Abstract
Purpose
Acute respiratory distress syndrome (ARDS) is accompanied by a dysfunctional immune-inflammatory response following lung injury, including during coronavirus disease 2019 (COVID-19). Limited causal biomarkers exist for ARDS development. We sought to identify novel genetic susceptibility targets for ARDS to focus further investigation on their biological mechanism and therapeutic potential.
Methods
Meta-analyses of ARDS genome-wide association studies were performed with 1250 cases and 1583 controls in Europeans, and 387 cases and 387 controls in African Americans. The functionality of novel loci was determined in silico using multiple omics approaches. The causality of 114 factors potentially involved in ARDS development was assessed using Mendelian Randomization analysis.
Results
There was distinct genetic heterogeneity in ARDS between Europeans and African Americans. rs7967111 at 12p13.2 was functionally associated with ARDS susceptibility in Europeans (odds ratio = 1.38; P = 2.15 × 10–8). Expression of two genes annotated at this locus, BORCS5 and DUSP16, was dynamic but ultimately decreased during ARDS development, as well as downregulated in immune cells alongside COVID-19 severity. Causal inference implied that comorbidity of inflammatory bowel disease and elevated levels of C-reactive protein and interleukin-10 causally increased ARDS risk, while vitamin D supplementation and vasodilator use ameliorated risk.
Conclusion
Our findings suggest a novel susceptibility locus in ARDS pathophysiology that implicates BORCS5 and DUSP16 as potentially acting in immune-inflammatory processes. This locus warrants further investigation to inform the development of therapeutic targets and clinical care strategies for ARDS, including those induced by COVID-19.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Integration analyses of genome and transcriptome data reveal a novel functional locus possibly involved in the regulation of immune-inflammatory response in ARDS pathophysiology, and causal inference indicates several clinical interventions of ARDS development. Our findings inform further investigation of host–pathogen biology and therapeutic targets for ARDS, especially during the COVID-19 pandemic. |
Introduction
Acute respiratory distress syndrome (ARDS) is a type of hypoxemic respiratory failure in patients with conditions that predispose to or cause lung injury [1]. ARDS produces a high mortality rate (30–40%) in hospitals worldwide [2]. Incidence of this condition is increasing sharply with the ongoing coronavirus disease 2019 (COVID-19) pandemic [3], with a median onset time of 8–12 days after COVID-19 symptom onset [4, 5].
Multi-omics approaches identify biomarkers for disease etiology and therapeutics [6]. Previously, we summarized 201 ARDS genes from genome and transcriptome studies that are involved in key inflammatory pathways [7], supporting a fundamental understanding of ARDS development. However, single nucleotide polymorphisms (SNPs) related to ARDS derived from genome-wide association studies (GWAS) were confined to European populations, except for one study we launched in African Americans [8]. This narrow ancestry sampling and conventional analysis limit the discovery of biological clues about ARDS pathophysiology.
ARDS has heterogeneous etiology but is characterized by acute widespread inflammation in the lung commonly precipitated by sepsis or pneumonia [1, 9]. Similarly, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection induces aggressive inflammatory and immune responses that damage the respiratory tract, often causing ARDS as COVID-19 progresses [10, 11]. Recently, two COVID-19 GWAS preliminarily revealed host genetic components involved in the severe progression [12, 13], opening new avenues to elucidate COVID-19 pathophysiology, including ARDS development. Effective treatments for ARDS specifically caused by COVID-19, outside of supportive care interventions, remain elusive [14, 15]. Therefore, meeting the urgent need for effective ARDS therapies necessitates the identification of new therapeutic targets.
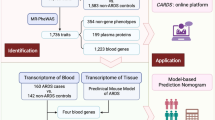
To investigate the genetic architecture and molecular factors related to ARDS development, we applied integrative omics approaches and performed Mendelian Randomization analysis to infer candidate causal factors (Fig. 1). Our findings aid in the understanding of the progressive course of ARDS and identify potential targets for therapeutic development.
Flowchart of study design. First, in the identification phase, ancestry-specific and trans-ancestry GWAS meta-analyses of ARDS were performed to compare two distinct ancestry populations to identify novel susceptibility genes for all-cause ARDS. Second, in the annotation phase, in silico analyses combined with transcriptome data were used to functionally annotate novel loci to decipher corresponding biological and genetic effects on ARDS pathophysiology. Last, in the application phase, a certain omics features and clinical observations were shared between all-cause ARDS and COVID-19 severity, suggesting clinical care of ARDS development, possibly as COVID-19 progresses. Flowchart was created online with BioRender.com
Methods
Study participants
Participants were recruited from the iSPAAR (Identification of SNPs Predisposing to Altered Acute Lung Injury Risk) consortium [16], MESSI (Molecular Epidemiology of Sepsis in the ICU) cohort [17, 18], and a study by Garcia et al. [8]. Details about population demographics and data cleaning are described in Supplementary Methods, Table E1, and Figure E1–E3.
Genetic association analysis and meta-analysis
GWAS was conducted separately for each ARDS cohort across European and African American ancestries. Logistic regression was performed in an additive genetic model with adjustments for age, sex, and top three population ancestry principal components. Odds ratios (ORs) and corresponding 95% confidence intervals (CIs) were calculated to estimate genetic effects. Association statistics from each ancestry group were combined separately using fixed meta-analysis with inverse-variance weighting of log-ORs via METAL [19], following the descriptor:./metal < script > , in which the commands of MARKER, ALLELE, WEIGHT, EFFECT, STDERR, FREQ, PROCESS, and PVAL were used to describe the input files across each study, and the commands of OUTFILE and ANALYSIS performed the final analysis.
RNA sequencing (RNA-Seq) analysis
Blood samples were collected from 160 ARDS cases and 142 controls (in-house dataset) for RNA-Seq analysis, of which 46 were matched to genotyping data for expression quantitative trait loci (QTL) analyses. Procedures of detection and quantitation are described in Supplementary Methods. Briefly, 19,898 protein-coding genes were identified for transcriptome analyses, including analyses of differential expression, pathway enrichment, and immune cell decomposition. Details for reanalysis of publicly supported COVID-19 single-cell RNA-Seq datasets are described in Supplementary Methods.
Mendelian randomization (MR) analysis
Two-sample MR analysis was conducted to obtain causal estimates of exposures on outcome using the TwoSampleMR R package [20]. Exposures empirically included 114 traits as key clinical issues of COVID-19, grouped by disease, hematology, biochemistry, coagulation, nutrition or habit, and treatment (Table E2). Genetic instruments of each exposure were obtained from GWAS summary statistics in Europeans (Supplementary Methods). The outcome was ARDS using GWAS summary statistics in Europeans, generated by meta-analysis. Inverse-variance-weighted (IVW), weighted median, and MR-Egger regression were primarily used to calculate effect size (β) and corresponding standard error (SE). The Wald ratio method was used if only one genetic instrument remained. Heterogeneity was estimated using MR-Egger and IVW methods. Directional pleiotropy was estimated via MR-Egger intercept test. Evidence score was calculated by aggregating the significance.
Statistical analysis
Analytical steps throughout the study are shown in Figure E2. In silico analyses are detailed in Supplementary Methods, including genetic heritability and correlation (via Genome-wide Complex Trait Analysis [GCTA], LD Score Regression [LDSC], and LD Hub), gene set analysis (via Multi-marker Analysis of GenoMic Annotation [MAGMA] and Data-driven Expression-Prioritized Integration for Complex Traits [DEPICT]), genetic function annotation (via Functional Annotation of Variants—Online Resource [FAVOR]), phenome-wide association studies annotation (PheWAS; via SAIGE UKB and PhenomeXcan), expression pattern analyses (via Human Protein Atlas [HPA], Cancer Cell Line Encyclopedia [CCLE], Database of Immune Cell Expression, Expression quantitative trait loci and Epigenomics [DICE], UCSC Cell Browser, and Gene Expression Omnibus [GEO]), immune cell estimation (via CIBERSORTx), and polygenic risk score (PRS). Linear regression analysis, Kruskal–Wallis test, Wilcoxon signed-rank test, and Spearman rank correlation analysis were conducted as appropriate. All statistical analyses were done in R (version 3.5.1).
Results
Genetic architecture of ARDS in European and African American populations
We first conducted ancestry-specific GWAS meta-analyses for ARDS. In iSPAAR and MESSI European cohorts comprising 1250 cases and 1583 controls with 5,749,543 common variants (Fig. E2), rs7967111 A > G had a marginal effect of genome-wide significance on increasing risk of ARDS (OR = 1.35, 95% CI = 1.21–1.51, P = 6.64 × 10–8; Table E3 and Fig. E4A). For African American ancestry, we included 387 cases and 387 controls from MESSI and Garcia et al. cohorts with 6,255,902 variants (Fig. E2) and observed a top signal of rs619652 A > G reaching a nominal significance (OR = 1.86, 95% CI = 1.48–2.35, P = 1.59 × 10–7; Table E3 and Fig. E4A). Globally, the estimated genetic heritability of ARDS was higher in Europeans (GCTA: h2SNP = 0.129, SE = 0.023; LDSC: h2SNP = 0.126, SE = 0.049) than African Americans (GCTA: h2SNP = 0.039, SE = 0.032).
At the single-variant level, no significant loci were shared across ancestries (empirical threshold at P < 1 × 10–4; Table E4 and Fig. E4B), even at the gene-set level analyzed via MAGMA (Fig. E4C). Nevertheless, gene set enrichment analysis via DEPICT identified 45 ancestry-shared reconstituted enrichment sets (Fig. E4D), such as immune response activation (top in Europeans). The genetic correlation of ARDS between the two populations was 0.266, but the large SE (0.589) caused by the small sample size limits interpretation. These findings suggest a distinct genetic heterogeneity of ARDS between ancestries, but some shared genetic components.
Thus, we performed a trans-ancestry GWAS meta-analysis. Unexpectedly, no loci achieved genome-wide significance, even at the empirical threshold of P < 1 × 10−6 (Fig. E5). Notably, rs7967111 from Europeans maintained a statistically significant association with ARDS (OR = 1.26, 95% CI = 1.14–1.39, P = 4.38 × 10–6), while rs619652 from African Americans was not validated in trans-ancestry populations and had dramatic heterogeneity—even its minor allele was exchanged from G in Europeans to A in African Americans (Table E3).
To confirm the genetic association of rs7967111, we performed sensitivity analysis by requiring individual genotyping data from two European cohorts. Intriguingly, rs7967111 exceeded genome-wide significance (P < 5 × 10–8), with OR of 1.38 (P = 2.15 × 10–8; Table 1) after adjusting for potential confounders. Thus, we retained rs7967111 for further analysis.
Functional analysis of rs7967111 at 12p13.2
Functional annotation showed that rs7967111 was located at the third intron of BORCS5 and approximately 20 kb downstream of DUSP16 within a high-LD region (Fig. E6A,B), harboring strong functional signals of enhancer activity, histone modification, and transcription factor binding calculated by FAVOR (Fig. E6C). Subsequently, genetic pleiotropy analysis indicated that rs7967111 may dysregulate BORCS5 and DUSP16 expression in blood (in-house dataset, Fig. 2a) and lung tissues (public datasets, Table E5) and alter neighboring methylation level at CpG site cg19207364, surrounding H3K27ac modification activity, and 15 blood metabolites abundance (Table E5). However, rs7967111 did not affect 11 ARDS protein biomarkers (Fig. E7). PheWAS for pleiotropy evaluation via SAIGE UKB and PhenomeXcan revealed that rs7967111 was dramatically associated with multiple phenotypes, especially in the respiratory system (Fig. E8), and both BORCS5 and DUSP16 were significantly correlated with more than 100 phenotypes (Fig. E9). These results suggest that rs7967111 at 12p13.2 serves as a novel functional locus in ARDS development.
Gene expression patterns across genotypes and groups. A eQTL analysis for genetic effects of rs7967111 on BORCS5 (left) and DUSP16 (right) expression. We used 46 pairs of genotyping and RNA-Seq data from in-house ARDS blood and calculated P-values using linear regression analysis. B Dynamic gene expression patterns in preclinical lipopolysaccharide (LPS)-induced lung injury models. Gene expression was extracted from Gene Expression Omnibus datasets of GSE9314 in mice and GSE5883 in humans. P-values were calculated by ANOVA test. C Correlation matrix plot for pairwise similarity (Spearman correlation) among candidate genes and abundances of 22 immune cell types, estimated from the blood transcriptome (B) using CIBERSORTx
Expression pattern and biological effect of susceptibility genes
BORCS5 and DUSP16 were expressed across tissues at both protein and RNA levels (HPA database, Fig. E10). In the in-house dataset of 160 ARDS cases and 142 controls, BORCS5 and DUSP16 expression were highly correlated (r = 0.76, P < 0.001; Fig. E11A) regardless of case status (Fig. E11B). Strong correlations were also identified in cell lines, particularly in lung cells (rall = 0.44, Pall < 0.001; rlung = 0.51, Plung < 0.001; CCLE dataset, Fig. E11C) and immune cells (r > 0.95, P < 0.001; DICE dataset, Fig. E11D).
Similar expression patterns were found at single-cell levels using the UCSC Cell Browser. BORCS5 (6.4% of cells) and DUSP16 (14.5%) were widely spread over nine cell types originating from lung tissues (Fig. E12A), especially enriched across primary bronchial epithelial cells (3.2% and 1.1%, respectively; Fig. E12B). Notably, BORCS5 and DUSP16 were highly expressed in pulmonary immune cells, including macrophages and dendritic cells (Fig. E12C).
Preclinical models of lung injury induced by lipopolysaccharide (LPS) retrieved from GEO were applied to assess the potential biological effects of candidate genes in ARDS pathobiology. Intriguingly, dynamic expression patterns of BORCS5 and DUSP16 were observed in both mouse lung tissues (GSE9314) and human lung microvascular endothelial cells (GSE5883) exposed to LPS over time (Fig. 2b). Both BORCS5 and DUSP16 showed greater expression in the first 4–8 h after LPS exposure, but then their expression decreased dramatically.
Immune characteristics of ARDS by transcriptome profiling
We next profiled the blood transcriptome for 160 ARDS cases and 142 controls. There yielded 142 differentially expressed genes (112 upregulated and 30 downregulated; Fig. E13A), and these represented enrichment for key immune pathways including Toll-like receptor signaling, CD8+ T cells, and immunodeficiency disease via gene set enrichment analysis (Fig. E13B–D). Further, among 22 immune cell types decomposed in the transcriptome via CIBERSORTx, 5 immune cell types were significantly increased in ARDS cases (Fig. E14). Remarkably, BORCS5 and DUSP16 were dramatically positively correlated with gamma delta T cells, M2 macrophages, and neutrophils, and negatively with CD8+ T cells, regulatory T cells, monocytes, and resting mast cells (Fig. 2c). However, rs7967111 did not significantly influence these cell fractions (Fig. E15).
Transferability of all-cause ARDS findings
Given ARDS is a complication during the severe progression of COVID-19, we attempted to assess the transferability of all-cause ARDS findings (defined in this study) to understand COVID-19 severity. Genetically, three severe COVID-19 relevant SNPs (i.e., rs657152, rs10735079, and rs2109069) [12, 13] displayed consistent associations with all-cause ARDS development in this study, of which 2 were nominally significant (P = 0.040, 0.013, and 0.045, respectively; Fig. E16). Intriguingly, the PRS calculated by COVID-19 severity GWAS was significantly higher in ARDS cases than controls (Fig. 3a). In single-cell transcriptome profiles, BORCS5 and DUSP16 expression were significantly decreased or on a downward trend aligning with severe progression of COVID-19 in dendritic cells derived from upper respiratory tract samples (Fig. 3b) or peripheral blood mononuclear cells (Fig. 3c) [21, 22].
Omics transferability assessment of all-cause ARDS. A Differential polygenic risk score (PRS) between ARDS cases and controls. PRS was calculated via weighting effect size derived from three severe COVID-19 GWAS assigned to each ARDS case and control in Europeans. P-value was calculated via t-test. B,C Differential expression analysis for BORCS5 and DUSP16 in dendritic cells derived from two single-cell RNA-Seq datasets of upper respiratory tract samples [21] (B) and peripheral blood mononuclear cells [22] (C) in COVID-19 patients and healthy controls, respectively. Dots represent cells expressing candidate genes and are colored by the severity of COVID-19. Healthy samples from (B) dataset were removed because only one cell expressed candidate genes. P-values were calculated from a Wilcoxon test. The y-axis is on a log-10 scale to show gene expression, and “n” indicates the number of cells detected with candidate gene expression. The solid line of each plot indicates the median of gene expression for each group, and box edges mark lower and upper quartiles of gene expression
Putative causal relationship between traits and ARDS
Moreover, we estimated causal effects of 114 candidate risk factors derived from clinical observations of COVID-19 under the MR analytic framework (Fig. 4a). Six associations with evidence scores ≥ 2 are shown in Fig. 4b. Notably, inflammatory bowel disease (IBD) and immune/inflammatory biomarkers [i.e., C-reactive protein (CRP), interleukin-10 (IL-10), and immunoglobulin G index levels in cerebrospinal fluid] were causally associated with increased risk of ARDS development (all β > 0; PIVW = 0.027, 0.015, 0.002, and < 0.001, respectively), while daily supplements of vitamin D (βMR-Egger = −25.80, PMR-Egger = 0.001) and use of vasodilators to treat cardiac diseases (βIVW = −0.25, PIVW = 0.031) were associated with decreased likelihood of severe progression (Table E2). Intriguingly, 47 of 855 traits were genetically correlated with ARDS in Europeans using LD Hub (Table E6), including IBD.
Results of Mendelian Randomization analysis for all-cause ARDS. a Schematic diagram for assumptions of Mendelian Randomization analysis. Assumption 1: genetic instruments selected should be robustly associated with the exposure, usually underlying P < 5 × 10–8 (P < 1 × 10–6 or P < 1 × 10–4 as suggestive significance level); Assumption 2: genetic variants should not be associated with potential confounders; and Assumption 3: genetic variants of an exposure should affect the outcome risk only through the risk factor, not via alternative pathways. b Scatter plots for putative causal associations between six diseases/traits and all-cause ARDS under evidence score of ≥ 2. P-values were calculated with IVW or MR-Egger regression
Discussion
To our knowledge, this is the first study to integrate trans-ancestry GWAS in Europeans and African Americans, transcriptome analyses across bulk tissues and single cells, and preclinical models by mice and cells, to reveal a novel locus at 12p13.2 harboring rs7967111 as genetic predictor of ARDS susceptibility, BORCS5 and DUSP6 involved in heterogeneous etiologies of ARDS, and potential clinical interventions of ARDS development.
Initially, we launched a large-scale ARDS GWAS analysis comprising European and African American populations. Although a distinct genetic architecture of ARDS exists across ancestries, gene set-based inference eliminated ancestry specificity, which showed 45 shared functional elements in ARDS etiology, including immune response. Trans-ancestry GWAS meta-analysis improves the resolution of genetic effects on phenotypes [23]. However, we did not observe signals passing genome-wide significance when combining the two ancestries, possibly because of the limited sample size and varied ethnic background of African Americans. Thus, a large-scale non-European population for ARDS genomic studies was advocated to further maximize genetic discovery and reduce health disparities.
Nevertheless, rs7967111 at 12p13.2 presented a significant genome-wide effect on increasing ARDS risk after sensitivity analyses. The subsequent pleiotropic evaluation indicated that rs7967111 was a potential causal variant of ARDS by disturbing a cascade of DNA methylation, histone modification, gene expression, and metabolites. PheWAS is well suited to facilitate pleiotropic evaluation of novel risk SNPs on phenotypes [24]. Our analyses further indicated that rs7967111 and the annotated genes BORCS5 and DUSP16 displayed biological roles in multiple traits or diseases, especially those referring to the lung.
The following transcriptome analyses spanning bulk tissues and single cells dissected their functions in ARDS that BORCS5 and DUSP16 were not only highly correlated mutually in pulmonary immune cells but also influenced cell fractions of immune CD8+ T cells, regulatory T cells, and macrophages in blood, separately from genetic effects of rs7967111. Notably, preclinical models analyses revealed that BORCS5 and DUSP16 expression exhibit dynamic change but ultimately decrease during ARDS development. Physiologically, BORCS5 recruits ARL8B to promote lysosome movement and reduce cell spreading and migration [25, 26]; and DUSP16 inactivates MAP kinases to regulate cellular senescence important for immune responses [27, 28]. These comprehensive analyses indicate that rs7967111 was a functional genetic predictor for ARDS susceptibility via disrupting both BORCS5 and DUSP16 expression, among which the latter served as regulatory factors affecting immune response in ARDS pathophysiology under certain causes.
Emerging studies propose that it is reasonable to borrow concepts from other-cause ARDS pathogenesis to inform evolving understanding of COVID-19 ARDS in severe progression while awaiting more disease-focused data [29]. Clinically, patients who progress to severe COVID-19 meet ARDS diagnosis criteria [30], and represent a specific subset of all-cause ARDS with clinical similarity in respiratory mechanics, therapeutic response, and prognosis [31, 32]. Therefore, we hypothesized whether the available data on all-cause ARDS could glean insights into ARDS development as COVID-19 progresses. In this study, we observed that several genetic components in COVID-19 severity [12, 13] were shared in all-cause ARDS and the corresponding PRS could distinguish ARDS cases from controls. Biologically, an intriguing observation from single-cell transcriptome data showed gradually reduced expression of both BORCS5 and DUSP16 in dendritic cells correlated with COVID-19 severity, which was in accordance with the expression pattern in preclinical models of ARDS development. Together, these features may suggest mechanisms that BORCS5 and DUSP16 may be involved in COVID-19 severity as similar to the immune-inflammation response in ARDS pathophysiology.
Furthermore, we observed some clinically instructive results by MR analysis. Elevated levels of CRP and IL-10 were commonly happened in severe COVID-19 patients [33], and IBD patients were at higher risk of developing severity [34]; in contrast, vasodilator use and vitamin D supplements might reduce the risk of COVID-19 infections and mortality [35, 36]. These clinical observations were consistent with their causal associations with ARDS development, which may guide the ARDS care management, especially during COVID-19 severe progression.
We acknowledge several limitations of this study. First, the transcriptome for all-cause ARDS was derived from blood, possibly explaining non-significant differential expression of BORCS5 and DUSP16. Further studies applying lung-specific transcriptomes of ARDS (including COVID-19 ARDS) will inform future treatment strategies. Second, it is necessary to perform large-scale ARDS GWAS in diverse populations to confirm our findings, especially for the transferability of all-cause ARDS to COVID-19 ARDS in severe progression. The generalizability of these results warrants further omics validation by recruiting larger populations of patients with COVID-19 with and without ARDS. Third, MR associations are subject to pleiotropic effects of genetic instruments, such as treatments in IBD and all-cause ARDS, which might be associated with some unexpected hidden confounders in GWAS summary statistics calculations. Therefore, well-powered randomized trials are needed to conclusively evaluate this association.
In summary, we uncovered a novel locus at 12p13.2 encompassing rs7967111 and functional involvement of BORCS5 and DUSP16 in ARDS development via immune-inflammatory processes. A certain extent of shared molecular characteristics and clinical observations indicate the leverage of findings in predicting ARDS susceptibility, elucidating ARDS pathophysiology, and informing treatment and intervention strategies for ARDS development, including those induced by COVID-19.
Availability of data and materials
For the iSPAAR consortium dataset, the genotype data and relevant covariate information (age, sex, ancestry, principal components, etc.) are deposited in dbGaP under accession codes phs000631.v1.p1. For MESSI and the African American dataset, dbGaP submission is forthcoming in accordance with the NIH genomic data sharing policy. In advance of their availability on dbGaP, full summary statistics are available on request to the authors.
Code availability
Not applicable.
References
Fan E, Brodie D, Slutsky AS (2018) Acute respiratory distress syndrome: advances in diagnosis and treatment. JAMA 319(7):698–710. https://doi.org/10.1001/jama.2017.21907
Bellani G, Laffey JG, Pham T, Fan E, Brochard L, Esteban A, Gattinoni L, van Haren F, Larsson A, McAuley DF, Ranieri M, Rubenfeld G, Thompson BT, Wrigge H, Slutsky AS, Pesenti A (2016) Epidemiology, patterns of care, and mortality for patients with acute respiratory distress syndrome in intensive care units in 50 countries. JAMA 315(8):788–800. https://doi.org/10.1001/jama.2016.0291
Network C-IGobotR, the C-ICUI (2020) Clinical characteristics and day-90 outcomes of 4244 critically ill adults with COVID-19: a prospective cohort study. Intensive Care Med. https://doi.org/10.1007/s00134-020-06294-x
Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, Wang B, Xiang H, Cheng Z, Xiong Y, Zhao Y, Li Y, Wang X, Peng Z (2020) Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. Jama. https://doi.org/10.1001/jama.2020.1585
Li X, Ma X (2020) Acute respiratory failure in COVID-19: is it “typical” ARDS? Crit Care 24(1):198. https://doi.org/10.1186/s13054-020-02911-9
Karczewski KJ, Snyder MP (2018) Integrative omics for health and disease. Nat Rev Genet 19(5):299–310. https://doi.org/10.1038/nrg.2018.4
Lynn H, Sun X, Casanova N, Gonzales-Garay M, Bime C, Garcia JGN (2019) Genomic and genetic approaches to deciphering acute respiratory distress syndrome risk and mortality. Antioxid Redox Signal 31(14):1027–1052. https://doi.org/10.1089/ars.2018.7701
Bime C, Pouladi N, Sammani S, Batai K, Casanova N, Zhou T, Kempf CL, Sun X, Camp SM, Wang T, Kittles RA, Lussier YA, Jones TK, Reilly JP, Meyer NJ, Christie JD, Karnes JH, Gonzalez-Garay M, Christiani DC, Yates CR, Wurfel MM, Meduri GU, Garcia JGN (2018) Genome-wide association study in African Americans with acute respiratory distress syndrome identifies the selectin P ligand gene as a risk factor. Am J Respir Crit Care Med 197(11):1421–1432. https://doi.org/10.1164/rccm.201705-0961OC
Matthay MA, Arabi YM, Siegel ER, Ware LB, Bos LDJ, Sinha P, Beitler JR, Wick KD, Curley MAQ, Constantin JM, Levitt JE, Calfee CS (2020) Phenotypes and personalized medicine in the acute respiratory distress syndrome. Intensive Care Med. https://doi.org/10.1007/s00134-020-06296-9
Zou L, Ruan F, Huang M, Liang L, Huang H, Hong Z, Yu J, Kang M, Song Y, Xia J, Guo Q, Song T, He J, Yen HL, Peiris M, Wu J (2020) SARS-CoV-2 viral load in upper respiratory specimens of infected patients. N Engl J Med 382(12):1177–1179. https://doi.org/10.1056/NEJMc2001737
Cevik M, Kuppalli K, Kindrachuk J, Peiris M (2020) Virology, transmission, and pathogenesis of SARS-CoV-2. BMJ 371:m3862. https://doi.org/10.1136/bmj.m3862
Ellinghaus D, Degenhardt F, Bujanda L, Buti M, Albillos A, Invernizzi P, Fernandez J, Prati D, Baselli G, Asselta R, Grimsrud MM, Milani C, Aziz F, Kassens J, May S, Wendorff M, Wienbrandt L, Uellendahl-Werth F, Zheng T, Yi X, de Pablo R, Chercoles AG, Palom A, Garcia-Fernandez AE, Rodriguez-Frias F, Zanella A, Bandera A, Protti A, Aghemo A, Lleo A, Biondi A, Caballero-Garralda A, Gori A, Tanck A, Carreras Nolla A, Latiano A, Fracanzani AL, Peschuck A, Julia A, Pesenti A, Voza A, Jimenez D, Mateos B, Nafria Jimenez B, Quereda C, Paccapelo C, Gassner C, Angelini C, Cea C, Solier A, Pestana D, Muniz-Diaz E, Sandoval E, Paraboschi EM, Navas E, Garcia Sanchez F, Ceriotti F, Martinelli-Boneschi F, Peyvandi F, Blasi F, Tellez L, Blanco-Grau A, Hemmrich-Stanisak G, Grasselli G, Costantino G, Cardamone G, Foti G, Aneli S, Kurihara H, ElAbd H, My I, Galvan-Femenia I, Martin J, Erdmann J, Ferrusquia-Acosta J, Garcia-Etxebarria K, Izquierdo-Sanchez L, Bettini LR, Sumoy L, Terranova L, Moreira L, Santoro L, Scudeller L, Mesonero F, Roade L, Ruhlemann MC, Schaefer M, Carrabba M, Riveiro-Barciela M, Figuera Basso ME, Valsecchi MG, Hernandez-Tejero M, Acosta-Herrera M, D’Angio M, Baldini M, Cazzaniga M, Schulzky M, Cecconi M, Wittig M, Ciccarelli M, Rodriguez-Gandia M, Bocciolone M, Miozzo M, Montano N, Braun N, Sacchi N, Martinez N, Ozer O, Palmieri O, Faverio P, Preatoni P, Bonfanti P, Omodei P, Tentorio P, Castro P, Rodrigues PM, Blandino Ortiz A, de Cid R, Ferrer R, Gualtierotti R, Nieto R, Goerg S, Badalamenti S, Marsal S, Matullo G, Pelusi S, Juzenas S, Aliberti S, Monzani V, Moreno V, Wesse T, Lenz TL, Pumarola T, Rimoldi V, Bosari S, Albrecht W, Peter W, Romero-Gomez M, D’Amato M, Duga S, Banales JM, Hov JR, Folseraas T, Valenti L, Franke A, Karlsen TH, GG Severe Covid (2020) Genomewide association study of severe Covid-19 with respiratory failure. N Engl J Med. https://doi.org/10.1056/NEJMoa2020283
Pairo-Castineira E, Clohisey S, Klaric L, Bretherick AD, Rawlik K, Pasko D, Walker S, Parkinson N, Fourman MH, Russell CD, Furniss J, Richmond A, Gountouna E, Wrobel N, Harrison D, Wang B, Wu Y, Meynert A, Griffiths F, Oosthuyzen W, Kousathanas A, Moutsianas L, Yang Z, Zhai R, Zheng C, Grimes G, Beale R, Millar J, Shih B, Keating S, Zechner M, Haley C, Porteous DJ, Hayward C, Yang J, Knight J, Summers C, Shankar-Hari M, Klenerman P, Turtle L, Ho A, Moore SC, Hinds C, Horby P, Nichol A, Maslove D, Ling L, McAuley D, Montgomery H, Walsh T, Pereira A, Renieri A, Gen OI, Investigators I, Initiative C-HG, Me I, Investigators B, Gen CI, Shen X, Ponting CP, Fawkes A, Tenesa A, Caulfield M, Scott R, Rowan K, Murphy L, Openshaw PJM, Semple MG, Law A, Vitart V, Wilson JF, Baillie JK (2020) Genetic mechanisms of critical illness in Covid-19. Nature. https://doi.org/10.1038/s41586-020-03065-y
Ramanathan K, Antognini D, Combes A, Paden M, Zakhary B, Ogino M, MacLaren G, Brodie D, Shekar K (2020) Planning and provision of ECMO services for severe ARDS during the COVID-19 pandemic and other outbreaks of emerging infectious diseases. Lancet Respir Med 8(5):518–526. https://doi.org/10.1016/S2213-2600(20)30121-1
Goligher EC, Ranieri VM, Slutsky AS (2021) Is severe COVID-19 pneumonia a typical or atypical form of ARDS? And does it matter? Intensive Care Med 47(1):83–85. https://doi.org/10.1007/s00134-020-06320-y
National Heart L, Blood Institute Acute Respiratory Distress Syndrome Clinical Trials N, Rice TW, Wheeler AP, Thompson BT, Steingrub J, Hite RD, Moss M, Morris A, Dong N, Rock P (2012) Initial trophic vs full enteral feeding in patients with acute lung injury: the EDEN randomized trial. Jama 307(8):795–803. https://doi.org/10.1001/jama.2012.137
Reilly JP, Wang F, Jones TK, Palakshappa JA, Anderson BJ, Shashaty MGS, Dunn TG, Johansson ED, Riley TR, Lim B, Abbott J, Ittner CAG, Cantu E, Lin X, Mikacenic C, Wurfel MM, Christiani DC, Calfee CS, Matthay MA, Christie JD, Feng R, Meyer NJ (2018) Plasma angiopoietin-2 as a potential causal marker in sepsis-associated ARDS development: evidence from Mendelian randomization and mediation analysis. Intensive Care Med 44(11):1849–1858. https://doi.org/10.1007/s00134-018-5328-0
Su L, Zhai R, Sheu CC, Gallagher DC, Gong MN, Tejera P, Thompson BT, Christiani DC (2009) Genetic variants in the angiopoietin-2 gene are associated with increased risk of ARDS. Intensive Care Med 35(6):1024–1030. https://doi.org/10.1007/s00134-009-1413-8
Willer CJ, Li Y, Abecasis GR (2010) METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26(17):2190–2191. https://doi.org/10.1093/bioinformatics/btq340
Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D, Laurin C, Burgess S, Bowden J, Langdon R, Tan VY, Yarmolinsky J, Shihab HA, Timpson NJ, Evans DM, Relton C, Martin RM, Davey Smith G, Gaunt TR, Haycock PC (2018) The MR-Base platform supports systematic causal inference across the human phenome. eLife 7. https://doi.org/10.7554/eLife.34408
Chua RL, Lukassen S, Trump S, Hennig BP, Wendisch D, Pott F, Debnath O, Thurmann L, Kurth F, Volker MT, Kazmierski J, Timmermann B, Twardziok S, Schneider S, Machleidt F, Muller-Redetzky H, Maier M, Krannich A, Schmidt S, Balzer F, Liebig J, Loske J, Suttorp N, Eils J, Ishaque N, Liebert UG, von Kalle C, Hocke A, Witzenrath M, Goffinet C, Drosten C, Laudi S, Lehmann I, Conrad C, Sander LE, Eils R (2020) COVID-19 severity correlates with airway epithelium-immune cell interactions identified by single-cell analysis. Nat Biotechnol 38(8):970–979. https://doi.org/10.1038/s41587-020-0602-4
Lee JS, Park S, Jeong HW, Ahn JY, Choi SJ, Lee H, Choi B, Nam SK, Sa M, Kwon JS, Jeong SJ, Lee HK, Park SH, Park SH, Choi JY, Kim SH, Jung I, Shin EC (2020) Immunophenotyping of COVID-19 and influenza highlights the role of type I interferons in development of severe COVID-19. Sci Immunol 5 (49). https://doi.org/10.1126/sciimmunol.abd1554
Evangelou E, Ioannidis JP (2013) Meta-analysis methods for genome-wide association studies and beyond. Nat Rev Genet 14(6):379–389. https://doi.org/10.1038/nrg3472
Hall MA, Verma A, Brown-Gentry KD, Goodloe R, Boston J, Wilson S, McClellan B, Sutcliffe C, Dilks HH, Gillani NB, Jin H, Mayo P, Allen M, Schnetz-Boutaud N, Crawford DC, Ritchie MD, Pendergrass SA (2014) Detection of pleiotropy through a phenome-wide association study (PheWAS) of epidemiologic data as part of the Environmental Architecture for Genes Linked to Environment (EAGLE) study. PLoS Genet 10(12):e1004678. https://doi.org/10.1371/journal.pgen.1004678
Pu J, Schindler C, Jia R, Jarnik M, Backlund P, Bonifacino JS (2015) BORC, a multisubunit complex that regulates lysosome positioning. Dev Cell 33(2):176–188. https://doi.org/10.1016/j.devcel.2015.02.011
Niwa S, Tao L, Lu SY, Liew GM, Feng W, Nachury MV, Shen K (2017) BORC regulates the axonal transport of synaptic vesicle precursors by activating ARL-8. Curr Biol 27(17):2569–2578 e2564. https://doi.org/10.1016/j.cub.2017.07.013
Zhang H, Zheng H, Mu W, He Z, Yang B, Ji Y, Hui L (2015) DUSP16 ablation arrests the cell cycle and induces cellular senescence. FEBS J 282(23):4580–4594. https://doi.org/10.1111/febs.13518
Low HB, Zhang Y (2016) Regulatory roles of MAPK phosphatases in cancer. Immune Netw 16(2):85–98. https://doi.org/10.4110/in.2016.16.2.85
Torres Acosta MA, Singer BD (2020) Pathogenesis of COVID-19-induced ARDS: implications for an ageing population. Eur Respir J 56(3). https://doi.org/10.1183/13993003.02049-2020
Berlin DA, Gulick RM, Martinez FJ (2020) Severe Covid-19. N Engl J Med. https://doi.org/10.1056/NEJMcp2009575
Chiumello D, Busana M, Coppola S, Romitti F, Formenti P, Bonifazi M, Pozzi T, Palumbo MM, Cressoni M, Herrmann P, Meissner K, Quintel M, Camporota L, Marini JJ, Gattinoni L (2020) Physiological and quantitative CT-scan characterization of COVID-19 and typical ARDS: a matched cohort study. Intensive Care Med. https://doi.org/10.1007/s00134-020-06281-2
Brault C, Zerbib Y, Kontar L, Fouquet U, Carpentier M, Metzelard M, Soupison T, De Cagny B, Maizel J, Slama M (2020) COVID-19- versus non-COVID-19-related acute respiratory distress syndrome: differences and similarities. Am J Respir Crit Care Med 202(9):1301–1304. https://doi.org/10.1164/rccm.202005-2025LE
Tay MZ, Poh CM, Renia L, MacAry PA, Ng LFP (2020) The trinity of COVID-19: immunity, inflammation and intervention. Nat Rev Immunol. https://doi.org/10.1038/s41577-020-0311-8
Derikx L, Lantinga MA, de Jong DJ, van Dop WA, Creemers RH, Romkens TEH, Jansen JM, Mahmmod N, West RL, Tan A, Bodelier AGL, Gorter MHP, Boekema PJ, Halet ERC, Horjus CS, van Dijk MA, Hirdes MMC, Epping Stippel LSM, Jharap B, Lutgens M, Russel MG, Gilissen LPL, Nauta S, van Bodegraven AA, Hoentjen F (2020) Clinical outcomes of COVID-19 in patients with inflammatory bowel disease: a nationwide cohort study. J Crohns Colitis. https://doi.org/10.1093/ecco-jcc/jjaa215
Shoaib MH, Ahmed FR, Sikandar M, Yousuf RI, Saleem MT (2021) A journey from SARS-CoV-2 to COVID-19 and beyond: a comprehensive insight of epidemiology, diagnosis, pathogenesis, and overview of the progress into its therapeutic management. Front Pharmacol 12:576448. https://doi.org/10.3389/fphar.2021.576448
Hu J, Zhang L, Lin W, Tang W, Chan FKL, Ng SC (2021) Review article: probiotics, prebiotics and dietary approaches during COVID-19 pandemic. Trends Food Sci Technol 108:187–196. https://doi.org/10.1016/j.tifs.2020.12.009
Acknowledgements
The authors acknowledge the patients of the ARDS cohorts who graciously agreed to participate in this research study. They also acknowledge the nurses, physicians, and staff in the medical and surgical ICUs who participated in the clinical care of the enrolled patients. They also thank Dr. Mark M. Wurfel (University of Washington, Seattle, WA, USA) for his efforts in co-leading the iSPAAR cohort.
Funding
This study was supported by the US National Institutes of Health (R01HL060710, R56HL134356, and ES000002 to D.C.C.) and the Special Program of the National Natural Science Foundation of China (NSFC) on tracing, pathogenesis, prevention, and treatment for COVID-19 (82041024 to F.C.).
Author information
Authors and Affiliations
Contributions
M.L.D. and D.C.C. conceived the study design and had full access to the data. M.L.D. took responsibility for the integrity of data and accuracy of the analyses. D.C.C., J.G.G., J.D.C., Z.Z.Z., P.T., N.J.M., and M.L.W. organized and entered data. M.L.D., J.Y.X., G.S.C., S.P.S., X.S.D., H.L., and J.N.H. contributed to data analyses. M.L.D., J.Y.X., Q.Y.Y., Z.D.Z., F.C., M.L.W., and X.H.L. contributed to data interpretation. M.L.D. and D.C.C. drafted the manuscript. All authors made significant contributions to the final manuscript and approve its submission.
Corresponding authors
Ethics declarations
Conflicts of interest
The authors state that there is no conflict of interest.
Ethical approval
The study protocol of each cohort was approved by the institutional review board at each participating site.
Consent to participate
All participants provided written consent.
Consent for publication
All authors approve this study submission.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Du, M., Garcia, J.G.N., Christie, J.D. et al. Integrative omics provide biological and clinical insights into acute respiratory distress syndrome. Intensive Care Med 47, 761–771 (2021). https://doi.org/10.1007/s00134-021-06410-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00134-021-06410-5