Abstract
Modification of the cancer-associated chromatin landscape in response to therapeutic DNA damage influences gene expression and contributes to cell fate. The central histone mark H2Bub1 results from addition of a single ubiquitin on lysine 120 of histone H2B and is an important regulator of gene expression. Following treatment with a platinum-based chemotherapeutic, there is a reduction in global levels of H2Bub1 accompanied by an increase in levels of the tumor suppressor p53. Although total H2Bub1 decreases following DNA damage, H2Bub1 is enriched downstream of transcription start sites of specific genes. Gene-specific H2Bub1 enrichment was observed at a defined group of genes that clustered into cancer-related pathways and correlated with increased gene expression. H2Bub1-enriched genes encompassed fifteen p53 target genes including PPM1D, BTG2, PLK2, MDM2, CDKN1A and BBC3, genes related to ERK/MAPK signalling, those participating in nucleotide excision repair including XPC, and genes involved in the immune response and platinum drug resistance including POLH. Enrichment of H2Bub1 at key cancer-related genes may function to regulate gene expression and influence the cellular response to therapeutic DNA damage.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chromatin remodelling after DNA damage influences gene transcription and may ultimately determine cell fate as part of the DNA damage response (DDR). Knowledge of the DDR is essential to understanding the way malignant cells respond to DNA damage induced by chemotherapeutic agents that aim to disable cell survival pathways and promote apoptosis. Innate or acquired resistance to many of the standard-of-care chemotherapeutic agents, including platinum-drugs such as carboplatin, needs to be overcome to maximise the full potential of these drugs for the treatment of cancer patients [1, 2]. Maintenance of structure and function of the epigenome is important for the ability of cells to survive DNA damage and changes to chromatin marks have been linked to chemoresistance [3,4,5,6,7,8,9,10,11,12,13,14].
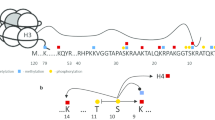
Ubiquitin is emerging as a protein with key roles in modelling the cancer epigenome through post-translational modification (PTM) of specific histone lysines [15, 16]. Histones are the most abundantly monoubiquitinated protein conjugates in the nucleus of mammalian cells [17]. At 8.5 kDa, the addition of a single ubiquitin to a histone lysine residue is one of the largest histone PTMs. Monoubiquitination of histone H2B at lysine 120, H2Bub1, promotes an open chromatin configuration, as does monoubiquitination on histone H4 at position 31 (H4ub31) [18]. H2Bub1 physically separates the chromatin strands and promotes histone eviction from the nucleosome, creating a chromatin structure that is more accessible to transcription factors and DNA repair proteins [19]. Beyond the steric effect of the bulky ubiquitin molecule, it is apparent that both the nucleosomal position and properties inherent to this specific histone PTM are important for H2Bub1 function [20]. At a different position in the nucleosome, monoubiquitination of histone H2A at lysine 119 by ring finger ligases in polycomb repressor complex 1 (PRC1) leads to the opposite effect, resulting in compacted, transcriptionally silenced chromatin [21].
H2Bub1 recruits other chromatin remodelling factors, highlighting it as a central regulator of histone cross-talk [22,23,24,25,26,27]. These factors include subunits of the SWI/SNF chromatin remodelling complex (BRG1, BRM, BAF155 and BAF170) [28], the WDR82 subunit of the SET1A/B COMPASS complex required for di- and tri-methylation of histone H3 lysine 4 [29,30,31,32] and the non-SET domain DOTL1 methyltransferase that methylates histone H3 lysine 79 [24, 33]. In response to stimuli such as DNA damage, cyclin-dependent kinase 9 (CDK9) phosphorylates the E2 ubiquitin-conjugating enzyme UBE2A and the Ser2 residue in the carboxy-terminal domain (CTD) of RNA polymerase II (Pol II). This creates a binding pocket for the linker protein WAC that interacts with Pol II and the E3 ubiquitin ligase heterodimeric complex RNF20-RNF40. The E3 ubiquitin ligase RNF20-RNF40 complex both monoubiquitinates histone H2B lysine 120 (H2Bub1), and interacts with the human PAF1 transcriptional complex that in turn associates with RNA Pol II to promote transcriptional elongation (reviewed in [15]). We have previously shown the importance of RNF20 and RNF40 in maintaining global levels of H2Bub1 [34]. The presence of chromatin-associated H2Bub1 has been linked with genes that are highly expressed [35], although this correlation is not absolute [36].
Here, we have used wild-type (wt) p53 cancer cell line models to map the localisation of H2Bub1-enrichment in response to DNA damage with a platinum-based chemotherapeutic agent. We have correlated H2Bub1 enrichment with gene expression levels in response to treatment with cisplatin, thereby identifying genes and pathways involved in the determination of H2Bub1-related cell fate following DNA damage.
Materials and methods
Cell culture
A2780, MCF7 and HEY1 cells were used as models of wt p53 cell lines. MCF7 is a breast cancer cell line [37] and HEY1 and A2780 are of ovarian cancer origin [38, 39]. The high-grade serous ovarian cancer (HGSOC) cell lines OVCAR-3 (p.Arg248Gln) and Kuramochi (p.Asp281Tyr) were used as models of mutant p53 [38]. The ovarian and non-small cell lung cancer p53 null cell lines SKOV3 [40] and H1299 [41], respectively c.267del1 (p.S90fsX33) and p53 gene deletion c.(del), were also employed. A2780 was sourced from Sigma-Aldrich Pty. Ltd. (Sydney, NSW, Australia); MCF7, OVCAR-3, SKOV3 and H1299 were from the American Type Culture Collection (ATCC; Manassas, VA, USA) and Kuramochi was sourced from the Japanese Collection of Research Bioresources (JCRB) Cell Bank (Ibaraki city, Osaka, Japan). All cell lines were cultured at 37 °C in RPMI 1640 medium (Invitrogen, Carlsbad, CA, USA) supplemented with 10% foetal bovine serum (SAFC Bioscience, Brooklyn, VIC, Australia) in a humidified incubator containing 5% CO2 (Thermo, Waltham, MA, USA). Media for MCF7 was additionally supplemented with 1% insulin (Thermo Fisher Scientific Australia Pty. Ltd, Scoresby, VIC, Australia). Cell line authentication was performed by CellBank Australia (Westmead, NSW, Australia). Cisplatin (#P4394), nutlin-3a (#SML0580) and 5,6-Dichlorobenzimidazole 1-β-D-ribofuranoside (DRB, #D1916) were purchased from Sigma-Aldrich, Sydney, NSW, Australia.
Cloning and mutagenesis
Full length wt TP53 sequence was PCR amplified from HEK293 cells, cloned into the HindIII/XhoI sites of the mammalian expression vector pcDNA4/TO (Life Technologies, Mulgrave, VIC, Australia) and verified by sequence analysis (Australian Genome Research Facility, Westmead, NSW, Australia). Plasmid DNA was propagated in JM109 Escherichia coli cells and purified using the Wizard® Plus SV Miniprep DNA Purification System (Promega, Alexandria, NSW, Australia) or Qiagen Plasmid plus Midi kit (Qiagen, Chadstone, VIC, Australia). The gain of function TP53 mutation identified in OVCAR-3 cells, p.Arg248Gln, was engineered using forward and reverse primers (mutated nucleotide is bolded and underlined with original sequence noted in brackets); for: 5′-CCTGCATGGGCGGCATGAACCA(G → A)GAGGCCCATCCTCACCATCATC-3′, and rev: 5′-GATGATGGTGAGGATGGGCCTCT(C → T)GGTTCATGCCGCCCATGCAGG-3′. Plasmid DNA was transfected into cells using X-tremeGENE™ 9 following the manufacturer’s protocols (Sigma-Aldrich, Sydney, NSW, Australia).
Western blotting
Cells were washed with cold PBS, and lysed using urea buffer (20 mM NaH2PO4, 0.5 M NaCl, 20 mM imidazole, 8 M urea, 0.5% triton, 20 mM Tris, pH 8, 0.5 mM DTT, 0.5 mM iodacetamide). Protein loading buffer (6% w/v SDS, 40% w/v sucrose, 20 mM Tris, pH 6.8, 0.15% w/v bromophenol blue) was added at a 1:3 ratio to urea buffer. Extracts were sonicated for 30 s and denatured at 95 °C for 5 min before being separated on 4–12% Bis–Tris gels (Life Technologies, Thornton, NSW, Australia) at 180 V for 1 h and undergoing wet transfer at 100 V for 2 h (Bio-Rad Laboratories, Gladesville, NSW, Australia) onto nitrocellulose membrane (Amersham™ Protran® Supported Western Blotting Membrane, GE Healthcare, Sigma-Aldrich). Membranes were blocked in 5% skim milk and immunoblotted overnight with one of the following primary antibodies: anti-monoubiquitinated H2B-K120 (#MM0029; Medimabs, Montreal, Canada); anti-Histone H2B (#AB52484; Abcam, Cambridge, MA, USA); GAPDH (#14C10) and phosphorylated p53 (Ser15) (#9284; Cell Signaling Technology, Danvers, MA, USA) or γH2AX (#05-636; Merck Millipore, Bayswater, VIC, Australia). Membranes were probed with peroxidase labelled secondary antibodies (anti-rabbit IgG, #NA934V; or, anti-mouse IgG, #NA931V, GE Healthcare Biosciences, NSW, Australia) for 1 h at room temperature. Chemiluminescent signal was detected by SuperSignal ECL Dura reagent (Pierce, Rockford, IL, USA) and visualized using the Fujifilm LAS-4000 imaging system (Berthold Australia, Bundoora, VIC, Australia). Quantitation by densitometry was undertaken using Image J version 1.47v (National Institute of Health, USA) or Multi Gauge 3.0 software (Fujifilm Australia Pty. Ltd., Brookvale, NSW, Australia).
RNA isolation and gene expression analyses
Total RNA was extracted using the RNeasy Mini Kit (Qiagen, Chadstone, VIC, Australia) and automated using a QIAcube (Qiagen, Chadstone, VIC, Australia) and cDNA generated using Maxima H Minus First Strand cDNA Synthesis Kit (Thermo Fisher Scientific Australia Pty. Ltd, Scoresby, VIC, Australia). Gene expression was analyzed in triplicate using TaqMan assays: TP53, Hs01034249_m1; CDKN1A, Hs00355782_m1; BBC3, Hs00248075_m1; MDM2, Hs00242813_m1; BAX, Hs00180269_m1; GADD45A, Hs00169255_m1; PLK2, Hs00198320_m1 and HMBS, Hs00609297_m1 (Life Technologies, Mulgrave, VIC, Australia) and the iTaq Universal Probes Supermix (Bio-Rad Laboratories, Gladesville, NSW, Australia). Reagents were dispensed using an epMotion 5070 robot (Eppendorf South Pacific Pty Ltd., North Ryde, NSW, Australia), run on a 7900HT Fast Real-Time PCR System and data analyzed using RQ manger software (Thermo Fisher Scientific Australia Pty. Ltd, Scoresby, VIC, Australia).
RNA sequencing
A2780 cells were treated with 9.9 µM cisplatin [IC75 dose] or vehicle control (0.9% saline) for 24 h and RNA was extracted as described above with an additional on-column DNase digest step. RNA samples were run on the Agilent 2100 bioanalyzer RNA 6000 Nano Labchip and samples which passed quality control were processed using TruSeq Stranded mRNA-seq sample preparation, followed by running on the HiSeq 2500 system (Illumina) with the 100 bp SR Rapid run protocol, giving an output of 350 M reads per run (Ramaciotti Centre for Genomics, Kensington, NSW, Australia), yielding approximately 30 M reads per replicate sample. RNA-seq data was validated by qRT-PCR for selected highly expressed genes. Read alignment and junction finding was performed using STAR (Version 2.5.1b) [42] and differential gene expression determined using Cuffdiff2 (Version 2.2.1) [43].
ChIP and ChIP-sequencing
A2780 cells (seven T175 flasks for ChIP-seq or three T175 flasks for ChIP-qPCR grown to 70% confluence) were treated with 9.9 µM cisplatin, or saline control, for 24 h before being cross-linked for 10 min using 1% formaldehyde (Thermo Fisher Scientific Australia, Scoresby, VIC, Australia). Formaldehyde was quenched using a final concentration of 0.125 M glycine (Sigma-Aldrich, Castle Hill, NSW, Australia) and cell numbers were normalized between cisplatin-treated and vehicle control. Cells were lysed in Farnham Lysis Buffer (5 mM PIPES pH 8, 85 mM KCl, 0.5% NP-40), before nuclei were isolated and lysed in RIPA buffer (1 × PBS, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS) and sonicated in an ice bath for 7–10 cycles of 10 s (10 pulses of 0.9 s on to 0.1 s off) at 30% amplitude using a Branson Sonifier SFX550 (Branson Ultrasonics, Danbury, CT, USA). Fragment size of the sonicated chromatin was checked using a 1.2% agarose gel. One percent of the sonicated chromatin was put aside to act as the input control. The remaining sample was incubated with either 3.5 µg (for ChIP-qPCR) or 5 µg (for ChIP-seq) of the H2Bub1 antibody (#05–1312, Merck Millipore, Bayswater, VIC, Australia) or mouse IgG isotype control (#31202, Thermo Fisher Scientific Scoresby, VIC, Australia), bound to 125 µl (ChIP-qPCR) or 150 µl (ChIP-seq) of M-280 sheep anti-mouse IgG (#11202D) or sheep anti-rabbit IgG (#11204D) conjugated Dynabeads (Thermo Fisher Scientific Australia, Scoresby, VIC, Australia) overnight on a rotator at 4 °C.
Chromatin bound antibody beads were washed six times with 500 mM LiCl IP Wash Buffer (100 mM Tris pH 7.5, 500 mM LiCl, 1% NP-40, 1% sodium deoxycholate) on a rotator at 4 °C and once with TE Buffer. Beads were resuspended in 200 µl IP Elution Buffer (1% SDS, 0.1 M NaHCO3) and incubated for 65 °C for 1 h on a shaking Thermomixer before beads were removed from the IP samples. IP Elution Buffer was added to input control samples and together with the IP samples were reverse cross-linked overnight at 65 °C. DNA was purified using a DNA Clean & Concentrator-25 column (#D4004; Zymo Research, Irvine, CA, USA). For ChIP-seq, the concentration of purified DNA was measured using Qubit Fluorometric quantitation (Life Technologies Australia Pty. Ltd., Mulgrave, VIC, Australia) and quality assessment performed with a 2100 Bioanalyzer (Agilent Technologies, Santa Clare, CA, USA) at the Ramaciotti Centre for Genomics (University of New South Wales, Randwick, Australia). TruSeq ChIP-seq library prep (Illumina) was used to prepare libraries from 5 ng of DNA. Sample preparation was performed as per the manufacturer’s instructions except that the number of PCR cycles was reduced from 18 to 16 PCR cycles and size selection of the DNA was performed using the Pippin 2% agarose cassette (Sage Science Inc., Beverly, MA, USA) selecting for 130–380 bp fragments (target 255 bp). The libraries were pooled at equal ratio and sequenced in one NextSeq 500 1 × 75 bp High Output flowcell (Illumina Inc., San Diego, CA, USA) by the Ramaciotti Centre for Genomics and data received as FASTQ files.
The human GRCh37/hg19 reference sequence was utilised for alignment and the UCSC hg19 annotation was used for feature recognition. Following trimming, 67 bp reads were aligned using Bowtie2 (Version 2.2.6) [44] yielding approximately 70 M sequences per sample and genomic regions where H2Bub1 was localised were determined using HOMER (https://homer.ucsd.edu/homer/) [45], and a histone-specific, variable length, regional binding approach. Increased H2Bub1 DNA binding between cisplatin and saline treated conditions was accomplished using HOMER, requiring at least a fourfold increase in tags (treated versus saline) and a Poisson p value over saline of 0.0001. The reverse approach was used to identify binding that was higher in saline versus cisplatin. Prior to pathway analysis, regions of robust H2Bub1 DNA binding were selected using a HOMER peak score cut-off ≥ 100, where peak score quantitates position-adjusted reads from the peak region. This cut-off was chosen by correlating the score with visual inspection of BED files uploaded to the UCSC Genome Browser. Tag counts were normalized by total tag count per sample.
Chromatin immunoprecipitation qPCR (ChIP-qPCR)
ChIP-qPCR was used to validate H2Bub1 enrichment at specific gene sequences in the discovery cell line A2780 and to expand analyzes to the additional TP53 wt line MCF7 and mutant (OVCAR-3 and Kuramochi). Cell lines were treated with cisplatin for 24 h at doses equivalent to IC75 (A2780, MCF7) or IC80 (OVCAR-3, Kuramochi). Furthermore, ChIP-qPCR was used to assess H2Bub1 enrichment at specific genes following transfection of mutant (p.Arg248Gln) or wt TP53 into the p53 null cell line SKOV3. Primer design was informed by H2Bub1 enrichment in cisplatin-treated A2780 cells, both within the gene sequence (enriched) and downstream of gene sequences (not-enriched). Primers to GAPDH and SPATA17 (intergenic region 1) sequences have previously been published as, respectively, positive and negative regions of H2Bub1 enrichment [46]. Primers designed both within the coding sequence (+) and downstream (−) of the p53 target genes BAX, MDM2, BBC3, CDKN1A, GADD45A and PLK2 were used to assess H2Bub1 enrichment by qRT-PCR following ChIP. Primer sequences are listed in Suppl. Table 1. IgG controls were used to demonstrate specificity of the H2Bub1 antibody for immunoprecipitation. qRT-PCR was performed in triplicate using specific primer pairs and the SensiMix SYBR Hi-ROX Kit (Bioline, Eveleigh, NSW, Australia).
The MDM2 antagonist nutlin-3a was used to determine the likelihood that changes in H2Bub1 enrichment were related to the presence of p53. A2780 cells were treated with IC75 (21.3 µM) levels of nutlin-3a for 24 h, followed by ChIP-qPCR as above. Further, to determine whether the presence of H2Bub1 is dependent upon transcription, an inhibitor of RNA pol II transcriptional elongation, DRB, was used. A2780 cells were treated with IC75 doses of cisplatin for 16 h after which time the drug was removed and the cells treated with 50 µM of DRB for 5 h. Cells were fixed and ChIP-qPCR performed as above.
Statistical analysis
IBM SPSS software version 22.0 (SPSS Australasia Pty Ltd., Chatswood, NSW, Australia) was used for statistical analyzes. ChIP-qPCR, gene expression and Western blot data are expressed as the mean ± SEM from at least three independent experiments. One-way ANOVA (posthoc Tukey HSD test) or one sample t tests were used to determine statistical significance. For all analyzes, p < 0.05 was considered to be statistically significant.
Results
Global levels of H2Bub1 decrease in response to DNA damage in the presence of wild-type p53
Wild-type p53 cell lines were treated with cisplatin corresponding to either an IC75 (A2780) or IC80 (HEY and MCF7) dose (Suppl. Fig. 1) and cell lysates harvested at 0, 4, 8, 18 and 24 h. Strong expression of the double strand break (DSB) marker γ-H2AX was present in all cell lines from 18 h, with A2780 showing earlier presence of this histone mark (Fig. 1a–c). Largely in parallel with γ-H2AX, stabilized p53, indicated by serine15 (Ser15) phosphorylation, emerged in response to DNA damage. While levels of total H2B and the reference protein GAPDH remained constant, levels of H2Bub1 significantly decreased over time in all wt p53 cells (Fig. 1a–c). This was in contrast to the p53 null cell lines SKOV3 and H1299 that showed no significant change in H2Bub1 levels in response to an IC80 dose of cisplatin over 24 h, despite clear induction of the DSB marker γH2AX (Fig. 1d, e Suppl. Figure 1). Transient transfection of wt p53 into both p53 null cell lines increased expression levels of phosphor-p53 and the p21 protein and significantly reduced H2Bub1 levels (Fig. 1f, g). Taken together, these data suggest that increased levels of wt p53 or its Ser15 phosphoform, whether in response to DNA damage or transient transfection, is associated with the reduction of total levels of H2Bub1.
Regulation of H2Bub1 levels in response to DNA damage is influenced by the presence of wild-type p53. a Representative immunoblots and quantitation for A2780 (N = 4), b MCF7 (N = 4), and c HEY (N = 4). The H2Bub1:TH2B ratio was normalized within experiments to the vehicle control (s; saline treated at 4 h) and shown as the mean ± SEM. A2780 and MCF7 cells showed significant reduction in H2Bub1 at 18 and 24 h. HEY1 cells show significant reduction in H2Bub1 at 24 h. In p53 null cell lines d SKOV3 (n = 5) and e H1299 (n = 4), representative immunoblots and data from pooled analyzes of p53 null cell lines treated with an IC80 dose of cisplatin (Suppl. Figure 1) show constant H2Bub1 levels over 24 h. f SKOV3 (n = 6) and g H1299 (n = 4) cells show a decrease in H2Bub1 levels when transfected with wt p53 relative to the empty vector (ev) control 48 h after transfection. In all cases, the H2Bub1:TH2B ratio was normalized within experiments to either the vehicle control (S; saline treated at 4 h) or ev control (pcDNA4) and shown as the mean ± SEM. *p < 0.05, **p ≤ 0.005 (one-way ANOVA, Tukey’s post hoc test or one-sample t test). p-p53 p53 phosphorylation at serine 15, TH2B total histone H2B
Epigenomic distribution of chromatin-bound H2Bub1 is enriched at a subset of genes in response to DNA damage
To determine the genomic distribution of chromatin-bound H2Bub1 we performed H2Bub1 ChIP-seq on the A2780 cell line. Analysis of chromatin immunoprecipitation sequencing (ChIP-seq) data showed that the broad distribution of H2Bub1 was similar between saline and cisplatin-treated cells, with most H2Bub1 found at introns (43.7–45.7%), followed by exons (21.6–21.9%), 5′-UTR (11.2–12.1%) and promoters (9.5–9.7%). Lower amounts of chromatin-bound H2Bub1 were found in saline treated cells at transcription termination sites (TTS; 3.8%), followed by 3′-UTR (3.4%), intergenic regions (2.5%) and non-coding (ncRNA; 1.2%). In cisplatin-treated cells, H2Bub1 was found at intergenic (4.5%), TTS (4.1%), 3′-UTR (4.0%) and ncRNA (1.2%) regions (Fig. 2a, b).
Distribution of H2Bub1 enrichment throughout the genome, including gene specific H2Bub1 enrichment in response to DNA damage. a Genome ontology and intragenic distribution of chromatin bound H2Bub1 in cells with and without DNA damage. Genomic regions of H2Bub1 enrichment identified using HOMER in saline treated A2780 cells and b A2780 cells treated with an IC75 dose of cisplatin (Suppl. Figure 1). c H2Bub1-enrichment is observed up to 10,000 nucleotides downstream of transcription start sites (TSS) in response to cisplatin treatment (TTS, transcription termination site; UTR, untranslated region; ncRNA, non-coding RNA). d Heatmap of H2Bub1-enrichment patterns in saline versus cisplatin-treated (IC75 dose) A2780 cells (log10 tags normalized to total tag count per sample) showing fifty-eight H2Bub1-enriched protein coding genes clustered into pathways that appeared more than once (Genecards). e Heatmap showing sixty-one H2Bub1-enriched protein coding genes that did not cluster into pathways based on Genecards. f Heatmap showing H2Bub1-depletion in response to cisplatin was seen in only 6 protein-coding and one ncRNA (PTPRG-AS1). g Heatmap showing H2Bub1-enrichment in response to cisplatin was observed in 12 ncRNA and the small nucleolar RNA SNORA16A (S, saline; C, cisplatin treated)
Despite the overall decrease in H2Bub1 in response to DNA damage, H2Bub1 was enriched in a subset of genes in cisplatin-treated cells compared to saline treated, up to 10,000 nucleotides downstream of transcription start sites (TSS; Fig. 2c). Using a cut-off of peak scores > 100 and at least fourfold enrichment, 132 unique genes were found to be enriched for chromatin-bound H2Bub1 in cisplatin versus saline treated cells, and seven genes showed loss of H2Bub1 enrichment (Fig. 2d–g). Of the 132 H2Bub1-enriched genes, 119 were protein coding, twelve were ncRNA and one was the small nucleolar RNA SNORA16A. For the seven genes showing loss of H2Bub1 enrichment under conditions of DNA damage, six were protein coding and one, PTPRG-AS1, was a lncRNA (Fig. 2f).
Genes enriched for chromatin-bound H2Bub1 following treatment with cisplatin cluster into known cancer-associated pathways
Fifty-eight of the 132 H2Bub1-enriched genes identified in response to cisplatin treatment were included in pathways containing at least two of these genes (Table 1, Fig. 2d, Suppl. Figure 2). Based on Genecard Pathways, the p53 pathway contained the highest number of H2Bub1-enriched genes (N = 15), suggesting that enrichment of H2Bub1 in response to DNA damage may be an important mechanism enabling p53 signalling. STRING analysis of these 132 H2Bub1-enriched genes identified p53 signalling as the top KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway (False Discovery Rate (FDR) 2.82 × 10–5; Suppl. Figure 3). Canonical p53 target genes, including CDKN1A and MDM2, all showed enrichment of H2Bub1 in exonic and intronic regions in response to cisplatin. No enrichment of H2Bub1 was seen for TP53 itself visualized using the UCSC Genome Browser (Fig. 3). This is in line with elevated levels of p53 in response to DNA damage being predominantly due to protein stabilization through post-translational modification, while increased expression of p53 target genes is a transcriptional event (reviewed in [47]).
Wild-type p53 cells treated with cisplatin show H2Bub1 enrichment in p53 target genes. a University of California Santa Cruz (UCSC) genome browser hg19 images of representative p53 target genes CDKN1A and b MDM2 (A2780 cells treated with saline or cisplatin). RNA-seq and ChIP-seq signals are shown, as is the layered image of the active transcription mark H3K27Ac of seven cell lines from ENCODE. Exonic regions are marked by boxes in RefSeq tracks. c UCSC genome browser hg19 image of TP53, comparing H2Bub1-enriched chromatin in response to DNA damage as above
Cisplatin-induced H2Bub1 enrichment was validated for a selection of p53 target genes BAX, MDM2, BBC3, CDKN1A, GADD45A and PLK2 in the wt p53 cell line A2780 and the mutant p53 line OVCAR-3 using quantitative real-time PCR (qRT-PCR; Fig. 4a–d). For all genes analyzed, H2Bub1 showed significant enrichment in cisplatin-treated wt p53 cells, in the case of BBC3 being enriched tenfold over saline treated cells (Fig. 4a). While some H2Bub1 enrichment was observed in mutant p53 cells in response to cisplatin, the degree of enrichment was two-fold or less, and in the case of MDM2, a small decrease in H2Bub1 was observed in cisplatin-treated cells (Fig. 4c). H2Bub1 enrichment in response to cisplatin was associated with increased expression of p53 target genes in wt p53 cells, in the case of CDKN1A displaying 38-fold higher expression than in saline treated cells (Fig. 4b). While some increase in gene expression in response to cisplatin in p53 mutant cells was seen, significant changes were observed at only two-fold or lower levels (Fig. 4d). Similar results were observed in the additional p53 wt and mutant cell lines, respectively MCF7 and Kuramochi (Suppl. Figure 4). These data support our ChIP-seq data that identified genes involved in the p53 signalling pathway as H2Bub1 enriched in response to cisplatin in wt p53 cells.
H2Bub1 enrichment in p53 target genes in the presence of wild-type or mutant p53. a Increased H2Bub1 enrichment in exons and introns of p53 target genes in response to cisplatin is observed in A2780 wild-type p53 cells treated with saline (s) or cisplatin (c; IC75 dose) for 24 h following ChIP with an anti-H2Bub1 antibody or IgG followed by qRT-PCR using either primers within (+) or downstream (−) of the coding regions of p53 target genes. Data within replicates was normalized to the experimental mean and presented as mean ± SEM (N = 3) analyzed using one-way ANOVA with Tukey’s post hoc test. b p53 target genes under the same conditions as above showed increased expression following treatment with cisplatin (N = 4; data expressed relative to saline treated cells (one-sample t test); GOI gene of interest. c OVCAR-3 p53 mutant cells treated with saline or cisplatin (IC80 dose; Suppl. Figure 1) for 24 h underwent ChIP and qRT-PCR as above (N = 3). d Expression of p53 target genes in OVCAR-3 cells in response to an IC80 dose of cisplatin as above (N = 3). (*p < 0.05, **p < 0.005). e Transfection of wt p53 into p53 null SKOV-3 cells leads to increased H2Bub1-enrichment at the coding region of p53 target genes relative to transfection with mutant p53, in parallel with increased gene expression. Transfected cells underwent ChIP with anti-H2Bub1 or IgG antibodies followed by qRT-PCR using either primers within (+) or downstream (−) of the coding regions of p53 target genes. Data within replicates was normalized to the experimental mean and presented as mean ± SEM (N = 4) analyzed using one-way ANOVA with Tukey’s post hoc test. f Expression of p53 target genes in transfected cells (N = 4). Data expressed relative to saline treated cells (one sample t test), (*p < 0.05, **p < 0.005). g Treatment of A2780 cells with the MDM2 inhibitor nutlin-3a leads to H2Bub1-enrichment at the coding region of p53 target genes. A2780 wt p53 cells treated with vehicle (DMSO) or nutlin-3a (21.3 µM, correlating with an IC75 dose) for 24 h underwent ChIP with an anti-H2Bub1 antibody or IgG followed by qRT-PCR using either primers within (+) or downstream (−) of the coding regions of p53 target genes. Data within replicates was normalized to the experimental mean and presented as mean ± SEM (N = 3) analyzed using one-way ANOVA with Tukey’s post hoc test. h Expression of p53 target genes in A2780 cells in response to an IC75 dose of nutlin-3a, or vehicle (DMSO) for 24 h (N = 4). Data expressed relative to DMSO treated cells (one sample t test), (*p < 0.05, **p ≤ 0.005)
Functional analysis of the role of p53 on H2Bub1 chromatin modification
Next, to interrogate the role of p53 in H2Bub1 genomic distribution, we transfected either wt or mutant (p.Arg248Gln) p53 into SKOV3 p53 null cells followed by H2Bub1 ChIP and qRT-PCR. Our data revealed that the p53 target genes MDM2, BBC3 and CDKN1A showed significant enrichment of chromatin-bound H2Bub1 in wt p53 transfected cells relative to mutant p53 transfected cells. Only PLK2 showed a decrease in H2Bub1 enrichment in response to DNA damage in wt p53 transfected cells (Fig. 4e). Furthermore, all p53 target genes analyzed showed higher gene expression in p53 wt transfected cells relative to p53 mutant cells (Fig. 4f). This data provides additional evidence that H2Bub1 enrichment correlating with high levels of gene expression of p53 target genes is promoted by wt p53.
Finally, we sought to gain additional evidence that the phenomenon observed was directly related to p53 rather than a generalised effect of DNA damage. We treated A2780 cells with the MDM2 inhibitor nutlin-3a that blocks polyubiquitination of p53, preventing its degradation via the 26S proteasome and resulting in higher levels of p53 in the cell. This treatment also led to H2Bub1 enrichment at p53 target genes, between 2.7 to 7.9 fold (Fig. 4g), alongside increased expression of the same genes (Fig. 4h). Taken together, these data provides further evidence that H2Bub1-enrichment in the coding regions of p53 target genes is a key feature of wt p53 signalling.
Other H2Bub1-enriched genes in response to DNA damage included those that clustered into involvement with the immune system or immune response (N = 10), ERK/MAPK signalling-related (N = 8), metabolic pathways (N = 8), membrane-trafficking (N = 7), chromatin-regulation pathways (N = 2), nucleotide excision repair pathway (NER; N = 2), sphingolipid signalling (N = 2), Hippo signalling (N = 2) and circadian rhythm associated (N = 2) (Table 1). POLH, functioning in platinum drug resistance and DNA repair pathways, and HSPA4L, functioning in senescence pathways, were also enriched for H2Bub1 in response to DNA damage. The six protein-coding genes and one lncRNA showing reduction of H2Bub1 in response to DNA damage (AKNA, C1QL3, DDR2, FAM129A, PTPRG-AS1, SLC7A1 and TBL1X) did not show any functional enrichment (STRING analysis, data not shown).
Enrichment of chromatin-bound H2Bub1 correlates with transcriptional elongation and gene expression levels following DNA damage
RNA sequencing was used to profile gene expression 2, 6 and 24 h post treatment of A2780 cells with 9.9 µM cisplatin. Gene expression at these time points is presented as heatmaps for the 119 protein coding genes that showed H2Bub1 enrichment in response to cisplatin (Fig. 5a, b), as well as the 6 protein coding genes where H2Bub1 was depleted in response to DNA damage (Fig. 5c). At 24 h, there were observed similarities between H2Bub1 enrichment and increased gene expression relative to saline treated cells (85%, 101/119; Fig. 5d), although some genes with H2Bub1 enrichment in response to cisplatin showed unchanged or decreased levels of gene expression (15%). Of the 6 genes that showed loss of H2Bub1 in response to cisplatin, 5 showed decreased gene expression and one expressed at very low levels, C1QL3, remained constant (Fig. 5e).
Correlation of H2Bub1 enriched genes and gene expression in response to DNA damage. a Heatmaps of log2 fold-change gene expression of H2Bub1-enriched genes after treatment with cisplatin at 2, 6 and 24 h relative to saline treated cells. Expression of H2Bub1-enriched genes post treatment of A2780 cells with an IC75 dose of cisplatin that clustered into cancer-related pathways (based on Genecards). b Expression of H2Bub1-enriched genes that did not cluster into pathways (based on Genecards). Five genes with low expression were excluded (NRG3, KIAA1211L, CPEB4, PRCD, ARHGAP30). c Expression of genes where H2Bub1-enrichment was decreased in response to cisplatin treatment. C1QL3 was excluded due to very low levels of gene expression. cisp cisplatin. d Venn diagrams displaying overlap of H2Bub1 patterns and the direction of gene expression in response to cisplatin at 2, 6 and 24 h in A2780 cells. Genes displaying enriched H2Bub1 at exons and introns (green circles) in response to treatment with an IC75 dose of cisplatin (N = 119) correlated with increased gene expression (overlap of pink and green circles) at, 2 h, 39% (46/119), 6 h, 66% (79/119), and 24 h, 85% (101/119). ^Genes where expression was not increased. e Conversely, genes where H2Bub1 levels were decreased at their coding regions (salmon circle) in response to treatment with cisplatin showed decreased gene expression (overlap of salmon and yellow circles) at 2 h, 83% (5/6), 6 h, 83% (5/6), and 24 h, 83% (5/6)
To prove that the presence of H2Bub1 is reliant on transcriptional elongation, A2780 cells were treated with the transcriptional inhibitor DRB. A dose–response curve determined the optimal level of DRB to inhibit transcription (data not shown). To demonstrate that the selected dose of 50 µM DRB was sufficient to inhibit transcription in A2780 cells following DNA damage with cisplatin, we determined that levels of MDM2 were decreased in DRB versus vehicle treated cells (Suppl. Figure 5a). H2Bub1 ChIP-qPCR under the same conditions showed a highly significant decrease in H2Bub1enrichment in the coding region of all p53 target genes assessed following treatment with DRB (Suppl. Figure 5b). These results clearly demonstrate that H2Bub1 enrichment at p53 target genes only occurs in the presence of transcriptional elongation.
Discussion
The changing chromatin landscape of cancer cells in response to therapeutic DNA damage influences gene expression [48, 49]. H2Bub1 is a central histone modification associated with open chromatin configuration and promotion of transcriptional elongation. Regions of H2Bub1 enrichment regulate the accessibility of genomic sites. We have used cancer cell line models to show that many of these H2Bub1-enriched genes in response to DNA damage are those that contribute to the determination of cell fate. DNA damage with the platinum drug cisplatin led to a decrease of global levels of H2Bub1 in wt p53 cell lines, in parallel with increased levels of phosphorylated p53. This parallels a previous report where decreased global levels of H2Bub1 were seen after treatment of the p53 wt osteosarcoma cell line U2OS with the chemotherapeutic agent doxorubicin [35]. Treatment with the transcriptional inhibitors actinomycin D, 5,6-dichloro-β-d-ribofuranosylbenzimidazole (DRB), and α-amanitin in HeLa cells has also been reported to lead to decreased levels of global H2Bub1 [50]. In the context of DNA damage, we have clearly shown that inhibition of transcriptional elongation with DRB leads to reduced H2Bub1 at the coding region of p53 target genes. Interestingly, we did not observe decreases in global levels of H2Bub1 in response to cisplatin in cell lines that are null for the p53 transcription factor; however, transfection of wt p53 into p53 null cells led to a significant reduction in global H2Bub1. Taken together, this suggests that the induction and/or phosphorylation of wt p53 is correlated with loss of global levels of H2Bub1.
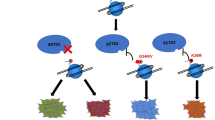
We used chromatin immunoprecipitation followed by massively parallel sequencing (ChIP-seq) to show that while global levels of H2Bub1 decreased in response to DNA damage, specific H2Bub1-enrichment was identified up to 10,000 nucleotides downstream of the TSS in a specific subset of genes and non-coding RNA. H2Bub1-enrichment peaked at around 1000 nucleotides downstream of the TSS. This suggests that in response to DNA damage, loss of H2Bub1 might function to shut down expression of many genes that are normally transcriptionally active but not required to manage the response to DNA damage. At the same time, H2Bub1 becomes enriched at specific genes, allowing their increased expression via transcriptional elongation and so facilitating their function in the DNA damage response (Fig. 6).
Schematic outlining the enrichment of H2Bub1 in subsets of genes in response to DNA damage facilitating transcriptional elongation. Chromatin within genes, indicated by the red line, remains tightly wound around the nucleosome under basal conditions. In response to DNA damage, indicated by yellow lightning bolt symbols, the H2B histone tail is monoubiquitinated at lysine 120 (H2Bub1, indicated by a red circular form), leading to a more open chromatin accessible by RNA polymerase II, promoting transcriptional elongation (indicated by a green arrow) of genes involved in determining cell fate
Of note, the genome-wide occupancy of H2Bub1-enrichment remained largely unchanged when viewed as proportions of coding and non-coding regions both basally and in response to cisplatin. The majority of enriched H2Bub1 resided in intronic regions, 45.7% in vehicle treated versus 43.7% in cisplatin-treated cells. The next most frequently enriched regions were exons, with 21.6% in vehicle treated versus 21.9% in cisplatin-treated cells, followed by 5′-UTR, promoters, TSSs, 3′-UTR, intergenic regions and non-coding RNA in saline treated cells; and 5′-UTR, promoters, intergenic regions, TTSs, 3′-UTR and non-coding RNA in cisplatin-treated cells. The role of H2Bub1 in introns remains speculative, although it is possible that its presence facilitates access to DNA for regulatory factors, including for splicing. In undifferentiated human teratocarcinoma NCCIT cells, H2Bub1 was present at the 5′ regions of basally transcribed genes, as well as at intron–exon boundaries of highly expressed exons and in skipped exons [51]. Interestingly, in a recent study using ATAC sequencing (Assay for Transposase-Accessible Chromatin sequencing), a more accessible chromatin structure was observed in promoter regions following depletion of the H2Bub1 writer RNF20 [52]. Hooda and colleagues used an elegant model of normal fallopian tube secretory epithelial cells artificially immortalized by the SV40 large T antigen. SV40 transformed cells are known to contain high levels of stabilized p53 but with abrogated wild-type p53 function [53]. It is therefore difficult to directly compare the results of that study with those that we report in a wild-type p53 cancer cell line model in the context of H2Bub1 localisation.
Using stringent conditions of peak scores ≥ 100 and a four-fold increase in cisplatin versus saline tags, a small set of 119 coding genes, 12 non-coding genes and a single snoRNA (SNORA16A) were identified as H2Bub1-enriched in response to DNA damage. Based on Genecard pathways, H2Bub1-enriched genes clustered into key cancer-associated pathways, with the p53 signalling pathway identified as the most highly enriched pathway by KEGG (Kyoto Encyclopedia of Genes and Genomes) analysis. Genes that were H2Bub1-enriched in p53 signalling included PPM1D, MDM2, CDKN1A, BBC3 and GADD45A. CDKN1A has previously shown to be H2Bub1-enriched in the presence of the H2Bub1 E3 ligase RNF20 compared to down-regulated RNF20 [35]. This is consistent with the physical association of p53 with both RNF20 and RNF40 [54]. Of note, in the current study, the TP53 gene itself was not H2Bub1-enriched in response to DNA damage. It is long established that stabilization of p53 leading to increased levels of this tumor suppressor protein is due to post-translational modifications including acetylation and phosphorylation, rather than being predominantly a transcriptional event [54, 55].
To demonstrate that H2Bub1-enrichment in p53 transcriptional targets was specific to p53 and not purely an effect of DNA damage, we treated A2780 cells with the MDM2 inhibitor nutlin-3a that prevents polyubiquitination of p53 by its E3 ligase MDM2 stopping its subsequent degradation via the 26S proteasome. Using ChiP-qPCR and primers designed to the exons and downstream regions of the p53 target genes CDKN1A, MDM2, BAX, BBC3, PLK2 and GADD45A we demonstrated H2Bub1 enrichment in the coding regions of p53 gene targets in nutlin-3a treated cells. Therefore, by three independent processes that increase levels of wt p53 in the cell, namely induction of DNA damage, wt p53 transfection and treatment with nutlin-3a, we confirmed H2Bub1-enrichment at p53 target genes. Similar levels of H2Bub1 enrichment in p53 target genes were not seen in cell lines that harboured endogenous p53 mutations or when mutant p53 was transfected into p53 null cells, highlighting the correlation between H2Bub1 enrichment and wt p53.
Other H2Bub1-enriched genes included those encoding proteins involved in the immune response (e.g. PNP and TRIM2), ERK/MAPK signalling (e.g. FOS and SOD1), sphingolipid signalling (S1PR3 and CERS6), the Hippo signalling pathway (PARD6G and AMOTL1), nucleotide excision repair (NER; POLR2A and XPC), trafficking (e.g. GOSR1 and AP3S1) and metabolism (e.g. SDHA and ALDH7A1). Furthermore, POLH that codes for DNA polymerase η (pol η) known to be involved in resistance to platinum based drugs, was H2Bub1-enriched in response to DNA damage [56].
Like POLH1, expression of several H2Bub1-enriched genes and pathways have already been linked specifically with cisplatin. For example, cisplatin is known to upregulate the mitogen-activated protein kinase (MAPK) signalling pathway that includes extracellular signal-regulated kinase (ERK), promoting the survival of cancer cells and cisplatin resistance [57,58,59]. Of the genes shown to be H2Bub1-enriched in response to DNA damage that form part of this pathway, a number have been linked to cisplatin sensitivity, including EPHA2 [60], FOS [61] and SOD1 [62]. Furthermore, overexpression of the NER gene XPC has also been linked to increased cisplatin resistance [63].
RNA sequencing data at 24 h post treatment with cisplatin showed that 85% (101 of 119) of genes that were H2Bub1-enriched also showed increased gene expression above basal levels, suggesting strong correlation between H2Bub1-enrichment and increased gene expression in line with the role of H2Bub1 in transcriptional elongation [64]. None of the H2Bub1 depleted genes were related by common pathways. An additional 12 non-coding RNA, including one microRNA, three long non-coding RNA, and a single snoRNA (SNORA16A) were found to be H2Bub1-enriched in response to DNA damage, while six protein coding and one non-coding RNA were depleted in H2Bub1 in response to DNA damage. Of the H2Bub1-enriched long non-coding RNA, two are known to be induced by p53, specifically LINC01021 [65] and LINC-PINT [66]. Therefore, it is apparent that H2Bub1-enrichment correlates with both p53 dependent and p53 independent pathways following DNA damage.
It is possible that if cells were to be treated with a DNA damaging agent other than cisplatin, different patterns of H2Bub enrichment might be seen. This concept of alternative patterns of H2Bub1 in response to other agents or processes resulting in DNA damage would appear to be the case in recent studies investigating the accumulation of H2Bub1 at sites of DSBs. Studies have shown that H2Bub1 accumulates at sites of DSBs in response to the radiomimetic drug neocarzinostatin or ionising radiation [50, 67]. In contrast, a recent study used 4-hydroxytamoxifen to examine chromatin modifications at sites of DSBs reported loss of H2Bub1 at these sites, identifying a switch to enrichment of acetylation (H2BK120ac) with likely involvement of the chromatin modifying complex SAGA [68, 69]. It remains to be observed whether differences at histone H2B lysine 120 in response to different DNA damaging agents might also be seen in transcriptional responses such as those reported in this study.
In summary, we have shown H2Bub1 enrichment at important cancer-related genes following DNA damage with cisplatin, including those participating in p53 signalling and involved in resistance to cisplatin. Further, our data suggest that for a subset of genes, this phenomenon is assisted by a functional wild-type p53, which, when absent, results in inhibition of this process. These findings support the growing body of evidence that is establishing H2Bub1 as a key histone modifier influencing gene transcription in both the cancer and the cancer therapeutic chromatin landscape.
References
Apps MG, Choi EH, Wheate NJ (2015) The state-of-play and future of platinum drugs. Endocr Relat Cancer 22(4):R219–233. https://doi.org/10.1530/erc-15-0237
Bowtell DD, Bohm S, Ahmed AA, Aspuria PJ, Bast RC Jr, Beral V, Berek JS, Birrer MJ, Blagden S, Bookman MA, Brenton JD, Chiappinelli KB, Martins FC, Coukos G, Drapkin R, Edmondson R, Fotopoulou C, Gabra H, Galon J, Gourley C, Heong V, Huntsman DG, Iwanicki M, Karlan BY, Kaye A, Lengyel E, Levine DA, Lu KH, McNeish IA, Menon U, Narod SA, Nelson BH, Nephew KP, Pharoah P, Powell DJ Jr, Ramos P, Romero IL, Scott CL, Sood AK, Stronach EA, Balkwill FR (2015) Rethinking ovarian cancer II: reducing mortality from high-grade serous ovarian cancer. Nat Rev Cancer 15(11):668–679. https://doi.org/10.1038/nrc4019
Chapman-Rothe N, Curry E, Zeller C, Liber D, Stronach E, Gabra H, Ghaem-Maghami S, Brown R (2013) Chromatin H3K27me3/H3K4me3 histone marks define gene sets in high-grade serous ovarian cancer that distinguish malignant, tumor-sustaining and chemo-resistant ovarian tumor cells. Oncogene 32(38):4586–4592. https://doi.org/10.1038/onc.2012.477
Liu D, Zhang XX, Li MC, Cao CH, Wan DY, Xi BX, Tan JH, Wang J, Yang ZY, Feng XX, Ye F, Chen G, Wu P, Xi L, Wang H, Zhou JF, Feng ZH, Ma D, Gao QL (2018) C/EBPbeta enhances platinum resistance of ovarian cancer cells by reprogramming H3K79 methylation. Nat Commun 9(1):1739. https://doi.org/10.1038/s41467-018-03590-5
Long J, Fang WY, Chang L, Gao WH, Shen Y, Jia MY, Zhang YX, Wang Y, Dou HB, Zhang WJ, Zhu J, Liang AB, Li JM, Hu J (2017) Targeting HDAC3, a new partner protein of AKT in the reversal of chemoresistance in acute myeloid leukemia via DNA damage response. Leukemia 31(12):2761–2770. https://doi.org/10.1038/leu.2017.130
Cacan E (2016) Histone deacetylase-1-mediated suppression of FAS in chemoresistant ovarian cancer cells. Anticancer Res 36(6):2819–2826
Cacan E (2017) Epigenetic regulation of RGS2 (regulator of G-protein signaling 2) in chemoresistant ovarian cancer cells. J Chemother 29(3):173–178. https://doi.org/10.1080/1120009x.2016.1277007
Castilho RM, Squarize CH, Almeida LO (2017) Epigenetic modifications and head and neck cancer: implications for tumor progression and resistance to therapy. Int J Mol Sci. https://doi.org/10.3390/ijms18071506
Gardner EE, Lok BH, Schneeberger VE, Desmeules P, Miles LA, Arnold PK, Ni A, Khodos I, de Stanchina E, Nguyen T, Sage J, Campbell JE, Ribich S, Rekhtman N, Dowlati A, Massion PP, Rudin CM, Poirier JT (2017) Chemosensitive relapse in small cell lung cancer proceeds through an EZH2-SLFN11 axis. Cancer Cell 31(2):286–299. https://doi.org/10.1016/j.ccell.2017.01.006
Huo H, Magro PG, Pietsch EC, Patel BB, Scotto KW (2010) Histone methyltransferase MLL1 regulates MDR1 transcription and chemoresistance. Cancer Res 70(21):8726–8735. https://doi.org/10.1158/0008-5472.Can-10-0755
Li J, Yu B, Deng P, Cheng Y, Yu Y, Kevork K, Ramadoss S, Ding X, Li X, Wang CY (2017) KDM3 epigenetically controls tumorigenic potentials of human colorectal cancer stem cells through Wnt/beta-catenin signalling. Nat Commun 8:15146. https://doi.org/10.1038/ncomms15146
Lu C, Yang D, Sabbatini ME, Colby AH, Grinstaff MW, Oberlies NH, Pearce C, Liu K (2018) Contrasting roles of H3K4me3 and H3K9me3 in regulation of apoptosis and gemcitabine resistance in human pancreatic cancer cells. BMC Cancer 18(1):149. https://doi.org/10.1186/s12885-018-4061-y
Samulitis BK, Pond KW, Pond E, Cress AE, Patel H, Wisner L, Patel C, Dorr RT, Landowski TH (2015) Gemcitabine resistant pancreatic cancer cell lines acquire an invasive phenotype with collateral hypersensitivity to histone deacetylase inhibitors. Cancer Biol Ther 16(1):43–51. https://doi.org/10.4161/15384047.2014.986967
Staberg M, Rasmussen RD, Michaelsen SR, Pedersen H, Jensen KE, Villingshoj M, Skjoth-Rasmussen J, Brennum J, Vitting-Seerup K, Poulsen HS, Hamerlik P (2018) Targeting glioma stem-like cell survival and chemoresistance through inhibition of lysine-specific histone demethylase KDM2B. Mol Oncol 12(3):406–420. https://doi.org/10.1002/1878-0261.12174
Cole AJ, Clifton-Bligh R, Marsh DJ (2015) Histone H2B monoubiquitination: roles to play in human malignancy. Endocr Relat Cancer 22(1):T19–33. https://doi.org/10.1530/erc-14-0185
Marsh DJ, Dickson KA (2019) Writing histone monoubiquitination in human malignancy—the role of RING finger E3 ubiquitin ligases. Genes 10(1):15. https://doi.org/10.3390/genes10010067
Clague MJ, Coulson JM, Urbe S (2008) Deciphering histone 2A deubiquitination. Genome Biol 9(1):202. https://doi.org/10.1186/gb-2008-9-1-202
Machida S, Sekine S, Nishiyama Y, Horikoshi N, Kurumizaka H (2016) Structural and biochemical analyzes of monoubiquitinated human histones H2B and H4. Open Biol. https://doi.org/10.1098/rsob.160090
Fierz B, Chatterjee C, McGinty RK, Bar-Dagan M, Raleigh DP, Muir TW (2011) Histone H2B ubiquitylation disrupts local and higher-order chromatin compaction. Nat Chem Biol 7(2):113–119. https://doi.org/10.1038/nchembio.501
Braun S, Madhani HD (2012) Shaping the landscape: mechanistic consequences of ubiquitin modification of chromatin. EMBO Rep 13(7):619–630. https://doi.org/10.1038/embor.2012.78
Simon JA, Kingston RE (2009) Mechanisms of polycomb gene silencing: knowns and unknowns. Nat Rev Mol Cell Biol 10(10):697–708. https://doi.org/10.1038/nrm2763
Marsh DJ, Shah JS, Cole AJ (2014) Histones and their modifications in ovarian cancer—drivers of disease and therapeutic targets. Front Oncol 4:144. https://doi.org/10.3389/fonc.2014.00144
Lee JS, Smith E, Shilatifard A (2010) The language of histone crosstalk. Cell 142(5):682–685. https://doi.org/10.1016/j.cell.2010.08.011
Min J, Feng Q, Li Z, Zhang Y, Xu RM (2003) Structure of the catalytic domain of human DOT1L, a non-SET domain nucleosomal histone methyltransferase. Cell 112(5):711–723
Briggs SD, Xiao T, Sun ZW, Caldwell JA, Shabanowitz J, Hunt DF, Allis CD, Strahl BD (2002) Gene silencing: trans-histone regulatory pathway in chromatin. Nature 418(6897):498. https://doi.org/10.1038/nature00970
Ng HH, Xu RM, Zhang Y, Struhl K (2002) Ubiquitination of histone H2B by Rad6 is required for efficient Dot1-mediated methylation of histone H3 lysine 79. J Biol Chem 277(38):34655–34657. https://doi.org/10.1074/jbc.C200433200
Sun ZW, Allis CD (2002) Ubiquitination of histone H2B regulates H3 methylation and gene silencing in yeast. Nature 418(6893):104–108. https://doi.org/10.1038/nature00883
Shema-Yaacoby E, Nikolov M, Haj-Yahya M, Siman P, Allemand E, Yamaguchi Y, Muchardt C, Urlaub H, Brik A, Oren M, Fischle W (2013) Systematic identification of proteins binding to chromatin-embedded ubiquitylated H2B reveals recruitment of SWI/SNF to regulate transcription. Cell Rep 4(3):601–608. https://doi.org/10.1016/j.celrep.2013.07.014
Kim J, Guermah M, McGinty RK, Lee JS, Tang Z, Milne TA, Shilatifard A, Muir TW, Roeder RG (2009) RAD6-Mediated transcription-coupled H2B ubiquitylation directly stimulates H3K4 methylation in human cells. Cell 137(3):459–471. https://doi.org/10.1016/j.cell.2009.02.027
Shilatifard A (2012) The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu Rev Biochem 81:65–95. https://doi.org/10.1146/annurev-biochem-051710-134100
Smith E, Shilatifard A (2010) The chromatin signaling pathway: diverse mechanisms of recruitment of histone-modifying enzymes and varied biological outcomes. Mol Cell 40(5):689–701. https://doi.org/10.1016/j.molcel.2010.11.031
Wu M, Wang PF, Lee JS, Martin-Brown S, Florens L, Washburn M, Shilatifard A (2008) Molecular regulation of H3K4 trimethylation by Wdr82, a component of human Set1/COMPASS. Mol Cell Biol 28(24):7337–7344. https://doi.org/10.1128/mcb.00976-08
van Welsem T, Korthout T, Ekkebus R, Morais D, Molenaar TM, van Harten K, Poramba-Liyanage DW, Sun SM, Lenstra TL, Srivas R, Ideker T, Holstege FCP, van Attikum H, El Oualid F, Ovaa H, Stulemeijer IJE, Vlaming H, van Leeuwen F (2018) Dot1 promotes H2B ubiquitination by a methyltransferase-independent mechanism. Nucleic Acids Res 46(21):11251–11261. https://doi.org/10.1093/nar/gky801
Dickson KA, Cole AJ, Gill AJ, Clarkson A, Gard GB, Chou A, Kennedy CJ, Henderson BR, Fereday S, Traficante N, Alsop K, Bowtell DD, deFazio A, Clifton-Bligh R, Marsh DJ (2016) The RING finger domain E3 ubiquitin ligases BRCA1 and the RNF20/RNF40 complex in global loss of the chromatin mark histone H2B monoubiquitination (H2Bub1) in cell line models and primary high-grade serous ovarian cancer. Hum Mol Genet 25(24):5460–5471. https://doi.org/10.1093/hmg/ddw362
Minsky N, Shema E, Field Y, Schuster M, Segal E, Oren M (2008) Monoubiquitinated H2B is associated with the transcribed region of highly expressed genes in human cells. Nat Cell Biol 10(4):483–488. https://doi.org/10.1038/ncb1712
Xie W, Nagarajan S, Baumgart SJ, Kosinsky RL, Najafova Z, Kari V, Hennion M, Indenbirken D, Bonn S, Grundhoff A, Wegwitz F, Mansouri A, Johnsen SA (2017) RNF40 regulates gene expression in an epigenetic context-dependent manner. Genome Biol 18(1):32. https://doi.org/10.1186/s13059-017-1159-5
Soule HD, Vazguez J, Long A, Albert S, Brennan M (1973) A human cell line from a pleural effusion derived from a breast carcinoma. J Natl Cancer Inst 51(5):1409–1416
Domcke S, Sinha R, Levine DA, Sander C, Schultz N (2013) Evaluating cell lines as tumor models by comparison of genomic profiles. Nat Commun 4:2126. https://doi.org/10.1038/ncomms3126
Anglesio MS, Wiegand KC, Melnyk N, Chow C, Salamanca C, Prentice LM, Senz J, Yang W, Spillman MA, Cochrane DR, Shumansky K, Shah SP, Kalloger SE, Huntsman DG (2013) Type-specific cell line models for type-specific ovarian cancer research. PLoS One 8(9):e72162. https://doi.org/10.1371/journal.pone.0072162
Ikediobi ON, Davies H, Bignell G, Edkins S, Stevens C, O'Meara S, Santarius T, Avis T, Barthorpe S, Brackenbury L, Buck G, Butler A, Clements J, Cole J, Dicks E, Forbes S, Gray K, Halliday K, Harrison R, Hills K, Hinton J, Hunter C, Jenkinson A, Jones D, Kosmidou V, Lugg R, Menzies A, Mironenko T, Parker A, Perry J, Raine K, Richardson D, Shepherd R, Small A, Smith R, Solomon H, Stephens P, Teague J, Tofts C, Varian J, Webb T, West S, Widaa S, Yates A, Reinhold W, Weinstein JN, Stratton MR, Futreal PA, Wooster R (2006) Mutation analysis of 24 known cancer genes in the NCI-60 cell line set. Mol Cancer Ther 5(11):2606–2612. https://doi.org/10.1158/1535-7163.mct-06-0433
Mitsudomi T, Steinberg SM, Nau MM, Carbone D, D'Amico D, Bodner S, Oie HK, Linnoila RI, Mulshine JL, Minna JD et al (1992) p53 gene mutations in non-small-cell lung cancer cell lines and their correlation with the presence of ras mutations and clinical features. Oncogene 7(1):171–180
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29(1):15–21. https://doi.org/10.1093/bioinformatics/bts635
Trapnell C, Hendrickson DG, Sauvageau M, Goff L, Rinn JL, Pachter L (2013) Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat Biotechnol 31(1):46–53. https://doi.org/10.1038/nbt.2450
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9(4):357–359. https://doi.org/10.1038/nmeth.1923
Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, Cheng JX, Murre C, Singh H, Glass CK (2010) Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell 38(4):576–589. https://doi.org/10.1016/j.molcel.2010.05.004
Fuchs G, Hollander D, Voichek Y, Ast G, Oren M (2014) Cotranscriptional histone H2B monoubiquitylation is tightly coupled with RNA polymerase II elongation rate. Genome Res 24(10):1572–1583. https://doi.org/10.1101/gr.176487.114
Xu Y (2003) Regulation of p53 responses by post-translational modifications. Cell Death Differ 10(4):400–403. https://doi.org/10.1038/sj.cdd.4401182
Dabin J, Fortuny A, Polo SE (2016) Epigenome maintenance in response to DNA damage. Mol Cell 62(5):712–727. https://doi.org/10.1016/j.molcel.2016.04.006
Rossetto D, Truman AW, Kron SJ, Cote J (2010) Epigenetic modifications in double-strand break DNA damage signaling and repair. Clin Cancer Res 16(18):4543–4552. https://doi.org/10.1158/1078-0432.Ccr-10-0513
Moyal L, Lerenthal Y, Gana-Weisz M, Mass G, So S, Wang SY, Eppink B, Chung YM, Shalev G, Shema E, Shkedy D, Smorodinsky NI, van Vliet N, Kuster B, Mann M, Ciechanover A, Dahm-Daphi J, Kanaar R, Hu MC, Chen DJ, Oren M, Shiloh Y (2011) Requirement of ATM-dependent monoubiquitylation of histone H2B for timely repair of DNA double-strand breaks. Mol Cell 41(5):529–542. https://doi.org/10.1016/j.molcel.2011.02.015
Jung I, Kim SK, Kim M, Han YM, Kim YS, Kim D, Lee D (2012) H2B monoubiquitylation is a 5′-enriched active transcription mark and correlates with exon-intron structure in human cells. Genome Res 22(6):1026–1035. https://doi.org/10.1101/gr.120634.111
Hooda J, Novak M, Salomon MP, Matsuba C, Ramos RI, MacDuffie E, Song M, Hirsch MS, Lester J, Parkash V, Karlan BY, Oren M, Hoon DS, Drapkin R (2019) Early loss of Histone H2B monoubiquitylation alters chromatin accessibility and activates key immune pathways that facilitate progression of ovarian cancer. Cancer Res 79(4):760–772. https://doi.org/10.1158/0008-5472.CAN-18-2297
Technau A, Wolff A, Sauder C, Birkner N, Brandner G (2001) p53 in SV40-transformed DNA-damaged human cells binds to its cognate sequence but fails to transactivate target genes. Int J Oncol 18(2):281–286
Wu C, Cui Y, Liu X, Zhang F, Lu LY, Yu X (2020) The RNF20/40 complex regulates p53-dependent gene transcription and mRNA splicing. J Mol Cell Biol 12(2):113–124
Lavin MF, Gueven N (2006) The complexity of p53 stabilization and activation. Cell Death Differ 13(6):941–950. https://doi.org/10.1038/sj.cdd.4401925
Albertella MR, Green CM, Lehmann AR, O'Connor MJ (2005) A role for polymerase eta in the cellular tolerance to cisplatin-induced damage. Cancer Res 65(21):9799–9806. https://doi.org/10.1158/0008-5472.Can-05-1095
Roberts PJ, Der CJ (2007) Targeting the Raf-MEK-ERK mitogen-activated protein kinase cascade for the treatment of cancer. Oncogene 26(22):3291–3310. https://doi.org/10.1038/sj.onc.1210422
Wang J, Zhou JY, Wu GS (2007) ERK-dependent MKP-1-mediated cisplatin resistance in human ovarian cancer cells. Cancer Res 67(24):11933–11941. https://doi.org/10.1158/0008-5472.Can-07-5185
Persons DL, Yazlovitskaya EM, Cui W, Pelling JC (1999) Cisplatin-induced activation of mitogen-activated protein kinases in ovarian carcinoma cells: inhibition of extracellular signal-regulated kinase activity increases sensitivity to cisplatin. Clin Cancer Res 5(5):1007–1014
Hasegawa J, Sue M, Yamato M, Ichikawa J, Ishida S, Shibutani T, Kitamura M, Wada T, Agatsuma T (2016) Novel anti-EPHA2 antibody, DS-8895a for cancer treatment. Cancer Biol Ther 17(11):1158–1167. https://doi.org/10.1080/15384047.2016.1235663
Moorehead RA, Singh G (2000) Influence of the proto-oncogene c-fos on cisplatin sensitivity. Biochem Pharmacol 59(4):337–345
Kim KK, Han A, Yano N, Ribeiro JR, Lokich E, Singh RK, Moore RG (2015) Tetrathiomolybdate mediates cisplatin-induced p38 signaling and EGFR degradation and enhances response to cisplatin therapy in gynecologic cancers. Sci Rep 5:15911. https://doi.org/10.1038/srep15911
Zhang Y, Cao J, Meng Y, Qu C, Shen F, Xu L (2018) Overexpression of xeroderma pigmentosum group C decreases the chemotherapeutic sensitivity of colorectal carcinoma cells to cisplatin. Oncol Lett 15(5):6336–6344. https://doi.org/10.3892/ol.2018.8127
Pavri R, Zhu B, Li G, Trojer P, Mandal S, Shilatifard A, Reinberg D (2006) Histone H2B monoubiquitination functions cooperatively with FACT to regulate elongation by RNA polymerase II. Cell 125(4):703–717. https://doi.org/10.1016/j.cell.2006.04.029
Kaller M, Gotz U, Hermeking H (2017) Loss of p53-inducible long non-coding RNA LINC01021 increases chemosensitivity. Oncotarget 8(61):102783–102800. https://doi.org/10.18632/oncotarget.22245
Marin-Bejar O, Mas AM, Gonzalez J, Martinez D, Athie A, Morales X, Galduroz M, Raimondi I, Grossi E, Guo S, Rouzaut A, Ulitsky I, Huarte M (2017) The human lncRNA LINC-PINT inhibits tumor cell invasion through a highly conserved sequence element. Genome Biol 18(1):202. https://doi.org/10.1186/s13059-017-1331-y
Nakamura K, Kato A, Kobayashi J, Yanagihara H, Sakamoto S, Oliveira DV, Shimada M, Tauchi H, Suzuki H, Tashiro S, Zou L, Komatsu K (2011) Regulation of homologous recombination by RNF20-dependent H2B ubiquitination. Mol Cell 41(5):515–528. https://doi.org/10.1016/j.molcel.2011.02.002
Clouaire T, Rocher V, Lashgari A, Arnould C, Aguirrebengoa M, Biernacka A, Skrzypczak M, Aymard F, Fongang B, Dojer N, Iacovoni JS, Rowicka M, Ginalski K, Cote J, Legube G (2018) Comprehensive mapping of histone modifications at DNA double-strand breaks deciphers repair pathway chromatin signatures. Mol Cell 72(2):250–262.e256. https://doi.org/10.1016/j.molcel.2018.08.020
Helmlinger D, Tora L (2017) Sharing the SAGA. Trends Biochem Sci 42(11):850–861. https://doi.org/10.1016/j.tibs.2017.09.001
Acknowledgements
Dr Michael Hahn is acknowledged for the initial cloning of wt p53.
Funding
This project was supported by a Research Scholar Award from the Sydney Vital Translational Cancer Research Centre, Cancer Institute NSW [to A.J.C.] and project Grant from the Cancer Council NSW [RG13-10; to D.J.M.]. A.J.C. was supported by an Australian Postgraduate Award and Northern Clinical School Top-Up Scholarship. D.J.M. was supported by the Australian Research Council (ARC) (ARC Future Fellowship [FT100100489]) and National Health and Medical Research Council (NHMRC) (NHMRC Senior Research Fellowship [APP1004799]).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.

18_2020_3552_MOESM2_ESM.jpg
Suppl. Fig. 1 Cisplatin dose curves and maximal inhibitory concentrations (IC50, IC75, IC80) in µM for A2780, HEY1, MCF7, SKOV3, H1299, Kuramochi and OVCAR-3 (N = 3-4) (JPG 2248 kb)

18_2020_3552_MOESM3_ESM.jpg
Suppl. Fig. 2 University of California Santa Cruz (UCSC) genome browser hg19 images of representative H2Bub1-enriched genes in response to cisplatin. a, POLH1, b, EGR1 and c, TOB1. RNA-seq and ChIP-seq signals are shown, as is the layered image of the active transcription mark H3K27Ac of seven cell lines from ENCODE. Exonic regions are marked by boxes in RefSeq tracks. (JPG 3755 kb)

18_2020_3552_MOESM4_ESM.jpg
Suppl. Fig. 3 STRING analysis of 132 H2Bub1 enriched genes in A2780 cells following treatment with an IC75 dose of cisplatin for 24 h. Strings represent known interactions from either curated databases (aqua) or that have been experimentally determined (magenta); predicted interactions (red, green or blue), or other interactions from text mining (lime), co-expression (black) or protein homology (purple) (JPG 2921 kb)

18_2020_3552_MOESM5_ESM.jpg
Suppl. Fig. 4 Increased H2Bub1 enrichment in the coding region of p53 target genes in response to cisplatin [IC75 or IC80 dose levels] is observed in wild-type p53 cells (MCF7) compared to mutant p53 cells (Kuramochi). a, MCF7 wt p53 cells treated with saline (sal.) or cisplatin (cisp.; IC75 dose, Suppl. Fig. 1) for 24 h underwent ChIP with anti-H2Bub1 or IgG antibodies followed by qRT-PCR using either primers within (+) or downstream (-) of the coding regions of p53 target genes. Data within replicates was normalized to the experimental mean and presented as mean ± SEM (N = 5) analyzed using one-way ANOVA with Tukey’s post-hoc test. Right hand panel shows expression of p53 target genes in MCF7 cells in response to an IC75 dose of cisplatin (c), or saline (s) for 24 h (N = 4; data expressed relative to saline treated cells (one-sample t test); GOI, gene of interest). b, Kuramochi mutant p53 cells treated with saline or an IC80 dose of cisplatin (Suppl. Fig. 1) for 24 h underwent ChIP and qRT-PCR as above (N = 3). Right hand panel shows expression of p53 target genes in Kuramochi cells in response to an IC80 dose of cisplatin as above (N = 4). (* P < 0.05, ** P < 0.005) (JPG 1115 kb)

18_2020_3552_MOESM6_ESM.jpg
Suppl. Fig. 5 Inhibition of transcriptional elongation after DNA damage inhibits H2Bub1 enrichment at p53 target genes. a, Expression levels of MDM2 were assessed relative to the reference gene HMBS in A2780 cells treated with saline or IC75 doses of cisplatin (9.9 µM) for 16 h, followed by treatment with DMSO (vehicle) or 50 µM DRB for 5 h. Data within replicates (N=3) was normalized to the saline DMSO treatment and presented as mean ± SEM analyzed using one-way ANOVA with Tukey’s post-hoc test. Specific reduction in MDM2 levels was observed in cells treated with cisplatin and DRB compared to cells treated with cisplatin and DMSO (vehicle) (** P < 0.001). b, ChIP-qPCR data showing that H2Bub1 enrichment is significantly decreased in A2780 cells following DNA damage with cisplatin and treatment with the transcriptional inhibitor DRB in p53 target genes (+, primers within the coding region; -, primers downstream of the coding region). Data within replicates was normalized to the experimental mean and presented as mean ± SEM (N = 3) analyzed using one-way ANOVA with Tukey’s post-hoc test (** P < 0.001) (JPG 705 kb)
Rights and permissions
About this article
Cite this article
Cole, A.J., Dickson, KA., Liddle, C. et al. Ubiquitin chromatin remodelling after DNA damage is associated with the expression of key cancer genes and pathways. Cell. Mol. Life Sci. 78, 1011–1027 (2021). https://doi.org/10.1007/s00018-020-03552-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-020-03552-5