Abstract
Seven strains that can degrade nitrobenzene at low temperature were isolated from the sediments of a nitrobenzene-polluted river. One of the strains, NB-1, can mineralize 20 mg/L nitrobenzene completely from 2.5 to 35°C with an optimum temperature of 25°C. NB-1 was identified as Pseudomonas putida according to its morphology, biochemical properties, and 16S rDNA sequence analysis. The strain could tolerate and degrade 120 mg/L nitrobenzene within 175 h at 5°C. It can be used for the bioremediation of nitrobenzene-contaminated environment.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Nitrobenzene (NB) is one of the top 50 industrial chemicals produced in United States; about 19 million pounds of NB is released into the environment annually due to its usage, leakage or industrial accidents (Storck et al. 1996; Haigler and Spain 1991). For example, the Songhuajiang River in northeast China suffered a major water pollution incident owing to the explosion of a petrochemical plant at the upper reaches on November 13, 2005. More than 500,000 tons of NB was discharged into Songhuajiang River, resulting in a highly NB contaminated environment (http://www.chinaview.cn, 2005).
For the remediation of large-scale contaminated environment, biological methods are usually preferred. Major advantages of bioremediation are lower capital costs and the ability to perform the task on site. Bioremediation process requires specific bacteria to biodegrade the contaminants. Aerobic bacteria that are capable of utilizing NB as sole carbon, nitrogen, and energy sources have been isolated from contaminated soils and groundwaters, such as Acidovorax sp. (Lessner et al. 2003), Comamonas sp. (Nishino and Spain 1995) and Pseudomonas sp. (Zhao and Ward 2000).
However, the above mentioned microorganisms are all mesophilic, which show optimal activity at temperatures higher than those found in nature and they cannot grow at low temperature. There is surprisingly little information about the degradative potential of cold-adapted culture for the degradation of NB. When real bioremediation is concerned, environmental temperature must be considered and the activity of microorganisms adapted to low temperatures and able to degrade pollutants is required (Philipp et al. 2005).
Therefore it was the objective of this study to detect, isolate, and identify cold-adapted bacteria from nitrobenzene contaminated sediments (Songhuajiang River) and to evaluate their potential for the biodegradation of nitrobenzene at low temperature.
Materials and Methods
All microbial enrichment and isolation procedures were performed in media with inorganic culture media base of the following composition (in grams per liter): Na2HPO4·12H2O, 3.8; KH2PO4, 1; NaCl, 1; NH4Cl, 0.1; MgSO4, 0.2. The sediment samples were collected from the Songhuajiang river on January of 2006. Six grams of samples were mixed with inorganic culture media (100 mL) for 2 min. The mixture (10 mL) was added into the flask containing 100 mL inorganic culture media and 20 mg/L nitrobenzene. The mixture culture was placed on a shaker (120 rpm) at 30°C. The biodegradation of NB in the flask was monitored periodically with gas chromatography (GC) with NB repeatedly added into the flask until the degradation reached a stable level.
At stable degradation of nitrobenzene, the mixed culture was diluted in inorganic culture media and plated onto media with nutrient agar and 100 mg/L nitrobenzene to obtain single colonies (the plates were incubated at 30°C for 24 h). The loopfuls of single colony from each plate were streaked onto other fresh agar plates containing nitrobenzene to check for purity. Individual colonies were then isolated, tested for their degradation activity and stored at −80°C in 15% glycerol.
The nitrobenzene degrading bacteria was identified according to the criteria of Bergey’s manual of determinative bacteriology and by sequencing the partial 16S rDNA gene. The sequence was performed at State Key Laboratory of Microbial Resources (Chinese Academy of Science, China). PCR amplifications were performed using the reverse primer 5′-TACGG TACCT TGTTA CGACTT-3′ and the forward primer 5′-AAGAG TTTGA TCTGG CTCAG-3′ for 16S rDNA. The 16s rDNA sequence was compared against the GenBank database using DNAstar software.
Nitrobenzene biodegradation by the isolated bacteria was firstly studied at different temperatures, i.e. 0, 2.5, 5, 15, 25, 35, 45°C to find out the optimum cell growth and nitrobenzene degradation temperature. Each 300 mL flask contained 150 inorganic culture media with cell suspensions containing 107 cells/mL, and then 20 mg/L nitrobenzene was added to serve as the sole carbon and energy source. After sealing with cotton stoppers, the flasks were shaken at 120 rpm and samples were taken at appropriate time to observe nitrobenzene removal. Control experiments were also performed where necessary. Similar experiments were performed to find the effect of initial nitrobenzene concentration (20, 40, 80, 120 and 160 mg/L) and pH (4, 5, 6, 7, 8, and 9) on nitrobenzene degradation at 5°C.
Cell concentration was determined by measuring the optical density (OD) at an absorbance of 560 nm using a Shimadzu UV–Visible Spectrophotometer UV-1601. For analysis of NB concentrations, the sample was centrifuged and extracted with methyl-t-butyl ether. A 1 μL extract was then analyzed for nitrobenzene by using a capillary gas chromatograph (Aglient, Model 6890N) with an ECD detector. The Dissolved oxygen carbon (DOC) was analyzed using a Shimadzu TOC analyzer (TOC-5000A).
Results and Discussion
The sediments were collected during the coldest month and therefore appeared more likely to contain psychrotrophic strains. The samples were inoculated in the medium containing nitrobenzene. Initially about 14 isolates were grown out of which seven were distinct according to their morphology. Thus, the soil samples appeared to contain a wide range of nitrobenzene-degrading strains in which the influence of cold temperature was subsequently tested. The detailed morphological and nitrobenzene-degrading characteristics of the seven isolates were given in Table 1. As can be seen, strain NB-1, 3, 4, 6 exhibited more growth and faster degradation in nitrobenzene than strain NB-2, 5, 7.
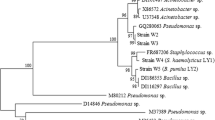
Growth and nitrobenzene-degrading profiles of NB-1 and NB-5 were shown in Fig. 1. Both strains can utilize NB as the sole carbon and energy source. NB-1 showed faster growth and nitrobenzene degradation rates than NB-5. It took about 43 and 70 h for NB-1 and NB-5 to degrade 90% of the nitrobenzene respectively. Therefore, NB-1 was selected as the representative bacteria for the biodegradation of nitrobenzene in the following study. Upon comparison of the partial 16S rDNA gene sequence (1,492 bases) obtained from strain NB1 with sequences from the Genebank Database, the highest degree of identity was obtained with the 16S rDNA gene sequence of a Pseudomonas putida. Analysis of the 16S rDNA gene sequence and phenotypic analysis (Table 2) suggested that this strain (NB1) was a Pseudomonas putida one.
Figure 2 shows the growth rate of NB-1 and nitrobenzene degradation rate with different temperatures (0–45°C). The optimum growth temperature of Pseudomonas putida was around 25°C by utilizing 20 mg/L of nitrobenzene as sole substrate. However, complete growth inhibition of Pseudomonas putida occurred at 0 and 45°C. Eighty-five percent of dissolved organic carbon was removed after the degradation experiment, showing the ability of the bacteria to mineralize nitrobenzene.
In cold and temperature climates, environment temperatures usually are subjected to important daily and seasonal fluctuations. Cold-adapted microorganisms are potentially interesting for use when soil and wastewater are treated in areas that experience low temperatures for large parts of the year. These organisms can be divided into psychrophilic and psychrotrophic (or psychrotolerant) organisms according to definitions used by Morita (1975). Psychrophiles are organisms that have minimum, optimum and maximum growth temperatures of ≤0, ≤15, ≤20°C, respectively, while the corresponding temperatures for psychroptrophs are 0–5, >15 and >20°C (Morita 1975; Margesin and Schinner 1997). These ranges show that the most interesting organisms, in many cases, are the psychrotrophs since they are active also at temperatures above 20°C. Consequently, they may be more active than the psychrophiles for a longer period each year in many environments. Our result reveals that NB1 is a psychrotrophic bacterium, having a potential for the bioremediation of nitrobenzene contaminated environment.
Figure 3 shows the temporal profiles of nitrobenzene at different initial concentrations at 5°C. Inoculated bacteria can degrade nitrobenzene till concentration of 120 mg/L with a prolonged lag phase of 22, 46 and 75 h. At 160 mg/L, only 8% of nitrobenzene was removed within 165 h. Inhibition was severe at this concentration and bacteria cannot degrade the nitrobenzene at the time course.
The removal of 20 mg/L of nitrobenzene by NB-1 with different initial pH value at 5°C at 30 and 60 h is shown in Fig. 4. It is clearly that the optimum pH for NB removal was between 6 and 9. NB-1 was also observed to degrade NB maximally at pH 7.0, showing that a neutral to slightly alkaline pH may be required for NB degradation. This must be considered during bioremediation of nitrobenzene contaminated environment.
While NB-degrading pure cultures have been isolated from nitroaromatic-contaminated sites, NB tends to be persist in the environment. The estimated half-life of NB in environment such as surface water, ground water, and soil ranged from several days to more than 625 days (Mackey et al. 1995). Our results show that cold-adapted nitrobenzene degrader can be readily isolated, which may be of interest in view of environment bioremediation.
References
Haigler BE, Spain JC (1991) Biotransformation of nitrobenzene by bacteria containing toluene degradative pathways. Appl Environ Microbiol 57(11):3156–3162
Hughes JB, Beckles DM, Chandra SD, Ward CH (1997) Utilization of bioremediation processes for the treatment of PAH-contaminated sediments. J Ind Microbiol Biotechnol 18: 152–160
Lessner DJ, Parales RE, Narayan S, Gibson DT (2003) Expression of nitroarene dioxygenase genes in Comamonas sp. strain JS765 and Acidovorax sp. strain JS42 is induced by multiple aromatic compounds. J Bacteriol 185:3895–3904
Mackey QD, Shin WY, Ma KC (1995) Illustrated handbook of physical properties and environmental fate for organic chemicals. Lewis publishers, Boca Raton
Margesin R, Schinner F (1997) Bioremediation of diesel oil-contaminated Alpine soils at low temperatures. Appl Microbiol Biotechnol 47: 462–468
Morita RY (1975) Psychrophilic bacteria. Bacteriol Rev 39: 144–167
Nishino SF, Spain JC (1995) Oxidative pathway for the biodegradation of nitrobenzene by Comamonas sp. strain JS765. Appl Environ Microbiol 61:2308–2313
Philipp B, Pierre-Alain F, Nicole N, Franz S, Rosa M (2005) Biodegradation of phenol and phenol-related compounds by psychrophilic and cold-tolerant alpine yeasts. Chemosphere 59:909–918
Storck WJ, Layman PL, Reisch MS, Thayer AM (1996) Facts and figures for the chemical industry. Chem Eng News 24:38–79
www.chinaview.cn (2005) Major river pollution confirmed in northeast China, 2005–11—23,16:14:42, http://www.news.xinhuanet.com/english/2005-11/23/content_3823846.htm
Zhao JS, Owen PW (2000) Cometabolic biotransformation of nitrobenzene by 3-nitrophenol degrading Pseudomonas putida. Can J Microbiol 46(7):643–648
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, Y., Hu, H. & Wu, Q. Isolation and Characterization of Psychrotrophic Nitrobenzene-Degrading Strains from River Sediments. Bull Environ Contam Toxicol 79, 340–344 (2007). https://doi.org/10.1007/s00128-007-9239-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00128-007-9239-y