Bezemer ID, Doggen CJ, Vos HL, Rosendaal FR (2007) No association between the common MTHFR 677C->T polymorphism and venous thrombosis: results from the MEGA study. Arch Intern Med 167:497–501 (also included in cumulative thesis: Bezemer ID (2009) Genetic variation and susceptibility to venous thrombosis, Leiden University)
Bolt HM, Golka K (2007) The debate on carcinogenicity of permanent hair dyes: new insights. Crit Rev Toxicol 37:521–536
Bolt HM, Thier R (2006) Relevance of the deletion polymorphisms of the glutathione S-transferases GSTT1 and GSTM1 in pharmacology and toxicology. Curr Drug Metab 7:613–628
Brockmöller J, Cascorbi I, Kerb R, Roots I (1996) Combined analysis of inherited polymorphisms in arylamine N-acetyltransferase 2, glutathione S-transferases M1 and T1, microsomal epoxide hydrolase, and cytochrome P450 enzymes as modulators of bladder cancer risk. Cancer Res 56:3915–3925
Browning SR (2008) Missing data imputation and haplotype phase inference for genome-wide association studies. Hum Genet 124:439–450
Browning SR, Browning BL (2007) Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am J Hum Genet 81:1084–1097
Browning BL, Browning SR (2009) A unified approach to genotype imputation and haplotype-phase inference for large data sets of trios and unrelated individuals. Am J Hum Genet 84:210–223
Cadenas C, Franckenstein D, Schmidt M et al (2010) Role of thioredoxin reductase 1 and thioredoxin interacting protein in prognosis of breast cancer. Breast Cancer Res 12:R44
Cai DW, Liu XF, Bu RG et al (2009) Genetic polymorphisms of MTHFR and aberrant promoter hypermethylation of the RASSF1A gene in bladder cancer risk in a Chinese population. J Int Med Res 37:1882–1889
Carmo H, Brulport M, Hermes M et al (2006) Influence of CYP2D6 polymorphism on 3,4-methylenedioxymethamphetamine (‘Ecstasy’) cytotoxicity. Pharmacogenet Genomics 16:789–799
Carr SM, Marshall HD, Duggan AT et al (2008) Phylogeographic genomics of mitochondrial DNA: highly-resolved patterns of intraspecific evolution and a multi-species, microarray-based DNA sequencing strategy for biodiversity studies. Comp Biochem Physiol Part D Genomics Proteomics 3:1–11
Cascorbi I, Roots I, Brockmöller J (2001) Association of NAT1 and NAT2 polymorphisms to urinary bladder cancer: significantly reduced risk in subjects with NAT1*10. Cancer Res 61:5051–5056
Castillejo A, Mata-Balaguer T, Guarinos C et al (2009) The Int7G24A variant of transforming growth factor-beta receptor type I is a risk factor for colorectal cancer in the male Spanish population: a case–control study. BMC Cancer 9:406
Chang CH, Chiu CF, Wang HC et al (2009a) Significant association of ERCC6 single nucleotide polymorphisms with bladder cancer susceptibility in Taiwan. Anticancer Res 29:5121–5124
Chang CH, Chiu CF, Liang SY et al (2009b) Significant association of Ku80 single nucleotide polymorphisms with bladder cancer susceptibility in Taiwan. Anticancer Res 29:1275–1279
Chang CH, Wang RF, Tsai RY et al (2009c) Significant association of XPD codon 312 single nucleotide polymorphism with bladder cancer susceptibility in Taiwan. Anticancer Res 29:3903–3907
Chang CH, Chang CL, Tsai CW et al (2009d) Significant association of an XRCC4 single nucleotide polymorphism with bladder cancer susceptibility in Taiwan. Anticancer Res 29:1777–1782
Chao C, Zhang ZF, Berthiller J, Boffetta P, Hashibe M (2006) NAD(P)H: quinone oxidoreductase 1 (NQO1) Pro187Ser polymorphism and the risk of lung, bladder, and colorectal cancers: a meta-analysis. Cancer Epidemiol Biomarkers Prev 5:979–987
Chen WC, Wu HC, Hsu CD, Chen HY, Tsai FJ (2002) p21 gene codon 31 polymorphism is associated with bladder cancer. Urol Oncol 7:63–66
Chen T, Jackson C, Costello B et al (2004) An intronic variant of the TGFBR1 gene is associated with carcinomas of the kidney and bladder. Int J Cancer 112:420–425
Chen M, Cassidy A, Gu J et al (2009) Genetic variations in PI3K-AKT-mTOR pathway and bladder cancer risk. Carcinogenesis 12:2047–2052
Cheung AL, Deng W (2008) Telomere dysfunction, genome instability and cancer. Front Biosci 13:2075–2090
Chiu CF, Tsai MH, Tseng HC et al (2008) A novel single nucleotide polymorphism in XRCC4 gene is associated with oral cancer susceptibility in Taiwanese patients. Oral Oncol 44:898–902
Choi JY, Lee KM, Cho SH et al (2003) CYP2E1 and NQO1 genotypes, smoking and bladder cancer. Pharmacogenetics 13:349–355
Choudhury A, Elliott F, Iles MM et al (2008) Analysis of variants in DNA damage signalling genes in bladder cancer. BMC Med Genet 9:69
Cox DG, Hankinson SE, Kraft P, Hunter DJ (2004) No association between GPX1 Pro198Leu and breast cancer risk. Cancer Epidemiol Biomarkers Prev 1821–1822
de Verdier PJ, Sanyal S, Bermejo JL, Steineck G, Hemminki K, Kumar R (2010) Genotypes, haplotypes and diplotypes of three XPC polymorphisms in urinary-bladder cancer patients. Mutat Res 694:39–44
Dominguez-Sola D, Ying CY, Grandori C, Ruggiero L, Chen B, Li M, Galloway DA, Gu W, Gautier J, Dalla-Favera R (2007) Non-transcriptional control of DNA replication by c-Myc. Nature 448:445–451
Federico A, Pallante P, Bianco M et al (2009) Chromobox protein homologue 7 protein, with decreased expression in human carcinomas, positively regulates E-cadherin expression by interacting with the histone deacetylase 2 protein. Cancer Res 69:7079–7087
Figueroa JD, Malats N, Real FX et al (2007) Genetic variation in the base excision repair pathway and bladder cancer risk. Hum Genet 121:233–242
Figueroa JD, Malats N, García-Closas M et al (2008) Bladder cancer risk and genetic variation in AKR1C3 and other metabolizing genes. Carcinogenesis 29:1955–1962
Florl AR, Schulz WA (2008) Chromosomal instability in bladder cancer. Arch Toxicol 82:173–182
Frank B, Hemminki K, Shanmugam KS et al (2005) Association of death receptor 4 haplotype 626C–683C with an increased breast cancer risk. Carcinogenesis 26:1975–1977
Galli P, Cadoni G, Volante M et al (2009) A case-control study on the combined effects of p53 and p73 polymorphisms on head and neck cancer risk in an Italian population. BMC Cancer 9:137
Gangwar R, Ahirwar D, Mandhani A, Mittal RD (2009a) Do DNA repair genes OGG1, XRCC3 and XRCC7 have an impact on susceptibility to bladder cancer in the North Indian population? Mutat Res 680:56–63
Gangwar R, Mandhani A, Mittal RD (2009b) Caspase 9 and caspase 8 gene polymorphisms and susceptibility to bladder cancer in north Indian population. Ann Surg Oncol 16:2028–2034
Gangwar R, Mandhani A, Mittal RD (2011) Functional polymorphisms of cyclooxygenase-2 (COX-2) gene and risk for urinary bladder cancer in North India. Surgery 149:126–134
García-Closas M, Malats N, Real FX et al (2006) Genetic variation in the nucleotide excision repair pathway and bladder cancer risk. Cancer Epidemiol Biomarkers Prev 15:536–542
García-Closas M, Malats N, Real FX et al (2007) Large-scale evaluation of candidate genes identifies associations between VEGF polymorphisms and bladder cancer risk. PLoS Genet 3:e29
García-Closas M, Malats N, Silverman D et al (2005) NAT2 slow acetylation, GSTM1 null genotype, and risk of bladder cancer: results from the Spanish Bladder Cancer Study and meta-analyses. Lancet 366(9486):649–659
Gaudet MM, Gammon MD, Santella RM et al (2005) MnSOD Val-9Ala genotype, pro- and anti-oxidant environmental modifiers, and breast cancer among women on Long Island, New York. Cancer Causes Control 16:1225–1234
Gehrmann M, Schmidt M, Brase JC, Roos P, Hengstler JG (2008) Prediction of paclitaxel resistance in breast cancer: is CYP1B1*3 a new factor of influence? Pharmacogenomics 9:969–974
Golka K, Prior V, Blaszkewicz M et al (1996) Occupational history and genetic N-acetyltransferase polymorphism in urothelial cancer patients of Leverkusen, Germany. Scand J Work Environ Health 22:332–338
Golka K, Kopps S, Myslak ZW (2004) Carcinogenicity of azo colorants: influence of solubility and bioavailability. Toxicol Lett 151:203–210
Golka K, Hermes M, Selinski S et al (2009) Susceptibility to urinary bladder cancer: relevance of rs9642880[T], GSTM1 0/0 and occupational exposure. Pharmacogenet Genomics 19:903–906
Gómez-Baldó L, Schmidt S, Maxwell CA et al. (2010) TACC3-TSC2 maintains nuclear envelope structure and controls cell division. Cell Cycle 9(6) (Epub ahead of print)
Halperin E, Stephan DA (2009a) SNP imputation in association studies. Nat Biotechnol 27:349–351
Halperin E, Stephan DA (2009b) Maximizing power in association studies. Nat Biotechnol 27:255–256
Hao K, Chudin E, McElwee J, Schadt EE (2009) Accuracy of genome-wide imputation of untyped markers and impacts on statistical power for association studies. BMC Genet 10:27
Haq I, Chappell S, Johnson SR et al (2010) Association of MMP-2 polymorphisms with severe and very severe COPD: a case control study of MMPs-1, 9 and 12 in a European population. BMC Med Genet 11:7
Harries LW, Stubbins MJ, Forman D, Howard GC, Wolf CR (1997) Identification of genetic polymorphisms at the glutathione S-transferase Pi locus and association with susceptibility to bladder, testicular and prostate cancer. Carcinogenesis 18:641–644
Hazra A, Chamberlain RM, Grossman HB, Zhu Y, Spitz MR, Wu X (2003) Death receptor 4 and bladder cancer risk. Cancer Res 63:1157–1159
Hebbring SJ, Adjei AA, Baer JL et al (2007) Human SULT1A1 gene: copy number differences and functional implications. Hum Mol Genet 16:463–470
Hellwig B, Hengstler JG, Schmidt M, Gehrmann MC, Schormann W, Rahnenführer J (2010) Comparison of scores for bimodality of gene expression distributions and genome-wide evaluation of the prognostic relevance of high-scoring genes. BMC Bioinformatics 11:276
Hengstler JG, Arand M, Herrero ME, Oesch F (1998) Polymorphisms of N-acetyltransferases, glutathione S-transferases, microsomal epoxide hydrolase and sulfotransferases: influence on cancer susceptibility. Recent Results Cancer Res 154:47–85
Hengstler JG, Utesch D, Steinberg P et al (2000) Cryopreserved primary hepatocytes as a constantly available in vitro model for the evaluation of human and animal drug metabolism and enzyme induction. Drug Metab Rev 32:81–118
Hewitt NJ, Lechón MJ, Houston JB et al (2007) Primary hepatocytes: current understanding of the regulation of metabolic enzymes and transporter proteins, and pharmaceutical practice for the use of hepatocytes in metabolism, enzyme induction, transporter, clearance, and hepatotoxicity studies. Drug Metab Rev 39:159–234
Hindorff LA, Sethupathy P, Junkins HA et al (2009) Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci USA 106:9362–9367
Huang L, Li Y, Singleton AB, Hardy JA, Abecasis G, Rosenberg NA, Scheet P (2009) Genotype-imputation accuracy across worldwide human populations. Am J Hum Genet 84:235–250
Human Arylamine N-Acetyltransferase Gene Nomenclature (2010) http://louisville.edu/medschool/pharmacology/consensus-human-arylamine-n-acetyltransferase-gene-nomenclature/
Hung RJ, Boffetta P, Brennan P et al (2004) GST, NAT, SULT1A1, CYP1B1 genetic polymorphisms, interactions with environmental exposures and bladder cancer risk in a high-risk population. Int J Cancer 110:598–604
Johne A, Roots I, Brockmöller J (2003) A single nucleotide polymorphism in the human H-ras proto-oncogene determines the risk of urinary bladder cancer. Cancer Epidemiol Biomarkers Prev 12:68–70
Keegan K, Johnson DE, Williams LT, Hayman MJ (1991) Isolation of an additional member of the fibroblast growth factor receptor family, FGFR-3. Proc Nat Acad Sci USA 88:1095–1099
Kempkes M, Golka K, Reich S, Reckwitz T, Bolt HM (1996) Glutathione S-transferase GSTM1 and GSTT1 null genotypes as potential risk factors for urothelial cancer of the bladder. Arch Toxicol 71:123–126
Kiemeney LA, van Houwelingen KP, Bogaerts M et al (2006) Polymorphisms in the E-cadherin (CDH1) gene promoter and the risk of bladder cancer. Eur J Cancer 42:3219–3227
Kiemeney LA, Thorlacius S, Sulem P et al (2008) Sequence variant on 8q24 confers susceptibility to urinary bladder cancer. Nat Genet 40:1307–1312
Kiemeney LA, Sulem P, Besenbacher S et al (2010) A sequence variant at 4p16.3 confers susceptibility to urinary bladder cancer. Nat Genet 42:415–419
Koff A, Cross F, Fisher A et al (1991) Human cyclin E, a new cyclin that interacts with two members of the CDC2 gene family. Cell 66:1217–1228
Kotsopoulos J, Vitonis AF, Terry Kl et al (2009) Coffee intake, variants in genes involved in caffeine metabolism, and the risk of epithelial ovarian cancer. Cancer Causes Control 20:335–344
Lee JM, Park S, Shin DJ et al (2009) Relation of genetic polymorphisms in the cytochrome P450 gene with clopidogrel resistance after drug-eluting stent implantation in Koreans. Am J Cardiol 104:46–51
Lefkimmiatis K, Caratozzolo MF, Merlo P et al (2009) p73 and p63 sustain cellular growth by transcriptional activation of cell cycle progression genes. Cancer Res 69:8563–8571
Lehmann ML, Selinski S, Blaszkewicz M et al (2010) Rs710521[A] on chromosome 3q28 close to TP63 is associated with increased urinary bladder cancer risk. Arch Toxicol 84:967–978
Leibovici D, Grossman HB, Dinney CP et al (2005) Polymorphisms in inflammation genes and bladder cancer: from initiation to recurrence, progression, and survival. J Clin Oncol 23:5746–5756
Li C, Wu W, Liu J et al (2006) Functional polymorphisms in the promoter regions of the FAS and FAS ligand genes and risk of bladder cancer in south China: a case–control analysis. Pharmacogenet Genomics 16:245–251
Li CP, Zhu YJ, Chen R et al (2007) Functional polymorphisms of JWA gene are associated with risk of bladder cancer. J Toxicol Environ Health A 70:876–884
Li DB, Wei X, Jiang LH, Wang Y, Xu F (2010) Meta-analysis of epidemiological studies of association of P53 codon 72 polymorphism with bladder cancer. Genet Mol Res 9:1599–1605
Li Y, Zheng T, Kilfoy BA et al (2010a) Genetic polymorphisms in cytochrome P450s, GSTs, NATs, alcohol consumption and risk of non-Hodgkin lymphoma. Am J Hematol 85:213–215
Li Y, Willer CJ, Ding J, Scheet P, Abecasis GR (2010b) MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet Epidemiol 34:816–834
Lin GF, Guo WC, Chen JG et al (2005) An association of UDP-glucuronosyltransferase 2B7 C802T (His268Tyr) polymorphism with bladder cancer in benzidine-exposed workers in China. Toxicol Sci 85:502–506
Liu Z, Li G, Wei S et al (2010) Genetic variations in TERT-CLPTM1L genes and risk of squamous cell carcinoma of the head and neck. Carcinogenesis 31:1977–1981
Lo YL, Jou YS, Hsiao CF et al (2009) A polymorphism in the APE1 gene promoter is associated with lung cancer risk. Cancer Epidemiol Biomarkers Prev 18:223–229
Lu C, Dong J, Ma H et al (2009) CCND1 G870A polymorphism contributes to breast cancer susceptibility: a meta-analysis. Breast Cancer Res Treat 116:571–575
Maalej A, Petit-Teixeira E, Michou L, Rebai A, Cornelis F, Ayadi H (2005) Association study of VDR gene with rheumatoid arthritis in the French population. Genes Immun 6:707–711
Manchanda PK, Bid HK, Mittal RD (2006) Association of urokinase gene 3′-UTR T/C polymorphism with bladder cancer. Urol Int 77:81–84
Marchini J, Howie B (2008) Comparing algorithms for genotype imputation. Am J Hum Genet 83:535–539
Marchini J, Howie B, Myers S, McVean G, Donnelly P (2007) A new multipoint method for genome-wide association studies by imputation of genotypes. Nat Genet 39:906–913
Marsh HP, Haldar NA, Bunce M et al (2003) Polymorphisms in tumour necrosis factor (TNF) are associated with risk of bladder cancer and grade of tumour at presentation. Br J Cancer 89:1096–1110
Mateuca RA, Roelants M, Iarmarcovai G et al (2008) hOGG1(326), XRCC1(399) and XRCC3(241) polymorphisms influence micronucleus frequencies in human lymphocytes in vivo. Mutagenesis 23:35–41
Matullo G, Guarrera S, Sacerdote C et al (2005) Polymorphisms/haplotypes in DNA repair genes and smoking: a bladder cancer case–control study. Cancer Epidemiol Biomarkers Prev 14:2569–2578
Michiels S, Laplanche A, Boulet T et al (2009) Genetic polymorphisms in 85 DNA repair genes and bladder cancer risk. Carcinogenesis 30:763–768
Mittal RD, Manchanda PK, Bhat S, Bid HK (2007) Association of vitamin-D receptor (Fok-I) gene polymorphism with bladder cancer in an Indian population. BJU Int 99:933–937
Mølle I, Ostergaard M, Melsvik D, Nyvold CG (2008) Infectious complications after chemotherapy and stem cell transplantation in multiple myeloma: implications of Fc gamma receptor and myeloperoxidase promoter polymorphisms. Leuk Lymphoma 49:1116–1122
Moore LE, Malats N, Rothman N et al (2007) Polymorphisms in one-carbon metabolism and trans-sulfuration pathway genes and susceptibility to bladder cancer. Int J Cancer 120:2452–2458
Mouse Genome Sequencing Consortium, Waterston RH, Lindblad-Toh K, Birney E et al (2002) Initial sequencing and comparative analysis of the mouse genome. Nature 420(6915):520–562
Narter KF, Agachan B, Sozen S, Cincin ZB, Isbir T (2010) CCR2-64I is a risk factor for development of bladder cancer. Genet Mol Res 9:685–692
NCBI Single Nucleoide Polymorphism (2010) http://www.ncbi.nlm.nih.gov/projects/SNP/snp_ref.cgi?geneId=1580
Nicolae DL (2006) Testing untyped alleles (TUNA)—applications to genome-wide association studies. Genet Epidemiol 30:718–727
Nonomura N, Tokizane T, Nakayama M et al (2006) Possible correlation between polymorphism in the tumor necrosis factor-beta gene and the clinicopathological features of bladder cancer in Japanese patients. Int J Urol 13:971–976
Nyquist PA, Winkler CA, McKenzie LM, Yanek LR, Becker LC, Becker DM (2009) Single nucleotide polymorphisms in monocyte chemoattractant protein-1 and its receptor act synergistically to increase the risk of carotid atherosclerosis. Cerebrovasc Dis 28:124–130
Onat OE, Tez M, Ozçelik T, Törüner GA (2006) MDM2 T309G polymorphism is associated with bladder cancer. Anticancer Res 26:3473–3475
Oscarson M (2001) Genetic polymorphisms in the cytochrome P450 2A6 (CYP2A6) gene: implications for interindividual differences in nicotine metabolism. Drug Metab Dispos 29:91–95
Ott N, Geddert H, Sarbia M (2008) Polymorphisms in methionine synthase (A2756G) and cystathionine beta-synthase (844ins68) and susceptibility to carcinomas of the upper gastrointestinal tract. J Cancer Res Clin Oncol 134:405–410
Ouerhani S, Oliveira E, Marrakchi R et al (2007) Methylenetetrahydrofolate reductase and methionine synthase polymorphisms and risk of bladder cancer in a Tunisian population. Cancer Genet Cytogenet 176:48–53
Pabalan N, Francisco-Pabalan O, Sung L, Jarjanazi H, Ozcelik H (2010) Meta-analysis of two ERCC2 (XPD) polymorphisms, Asp312Asn and Lys751Gln, in breast cancer. Breast Cancer Res Treat 124:531–541
Park JY, Lee WK, Jung DK et al (2009) Polymorphisms in the FAS and FASL genes and survival of early stage non-small cell lung cancer. Clin Cancer Res 15:1794–1800
Park SL, Bastani D, Goldstein BY et al (2010) Associations between NBS1 polymorphisms, haplotypes and smoking-related cancers. Carcinogenesis 31:1264–1271
Parkin DM (2008) The global burden of urinary bladder cancer. Scand J Urol Nephrol Suppl 12–20
Pavanello S, Mastrangelo G, Placidi D et al (2010) CYP1A2 polymorphisms, occupational and environmental exposures and risk of bladder cancer. Eur J Epidemiol 25:491–500
Paz-y-Miño C, Muñoz MJ, López-Cortés A et al (2010) Frequency of polymorphisms pro198leu in GPX-1 gene and ile58thr in MnSOD gene in the altitude Ecuadorian population with bladder cancer. Oncol Res 18:395–400
Pei YF, Li J, Zhang L, Papasian CJ, Deng HW (2008) Analyses and comparison of accuracy of different genotype imputation methods. PLoS One 3(10):e3551
Pei YF, Zhang L, Li J, Deng HW (2010) Analyses and comparison of imputation-based association methods. PLoS One 5(5):e10827
Pharmacogenomics Knowledge Base (2010) http://www.pharmgkb.org/
Pinto LA, Depner M, Klopp N et al (2010) MMP-9 gene variants increase the risk for non-atopic asthma in children. Respir Res 11:23
Pittman AM, Broderick P, Sullivan K et al (2008) CASP8 variants D302H and −652 6N ins/del do not influence the risk of colorectal cancer in the United Kingdom population. Br J Cancer 98:1434–1436
Platek ME, Shields PG, Marian C et al (2009) Alcohol consumption and genetic variation in methylenetetrahydrofolate reductase and 5-methyltetrahydrofolate-homocysteine methyltransferase in relation to breast cancer risk. Cancer Epidemiol Biomarkers Prev 18:2453–2459
Prieto-Castelló MJ, Cardona A, Marhuenda D, Roel JM, Corno A (2010) Use of the CYP2E1 genotype and phenotype for the biological monitoring of occupational exposure to styrene. Toxicol Lett 192:34–39
Purcell S, Neale B, Todd-Brown K et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
Puthothu B, Bierbaum S, Kopp MV et al (2009) Association of TNF-alpha with severe respiratory syncytial virus infection and bronchial asthma. Pediatr Allergy Immunol 20:157–163
Rafnar T, Sulem P, Stacey SN et al (2009) Sequence variants at the TERT-CLPTM1L locus associate with many cancer types. Nat Genet 41:221–227
Reddy BM, Reddy ANS, Nagaraja T et al (2006) Single nucleotide polymorphisms of the alcohol dehydrogenase genes among the 28 caste and tribal populations of India. Int J Hum Genet 6:309–316
Reddy P, Naidoo RN, Robins TG et al (2010) GSTM1, GSTP1, and NQO1 polymorphisms and susceptibility to atopy and airway hyperresponsiveness among South African schoolchildren. Lung 188:409–414
Reif DM (2006) Integrated analysis of genetic and proteomic data. Dissertation, Nashville, TN
Ricceri F, Guarrera S, Sacerdote C et al (2010) ERCC1 haplotypes modify bladder cancer risk: a case–control study. DNA Repair 9:191–200
Rothman N, García-Closas M, Chatterjee N et al (2010) A multi-stage genome-wide association study of bladder cancer identifies multiple susceptibility loci. Nat Genet 42:978–984
Rouguieg K, Picard N, Sauvage FL, Gaulier JM, Marquet P (2010) Contribution of the different UDP-glucuronosyltransferase (UGT) isoforms to buprenorphine and norbuprenorphine metabolism and relationship with the main UGT polymorphisms in a bank of human liver microsomes. Drug Metab Dispos 38:40–45
Safarinejad MR, Shafiei N, Safarinejad S (2010) Genetic susceptibility of methylenetetrahydrofolate reductase (MTHFR) gene C677T, A1298C, and G1793A polymorphisms with risk for bladder transitional cell carcinoma in men. Med Oncol (29 Oct 2010, Epub ahead of print)
Safarinejad MR, Shafiei N, Safarinejad S (2011) The association between bladder cancer and a single nucleotide polymorphism (rs2854744) in the insulin-like growth factor (IGF) binding protein-3 (IGFBP-3) gene. Arch Toxicol (in press)
Saravana Devi S, Vinayagamoorthy N, Agrawal M et al (2008) Distribution of detoxifying genes polymorphism in Maharastrian population of central India. Chemosphere 700:1835–1839
Sasaki T, Horikawa M, Orikasa K et al (2008) Possible relationship between the risk of Japanese bladder cancer cases and the CYP4B1 genotype. Jpn J Clin Oncol 38:634–640
Savage SA, Hou L, Lissowska J et al (2006) Interleukin-8 polymorphisms are not associated with gastric cancer risk in a Polish population. Cancer Epidemiol Biomarkers Prev 15:589–591
Sayan BS, Sayan AE, Yang AL et al (2007) Cleavage of the transactivation-inhibitory domain of p63 by caspases enhances apoptosis. Proc Natl Acad Sci USA 104:10871–10876
Scheet P, Stephens M (2006) A fast and flexible statistical model for large-scale population genotype data: applications to inferring missing genotypes and haplotypic phase. Am J Hum Genet 78:629–644
Schnakenberg E, Breuer R, Werdin R, Dreikorn K, Schloot W (2000) Susceptibility genes: GSTM1 and GSTM3 as genetic risk factors in bladder cancer. Cytogenet Cell Genet 91:234–238
Servin B, Stephens M (2007) Imputation-based analysis of association studies: candidate regions and quantitative traits. PLoS Genet 3:e114
Shao J, Gu M, Xu Z, Hu Q, Qian L (2007) Polymorphisms of the DNA gene XPD and risk of bladder cancer in a Southeastern Chinese population. Cancer Genet Cytogenet 177:30–36
Shi WX, Chen SQ (2004) Frequencies of poor metabolizers of cytochrome P450 2C19 in esophagus cancer, stomach cancer, lung cancer and bladder cancer in Chinese population. World J Gastroenterol 10:1961–1963
Shih CM, Kuo WH, Lin CW et al (2011) Association of polymorphisms in the genes of the urokinase plasminogen activation system with susceptibility to and severity of non-small cell lung cancer. Clin Chim Acta 412:194–198
Skjelbred CF, Saebø M, Wallin H et al (2006) Polymorphisms of the XRCC1, XRCC3 and XPD genes and risk of colorectal adenoma and carcinoma, in a Norwegian cohort: a case control study. BMC Cancer 6:67
Slattery ML, Wolff RK, Herrick JS, Caan BJ, Potter JD (2007) IL6 genotypes and colon and rectal cancer. Cancer Causes Control 18:1095–1105
Song DK, Xing DL, Zhang LR, Li ZX, Liu J, Qiao BP (2009) Association of NAT2, GSTM1, GSTT1, CYP2A6, and CYP2A13 gene polymorphisms with susceptibility and clinicopathologic characteristics of bladder cancer in Central China. Cancer Detect Prev 32:416–423
Srivastava DS, Mandhani A, Mittal RD (2008) Genetic polymorphisms of cytochrome P450 CYP1A1 (*2A) and microsomal epoxide hydrolase gene, interactions with tobacco-users, and susceptibility to bladder cancer: a study from North India. Arch Toxicol 82:633–639
Srivastava P, Gangwar R, Kapoor R, Mittal RD (2010a) Bladder cancer risk associated with genotypic polymorphism of the matrix metalloproteinase-1 and 7 in North Indian population. Dis Markers 29:37–46
Srivastava P, Mandhani A, Kapoor R, Mittal RD (2010b) Role of MMP-3 and MMP-9 and their haplotypes in risk of bladder cancer in North Indian cohort. Ann Surg Oncol 17:3068–3075
Stern MC, Lin J, Figueroa JD et al (2009) Polymorphisms in DNA repair genes, smoking, and bladder cancer risk: findings from the international consortium of bladder cancer. Cancer Res 69:6857–6864
Strassburg CP, Lankisch TO, Manns MP, Ehmer U (2008) Family 1 uridine-5′-diphosphate glucuronosyltransferases (UGT1A): from Gilbert’s syndrome to genetic organization and variability. Arch Toxicol 82:415–433
Su L, Zhou W, Asomaning K et al (2006) Genotypes and haplotypes of MMP-1, -3 and -12 genes and the risk of lung cancer. Proc Am Assoc Cancer Res 47:437
Sun P, Qiu Y, Zhang Z et al (2009) Association of genetic polymorphisms, mRNA expression of p53 and p21 with chronic benzene poisoning in a Chinese occupational population. Cancer Epidemiol Biomarkers Prev 18:1821–1828
Sun H, Qiao Y, Zhang X et al (2010) XRCC3 Thr241Met polymorphism with lung cancer and bladder cancer: a meta-analysis. Cancer Sci 101:1777–1782
Tang T, Cui S, Deng X et al (2010) Insertion/deletion polymorphism in the promoter region of NFKB1 gene increases susceptibility for superficial bladder cancer in Chinese. DNA Cell Biol 29:9–12
Tranah GJ, Giovannucci E, Ma J, Fuchs C, Hankinson SE, Hunter DJ (2004) Epoxide hydrolase polymorphisms, cigarette smoking and risk of colorectal adenoma in the Nurses’ Health Study and the Health Professionals Follow-up Study. Carcinogenesis 25:1211–1218
Tsironi EE, Pefkianaki M, Tsezou A et al (2009) Evaluation of MMP1 and MMP3 gene polymorphisms in exfoliation syndrome and exfoliation glaucoma. Mol Vis 15:2890–2895
van Dijk B, van Houwelingen KP, Witjes JA, Schalken JA, Kiemeney LA (2001) Alcohol dehydrogenase type 3 (ADH3) and the risk of bladder cancer. Eur Urol 40:509–514
Varghese JS, Easton DF (2010) Genome-wide association studies in common cancers—what have we learnt? Curr Opin Genet Dev 20:201–209
Vartanian J-P, Guetard D, Henry M, Wain-Hobson S (2008) Evidence for editing of human papillomavirus DNA by APOBEC3 in benign and precancerous lesions. Science 320:230–233
Verhaegh GW, Verkleij L, Vermeulen SH, den Heijer M, Witjes JA, Kiemeney LA (2008) Polymorphisms in the H19 gene and the risk of bladder cancer. Eur Urol 54:1118–1126
Verlaan M, Te Morsche RH, Roelofs HM et al (2004) Genetic polymorphisms in alcohol-metabolizing enzymes and chronic pancreatitis. Alcohol Alcohol 39:20–24
Visel A, Rubin EM, Pennacchio LA (2009) Genomic views of distant-acting enhancers. Nature 461(7261):199–205
Wang M, Zhang Z, Zhu H et al (2008) A novel functional polymorphism C1797G in the MDM2 promoter is associated with risk of bladder cancer in a Chinese population. Clin Cancer Res 14:3633–3640
Wang HC, Liu CS, Chiu CF et al (2009a) Significant association of DNA repair gene Ku80 genotypes with breast cancer susceptibility in Taiwan. Anticancer Res 29:5251–5254
Wang M, Zhang Z, Tian Y, Shao J, Zhang Z (2009b) A six-nucleotide insertion-deletion polymorphism in the CASP8 promoter associated with risk and progression of bladder cancer. Clin Cancer Res 15:2567–2572
Wang M, Wang M, Cheng G, Zhang Z, Fu G, Zhang Z (2009c) Genetic variants in the death receptor 4 gene contribute to susceptibility to bladder cancer. Mutat Res 661:85–92
Wang M, Qin C, Zhu J et al (2010a) Genetic variants of XRCC1, APE1, and ADPRT genes and risk of bladder cancer. DNA Cell Biol 29:303–311
Wang M, Wang M, Yuan L et al (2010b) A novel XPF -357A>C polymorphism predicts risk and recurrence of bladder cancer. Oncogene 29:1920–1928
Wang S, Tang J, Wang M, Yuan L, Zhang Z (2010c) Genetic variation in PSCA and bladder cancer susceptibility in a Chinese population. Carcinogenesis 31:621–624
Watabe T, Lin M, Ide H et al (2002) Growth, regeneration, and tumorigenesis of the prostate activates the PSCA promoter. Proc Natl Acad Sci USA 99:401–406
Wen H, Ding Q, Fang ZJ, Xia GW, Fang J (2009) Population study of genetic polymorphisms and superficial bladder cancer risk in Han-Chinese smokers in Shanghai. Int Urol Nephrol 41:855–864
Wolf S, Mertens D, Pscherer A et al (2006) Ala228 variant of trail receptor 1 affecting the ligand binding site is associated with chronic lymphocytic leukemia, mantle cell lymphoma, prostate cancer, head and neck squamous cell carcinoma and bladder cancer. Int J Cancer 118:1831–1835
Wu X, Gu J, Grossman HB et al (2006) Bladder cancer predisposition: a multigenic approach to DNA-repair and cell-cycle-control genes. Am J Hum Genet 78:464–479
Wu X, Ye Y, Kiemeney LA, Sulem P et al (2009) Genetic variation in the prostate stem cell antigen gene PSCA confers susceptibility to urinary bladder cancer. Nat Genet 41:991–995
Yang H, Gu J, Lin X et al (2008) Profiling of genetic variations in inflammation pathway genes in relation to bladder cancer predisposition. Clin Cancer Res 14:2236–2244
Yang JJ, Ko KP, Cho LY et al (2009) The role of TNF genetic variants and the interaction with cigarette smoking for gastric cancer risk: a nested case–control study. BMC Cancer 9:238
Yang W, Qi Q, Zhang H et al (2010) p21 Waf1/Cip1 polymorphisms and risk of esophageal cancer. Ann Surg Oncol 17:1453–1458
Yu Z, Schaid DJ (2007) Methods to impute missing genotypes for population data. Hum Genet 122:495–504
Yuan L, Gu X, Shao J et al (2010) Cyclin D1 G870A polymorphism is associated with risk and clinicopathologic characteristics of bladder cancer. DNA Cell Biol 29:611–617
Zhang Y, Jin M, Liu B et al (2008a) Association between H-RAS T81C genetic polymorphism and gastrointestinal cancer risk: a population based case–control study in China. BMC Cancer 8:256
Zhang Z, Wang S, Wang M, Tong N, Fu G, Zhang Z (2008b) Genetic variants in RUNX3 and risk of bladder cancer: a haplotype-based analysis. Carcinogenesis 29:1973–1978
Zheng L, Wang Y, Schabath MB, Grossman HB, Wu X (2003) Sulfotransferase 1A1 (SULT1A1) polymorphism and bladder cancer risk: a case–control study. Cancer Lett 202:61–69
Zhou P, Li JP, Zhang C (2011) Polymorphisms of tumor necrosis factor-alpha and breast cancer risk: appraisal of a recent meta-analysis. Breast Cancer Res Treat 126:253–256
Acknowledgments
We thank Ms. Susanne Lindemann for helpful bibliographic work.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Golka, K., Selinski, S., Lehmann, ML. et al. Genetic variants in urinary bladder cancer: collective power of the “wimp SNPs”. Arch Toxicol 85, 539–554 (2011). https://doi.org/10.1007/s00204-011-0676-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-011-0676-3