Abstract
Oncogenic mutants of c-Kit are often found in mastocytosis, gastrointestinal stromal tumors and acute myeloid leukemia. The activation mechanism of the most commonly occurring mutation, D816V in exon 17 of c-Kit, has been well-studied while other mutations remain fairly uncharacterized in this respect. In this study, we show that the constitutive activity of the exon 11 mutant V560D is weaker than the D816V mutant. Phosphorylation of downstream signaling proteins induced by the ligand for c-Kit, stem cell factor, was stronger in c-Kit/V560D expressing cells than in cells expressing c-kit/D816V. Although cells expressing c-Kit/V560D showed increased ligand-independent proliferation and survival compared to wild-type c-Kit-expressing cells, these biological effects were weaker than in c-Kit/D816V-expressing cells. In contrast to cells expressing wild-type c-Kit, cells expressing c-Kit/V560D were independent of Src family kinases for downstream signaling. However, the independence of Src family kinases was not due to a Src-like kinase activity that c-Kit/D816V displayed. Point mutations that selectively block the association of PI3 kinase with c-Kit/V560D inhibited ligand-independent activation of the receptor, while inhibition of the kinase activity of PI3 kinase with pharmacological inhibitors did not affect the kinase activity of the receptor. This suggests a lipid kinase-independent key role of PI3 kinase in c-Kit/V560D-mediated oncogenic signal transduction. Thus, PI3 kinase is an attractive therapeutic target in malignancies induced by c-Kit mutations independent of its lipid kinase activity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The type III receptor tyrosine kinase c-Kit belongs to the same subfamily as the platelet-derived growth factor (PDGF) receptors, Fms-like tyrosine kinase 3 (FLT3), and the macrophage colony stimulating factor (M-CSF) receptor. C-Kit plays important roles in hematopoiesis, melanogenesis, and gametogenesis under normal physiological conditions [1]. Binding of its ligand, stem cell factor (SCF) induces dimerization of c-Kit followed by activation of its intrinsic tyrosine kinase activity. This leads to phosphorylation of key tyrosine residues that constitute binding sites for Src homology 2 domain containing signaling proteins. These events result in the phosphorylation and activation of downstream signaling molecules that mediate the biological effects, such as cell proliferation, survival and migration.
Mutations of c-Kit have been identified in around 90 % of mastocytosis and 80 % of gastrointestinal stromal tumor (GIST) cases. It also exists, less commonly, in sub-group of acute myeloid leukemia (AML), in melanoma and in germ cell tumors such as seminomas and ovarian dysgerminomas [1]. Ligand-independent activation of oncogenic mutants of receptor tyrosine kinases has long been recognized as important mediators of oncogenesis. However, we and others have shown that some oncogenic mutants of receptor tyrosine kinases, in addition to being constitutively active, also gain altered substrate specificity [2, 3].
The Src family kinases are a group of non-receptor tyrosine kinases that are important signal mediators in receptor signaling. Activation of wild-type c-Kit is partially dependent on the activity of Src family kinases, and inhibition of Src family kinases leads to decreased c-Kit activation [4]. The D816V mutation is the most commonly identified and well characterized mutation of c-Kit in malignancies. We have previously found that c-Kit/D816V gains a Src family kinase-like activity and thus circumvents a requirement of Src family kinases for its activation and cell transformation [2]. This suggests that oncogenic c-Kit mutations, in addition to being ligand-independent, have different mechanisms of activation and signal transduction compared to wild-type c-Kit.
PI3 kinases are intracellular lipid kinases that are involved in receptor signal transduction under both normal and pathological situations. Unlike Src family kinases, PI3 kinases play an important role in c-Kit/D816V-mediated cell transformation which has been shown to be independent of its lipid kinase activity [5].
Since there is a difference between the way wild-type c-Kit and c-Kit-D816V utilize Src family kinases and PI3 kinases for receptor activation and downstream cellular signaling, it is of interest to investigate the role of these two kinase families in the signal transduction and cell transformation mediated by other oncogenic c-Kit mutants. By introducing mutations in c-Kit and by using selective inhibitors, we observed that the commonly identified exon 11 mutation V560D of c-Kit is not dependent on Src family kinases for its signaling. In contrast, the ligand-independent activation of c-Kit/V560D is totally dependent on an intact binding site for PI3 kinase in c-Kit but is independent of its lipid kinase activity. These results confirm that oncogenic c-Kit mutants have different mechanisms of both activation and signaling than wild-type c-Kit.
Materials and methods
Kits and reagents
The PI3-kinase inhibitor LY294002 and Src Family Kinase inhibitor SU6656 were from Sigma-Aldrich (St. Louis, MO), another PI3 kinase inhibitor GDC0941 was from Apexbio (Boston, MA). Lipofectamine 2000 was from Life Technologies (Carlsbad, CA). Chemiluminescent HRP substrate was from Millipore (Billerica, MA). QuikChange mutagenesis kit was from Agilent Technologies (Santa Clara, CA). Annexin V-PE apoptosis detection kit was from BD Biosciences (San Jose, CA). [γ-32P]ATP and protein G-Sepharose beads were from GE Healthcare (Little Chalfont, UK). All the reagents and kits were used according to the manufacturer’s instructions.
Cytokines and antibodies
Recombinant human SCF was purchased from Prospec Tany (Rehovot, Israel). The rabbit antibody KitC1, recognizing the C-terminal tail of human c-Kit, was purified as described [6]. Antibodies recognizing pTyr-568, pTyr-703, pTyr-Y721, and pTyr-936 in human c-Kit were previously described [2]. Antibodies against Akt, pErk and SHP2 were from Santa Cruz Biotechnology (Texas, TX). pAkt antibody was from Epitomics (Burlingame, CA). β-actin antibody were from Sigma-Aldrich (St. Louis, MO). SHC antibody was from BD Biosciences (San Jose, CA). Phosphotyrosine antibody 4G10 and Gab2 antibody was from Millipore (Billerica, MA). PE labeled c-Kit antibody (104D2) was from Biolegend (San Diego, CA). Horseradish peroxidase-conjugated anti-rabbit and anti-mouse secondary antibodies were from Life Technologies (Carlsbad, CA).
Cell culture
EcoPack virus packaging cell line (Clontech, Mountainview, CA) was grown in Dulbecco’s modified Eagle’s medium supplemented with 10 % fetal bovine serum, 100 units/ml penicillin and 100 µg/ml streptomycin. Ba/F3 cells (DSMZ, Braunschweig, Germany) were grown in RPMI 1640 medium supplemented with 10 % heat-inactivated fetal bovine serum, 100 units/ml penicillin and 100 µg/ml streptomycin, and 10 ng/ml recombinant murine IL-3 as recommended previously [7]. To establish Ba/F3 cell lines expressing c-Kit, EcoPack cells were transfected with either wild-type or mutant of c-Kit constructs in pMSCVpuro vector. Supernatants were collected to infect Ba/F3 cells followed by 2-weeks selection in 1.2 μg/ml puromycin. Expression levels of c-Kit were confirmed by flow cytometry and immunoblotting. C-Kit expressing Ba/F3 cells were grown in the same medium as untransfected Ba/F3 cells.
Cell survival and proliferation assay
Cell survival and proliferation assay was performed as described [5].
Cell stimulation, immunoprecipitation and Western blotting
Cell stimulation, immunoprecipitation and Western blotting were performed as described [5].
In vitro kinase activity assay
Ba/F3 cells were starved for 4 h in medium without serum and IL-3 and subsequently stimulated with SCF (100 ng/ml) for 5 min. Thereafter, cells were washed once in ice-cold PBS, lysed, immunoprecipitated with the KitC1 antibody, and processed for in vitro kinase assay as described [8] with the exception that Src optimal peptide [9] was used as substrate.
Results
The V560D mutant is less oncogenic than the D816V mutant of c-Kit
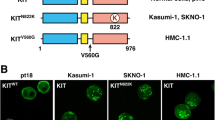
The exon 11 mutation V560D of c-Kit is frequently identified in GISTs [10] while the exon 17 mutation D816V of c-Kit is frequently identified in mastocytosis and less frequently in certain types of AML [1]. To compare the transforming ability of these two types of mutants, we transfected Ba/F3 cells, which lack endogenous c-Kit expression, with either wild-type c-Kit or c-Kit mutants. Cell surface expression of c-Kit was shown to differ between the three different mutant cell lines. While cells expressing wild-type c-Kit displayed the highest level of cell surface expression, c-Kit/V560D had intermediate surface expression, and cells expressing c-Kit/D816V displayed the lowest cell surface expression (Fig. 1a). C-Kit is expressed as a mature, fully glycosylated 145KD and an immature, partially glycosylated 125 kD protein [11], the ratio between the two c-Kit protein bands are also different between wild-type c-Kit and the two c-Kit mutants. The 145 kD band is dominant in cells expressing wild-type c-Kit while the two bands are more equal in intensity in the mutants (Fig. 1b). To investigate the oncogenic potential of these mutants, we examined cell survival and cell proliferation of transfected Ba/F3 cells in the presence or absence of SCF. Cell survival (Fig. 1c) and cell proliferation (Fig. 1d) of cells expressing wild-type c-Kit were dependent on the presence of SCF. In contrast, c-Kit/D816V supported both cell survival and cell proliferation independent of SCF stimulation. In comparison, Ba/F3 cells expressing c-Kit/V560D showed moderate cell survival and weak cell proliferation regardless of SCF stimulation, suggesting a weaker transforming ability of c-Kit/V560D compared with c-Kit/D816V.
Cell surface expression of c-Kit in Ba/F3 cells, cell survival and cell proliferation of Ba/F3 cells expressing wild-type c-Kit and c-Kit mutants. a Cell surface expression of c-Kit in Ba/F3 cells stably transfected with either wild-type c-Kit, c-Kit/V560D or c-Kit/816V was measured by flow cytometry (gray curve isotype control, black curve anti-c-Kit). b Ba/F3 cells expressing c-Kit were lysed and subjected to immunoprecipitation using a c-Kit antibody, and thereafter subjected to Western blotting using c-Kit antibody. The signal intensity of the 145 kD band and 125 kD band were quantified, the ratio between the 145 kD band and 125 kD band was calculated. c, d Ba/F3 cells expressing wild-type c-Kit, c-Kit/V560D or c-Kit/816V were analyzed for their proliferative response to SCF. Cells were washed free from IL-3, and subsequently seeded at a density of 70,000 cells/mL in 6-well plates. Thereafter cells were SCF-stimulated for 48 h (black column) or left untreated (empty column) as control. Living cells were counted under the microscope to examine cell proliferation (c) or stained with the apoptosis (d) detection kit (BD biosciences) and subsequently subjected to flow cytometry
The activation kinetics of c-Kit/V560D is different from that of c-Kit/D816V
To investigate the mechanism behind the weak transforming ability of c-Kit/V560D, c-Kit activation was compared between wild-type c-Kit, c-Kit/V560D and c-Kit/D816V expressing cells. Both c-Kit/V560D and c-Kit/D816V can induce ligand-independent activation of the receptor (Fig. 2a). It is well established that Src family kinases play a crucial role in the activation of wild-type c-Kit. The binding site of Src family kinases in c-Kit is Tyr568 [12]. By using a phospho-specific antibody, we show that Tyr568 is strongly phosphorylated in cells expressing c-Kit/V560D independent of SCF stimulation. In contrast, phosphorylation of the Grb2 binding sites Tyr703 and Tyr936 [13], and PI3 kinase binding site Tyr721 [14] was weaker in c-Kit/V560D than in c-Kit/D816V. In addition, the total phosphorylation of c-Kit/V560D was weaker than that of c-Kit/D816V. Similar to the receptor activation, the ligand-independent phosphorylation of downstream signaling molecules such as Akt and Erk was also weaker in c-Kit/V560D-expressing cells compared to c-Kit/D816V-expressing cells (Fig. 2b), although ligand stimulation transiently enhanced phosphorylation further in c-Kit/V560D expressing cells. These results suggest that c-Kit/V560D can weakly activate downstream signaling through Grb2 and PI3 kinase leading to weaker activation of Akt and Erk signaling as well as weaker cell proliferation and survival. The adapter proteins Gab2 and SHC are important intermediates in the activation of both Akt and Erk [15, 16], and the protein tyrosine phosphatase SHP2 is involved in Erk activation [17]. By using phosphotyrosine antibodies, c-Kit/V560D was shown to be able to phosphorylate Gab2, SHC and SHP2 even in the absence of ligand stimulation and similar to Akt and Erk phosphorylation, phosphorylation of these three proteins was weaker than that of c-Kit/D816V expressing cells (Fig. 2c).
Activation of wild-type c-Kit and c-Kit mutants, and downstream signaling pathways. a Western blot analysis using phospho-specific antibodies against known c-Kit phosphorylation sites. Ba/F3 cells expressing c-Kit were starved of serum and IL-3 for 4 h followed by SCF (100 ng/ml)-stimulation for 0, 2, 5, 15 or 30 min. Lysates were subjected to immunoprecipitation using a c-Kit antibody, and thereafter subjected to Western blotting using the following antibodies: pY568, pY703, pY721, pY936 and anti-phosphotyrosine antibodies. C-Kit antibody was used as control. b Phosphorylation of Akt and Erk in whole cell lysates (from the experiment 2A) was investigated by Western blot analysis using the following antibodies: anti-phosphoAkt (Ser473), anti-Akt, anti-phosphoErk (Thr202/Tyr204), anti-Erk and anti-β-actin. c Ba/F3 cells expressing c-Kit were starved of serum and IL-3 for 4 h followed by stimulation with SCF (100 ng/ml) for 2 min or left untreated. Lysates were subjected to immunoprecipitation using antibody against Gab2, SHC and SHP2, respectively. Tyrosine phosphorylation of Gab2, SHC and SHP2 was evaluated by Western blot analysis using anti-phosphotyrosine antibodies. Phosphorylation of Gab2, SHC and SHP2 was quantified and normalized against total protein
The kinase activity of c-Kit/V560D is not dependent on Src family kinases
Src family kinases are important for the activation of wild-type c-Kit [4]. We have previously found that the D816V mutation of c-Kit circumvents a requirement of Src family kinases for its activation of c-Kit and downstream signal transduction [2]. Thus, we sought to investigate whether the V560D mutant of c-Kit is still dependent on Src family kinases. By using a selective Src family kinase inhibitor, SU6656, we showed that both ligand-independent and ligand-dependent activation of c-Kit/V560D was independent on the kinase activity of Src family kinases (Fig. 3a). Likewise, phosphorylation of downstream signaling molecules such as Akt and Erk were not inhibited by the Src family kinase inhibitor SU6656 (Fig. 3b).
Src family kinases are not required for the activation of c-Kit/V560D and downstream signal transduction. a Ba/F3 cells expressing wild-type c-Kit, c-Kit/V560D and c-Kit/816V were starved of serum and IL-3 for 4 h, thereafter cells were pre-incubated with the Src inhibitor SU6656 (2 μm) for 30 min before SCF stimulation (100 ng/ml) for 2 min followed by immunoprecipitation using a c-Kit antibody. After SDS-PAGE and electrotransfer, membranes were probed with phosphotyrosine antibody and reprobed with c-Kit antibody. b Phosphorylation of Akt and Erk in whole cell lysates was investigated by Western blot analysis as in Fig. 2
C-Kit/V560D has not gained Src-like substrate specificity
We have previously found that c-Kit/D816V has gained substrate specificity similar to Src family kinases and that it thereby loses its dependence on Src family kinases for its activation and downstream signaling. Similar to c-Kit/D816V, the V560D mutant of c-Kit has also lost its dependence on Src family kinases for its activation and downstream signaling (Fig. 3). We, therefore, wanted to investigate whether also c-Kit/V560D, similar to c-Kit/D816V, has gained altered substrate specificity. Using an in vitro kinase assay utilizing the Src optimal peptide as a substrate, we could show that c-Kit/D816V displays substrate specificity similar to Src family kinases while both wild-type c-Kit and c-Kit/V560D do not show any such activity even in the presence of ligand (Fig. 4). These results indicate that the mechanism of activation and signaling is different between c-Kit/V560D and c-Kit/D816V.
C-Kit/V560D does not gain Src family kinase activity. Ba/F3 cells expressing wild-type c-Kit, c-Kit/V560D or c-Kit/816V were starved of serum and IL-3 for 4 h, stimulated with SCF (100 ng/ml) for 2 min. Cells were then lysed and immunoprecipitated using a c-Kit antibody. The immunoprecipitated c-Kit proteins were used in a kinase assay together with Src optimal peptide as substrate. the phosphorylation of the substrate by wild-type c-Kit or c-Kit/D816V was detected by Fuji FLA 3000. Signal intensity from multiple blots was quantified by the Multigauge software. Error bars indicate standard deviation
Ligand-independent activation of c-Kit/V560D is dependent on PI3 kinase in a lipid kinase-independent manner
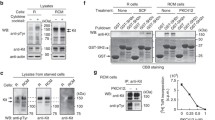
PI3 kinase plays an important role in the ligand-independent signaling by c-Kit/D816V, and c-Kit/D816V-mediated cell transformation is independent of its lipid kinase activity [5]. PI3 kinase binds to pY721 in c-Kit [18] and the binding requires the motif pYXXM in c-Kit [19]. By introducing the M724A mutation in c-Kit/V560D to block the direct PI3 kinase binding without affecting c-Kit tyrosine phosphorylation, we show that the ligand-independent activation of c-Kit/V560D is totally blocked although ligand can still activate the receptor (Fig. 5a). Ligand-dependent phosphorylation of PI3 kinase downstream signaling component Akt was partially reduced other than complete diminution, while PI3 kinase independent component Erk phosphorylation remained unchanged (Fig. 5b). These findings suggest that the c-Kit/V560D mutant can transduce signals without recruiting PI3 kinase but PI3 kinase association is required for constitutive activation of the receptor. In order to elucidate whether this effect is dependent on the lipid kinase activity of PI3 kinase, the effect of a PI3 kinase inhibitor on the activation of c-Kit/V560D was examined. The PI3 kinase inhibitor LY294002 did not block the activation of c-Kit/V560D independent of SCF stimulation (Fig. 5c). Furthermore, Akt phosphorylation was inhibited in the presence of LY294002 (Fig. 5d) indicating that PI3 kinase is activated by c-Kit/V560D. A cell proliferation assay in the presence of Src family kinase and PI3 kinase inhibitors further confirmed lipid kinase independent role of PI3 kinase in c-Kit/V560D signaling (Fig. 6).
Ligand-independent activation of c-Kit/V560D is dependent on PI3 kinase in a lipid kinase independent manner. a Ba/F3 cells expressing wild-type c-Kit, c-Kit/M724A, c-Kit/V560D or c-Kit/V560D/M724A were starved of serum and IL-3 for 4 h followed by SCF(100 ng/ml)-stimulation for 0, 2, 5, 15 or 30 min. Lysates were subjected to immunoprecipitation using a c-Kit antibody, and thereafter subjected to Western blotting using anti-phosphotyrosine antibodies and c-Kit antibodies. b Phosphorylation of Akt and Erk in whole cell lysates was investigated by Western blot analysis using the following antibodies: anti-phosphoAkt (Ser473), anti-Akt, anti-phosphoErk (Thr202/Tyr204), anti-Erk and anti-β-actin. c Ba/F3 cells expressing wild-type c-Kit or c-Kit/V560D were starved of serum and IL-3 for 4 h, thereafter cells were pre-incubated with the PI3 kinase inhibitor PY294002 (2 μM) for 30 min before SCF stimulation (100 ng/ml) for 2 min followed by immunoprecipitation using a c-Kit antibody. After SDS-PAGE and electrotransfer, membranes were probed with phosphor–tyrosine antibody and reprobed with c-Kit antibody. d Phosphorylation of Akt and Erk in whole cell lysates were investigated using Western blotting
Ba/F3 cells expressing wild-type c-Kit, c-Kit/V560D or c-Kit/D816V were analyzed for their proliferative response in the presence of Src family kinase inhibitor or PI3 kinase inhibitor. Cells were washed free from IL-3, and subsequently seeded at a density of 70,000 cells/mL in 6-well plates. Thereafter cells were SCF-stimulated for 48 h (black column) or left untreated (empty column) as control in the presence or absence of Src family kinase inhibitor SU6656 (2 µM) or PI3 kinase inhibitor LY294002 (10 µM) or GDC0941 (0.5 µM). Living cells were counted under the microscope to examine cell proliferation
Discussion
Mutations in c-Kit have been identified in exon 9, 11, 13 and 17 in different malignancies. In GISTs, most of c-Kit mutations were mapped to exon 11, and to a lesser extent also to exon 9 and 13 [20]. In mastocytosis, the D816V mutation in exon 17 of c-Kit is the most commonly identified mutation, found in about 80–90 % of patients [1]. The different mutants of c-Kit have different sensitivity to the current clinically approved c-Kit inhibitors. Mutations in exon 11 of c-Kit usually respond well to imatinib while c-Kit/D816V is insensitive to imatinib. After treatment with imatinib, relapsed GISTs many times gain secondary mutations in exon 13 and 17 of c-Kit that are resistant to imatinib [20, 21]. The difference in response of c-Kit mutations to inhibitors is most likely partially due to differences in activation mechanism. It is known that imatinib binds to the inactive kinase conformation of c-Kit [22], while c-Kit/D816V is constitutively in its active conformation [23] and thus does not bind imatinib. Moreover, different c-Kit mutants display considerable difference in intrinsic kinase activity. For example, c-Kit/D816V has a higher intrinsic kinase activity than c-Kit/V560D [21]. Therefore it is important to elucidate the detailed activation mechanism of these mutants to design more efficient drugs.
Since Src family kinases are very important in wild-type c-Kit signaling, it is tempting to consider them as therapeutic targets in tumors driven by c-Kit mutant. However, we have previously found that the c-Kit/D816V mutant circumvents a requirement for Src family kinases by displaying altered substrate specificity. For that reason, it is likely that Src family kinase inhibitor will probably not be useful for the treatment of malignancies expressing c-Kit/D816V. It is of importance to investigate whether the independency on Src family kinases of receptor activation and signaling is a common mechanism among other activating c-Kit mutants. By using inhibitors and generating oncogenic mutants of c-Kit, we demonstrate that the commonly occurring c-Kit mutation V560D in GISTs also is able to circumvent a requirement of Src family kinases for its signal transduction. These results further suggest that the use of Src family kinase inhibitors in the treatment of cancers carrying c-Kit mutations will not be a viable option.
PI3-kinases are lipid kinases that play important roles in cell survival, proliferation and metabolism. Overactivation of PI3 kinase contributes to transformation in several malignancies. As with Src family kinases, PI3-kinases are also attractive treatment targets for cancer. Various small molecule inhibitors have been developed and taken into clinical trials [24].
PI3 kinases contribute to wild-type c-Kit-mediated cell survival and proliferation. Ligand-dependent activation of wild-type c-Kit is not dependent on PI3-kinases [25]. In contrast, the most commonly identified c-Kit mutant, D816V, has been shown to be dependent on PI3 kinase for its transforming ability [18], thus making PI3-kinase an attractive target in the treatment of malignancies carrying the c-Kit/D816V mutant. In addition to its lipid kinase activity, we have recently demonstrated that the important role of PI3-kinase in c-Kit/D816V mediated cell transformation is independent on its lipid kinase activity [5]. Since currently available PI3-kinase inhibitors block the lipid kinase activity and no other functions of PI3-kinase, such as binding to other signal transduction molecules, they are probably of low clinical efficacy in this type of malignancies.
Mutations in the c-Kit gene lead to activation of its kinase activity by different mechanisms and display differential sensitivity to c-Kit inhibitors. It is interesting to know whether the dependence on PI3 kinases is a common mechanism among all c-Kit mutants. By introducing mutations in c-Kit that block the association of PI3-kinase with c-Kit, we showed that the ligand-independent activation of c-Kit/V560D is totally dependent on PI3-kinase. The loss of PI3-kinase binding completely abolishes ligand independent activation of c-Kit/V560D. As is the case with c-Kit/D816V, the contribution of PI3-kinases to ligand-independent activation of c-Kit/V560D is independent of its lipid kinase activity. These results suggest a new activity or function of PI3-kinase in addition to its lipid kinase activity.
Currently available PI3 kinase inhibitors are based on the inhibition of its lipid kinase activity. However, as we showed in this study, PI3-kinase inhibitors cannot inhibit the activation of c-Kit mutants and downstream signaling pathways and they will probably not be efficient in the treatment of malignancies carrying c-Kit mutations. To identify the novel activity of PI3 kinase and develop new inhibitors against the non-lipid kinase activity will be a new direction. The novel inhibitors of PI3 kinases might be useful in the treatment of malignancies driven by oncogenic mutants of c-Kit.
References
Lennartsson J, Rönnstrand L (2012) Stem cell factor receptor/c-Kit: from basic science to clinical implications. Physiol Rev 92:1619–1649. doi:10.1152/physrev.00046.2011
Sun J, Pedersen M, Rönnstrand L (2009) The D816V mutation of c-Kit circumvents a requirement for Src family kinases in c-Kit signal transduction. J Biol Chem 284:11039–11047. doi:10.1074/jbc.M808058200
Hayakawa F, Towatari M, Kiyoi H et al (2000) Tandem-duplicated Flt3 constitutively activates STAT5 and MAP kinase and introduces autonomous cell growth in IL-3-dependent cell lines. Oncogene 19:624–631. doi:10.1038/sj.onc.1203354
Voytyuk O, Lennartsson J, Mogi A et al (2003) Src family kinases are involved in the differential signaling from two splice forms of c-Kit. J Biol Chem 278:9159–9166. doi:10.1074/jbc.M211726200
Sun J, Mohlin S, Lundby A et al (2014) The PI3-kinase isoform p110delta is essential for cell transformation induced by the D816V mutant of c-Kit in a lipid-kinase-independent manner. Oncogene 33:5360–5369. doi:10.1038/onc.2013.479
Blume-Jensen P, Siegbahn A, Stabel S, Heldin CH, Rönnstrand L (1993) Increased Kit/SCF receptor induced mitogenicity but abolished cell motility after inhibition of protein kinase C. EMBO J 12:4199–4209
Kazi JU, Sun J, Rönnstrand L (2013) The presence or absence of IL-3 during long-term culture of Flt3-ITD and c-Kit-D816V expressing Ba/F3 cells influences signaling outcome. Exp Hematol 41:585–587. doi:10.1016/j.exphem.2013.03.005
Hansen K, Johnell M, Siegbahn A et al (1996) Mutation of a Src phosphorylation site in the PDGF beta-receptor leads to increased PDGF-stimulated chemotaxis but decreased mitogenesis. EMBO J 15:5299–5313
Songyang Z, Carraway KL 3rd, Eck MJ et al (1995) Catalytic specificity of protein-tyrosine kinases is critical for selective signalling. Nature 373:536–539. doi:10.1038/373536a0
Gounder MM, Maki RG (2011) Molecular basis for primary and secondary tyrosine kinase inhibitor resistance in gastrointestinal stromal tumor. Cancer Chemother Pharmacol 67(Suppl 1):S25–S43. doi:10.1007/s00280-010-1526-3
Brahimi-Adouane S, Bachet JB, Tabone-Eglinger S et al (2013) Effects of endoplasmic reticulum stressors on maturation and signaling of hemizygous and heterozygous wild-type and mutant forms of KIT. Mol Oncol 7:323–333. doi:10.1016/j.molonc.2012.10.008
Lennartsson J, Blume-Jensen P, Hermanson M, Pontén E, Carlberg M, Rönnstrand L (1999) Phosphorylation of Shc by Src family kinases is necessary for stem cell factor receptor/c-kit mediated activation of the Ras/MAP kinase pathway and c-fos induction. Oncogene 18:5546–5553. doi:10.1038/sj.onc.1202929
Thömmes K, Lennartsson J, Carlberg M, Rönnstrand L (1999) Identification of Tyr-703 and Tyr-936 as the primary association sites for Grb2 and Grb7 in the c-Kit/stem cell factor receptor. Biochem J 341(Pt 1):211–216
Serve H, Hsu YC, Besmer P (1994) Tyrosine residue 719 of the c-kit receptor is essential for binding of the P85 subunit of phosphatidylinositol (PI) 3-kinase and for c-kit-associated PI 3-kinase activity in COS-1 cells. J Biol Chem 269:6026–6030
Adams SJ, Aydin IT, Celebi JT (2012) GAB2—a scaffolding protein in cancer. Mol Cancer Res 10:1265–1270. doi:10.1158/1541-7786.MCR-12-0352
Wills MK, Jones N (2012) Teaching an old dogma new tricks: twenty years of Shc adaptor signalling. Biochem J 447:1–16. doi:10.1042/BJ20120769
Grossmann KS, Rosario M, Birchmeier C, Birchmeier W (2010) The tyrosine phosphatase Shp2 in development and cancer. Adv Cancer Res 106:53–89. doi:10.1016/S0065-230X(10)06002-1
Hashimoto K, Matsumura I, Tsujimura T et al (2003) Necessity of tyrosine 719 and phosphatidylinositol 3′-kinase-mediated signal pathway in constitutive activation and oncogenic potential of c-kit receptor tyrosine kinase with the Asp814Val mutation. Blood 101:1094–1102. doi:10.1182/blood-2002-01-0177
Backer JM, Myers MG Jr, Shoelson SE et al (1992) Phosphatidylinositol 3′-kinase is activated by association with IRS-1 during insulin stimulation. EMBO J 11:3469–3479
Corless CL, Barnett CM, Heinrich MC (2011) Gastrointestinal stromal tumours: origin and molecular oncology. Nat Rev Cancer 11:865–878. doi:10.1038/nrc3143
Gajiwala KS, Wu JC, Christensen J et al (2009) KIT kinase mutants show unique mechanisms of drug resistance to imatinib and sunitinib in gastrointestinal stromal tumor patients. Proc Natl Acad Sci USA 106:1542–1547. doi:10.1073/pnas.0812413106
Mol CD, Dougan DR, Schneider TR et al (2004) Structural basis for the autoinhibition and STI-571 inhibition of c-Kit tyrosine kinase. J Biol Chem 279:31655–31663. doi:10.1074/jbc.M403319200
Foster R, Griffith R, Ferrao P, Ashman L (2004) Molecular basis of the constitutive activity and STI571 resistance of Asp816Val mutant KIT receptor tyrosine kinase. J Mol Gr Model 23:139–152. doi:10.1016/j.jmgm.2004.04.003
Kurtz JE, Ray-Coquard I (2012) PI3 kinase inhibitors in the clinic: an update. Anticancer Res 32:2463–2470
Sun J, Pedersen M, Rönnstrand L (2008) Gab2 is involved in differential phosphoinositide 3-kinase signaling by two splice forms of c-Kit. J Biol Chem 283:27444–27451. doi:10.1074/jbc.M709703200
Acknowledgments
This research was supported by grants from the Swedish Cancer Society, the Swedish Research Council, the Strategic Cancer Research Program at Lund University, BioCARE, ALF governmental clinical grant, Stiftelsen Olle Engkvist Byggmästare, Alfred Österlund Foundation, Gunnar Nilsson Cancer Society, Kungliga Fysiografiska Sällskapet i Lund, Ollie and Elof Ericssons Stiftelse, Åke-Wiberg Stiftelse, Lars Hiertas Minne Stiftelse, Harald Jeanssons Stiftelse and Harald och Greta Jeanssons Stiftelse, Swedish Childhood Cancer Foundation, MAS Cancer Foundation and SUS Hospital Funds.
Conflict of interest
The authors declare that they have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lindblad, O., Kazi, J.U., Rönnstrand, L. et al. PI3 kinase is indispensable for oncogenic transformation by the V560D mutant of c-Kit in a kinase-independent manner. Cell. Mol. Life Sci. 72, 4399–4407 (2015). https://doi.org/10.1007/s00018-015-1944-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-015-1944-9