Abstract
Allosteric activators of human glucokinase (GK) had revealed significant hypoglycemic effects for therapy of type-2 diabetes (T2D) in animal as well as human models. Some newer N-benzimidazol-2yl substituted benzamide analogues were prepared and assessed for activation of GK accompanied by molecular docking investigations for predicting the bonding interactions of these derivatives with the residues in allosteric site of GK protein. Amongst the derivatives synthesized, compounds 2 and 7 strongly increased catalytic action of GK (GK activation fold >2.0 in comparison to control) in vitro. The results of in-vitro testing were supported by the molecular docking investigations of these analogues with GK protein’s allosteric site residues (showed appreciable H-bond interactions with Arg63 residue of GK). Derivatives investigated in present study afforded few lead compounds for the discovery of harmless and strong allosteric GK activating compounds for treating T2D.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
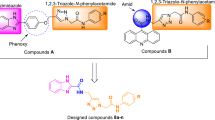
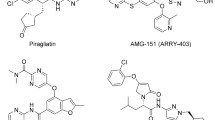
Type-2 diabetes (T2D) is a chronic disease of the food metabolic pathways owing to decreased insulin action resulting in hyperglycemia and prevalent amongst most of the patients suffering from diabetes [1,2,3]. Although ample types of oral antidiabetic agents are existing to be used for the management of T2D, no individual antidiabetic agent is valuable in attaining persistent homeostasis of plasma sugar within usual physiological range in majority of the persons suffering from T2D. Owing to the above points, these days doctors advise combination of hypoglycemic agents at an initial phase of T2D treatment. Additionally, overdose of hypoglycemic drugs may possibly result in serious hypoglycemia triggering brutal adverse reactions, and patients generally require urgent medical treatment [2, 4, 5]. Now-a-days, medicinal chemistry scientists are aiming at designing newer effective hypoglycemic agents having distinct mechanism of action at molecular level which could be used as single drug with improved safety [6, 7]. Glucokinase (GK) is a cytosolic enzyme that is primarily expressed in pancreatic β-cells and liver hepatocytes; as well as lifts up glucose transformation to glucose-6-phosphate with the aid of adenosinetriphosphate (ATP) [8, 9]. In beta-cells of pancreas gland, GK regulates glucose-instigated discharge of insulin and in liver hepatocytes of liver; it commands the breakdown of sugars. GK acts as an emergent medication focus for treatment and management of T2D due to its key function in controlling sugar breakdown. Small molecule activators of human GK are the unique class of therapeutically useful agents that allosterically activate GK and illustrate their plasma sugar lowering potential [6, 9,10,11,12]. Several GK activators had been progressed into clinical trials (phase II) including AZD6370, AZD1656, MK-0941, Piragliatin, and AMG151; even though strong decrease in blood sugar was observed, potential adverse reactions were also reported, such as hypoglycemia and elevated levels of triglycerides. Literature survey revealed that most of the drug discovery and development research associated with allosteric activators of human GK were mainly focused on the substituted benzamide analogues (specially bearing a hetero-aromatic ring connected to the benzamide NH- atom) probably owing towards their corresponding alignment outline and bonding interactions with the residues of allosteric binding site of GK protein [4, 6, 13,14,15,16,17,18,19,20,21,22,23,24,25,26,27]. Based on the above facts few newer N-benzimidazol-2-yl benzamide analogues were proposed as potential activators of human GK enzyme.
Materials and methods
Chemicals were acquired from reputed companies such as Spectrochem, SRL, S.D. Fine-Chem, Merck, Fisher Scientific and Sigma-Aldrich etc., and employed without purification. Melting points of the synthesized molecules were estimated by uncorrected Veego VMP-D melting point device. Silica Gel-G TLC was used for monitoring the progress of chemical reaction. Shimadzu FTIR spectrophotometer using potassium bromide pellet process was utilized for recording IR spectra. ‘Avance-II (Bruker) 400 MHz NMR spectrophotometer’ was employed for taking 1H-NMR and 13C-NMR spectra employing appropriate dutereated solvents. Results were represented in ‘parts per million’ (‘δ’, ppm) downfield to ‘Si(CH3)4’ (internal reference).
General procedure for preparation of designed molecules
Dry benzoic acid (1 mmol) was transferred to a flat base flask fixed on a magnetic agitator at constant temperature around 10 °C. Excess of HClSO3 (8.0 mL) was added carefully and observed to avoid any escape. When all the acid was liquefied and the exothermic response terminated, the flat bottom flask was heated at 70–80 °C using water bath for 2 h followed by cooling. The materials of flask were poured to crushed ice (150 g) cautiously with stirring and crystals of 3-(chlorosulphonyl)benzoic acid were filtered employing vacuum subsequent to cold water wash followed by air drying. The precipitates prepared above (1 mmol) were then reacted with the corresponding aliphatic and aromatic amines (1 mmol) under reflux using acetone till completion of reaction (observed using silica gel G TLC) following cooling and drying of the precipitates. The different sulphonamides prepared above (1 mmol) were refluxed for 3 h with sulfinyl chloride (1 mmol) and to receive equivalent acid chlorides excess sulfinyl chloride was withdrawn. Acid chlorides prepared above (1 mmol) were refluxed with 2-aminobenzimidazole (1.5 mmol). The end products (compounds 1–10) obtained by the evaporation of solvent were purified using recrystallization from ethanol [28,29,30].
N-(1H-Benzimidazol-2-yl)-3-(phenylsulfamoyl)benzamide (1)
Pale white solid; Yield: 72%; Mp (°C) 158–161; FTIR (KBr Pellets) ν cm−1: 3867.78 (NH str., CONH), 3737.50 (NH str., Benzamide), 3432.08.46 (NH str., SO2NH), 2973.38 (CH str., Aromatic), 1642.58 (Carbonyl C=O str., Benzamide), 1558.12 (NH bend, Aromatic amine), 1463.36 (C=N str., Aromatic, Benzimidazol-2-yl), 1417.54 (C=C str., Aromatic), 1296.70 (SO2 asym. str., SO2NH), 1100.00 (C–N str., Benzimidazol-2-yl), 1076.13 (SO2 sym. str., SO2NH, Sulphonamide), 752.08 (CH bend, Aromatic); 1H-NMR (δ ppm, DMSO-d6): 12.68 (s, 1H, NH, CONH), 8.02–8.64 (m, 4H, CH, C2, C4, C5 and C6 of C6H4CO), 7.36–8.46 (m, 4H, CH, Benzimidazol-2-yl), 6.38–7.19 (m, 5H, CH of C2, C3, C4, C5 and C6 of C6H5), 3.00 (s, 1H, NH, Benzimidazol-2-yl), 2.58 (s, 1H, NH, SO2NH); 13C-NMR (δ ppm, DMSO-d6): 168.67 (C=N), 164.15 (C=O), 151.78 (C), 136.16 (C), 134.89 (C), 131.78 (C), 130.13 (CH), 129.17 (CH), 128.41 (C), 128.02 (CH), 125.23 (CH), 123.67 (CH), 120.15 (CH), 119.34 (CH), 117.97 (CH), 115.42 (CH).
N-(1H-Benzimidazol-2-yl)-3-[(2-chloro-4-nitrophenyl)sulfamoyl]benzamide (2)
Yellowish brown solid; Yield: 78%; Mp (°C) 162–165; FTIR (KBr Pellets) ν cm−1: 3836.20 (NH str., CONH), 3446.91 (NH str., SO2-NH), 2928.28 (CH str., Aromatic), 1641.34 (C=O str., CONH), 1632.23 (NH bend, Ar–NH), 1551.35 (C=N str., Aromatic), 1464.11 (NO2 sym. str., NO2), 1413.94 (NO2 asym. str., NO2), 1299.66 (SO2 asym. str., SO2NH), 1100.00 (C–N str., Benzimidazol-2-yl), 1079.66 (SO2 sym. str., SO2NH), 684.36 (C–Cl str., Aromatic); 1H-NMR (δ ppm, DMSO-d6): 9.28 (s, 1H, NH, CONH), 8.67–8.43 (m, 4H, CH, C6H4CO), 7.34–7.91 (m, 4H, CH, Benzimidazol-2-yl), 7.08–7.24 (m, 3H, CH, C6H3ClNO2), 4.48 (s, 1H, NH, Benzimidazol-2-yl), 4.05 (s, 1H, NH, SO2NH); 13C-NMR (δ ppm, DMSO-d6): 169.78 (C=N), 164.58 (C=O), 147.46 (C), 139.06 (C), 137.12 (C), 135.03 (C), 134.45 (C), 132.08 (CH), 129.12 (CH), 128.69 (C), 127.58 (CH), 126.47 (C), 125.07 (CH), 124.17 (CH), 119.58 (CH), 118.14 (CH), 114.36 (CH), 112.94 (CH).
N-(1H-Benzimidazol-2-yl)-3-(benzylsulfamoyl)benzamide (3)
Light brown solid; Yield: 62%; Mp (°C) 160–163; FTIR (KBr Pellets) ν cm−1: 3755.80 (NH str., Benzamide), 3448.08 (NH str., SO2NH), 2996.40 (CH str., Aromatic), 2912.85 (CH str., Aliphatic), 1659.53 (C=O str., CONH), 1429.38 (NH bend, Ar–NH), 1311.51 (SO2 asym. str., SO2NH), 1100.00 (C–N str., Benzimidazol-2-yl), 1025.25 (SO2 sym. str., SO2NH), 696.02 (CH bend, Aromatic); 1H-NMR (δ ppm, DMSO-d6): 10.93 (s, 1H, NH, CONH), 8.20–8.51 (m, 4H, CH, C6H4CO), 7.70–8.14 (m, 4H, CH, Benzimidazol-2-yl), 7.12–7.58 (m, 5H, CH, C6 of C6H5), 6.64 (t, 1H, NH, Sulphonamide), 2.54 (s, 1H, NH, Benzimidazol-2-yl), 2.51 (d, 1H, CH, CH2); 13C-NMR (δ ppm, DMSO-d6): 175.18 (C=N), 164.76 (C=O), 153.46 (C), 141.24 (C), 140.15 (C), 135.14 (C), 135.03 (C), 132.26 (CH), 131.92 (CH), 128.08 (CH), 124.14 (CH), 122.16 (CH), 120.94 (CH), 119.04 (CH), 116.68 (CH), 45.88 (CH2).
N-(1H-Benzimidazol-2-yl)-3-(butylsulfamoyl)benzamide (4)
Light brown solid; Yield: 59%; Mp (°C) 156–158; FTIR (KBr Pellets) ν cm−1: 3754.38 (NH str., CONH, Benzamide), 3448.26 (NH str., SO2NH, Sulphonamide), 2930.77 (CH str., Aromatic), 2962.66 (CH str., Aliphatic), 1643.43 (C=O str., CONH, Benzamide), 1554.13 (NH bend, Ar–NH), 1464.82 (C=C str., Aromatic), 1415.35 (SO2 asym. str., SO2NH, Sulphonamide), 1100.00 (C–N str., Benzimidazol-2-yl), 1076.78 (SO2 sym. str., SO2NH); 1H-NMR (δ ppm, DMSO-d6): 8.59 (s, 1H, NH, CONH), 7.34–7.88 (m, 4H, CH, C6H4CO), 7.05–7.86 (m, 4H, CH, Benzimidazol-2-yl), 2.54 (t, 1H, NH, Sulphonamide), 2.50 (s, 1H, NH, Benzimidazol-2-yl), 1.31 (m, 9H, Butyl); 13C-NMR (δ ppm, DMSO-d6): 175.18 (C=N), 165.29 (C=O), 151.49 (C), 139.65 (C), 135.26 (C), 135.02 (C), 131.07 (CH), 128.44 (CH), 127.68 (CH), 124.59 (CH), 119.28 (CH), 118.93 (CH), 118.02 (CH), 115.02 (CH), 45.38 (CH2), 25.58 (CH2), 21.07 (CH2), 15.08 (CH3).
N-(1H-benzimidazol-2-yl)-3-(methylsulfamoyl)benzamide (5)
Dark brown solid; Yield: 52%; Mp (°C) 148–152; FTIR (KBr Pellets) ν cm−1: 3798.48 (NH str., CONH), 3448.44 (NH str., SO2NH), 3017.57 (CH str., Aromatic), 2966.14 (CH str., Alkyl), 1654.21 (Carbonyl str., CONH), 1598.09 (NH bend, Aromatic amine), 1544.68 (C=N str., Aromatic), 1388.45 (SO2 asym. str., SO2NH), 1189.77 (SO2 sym. str., SO2NH), 1100.00 (C–N str., Benzimidazol-2-yl), 789.65 (CH bend, Aromatic); 1H-NMR (δ ppm, DMSO-d6): 8.60 (s, 1H, NH, CONH), 7.49–7.98 (m, 4H, CH, C6H4CO), 7.10–7.58 (m, 4H, CH, C4, C5, C6 and C7 of Benzimidazol-2-yl), 5.20 (t, 1H, NH, Sulphonamide), 2.54 (s, 1H, NH, Benzimidazol-2-yl), 2.51 (s, 3H, Methyl); 13C-NMR (δ ppm, DMSO-d6): 174.27 (C=N), 165.16 (C=O), 152.86 (C), 141.06 (C), 138.84 (C), 135.38 (CH), 133.08 (C), 132.66 (CH), 127.58 (CH), 126.05 (CH), 122.98 (CH), 120.94 (CH), 118.18 (CH), 117.08 (CH), 31.14 (CH3).
N-(1H-Benzimidazol-2-yl)-3-[(2-methylphenyl)sulfamoyl]benzamide (6)
Light brown solid; Yield: 60%; Mp (°C) 162–165; FTIR (KBr Pellets) ν cm−1: 3791.96 (NH str., CONH, Benzamide), 3456.56 (NH str., SO2NH, Sulphonamide), 3013.67 (CH str., Aromatic), 2912.67 (CH str., Aliphatic), 1667.25 (NH str., SO2NH), 1604.66 (NH bend, Aromatic amine), 1578.56 (C=N str., Aromatic), 1345.34 (SO2 asym. str., SO2NH, Sulphonamide), 1103.78 (SO2 sym. str., SO2NH, Sulphonamide), 1100.00 (C–N str., Benzimidazol-2-yl), 850.55 (CH bend, Aromatic); 1H-NMR (δ ppm, DMSO-d6): 10.27 (s, 1H, NH, CONH), 8.01–8.46 (m, 4H, CH, C6H4CO), 7.34–8.02 (m, 4H, CH, Benzimidazol-2-yl), 7.04–7.39 (m, 4H, CH, C6H4CH3), 2.57 (s, 1H, NH, Benzimidazol-2-yl), 2.18 (s, 1H, NH, SO2NH), 2.14 (s, 3H, Methyl); 13C-NMR (δ ppm, DMSO-d6): 175.16 (C=N), 165.26 (C=O), 151.42 (C), 139.55 (C), 138.06 (C), 136.32 (C), 134.87 (CH), 133.34 (C), 129.67 (C), 129.12 (CH), 126.55 (CH), 125.12 (CH), 123.56 (CH), 121.68 (CH), 117.36 (CH), 17.32 (CH3).
N-(1H-Benzimidazol-2-yl)-3-[(4-bromophenyl)sulfamoyl]benzamide (7)
Grayish black solid; Yield: 74%; Mp (°C) 158–160; FTIR (KBr Pellets) ν cm−1: 3837.14 (NH str., Benzamide), 3732.98 (NH str., Benzamide), 3441.64 (NH str., SO2NH, Sulphonamide), 2974.87 (CH str., Aromatic carbon), 1641.67 (Carbonyl str., Benzamide), 1553.91 (NH bend, Aromatic amine), 1464.33 (C=N str., Aromatic), 1415.88 (SO2 asym. str., SO2NH), 1296.76 (SO2 sym. str., SO2NH), 1100.00 (C–N str., Benzimidazol-2-yl), 809.70 (CH bend, Aromatic), 753.82 (C–Br str., Aromatic); 1H-NMR (δ ppm, DMSO-d6): 9.49 (s, 1H, NH, CONH), 7.78–8.66 (m, 4H, CH, C6H4CO), 7.14–7.92 (m, 4H, CH, Benzimidazol-2-yl), 6.20–7.44 (m, 4H, CH, C6H4Br), 4.48 (s, 1H, NH, Benzimidazol-2-yl), 4.02 (s, 1H, NH, SO2NH); 13C-NMR (δ ppm, DMSO-d6): 175.28 (C=N), 165.82 (C=O), 152.47 (C), 142.05 (C), 139.10 (C), 136.97 (C), 136.26 (C), 132.98 (C), 132.02 (CH), 130.36 (CH), 130.01 (CH), 129.67 (CH), 127.12 (CH), 124.84 (CH), 122.04 (CH), 120.18 (CH), 116.45 (CH).
N-(1H-Benzimidazol-2-yl)-3-[(4-methylphenyl)sulfamoyl]benzamide (8)
Light brown solid; Yield: 63%; Mp (°C) 160–163; FTIR (KBr Pellets) ν cm−1: 3870.59 (NH str., Benzamide), 3755.40 (NH str., CONH), 3452.66 (NH str., SO2NH), 2997.49 (CH str., Aromatic), 1708.27 (C=O str., Amide), 1429.03 (NO2 sym. str., Aromatic nitro group), 1362.55 (NO2 asym. str., Aromatic nitro group), 1323.98 (SO2 asym. str., SO2NH), 1223.02 (SO2 sym. str., SO2NH), 1100.00 (C–N str., Benzimidazol-2-yl), 696.02 (CH bend, Aromatic); 1H-NMR (δ ppm, DMSO-d6): 7.95 (s, 1H, NH, CONH), 7.16–7.90 (m, 4H, CH, C6H4CO), 7.26–7.84 (m, 4H, CH, Benzimidazol-2-yl), 7.16–7.88 (m, 4H, CH, C6H4NO2), 5.00 (s, 1H, NH, Benzimidazol-2-yl), 2.50 (s, 1H, NH, Sulphonamide); 13C-NMR (δ ppm, DMSO-d6): 174.63 (C=N), 164.94 (C=O), 151.68 (C), 142.21 (C), 139.04 (C), 138.14 (C), 135.28 (C), 132.76 (C), 132.12 (CH), 128.69 (CH), 125.36 (CH), 122.09 (CH), 120.01 (CH), 119.25 (CH), 118.42 (CH), 117.68 (CH), 115.18 (CH), 24.14 (CH3).
N-(1H-Benzimidazol-2-yl)-3-[(4-nitrophenyl)sulfamoyl]benzamide (9)
Pale yellow solid; Yield: 74%; Mp (°C) 165–168; FTIR (KBr Pellets) ν cm−1: 3868.16 (NH str., CONH), 3754.28 (NH str., CONH), 3448.36 (NH str., SO2NH), 2930.77 (CH str., Aromatic), 1643.31 (C=O str., CONH), 1553.03 (NH bend), 1524.42 (NO2 sym. str.), 1464.83 (C=N str., Aromatic), 1441.52 (NO2 asym. str.), 1415.35 (CH bend, Aliphatic), 1300.62 (SO2 asym. str., SO2NH), 1100.00 (C–N str., Aromatic), 1076.79 (SO2 sym. str., SO2NH), 717.52 (CH bend, Aromatic); 1H-NMR (δ ppm, DMSO-d6): 10.47 (s, 1H, NH, CONH), 7.89–8.32 (m, 4H, CH, C6H4CO), 7.39–7.56 (m, 4H, CH, Benzimidazol-2-yl), 7.20–7.50 (m, 4H, CH, C6H4CH3), 2.50 (s, 1H, NH, Benzimidazol-2-yl); 13C-NMR (δ ppm, DMSO-d6): 175.56 (C=N), 165.08 (C=O), 153.84 (C), 143.80 (C), 139.46 (C), 138.86 (C), 137.47 (CH), 135.31 (C), 133.94 (C), 131.08 (CH), 129.95 (CH), 129.18 (CH), 126.02 (CH), 122.91 (CH), 121.27 (CH), 120.67 (CH), 119.37 (CH), 116.62 (CH).
N-(1H-benzimidazol-2-yl)-3-(propylsulfamoyl)benzamide (10)
Light brown solid; Yield: 48%; Mp (°C) 155–158; FTIR (KBr Pellets) ν cm−1: 3450.06 (NH str., CONH), 2996.68 (NH str., SO2NH), 2912.98 (CH str.), 1689.51 (C=O str., CONH), 1428.92 (NH bend), 1311.73 (C=N str., Aromatic), 1100.00 (C–N str., Benzimidazol-2-yl), 1023.65 (SO2 asym. str., SO2NH), 950.47 (SO2 sym. str., SO2NH), 696.27 (CH bend, Aromatic); 1H-NMR (δ ppm, DMSO-d6): 10.25 (s, 1H, NH, CONH), 8.37–8.71 (s, 3H, CH, C6H3CO), 7.56–7.94 (m, 4H, CH, Benzimidazol-2-yl), 2.61 (s, 1H, NH, Benzimidazol-2-yl), 2.58 (s, 1H, NH, SO2NH), 2.44 (m, 2H, Methylene), 2.05 (m, 2H, Methylene), 1.27 (t, 3H, methyl); 13C-NMR (δ ppm, DMSO-d6): 174.24 (C=N), 165.02 (C=O), 151.33 (C), 139.00 (C), 136.88 (C), 134.65 (C), 130.34 (CH), 129.17 (CH), 125.37 (CH), 124.92 (CH), 120.49 (CH), 118.78 (CH), 118.02 (CH), 117.33 (CH), 46.04 (CH2), 32.48 (CH2), 12.44 (CH3).
In silico estimation of pharmacokinetic properties
The designed molecules were analyzed for the prediction of pharmacokinetics using FAF-Drugs4 server; as well as accessed for their “drug likeness” potential utilizing Lipinski’s rule [27, 31, 32].
In vitro GK assay
GK activation potential of all the derivatives was assessed employing a combined response with glucose-6-phosphate dehydrogenase (G6PDH) using spectrometry. All the samples were made using DMSO and the in-vitro GK test was done in an ultimate quantity of ‘2000 µL’ comprising of 4-(2-Hydroxyethyl)-1-piperazineethanesulfonic acid (pH 7.4) (0.25 mM), 10 mM dextrose (10 mM), KCl (25 mM), MgCl2 (1 mM), 1,4-dithio-D-threitol (1 mM), 1 mM nicotinamide adenine dinucleotide, 1 mM ATP, G6PDH (2.5 U/mL), 0.5 µg GK, and derivatives to be tested (10 µM). Readings were taken at 340 nm following nurture time of 3 min and GK activation was computed in comparison to DMSO (activation of GK by DMSO alone was treated as 100%). All the results were represented as mean (n = 3) ± standard deviation (S.D.). The in vitro GK assay data for test groups was statistically analyzed by one-way ANOVA for comparison and significance from control group (value of p < 0.05) using GraphPad Prism (GraphPad Software Inc.) [22, 24, 33, 34].
Molecular docking investigations
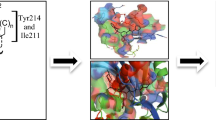
AutoDock Vina and AutoDock Tools were used for executing molecular docking of all the analogues in the allosteric location of the GK (PDB ID: ‘3IMX’) [35, 36]. The protocol followed for docking of designed analogues with GK is presented in Fig. 1 as previously reported [22, 24, 26, 30, 37, 38].
In silico prediction of toxicity
All samples were screened in silico to estimate the potential toxicity of such substances using online computer software pkCSM [39,40,41].
Results and discussion
Chemistry
3-(Chlorosulphonyl)-benzoic acid acquired through chlorosulphonation of benzoic acid was reacted with different amine derivatives to obtain various sulphonamide analogues which were then reacted with SO2Cl followed by reaction with 2-aminobenzimidazole to synthesize the desired compounds (Scheme 1) in good yield (Table 1).
A singlet signal comparable to one CONH proton at around δ 10 ppm was observed in the proton NMR spectra of the synthesized molecules verifying the existence of ‘benzamide link’ in synthesized compounds. The detection of singlet signal for one ‘NH proton’ of SO2NH functional moiety was recorded ~2.5 ppm, demonstrating the development of sulphonamide analogues by an interaction of sulphonyl chloride compounds with the respective amines. The presence of a singlet, two doublets and a triplet signal about δ 8 ppm referring to 1H-atoms at C2, C4, C5 and C6; correspondingly of the benzoic acid-derived phenyl moiety verified that the ‘benzamide link’ and the ‘sulphonamide linkage’ were meta to one another i.e., alienated by C2. Two doublets were recorded in the proton-NMR spectra of prepared compounds in 7–8 ppm range, apart from 2 ‘triplet signals’ pointing to four 1H-atoms (corresponding to aromatic CH), demonstrating that 2-aminobenzimidazole was combined with benzoyl chloride for development of benzamide derivatives. Occurrence of singlet signal at δ 5 ppm corresponding to NH of benzimidazol-2-yl further confirmed presence of benzimidazol-2-yl scaffold in synthesized compounds. In 13C-NMR pattern of the prepared molecules signals around δ 170 ppm indicated the existence of C=N bond in these derivatives and signals about δ 165 ppm illustrated occurrence of amide C=O bond therefore supporting development of benzamide linkage in produced derivatives. The FTIR spectrum of manufactured N-benzimidazol-2-yl substituted benzamide analogues demonstrated the occurrence of characteristics ‘stretching frequencies’ at 3300–3200, above 3000, 1400-1300/1200-1100 (asymmetric) and 3400–3100 cm−1 which correspond to amide NH, aromatic CH, SO2, sulphonamide NH functional groups respectively, therefore supporting the linkage of amide (CONH) and sulphonamide (SO2–NH) groups in developed molecules. Furthermore, the presence of carbonyl C=O stretching (1700–1600 cm−1) and NH-bending (1600 cm−1) vibrations in spectrum of molecules signified the occurrence of amide carbonyl and aromatic NH-functional group (Fig. 2).
Prediction of ADME properties
ADME parameters such as ‘molecular weight’ (MW), ‘partition coefficient’ (log P), distribution coefficient (log D), water solubility (log Sw), ‘topological polar surface area’ (tPSA), ‘H-bond acceptors’ (HBA), ‘H-bond donors’ (HBD), solubility (mg/L) and ‘number of rotatable bonds’ (NRB) were predicted for all the designed analogues. All of the designed analogues showed good pharmacokinetic parameters for oral bioavailability (Table 2) and drug-like properties as described using “Lipinski’s rule of five” (i.e., MW < 500 Da; log P < 5; HBA ≤ 10 and HBD: ≤ 5). All the designed molecules showed MW (330–472 Da), log P value (1.70–3.87), HBA (4–6) and HBD (3) within the range of ideal orally bioavailable drug candidate.
In vitro GK assay
The GK catalytic effect of the newly prepared compounds was assessed spectrometrically by calculating the absorbance around 340 nm through coupled interaction with G6PDH. Amongst the tested derivatives, two molecules (compounds 2 and 7) demonstrated peak GK activation (fold activation exceeding 2.0 in contrasts to control). Other compounds demonstrated modest activation (1.4 to 1.8-fold compared to control) of GK enzyme (Fig. 2).
Amongst the synthesized analogues, compounds bearing N-(2-chloro-4-nitrophenyl) sulphonamide moiety (2) and N-(4-bromophenyl) sulphonamide moiety (7) exhibited highest GK activity (GK activation fold of 2.11 and 2.07, respectively compared to control). Analogues having N-phenyl, N-2-methylphenyl and N-2- nitrophenyl sulphonamide moiety (1, 6 and 9) demonstrated 1.82, 1.81 and 1.78-fold GK activation, correspondingly. Synthesized compounds having N-4-methylphenyl and N-methyl sulphonamide moiety (8 and 10, respectively) exhibited moderate GK activity (1.67 and 1.64-fold GK activation, respectively). Analogues bearing N-benzyl and N-butyl sulphonamide moiety (3 and 4, respectively) exhibited ~1.5 times activation in comparison to control. Derivative having N-methyl sulphonamide moiety (5) showed lowest GK fold activation. Outcomes of the GK test demonstrated that replacement of substituted phenyl moiety substituted to SO2NH resulted in improved GK activity compared to those having alkyl group as can be observed from the GK activation potential of compounds 2, and 7. Substitution of aromatic moiety attached to SO2NH with alkyl chains have decreased the capacity for GK activation compared to compounds 4, 5 and 10 with aromatic moiety as shown by GK activity.
In silico docking investigations
In silico molecular docking investigations were performed to discover the connection and linking behaviors of developed molecules using AutoDock Vina in the allosteric location of GK (PDB ID: 3IMX). The reference GK activator (PDB entry: 3IMX) developed a similar linking sequence and transposed on bonding fashion of co-crystallized GK activator with ΔG of −9.0 kcal/mol validating accuracy of the docking methodology used. Most of the docked ligands showed appreciable linking in allosteric location of GK as established by examining their linkage interactions and ΔG of the paramount docked facades (Table 3). These compounds displayed strong H-bond interactions between ‘–NH’ of benzamide and ‘amide carbonyl’ of Arg63; and ‘N’ of the benzimidazol-2-yl ring and ‘amide –NH’ of Arg63 in GK protein’s allosteric position.
Super-positioning of the docking conformations of analogues 1, 2, 6 and 7, on that of reference ligand in the allosteric site of GK demonstrated that these molecules had the analogous binding and orientation arrangement in the allosteric site of GK enzyme as that of the x-ray crystallized effector (PDB entry: 3IMX) supporting the outcomes of in vitro GK test for these compounds (Fig. 3).
The docked conformations of analogues 1, 2, 6 and 7 demonstrated strong hydrogen bond interactions between ‘N’ of benzimidazol-2-yl group and amide –NH of Arg63 residues; and ‘benzamide –NH’ group and ‘backbone carbonyl’ of Arg63 in the allosteric location of GK with bond length in the range 2.9–3.1 Å; and 2.9–3.0 Å, respectively. Overall, the benzimidazol-2-yl moiety bonded to the benzamide ‘NH’ of these compounds protruded in the hydrophobic cavity displaying connections with the Val455 and Lys459 amino acid residues, along with Pro66 and Ile159 amino acid residues, aromatic moiety parceled in the cavity composed of Met210, Tyr214 and Val455 residues (Fig. 4) (Table 3).
In silico prediction of toxicity
The possible toxicity (mutagenic, cardiotoxicity, acute toxicity, hepatotoxicity, skin irritation, and chronic toxicity) for the optimized compounds was accessed using pkCSM online platform. Conferring to the results represented in Table 4; some of the compounds showed little toxicity probability. Mutagenicity was predicted for all the synthesized compounds. Cardiotoxicity (inhibition of hERG-I and hERG-II) was predicted for compounds 1 and 3. Hepatotoxicity was predicted for compounds 1, 3 and 8. In this perspective, the initial evaluation performed in silico using pkCMS online platform, can supplement forthcoming studies related to the safety of these compounds.
Conclusions
In summary, a series of novel N-benzimidazol-2-yl benzamide analogues were designed and prepared using structure-based drug design approach. Among these newly identified derivatives, analogues 2 and 7 unveiled maximum GK activation potential (>2.0-fold increase in GK catalytic activity in vitro). Outcomes of the in-vitro test were found to be analogous to the in-silico docking investigations with the GK enzyme. These analogues followed the ‘Lipinski’s rule of five’ for the “drug-like” characteristics. These newly developed compounds might assist in finding the lead analogues for discovery of strong and harmless activators of GK for T2D handling and management.
References
Kohei K. Pathophysiology of type 2 diabetes and its treatment policy. Jpn Med Assoc J. 2010;53:41–46.
Olokoba AB, Obateru OA, Olokoba LB. Type 2 diabetes mellitus: a review of current trends. Oman Med J. 2012;27:269–273. https://doi.org/10.5001/omj.2012.68
Beagley J, Guariguata L, Weil C, Motala AA. Global estimates of undiagnosed diabetes in adults. Diabetes Res Clin Pr. 2014;103:150–160. https://doi.org/10.1016/j.diabres.2013.11.001
Pal M. Recent advances in glucokinase activators for the treatment of type 2 diabetes. Drug Disco Today. 2009;14:784–792. https://doi.org/10.1016/j.drudis.2009.05.013
Grewal AS, Beniwal M, Pandita D, Sekhon BS, Lather M. Recent updates on peroxisome proliferator-activated receptor δ agonists for the treatment of metabolic syndrome. Med Chem. 2016;12:03–21. https://doi.org/10.2174/1573406411666150525105826
Grewal AS, Sekhon BS, Lather V. Recent updates on glucokinase activators for the treatment of type 2 diabetes mellitus. Mini Rev Med Chem. 2014;14:585–602. https://doi.org/10.2174/1389557514666140722082713
Grewal AS, Bhardwaj S, Pandita D, Lather V, Sekhon BS. Updates on aldose reductase inhibitors for management of diabetic complications and non-diabetic diseases. Mini Rev Med Chem. 2016;16:120–162. https://doi.org/10.2174/1389557515666150909143737
Pal M. Medicinal chemistry approaches for glucokinase activation to treat type 2 diabetes. Curr Med Chem. 2009;16:3858–3874. https://doi.org/10.2174/092986709789177993
Coghlan M, Leighton B. Glucokinase activators in diabetes management. Expert Opin Investig Drugs. 2008;17:145–167. https://doi.org/10.1517/13543784.17.2.145
Perseghin G. Exploring the in vivo mechanisms of action of glucokinase activators in type 2 diabetes. J Clin Endocrinol Metab. 2010;95:4871–4873. https://doi.org/10.1210/jc.2010-2049
Matschinsky FM, Zelent B, Doliba N, Li C, Vanderkooi JM, Naji A, et al. Glucokinase activators for diabetes therapy. Diabetes Care. 2011;34:S236–43. https://doi.org/10.2337/dc11-s236
Grewal AS, Lather V, Charaya N, Sharma N, Singh S, Kairys V. Recent developments in medicinal chemistry of allosteric activators of human glucokinase for type 2 diabetes mellitus therapeutics. Curr Pharm Des. 2020;26:2510–2552. https://doi.org/10.2174/1381612826666200414163148
Zhang L, Li H, Zhu Q, Liu J, Chen L, Leng Y, et al. Benzamide derivatives as dual-action hypoglycemic agents that inhibit glycogen phosphorylase and activate glucokinase. Bioorg Med Chem. 2009;17:7301–7312. https://doi.org/10.1016/j.bmc.2009.08.045
Pike KG, Allen JV, Caulkett PWR, Clarke DS, Donald CS, Fenwick ML, et al. Design of a potent, soluble glucokinase activator with increased pharmacokinetic half-life. Bioorg Med Chem Lett. 2011;21:3467–3470. https://doi.org/10.1016/j.bmcl.2011.03.093
Ericsson H, Sjoberg F, Heijer M, Dorani H, Johansson P, Wollbratt M, et al. The glucokinase activator AZD6370 decreases fasting and postprandial glucose in type 2 diabetes mellitus patients with effects influenced by dosing regimen and food. Diabetes Res Clin Pr. 2012;98:436–444. https://doi.org/10.1016/j.diabres.2012.09.025
Bowler JM, Hervert KL, Kearley ML, Miller BG. Small-molecule allosteric activation of human glucokinase in the absence of glucose. ACS Med Chem Lett. 2013;4:580–584. https://doi.org/10.1021/ml400061x
Sjostrand M, Ericsson H, Hartford M, Norjavaara E, Eriksson JW. Pharmacodynamic effects of the oral glucokinase activator AZD6370 after single doses in healthy volunteers assessed with euglycaemic clamp. Diabetes Obes Metab. 2013;15:35–41. https://doi.org/10.1111/j.1463-1326.2012.01672.x
Park K, Lee BM, Kim YH, Han T, Yi W, Lee DH, et al. Discovery of a novel phenylethyl benzamide glucokinase activator for the treatment of type 2 diabetes mellitus. Bioorg Med Chem Lett. 2013;23:537–542. https://doi.org/10.1016/j.bmcl.2012.11.018
Park K, Lee BM, Hyun KH, Lee DH, Choi HH, Kim H. et al. Discovery of 3-(4-methanesulfonylphenoxy)-N-[1-(2-methoxy-ethoxymethyl)-1H-pyrazol-3-yl]-5-(3-methylpyridin-2-yl)-benzamide as a novel glucokinase activator (GKA) for the treatment of type 2 diabetes mellitus. Bioorg Med Chem. 2014;22:2280–93. https://doi.org/10.1016/j.bmc.2014.02.009
Lei L, Liu Q, Liu S, Huan Y, Sun S, Chen Z, et al. Antidiabetic potential of a novel dual-target activator of glucokinase and peroxisome proliferator activated receptor-γ. Metab Clin Exp. 2015;64:1250–1261. https://doi.org/10.1016/j.metabol.2015.06.014
Wang Z, Shi X, Zhang H, Yu L, Cheng Y, Zhang H, et al. Discovery of cycloalkyl-fused N-thiazol-2-yl-benzamides as tissue non-specific glucokinase activators: design, synthesis, and biological evaluation. Eur J Med Chem. 2017;139:128–152. https://doi.org/10.1016/j.ejmech.2017.07.051
Charaya N, Pandita D, Grewal AS, Lather V. Design, synthesis and biological evaluation of novel thiazol-2-yl benzamide derivatives as glucokinase activators. Comput Biol Chem. 2018;73:221–229. https://doi.org/10.1016/j.compbiolchem.2018.02.018
McKerrecher D, Steven A. Design and development of the glucokinase activator AZD1656. In: Abdel-Magid AF, Pesti JA, Vaidyanathan R, (eds.) Complete Accounts of Integrated Drug Discovery and Development: Recent Examples from the Pharmaceutical Industry. Washington: American Chemical Society; 2018. https://doi.org/10.1021/bk-2018-1307.ch007
Grewal AS, Kharb R, Prasad DN, Dua JS, Lather V. N-Pyridin-2-yl benzamide analogues as allosteric activators of glucokinase: design, synthesis, in vitro, in silico and in vivo evaluation. Chem Biol Drug Des. 2019;93:364–372. https://doi.org/10.1111/cbdd.13423
Grewal AS, Dua JS, Prasad DN, Kharb R, Lather V. Design, synthesis and evaluation of novel 3,5-disubstituted benzamide derivatives as allosteric glucokinase activators. BMC Chem. 2019;13:2 https://doi.org/10.1186/s13065-019-0532-8
Grewal AS, Kharb R, Dua JS, Lather V. Molecular docking assessment of N-heteroaryl substituted benzamide derivatives as glucokinase activators. Asian J Pharm Pharm. 2019;5:129–136. https://doi.org/10.31024/ajpp.2019.5.1.18
Grewal AS, Arora S, Sharma N, Singh S. In silico docking studies of compounds from Persian shallot as allosteric glucokinase activators. Plant Arch. 2020;20:3768–3771.
Singh R, Lather V, Pandita D, Judge V, Arumugam KN, Grewal AS. Synthesis, docking and antidiabetic activity of some newer benzamide derivatives as potential glucokinase activators. Lett Drug Des Discov. 2017;14:540–553. https://doi.org/10.2174/1570180813666160819125342
Grewal AS, Lather V, Pandita D, Bhayana G. Synthesis, docking and evaluation of phenylacetic acid and trifluoromethylphenyl substituted benzamide derivatives as potential PPARδ agonists. Lett Drug Des Discov. 2017;14:1239–1251. https://doi.org/10.2174/1570180814666170327164443
Chauhan A, Grewal AS, Pandita D, Lather V. Novel cinnamic acid derivatives as potential PPARδ agonists for metabolic syndrome: design, synthesis, evaluation and docking studies. Curr Drug Disco Technol. 2020;17:338–347. https://doi.org/10.2174/1570163816666190314124543
Miteva M, Violas S, Montes M, Gomez D, Tuffery P, Villoutreix B. FAF-Drugs: free ADME/Tox filtering of compound collections. Nucl Acids Res. 2006;34:W738–44. https://doi.org/10.1093/nar/gkl065
Lagorce D, Bouslama L, Becot J, Miteva MA, Villoutreix BO. FAF-Drugs4: Free ADME-Tox filtering computations for chemical biology and early stages drug discovery. Bioinformatics. 2017;33:3658–3660. https://doi.org/10.1093/bioinformatics/btx491
Efanov AM, Barrett DG, Brenner MB, Briggs SL, Delaunois A, Durbin JD, et al. A novel glucokinase activator modulates pancreatic islet and hepatocyte function. Endocrinology. 2005;146:3696–3701. https://doi.org/10.1210/en.2005-0377
Futamura M, Hosaka H, Kadotani A, Shimazaki H, Sasaki K, Ohyama S, et al. An allosteric activator of glucokinase impairs the interaction of glucokinase and glucokinase regulatory protein and regulates glucose metabolism. J Biol Chem. 2006;281:37668–37674. https://doi.org/10.1074/jbc.M605186200
Trott O, Olson AJ. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J Comput Chem. 2010;31:455–461. https://doi.org/10.1002/jcc.21334
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexiblity. J Comput Chem. 2009;30:2785–2791. https://doi.org/10.1002/jcc.21256
Miteva M, Guyon F, Tufféry P. Frog2: Efficient 3D conformation ensemble generator for small compounds. Nucl Acids Res. 38:W622-7. https://doi.org/10.1093/nar/gkq325
Rathee D, Grewal AS, Dureja H, Lather V. Enzymatic inhibitory activity of iridoid glycosides from Picrorrhiza kurroa against matrix metalloproteinases: correlating in vitro targeted screening and docking. Comput Biol Chem. 2019;78:28–36. https://doi.org/10.1016/j.compbiolchem.2018.10.017
Pires DE, Blundell TL, Ascher DB. pkCSM: Predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J Med Chem. 2015;58:4066–4072. https://doi.org/10.1021/acs.jmedchem.5b00104
Salgueiro A, Folmer V, da Rosa H, Costa M, Boligon A, Paula F, et al. In vitro and in silico antioxidant and toxicological activities of achyrocline satureioides. J Ethnopharmacol. 2016;194:6–14. https://doi.org/10.1016/j.jep.2016.08.048
Pires DE, Kaminskas LM, Ascher DB. Prediction and optimization of pharmacokinetic and toxicity properties of the ligand. Methods Mol Biol. 2018;1762:271–284. https://doi.org/10.1007/978-1-4939-7756-7_14
Acknowledgements
The authors are thankful to Chitkara College of Pharmacy, Chitkara University, Punjab for their support and encouragement for this research work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Singh, S., Arora, S., Dhalio, E. et al. Design and synthesis of newer N-benzimidazol-2yl benzamide analogues as allosteric activators of human glucokinase. Med Chem Res 30, 760–770 (2021). https://doi.org/10.1007/s00044-020-02697-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-020-02697-z