Abstract
In order to develop potent histone deacetylase inhibitors, a virtual screening approach was performed to discover novel lead structures. A commercial database containing about 167,000 molecules was in silico filtered by rule of five, zinc-binding groups, pharmacophore models, and binding pattern analysis. At last, three molecules were selected for enzyme inhibition assay, and one compound 02 has IC50 of 1.6 μM against histone deacetylase 8 (HDAC8).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
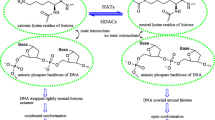
Histone deacetylases (HDACs) are responsible for deacetylating the N-terminal tails of histones, and the corresponding chromatin alterations regulate transcription and other nuclear events (Giandomenico et al., 2003; Gronroos et al., 2002; Jin et al., 2004; Minucci and Pelicci, 2006). Overexpression and aberrant recruitment of HDACs have a significant role in tumorigenesis. So far, 18 members of the HDAC family have been discovered and subdivided into 4 classes (De Ruijter et al., 2003; Gregoretti et al., 2004). Among these classes, class I HDACs (HDAC1, 2, 3, and 8) were proved to be the most correlative to the pathogenesis of cancer (Bernstein et al., 2000; Foglietti et al., 2006; Lagger et al., 2002; Sun and Hampsey, 1999).
HDAC inhibitors (HDACis) have the effects of apoptosis, cell cycle arrest, and differentiation (Insinga et al., 2005; Johnstone, 2002; Marks et al., 2001; Nebbioso et al., 2005). Suberoyl anilide hydroxamic acid (SAHA) (Richon et al., 1998) is the first HDACi approved by FDA for the treatment of cancer, and many promising HDACis are still under investigation in various stages of clinical trials (Zhang et al., 2008).
Virtual screening is a powerful method for discovering novel HDACis (Liu et al., 2010; Tang et al., 2009; Thangapandian et al., 2010). It is well known that the zinc-binding groups (ZBGs) in HDACis make significant contributions to the binding profile of HDACis around active site. In this article, a ZBG-based virtual screening was performed for searching novel HDAC8 inhibitor scaffold. The harvested candidates with reasonable physico-chemical properties were chosen for the further biological assay.
Materials and methods
Preparation of compound database
The SPECS chemical database (April 2009, downloaded from http://www.specs.net) was used in the current virtual screening process. All molecules in the database were converted to 3D conformations in UNITY database by CONCORD module in the SYBYL 7.3 software package. This database was filtered by “rule of five” (Lipinski et al., 2001), in which molecules having >5 number of H-bond donors, >10 of H-bond acceptors, >500 molecular weight, and >5 calculated LogP were excluded.
ZBG-based virtual screening
Hydrosulfide group, hydrazide group, and several other groups were defined as ZBG. Subsequently, the druglike database was screened by the defined ZBG criteria. As a result, the database was subdivided into several small databases. Obviously, these different sub-databases did not contain the same number of molecules. If there are more than 1,000 molecules in a sub-database, then this database will be screened by a pharmacophore model featured by a negative charge center and a hydrophobic point (Fig. 1a). If the number of molecules is still more than 1,000, then the refiltered database will be screened by a more restrict pharmacophore model (Fig. 1b) which contains three features (a negative charge center, a hydrophobic point, and an H-bond acceptor).
Docking study
The FlexX2 program (Rarey et al., 1997; 1996) in the SYBYL 7.0 package was used for the docking analysis. The holo-form crystal structure of HDAC8 (PDB entry: 1T64) was used as the receptor (Somoza et al., 2004), the derived sub-databases were docked into the active site of HDAC8, respectively. 30 docking patterns were generated for each molecule, and CSCORE was applied to evaluate their binding qualities. The ligand TSN (Fig. 2) in the crystal structure of HDAC8 functioning as a positive control was also docked to the active site. The scored molecules of each database were analyzed by the manual examination compared to TSN.
Activity assay
The method for in vitro activity assay has been described in our previous study (Zhang et al., 2010). In brief, the HDAC8 enzyme was expressed in Escherichia coli. Boc-Lys (acetyl)-AMC was used as the substrate of HDAC8. Vorinostat, the HDACi on market, was employed as a positive control. The compounds were serial diluted into six concentrations (100, 20, 4, 0.8, 0.16, and 0.032 μM/l) to probe their ability of inhibiting HDAC8 activity.
Result and discussion
Virtual screening
The number of molecules in the database was reduced from 167,475 to 157,822 by Lipinski’s rule of five. Afterward, the refined database was filtered by the defined ZBGs. There are 5,227 molecules containing carboxylic acid moiety, 2433 molecules having amide group, and 1864 molecules possessing carbamido substructure, respectively (Table 1). To save computational time, sub-databases containing these three groups were refiltered by a two-feature pharmacophore model. After this filtration, there are 39 molecules having amide group, 63 molecules having carbamido group, and 3,812 molecules containing carboxylic acid group, individually. Consequently, the 3,812 carboxylic acid group containing molecules were filtered by a three-feature pharmacophore, and only 411 molecules were survived for the docking study.
Docking
Totally 1,432 molecules were in silico screened by ZBG- and pharmacophore-based screening, following by docking analysis (Table 1). The ligand TSN in the active site of HDAC8 was first docked to the crystal structure. The docked conformation sharing very low RMSD value (1.72 Å) with the crystal structure of TSN suggested that this molecular docking method was reliable (Fig. 3).
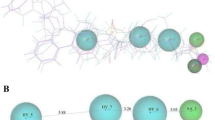
Only two molecules containing carboxylic acid groups had a higher score than the ligand TSN (Table 1). The top 100 scored molecules in the carboxylic acid group and hydrosulfide group databases were analyzed by visual examination. Our attention focused on three compounds 01 (database ID: AI/204-31755055), 02 (database ID: AK/968-41922428), and 03 (database ID: AA/516-31409020) (Fig. 2), although many molecules had good binding patterns in the active site of HDAC8. Figure 4 shows the location of these three molecules around the binding site. The carbonyl groups of 01 (Fig. 4a), the amino group and carbonyl group of 02 (Fig. 4b), and carboxyl group of 03 (Fig. 4c) stretched inside the channel to bind to the zinc ion. Multiple H-bond interactions can be found between the molecules and amino acid lined in the active site of HDAC8 (Fig. 5). Molecule 01 has H-bond interactions with residues Gly151 and Gly304 (Fig. 5a), in the meanwhile the amino group of molecule 02 functioning as an H-bond donor binds to His142 and His143 (Fig. 5b). The carbonyl group could contact Tyr306 as an H-bond acceptor. Moreover, molecule 03 has H-bond interactions with five residues (Fig. 5c). They are Asp101, His142, His143, Gly151, and Tyr306, respectively.
Activity assay
Three candidates were evaluated for their ability to inhibit HDAC8. Accordingly, only 02 showed potent inhibitory activity against HDAC8 with IC50 of 1.6 μM in comparison to Vorinostat (1.48 μM) (Fig. 6). This evidence demonstrates that the hydrazide group is one of the potent ZBGs. One case in point is molecule 02, which would be a lead compound in further chemical modification and optimization.
Conclusion
The current report disclosed a virtual screening approach for discovering novel HDACis. Various filter rules such as ZBGs, pharmacophore models, and docking methods were applied to minimize the candidate pool. The activities of four molecules were determined by in vitro enzyme inhibition assay. One molecule, 02, containing hydrazide group showed potent inhibitory activity against HDAC8. Our further study will be concentrated on chemical modification of this molecule.
References
Bernstein BE, Tong JK, Schreiber SL (2000) Genomewide studies of histone deacetylase function in yeast. Proc Natl Acad Sci USA 97:13708–13713
De Ruijter AJM, Van Gennip AH, Caron HN, Kemp S, Van Kuilenburg ABP (2003) Histone deacetylases (HDACs): characterization of the classical HDAC family. Biochem J 370:737–749
Foglietti C, Filocamo G, Cundari E, De Rinaldis E, Lahm A, Cortese R, Steinkuhler C (2006) Dissecting the biological functions of Drosophila histone deacetylases by RNA interference and transcriptional profiling. J Biol Chem 281:17968–17976
Giandomenico V, Simonsson M, Gronroos E, Ericsson J (2003) Coactivator-dependent acetylation stabilizes members of the SREBP family of transcription factors. Mol Cell Biol 23:2587–2599
Gregoretti IV, Lee YM, Goodson HV (2004) Molecular evolution of the histone deacetylase family: functional implications of phylogenetic analysis. J Mol Biol 338:17–31
Gronroos E, Hellman U, Heldin CH, Ericsson J (2002) Control of Smad7 stability by competition between acetylation and ubiquitination. Mol Cell 10:483–493
Insinga A, Monestiroli S, Ronzoni S, Gelmetti V, Marchesi F, Viale A, Altucci L, Nervi C, Minucci S, Pelicci PG (2005) Inhibitors of histone deacetylases induce tumor-selective apoptosis through activation of the death receptor pathway. Nat Med 11:71–76
Jin YH, Jeon EJ, Li QL, Lee YH, Choi JK, Kim WJ, Lee KY, Bae SC (2004) Transforming growth factor-beta stimulates p300-dependent RUNX3 acetylation, which inhibits ubiquitination-mediated degradation. J Biol Chem 279:29409–29417
Johnstone RW (2002) Histone-deacetylase inhibitors: novel drugs for the treatment of cancer. Nat Rev Drug Discov 1:287–299
Lagger G, O’Carroll D, Rembold M, Khier H, Tischler J, Weitzer G, Schuettengruber B, Hauser C, Brunmeir R, Jenuwein T, Seiser C (2002) Essential function of histone deacetylase 1 in proliferation control and CDK inhibitor repression. EMBO J 21:2672–2681
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46:3–26
Liu XH, Song HY, Zhang JX, Han BC, Wei XN, Ma XH, Cui WK, Chen YZ (2010) Identifying novel type ZBGs and nonhydroxamate HDAC inhibitors through a SVM-based virtual screening approach. Mol Inform 29:407–420
Marks PA, Rifkind RA, Richon VM, Breslow R, Miller T, Kelly WK (2001) Histone deacetylases and cancer: causes and therapies. Nat Rev Cancer 1:194–202
Minucci S, Pelicci PG (2006) Histone deacetylase inhibitors and the promise of epigenetic (and more) treatments for cancer. Nat Rev Cancer 6:38–51
Nebbioso A, Clarke N, Voltz E, Germain E, Ambrosino C, Bontempo P, Alvarez R, Schiavone EM, Ferrara F, Bresciani F et al (2005) Tumor-selective action of HDAC inhibitors involves TRAIL induction in acute myeloid leukemia cells. Nat Med 11:77–84
Rarey M, Kramer B, Lengauer T, Klebe G (1996) A fast flexible docking method using an incremental construction algorithm. J Mol Biol 261:470–489
Rarey M, Kramer B, Lengauer T (1997) Multiple automatic base selection: protein-ligand docking based on incremental construction without manual intervention. J Comput Aided Mol Des 11:369–384
Richon VM, Emiliani S, Verdin E, Webb Y, Breslow R, Rifkind RA, Marks PA (1998) A class of hybrid polar inducers of transformed cell differentiation inhibits histone deacetylases. Proc Natl Acad Sci USA 95:3003–3007
Somoza JR, Skene RJ, Katz BA, Mol C, Ho JD, Jennings AJ, Luong C, Arvai A, Buggy JJ, Chi E et al (2004) Structural snapshots of human HDAC8 provide insights into the class I histone deacetylases. Structure 12:1325–1334
Sun ZW, Hampsey M (1999) A general requirement for the Sin3-Rpd3 histone deacetylase complex in regulating silencing in Saccharomyces cerevisiae. Genetics 152:921–932
Tang H, Wang XS, Huang XP, Roth BL, Butler KV, Kozikowski AP, Jung M, Tropsha A (2009) Novel inhibitors of human histone deacetylase (HDAC) identified by QSAR modeling of known inhibitors, virtual screening, and experimental validation. J Chem Inf Model 49:461–476
Thangapandian S, John S, Sakkiah S, Lee KW (2010) Ligand and structure based pharmacophore modeling to facilitate novel histone deacetylase 8 inhibitor design. Eur J Med Chem 45:4409–4417
Zhang YJ, Fang H, Jiao J, Xu WF (2008) The structure and function of histone deacetylases: the target for anti-cancer therapy. Curr Med Chem 15:2840–2849
Zhang YJ, Feng JH, Liu CX, Zhang L, Jiao J, Fang H, Su L, Zhang XP, Zhang J, Li MY et al (2010) Design, synthesis and preliminary activity assay of 1,2,3,4-tetrahydroisoquinoline-3-carboxylic acid derivatives as novel Histone deacetylases (HDACs) inhibitors. Bioorg Med Chem 18:1761–1772
Acknowledgments
This study was supported by National Nature Science Foundation of China (Grant Nos. 30772654 and 36072541), Shandong Natural Science Foundation (No. JQ201019), Independent Innovation Foundation of Shandong University, IIFSDU (No. 2010JQ005), the Ph.D. Programs Foundation of Ministry of Education of PR China (No. 20060422029), and National High Technology Research and Development Program of China (Grant Nos. 2007AA02Z314 and 2009ZX09103-118).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, L., Li, M., Feng, J. et al. Discovery of a novel histone deacetylase 8 inhibitor by virtual screening. Med Chem Res 21, 152–156 (2012). https://doi.org/10.1007/s00044-010-9519-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-010-9519-7