Abstract
Objective
Histamine is an important mediator of biological functions and present in high amounts in inflammatory skin lesions which are characterised by a marked infiltration of myeloid derived cell populations. The aim of the study was to investigate the expression and function of histamine receptors, with a focus on the histamine H4 receptor (H4R) in detail during the differentiation process from monocytes to macrophages and on fully differentiated M1 macrophages.
Methods
Quantitative PCR, ELISA technique, and flow cytometry were applied to analyze expression levels of histamine receptors, of CXCL10, CCL4, CCL3, or IL-23 and of the macrophage differentiation marker CD68, respectively.
Results
We demonstrated that monocytes and fully differentiated M1 macrophages express H1R-, H2R-, and H4R mRNA which were differentially regulated during the differentiation process and in IFN-Ƴ and LPS classically activated M1 macrophages. The H3R mRNA was not expressed. During in vitro differentiation from monocytes to macrophages, the H4R agonist ST-1006 modified the M1 phenotype by up-regulating the macrophage differentiation marker CD68, by down-regulating the production of CXCL10, and by changing the morphology. In fully differentiated M1 macrophages, histamine or ST-1006 decreased the IFN-Ƴ- and LPS-induced CCL4 mRNA expression and protein production, whereas CCL3 or IL-23 production was not regulated via H4R.
Conclusions
We describe novel immunomodulatory functions of the H4R during the differentiation process of human monocyte-derived macrophages and in fully differentiated M1 macrophages. The down-regulation of Th1-related chemokines during the differentiation process or in classically activated macrophages via H4R may contribute to decreased migration of immune cells to the site of inflammation. This may have implications for the treatment of allergic diseases with H4R ligands regulating the dysbalance of Th2/Th1 polarizations in these disorders.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Monocyte-derived macrophages represent an essential component of the innate immune system and play substantial roles in the host defense by initiating a protective inflammatory response and restoring the homeostasis to avoid excessive tissue damage. The fate and functional diversity of tissue resident monocyte-derived macrophages strongly depend on chemokines, cytokines, and microbial products which constitute their local microenvironment. The resulting phenotypes of the different activated macrophages are commonly classified into M1, or referred as classically activated macrophages and M2, or referred as alternatively activated macrophages, thereby mirroring the Th1/Th2 polarization scheme in part. Th1-related cytokines like interferon gamma (IFN-Ƴ) as well as microbial stimuli such as lipopolysaccharides (LPS) polarize macrophages into an M1 phenotype, which is characterized by high expression levels of IL-12, IL-23, and low expression levels of IL-10 [1,2,3,4,5,6,7]. Macrophages accumulate in the dermis in acute or chronic inflammatory skin diseases such as atopic dermatitis (AD) or psoriasis and play a central role in regulating local inflammation by secreting cytokines and chemokines [8]. Chemokines are major players in inflammatory and in immune responses by supporting the selective recruitment and activation, e.g., of circulating monocytes or polarized T cells. On one hand, the cytokine–chemokine network created by macrophages is beneficial and contributes to the clearance of infectious agents, but on the other hand, it causes a pathological local response with persistent inflammation [9, 10].
Histamine, as a major mediator of allergic and inflammatory processes, is known to be present in lesions of chronic or relapsing inflammatory skin diseases such as AD and psoriasis [11, 12]. The biogenic amine can be generated and spontaneously secreted by numerous constitutive tissue- or immune cells. In the skin, it is stored and released in high concentrations from dermal mast cells and may modulate the phenotype and function of macrophages which are often located close to mast cells, e.g., in the perivascular dermis. We hypothesized that histamine, released during allergic reactions, contributes to the plasticity of differentiation and activation of professional phagocytes. Therefore, we investigated the functional role of histamine targeting M1 macrophages during the process of differentiation or on fully differentiated and classically IFN-Ƴ- and LPS-activated M1 macrophages.
The pleiotropic effects of histamine are mediated by activating one or more of four subtypes of histamine receptors which are differentially expressed on immune cells playing a role in inflammatory allergic conditions. The influence of histamine via H4R on cytokine or chemokine release in cells of the mononuclear lineage such as monocytes [13, 14], dendritic cells (DCs) [15], plasmacytoid dendritic cells (pDCs) [16], NK cells [17] or T cells [18, 19] has been described in the previous studies.
Here, we studied the expression levels of the histamine receptors during the differentiation process of monocyte-derived macrophages, on fully differentiated human M1 macrophages and on IFN-Ƴ and LPS classically activated M1 macrophages. To investigate the influence of histamine on the process of macrophage differentiation, we cultured monocytes during the development to macrophages in the presence of granulocyte-macrophage-colony-stimulating factor (GM-CSF) alone or in addition in the presence of histamine or the H4R agonist ST-1006. The expression of macrophage differentiation marker CD68, release of the Th1-related chemokine CXCL10, and assessment of F-actin-formation were measured in these experiments. Furthermore, we stimulated fully differentiated macrophages with histamine and an H4R specific agonist following activation with inflammatory stimuli such as IFN-Ƴ and LPS to analyze effects of histamine, in particular via the H4R, in activated M1 macrophages in a given inflammatory environmental context. We found that histamine via H4R influences the phenotype of M1 macrophages and reduces CXCL10 production [a chemokine, previously called IFN-γ-inducible protein 10 (IP-10)] when presented during the differentiation process from monocytes to macrophages. In activated human M1 macrophages, CCL4 production [a chemokine, previously called macrophage inflammatory protein-1β (MIP-1β)] was reduced by histamine and the H4R agonist.
Materials and methods
Differentiation of M1 macrophages
Residual blood samples from platelet (PLT) apheresis disposables used for routine PLT collection and of regular anonymous healthy donors served as source material for the isolation of human peripheral blood mononuclear cells (PBMCs). PBMCs were separated by density gradient centrifugation on lymphoprep (Pancoll, PAN-Biotech, Aidenbach, Germany). With a seeding density of 1 × 106 cells pro well the PBMCs were plated in three 24 well plates in Iscove’s Medium supplemented with AB serum (2,5% v/v). To attach the monocytes, the cells were incubated for 2 h at 5% C02 and 37 °C. The non-adherent cells were removed by vigorously washing the adherent cells three times with PBS. An appropriate amount of RPMI 1640, supplemented with 2 mM l-glutamine, 100 mg/ml penicillin/streptomycin, 12 mM Hepes, and 5% v/vFCS (PAN-Biotech; all other media components from Biochrom, Berlin, Germany) and 10 ng/ml GM-CSF (R&D,Wiesbaden, Germany) were added. The cells were incubated for 5 days at 5% C02 and 37 °C without medium change. The cells in the first 24 well plate were differentiated to M1 macrophages in the presence of granulocyte-macrophage colony-stimulating factor (GM-CSF), the cells in the second 24 well plate were differentiated in addition to GM-CSF with histamine, and in the third 24 well plate, the cells were differentiated in addition to GM-CSF with the H4R agonist ST-1006 including all changes of the medium. At day 5, another 50% by volume of fresh medium was added. At day 8, the respective media were completely changed, and at day 10, the fully differentiated M1 macrophages were used for experiments.
At day 10, macrophages appeared as adherent cells showing a typically morphology with a prominent nucleus, outspread cytoplasma, and a couple of pseudopodia. Fully differentiated macrophages are expected to be positive for the intracellularly expressed macrophage differentiation marker CD68.
Histamine receptor ligands
The following histamine receptor ligands were used in this study: Histamine (Alk-Scherax, Wedel, Germany) as agonist for all histamine receptors; 2-pyridylethylamine (Tocris Bioscience, Bristol, UK) as selective H1R agonist; amthamine (Tocris Bioscience, Bristol, UK) as selective H2R agonist; the H4R agonist ST-1006 (Institute of Pharmaceutical and Medicinal Chemistry, Heinrich Heine University, Duesseldorf, Germany) [20]; the selective H4R antagonist JNJ7777120 (Sigma Aldrich, Deisenhofen, Germany). All histamine receptor ligands were used at a concentration of 10 µM. In extensive previous dose finding studies, we could show that the concentration of 10 µM is optimal to demonstrate and reproduce robust H4R agonist mediated effects [21].
Microscopic imaging
Olympus IX 70 microscope (Olympus, Hamburg, Germany) with the help of the program cell P 3.4 (Olympus, Germany) was used to image the fully differentiated macrophages at day 10. We assessed cell morphology and cell shape from macrophages differentiated with GM-CSF versus macrophages which were generated with GM-CSF in the presence of histamine or ST-1006. Photomicrographs were taken at 20X magnification.
Assessment of F-actin formation
Nitrobenzoxadiazole–phallacidin (Invitrogen, Darmstadt, Germany) staining of macrophages differentiated with GM-CSF and macrophages which were generated with GM-CSF in the presence of histamine or ST-1006 was performed. Briefly, cells were carefully scraped from the culture plates and re-suspended in four plastic tubes at a concentration of 5 × 105 cells/ml in PBS lacking Ca2+ in each. This was done for the non-stimulated, histamine, or ST-1006 stimulated macrophages. Since the anaphylatoxin of the fifth component of complement (C5a) has been described to trigger actin polymerization on DCs, we used C5a as stimulus to induce formation of F-actin in monocyte-derived macrophages [22]. Macrophages were stimulated with C5a (1 µg/ml, Calbiochem, Darmstadt, Germany) for short periods of time (1, 30, and 60 s) at room temperature. Following stimulation, cells were immediately fixed after stimulation using 3.7% formaldehyde for 60 min. Nitrobenzoxadiazole–phallacidin staining was carried out as described previously [16].
Flow cytometric analysis
Fully differentiated M1 macrophages and macrophages treated with histamine or ST-1006 during the whole differentiation process were carefully scraped from the culture plates and seeded (5 × 105 cells per well) into 96 well plates. Fc receptors were blocked by incubation in a buffer containing 10 µg/ml heat-aggravated human immunoglobulin G (IgG) (Sigma, Deisenhofen, Germany). Cells were fixed, permeabilized using the BD Cytofix/Cytoperm fixation/permeabilization kit (BD Bioscience) and CD68, which is intracellularly expressed in cytoplasmic granules, was stained with anti-human CD68-APC (BioLegend), the respective isotype control was tested in parallel. Sample acquisition was performed by flow cytometry (FACS Calibur, Becton Dickinson, Heidelberg, Germany) and mean fluorescence intensities (MFI) were calculated by CellQuest Pro software (Becton Dickinson). To assess cell viability, macrophages were incubated with Annexin V-FITC and propidium iodide for 15 min (BD Bioscience) and analyzed by flow cytometry (FACS Calibur, Becton Dickinson).
RNA isolation and real-time quantitative LightCycler PCR
RNA isolation, cDNA synthesis, and PCR were performed according to the MIQE Guidelines. Fully differentiated macrophages were stimulated with 10 µM histamine, 10 µM ST-1006 for 24 h or left unstimulated. After 24 h, the cells were activated by the addition of IFN-Ƴ and LPS for further 24 h. For analysing the histamine receptor expression levels, total RNA was isolated from macrophages during the differentiation process and from fully differentiated or activated macrophages derived from healthy donors. RNA of all samples was immediately lysed with 200 µl RNA-Lysis buffer (Qiagen, Hilden, Germany) and stored at − 80 °C prior use. Total RNA was isolated including on-column digestion of with RNase-free DNase I using the RNeasy kit (Qiagen) according to the manufacturer’s instructions. Finally, the RNA was eluted in a volume of 30 µl. The RNA was checked for purity by ratio absorbance at 260/280 nm and the presence of co-purified contaminants were checked by ratio absorbance at 260/230 nm. The RNA was quantified using the NanoDrop Spectrophotometer (ThermoScientific, Massachusetts, USA). Almost equal amounts of RNA were introduced in the cDNA synthesis. The cDNA was synthesized by reverse transcription (QuantiTect reverse transcription kit, Qiagen, Germany). Real-time quantitative PCR was performed with evaluated Quantitect® primer assays for H1R (QT00199857), H2R (QT00210378), H3R (QT00210861), H4R (QT00032326), CCL4 (QT01008070) and ribosomal protein 20 (rps 20) (QT00079247) using SYBR® Green according to the manufacturer’s instructions (Qiagen, Hilden, Germany) using the LightCycler 1.5 and 480 (Roche, Mannheim, Germany).
The amount of the target mRNA relative to the amount of the reference rps 20 mRNA in the same sample was calculated using the comparative Ct method also known as the [delta] [delta] Ct method provided by the Relative Quantification Software (Roche Molecular Biochemicals). The Ct values of both the calibrator and the samples of interest are normalized to the appropriate endogenous housekeeping gene rps 20.
ELISA
Cell-free supernatants were taken from fully differentiated macrophages which were stimulated with histamine, 2-pyridylethylamine (H1R agonist), amthamine (H2R agonist) or ST-1006 for 24 h and activated by addition of IFN-Ƴ (200 ng/ml) and LPS (50 ng/ml) for further 24 h. For blocking experiments, the cells were treated with the H4R antagonist JNJ7777120 30 min before stimulation with receptor agonists. The chemokine- and cytokine production of CXCL10, CCL4, CCL3, and IL-23 were analyzed using commercially available ELISAs. The respective ELISAs were performed according to the manufacturer’s instructions (R&D Systems).
Statistics
For statistical analyses, the software GraphPad Prism Version 5.0 (San Diego, CA, USA) was used. Wilcoxon matched pairs test was performed and the median is shown in the graphs. A p value < 0.05 was regarded as statistically significant (p < 0.05 was labelled with *p < 0.01 was labelled with **p < 0.001 was labelled with ***).
Ethics
The investigation of the role of the histamine receptors in inflammatory diseases was approved by the local ethics committee of the Hannover Medical School (Vote 4253) and was conducted according to the declaration of Helsinki Principles.
Results
The H1R-, H2R-, and H4R mRNA are expressed on human monocytes and macrophages
The expression of histamine receptors on human monocytes and macrophages was examined at the mRNA level by LightCycler quantitative PCR. First, we analyzed the mRNA expression levels of histamine receptors in primary monocytes obtained from human PBMCs by 2 h adherence (day 0). Second, we tested fully differentiated macrophages at the end of the differentiation process after the cells had been cultured for 10 days. We detected H1R-, H2R-, and H4R mRNA expression in human primary monocytes, whereas we failed to detect the H3R mRNA expression (data not shown). Fully differentiated macrophages at day 10 showed significantly higher H1R- and H2R mRNA expression levels compared to monocytes after 2 h adherence (Fig. 1a, b). The mRNA expression of the H4R showed a tendency to be as well higher expressed in fully differentiated macrophages as compared to monocytes (Fig. 1c). These results indicate that the differentiation of monocytes to macrophages is associated with an overall increase of histamine receptor mRNA expression. In addition, the mRNA expression of the histamine receptors was investigated in fully differentiated M1 macrophages compared to M1 macrophages activated with IFN-Ƴ and LPS for 24 h. We observed that stimulation with LPS and IFN-Ƴ led to a significant decrease of H2R mRNA expression (Fig. 1e). In view to H1R (Fig. 1d) and H4R (Fig. 1f) we found a trend to lower mRNA expression levels in M1 macrophages after IFN-Ƴ and LPS stimulation.
H1R-, H2R-, and H4R mRNA are expressed in human monocytes and up-regulated in fully differentiated M1 macrophages. Primary human monocytes were obtained from PBMCs after 2 h adherence. M1 macrophages were differentiated from primary human monocytes in the presence of GM-CSF (10 ng/ml) for 10 days and then stimulated with IFN-Ƴ (200 ng/ml) and LPS (50 ng/ml) for 24 h or left unstimulated. A, B and C mRNA expression of the H1R, H2R and the H4R in monocytes obtained by adherence at day 0 (d 0) and in fully differentiated macrophages after a differentiation period of 10 days (d 10) are shown (a, b n = 11 experiments; c n = 8 experiments). d–f mRNA expression of the H1R, H2R and the H4R in fully differentiated- and in M1 macrophages activated with IFN-Ƴ (200 ng/ml) and LPS (50 ng/ml) for 24 h are depicted (d, e n = 12 experiments; f n = 6 experiments). Relative expression levels of the histamine receptor mRNA were assessed by real-time PCR and calculated by the [delta] [delta] Ct method. The values of both the calibrator and the samples are normalized to the housekeeping gene ribosomal protein 20 (rps20) and expressed as normalized ratio. The samples from one individual donor are connected by a line. Significant differences, as determined by the Wilcoxon signed rank test, are indicated as follows: *p < 0.05; **p < 0.01; medians are shown in the graphs. Ct method crossing point method, d day, d 0 monocytes after 2 h adherence, d 10 monocyte-derived macrophages after the differentiation period of 10 days, HXR histamine receptors, NS non-stimulated
The expression levels of macrophage differentiation marker CD68 are dose-dependently up-regulated when the H4R agonist ST-1006 is presented during the differentiation process of M1 macrophages
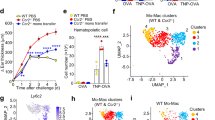
In vitro differentiation of monocytes in the presence of GM-CSF led to generation of M1 macrophages by up-regulating the differentiation marker CD68 (Fig. 2a). We differentiated monocytes to M1 macrophages with GM-CSF and in the presence of GM-CSF and histamine, 2-pyridylethylamine (H1R agonist), amthamine (H2R agonist), or ST-1006 in concentrations of 10 µM for all ligands. Expression levels of CD68 were measured by flow cytometry. We observed that the expression of CD68 was significantly up-regulated on fully differentiated M1 macrophages treated by ST-1006. The up-regulation of CD68 by ST-1006 occurs in a dose dependent manner which was most pronounced in the concentration of 10 µM (Fig. 2b). Treatment with 2-pyridylethylamine and amthamine had no effect on CD68 surface expression levels (Fig. 2c). In some cases, the expression levels of CD68 were also up-regulated by histamine (Figs. 2d, 3b). The cell viability was not affected by treatment with histamine (data not shown) or ST-1006 (Suppl. Figure 2) during the period of differentiation as assessed by Annexin V-FITC and Propidium Iodide staining.
Expression levels of the macrophage differentiation marker CD68 are up-regulated during differentiation. M1 macrophages treated with the H4R agonist ST-1006 during the differentiation process up-regulate CD68 expression in a dose dependent manner. Histamine and the H4R agonist ST-1006 decrease CXCL10 production in human M1 macrophages during this process. Primary monocytes were differentiated into M1 macrophages with GM-CSF (10 ng/ml) in the presence or absence of histamine, 2-pyridylethylamine (H1R agonist), amthamine (H2R agonist) or ST-1006 for 10 days. The expression of CD68 was analyzed by flow cytometry. a Expression of CD68 was measured at different days of the differentiation process as indicated. b Expression levels of CD68 on untreated cells and cells treated with ST-1006 in different concentrations as indicated are shown. c Expression levels of CD68 on untreated, 2-pyridylethylamine- (10 µM) or amthamine (10 µM) treated cells are shown. d Expression levels of CD68 on untreated and on histamine (10 µM) or ST-1006 (10 µM) treated cells are shown (only experiments where the non-stimulated samples showed mean fluorescence intensities < 450 were included in b–d). e Cell-free supernatants were taken at day 10 and the CXCL10 concentrations were analyzed by ELISA. Each dot of the NS samples represents one donor with paired observations in the stimulated groups. Significant differences, as determined by the Wilcoxon signed rank test, are indicated as follows: *p < 0.05; **p < 0.01; ***p < 0.001; medians are shown in the graphs. (a n = 6 experiments; b n = 8 experiments; c n = 8 experiments (cells were derived from the same donors as shown in b); d n = 13 experiments; e n = 8 experiments). NS non-stimulated, 2-Pyrid 2-pyridylethylamine (H1R agonist), Amth amthamine (H2R agonist), MFI mean fluorescence intensities
Expression levels of CD68 are up-regulated on M1 macrophages treated with histamine or the H4R agonist ST-1006 during the differentiation process. Primary monocytes were differentiated into M1 macrophages with GM-CSF (10 ng/ml) in the presence or absence of histamine (10 µM) or ST-1006 (10 µM) for 10 days. The expression of CD68 was analyzed by flow cytometry. A, macrophages were selected by an FSC/SSC gate. B, One representative histogram for histamine treated samples out of the 13 experiments which were shown in Fig. 2d, One representative histogram for ST-1006 treated samples out of the 13 experiments which were shown in Fig. 2d
The production of CXCL10 is reduced in M1 macrophages differentiated in the presence of histamine or the H4R agonist ST-1006
To assess a role for histamine or ST-1006 on chemokine production by M1 macrophages, we determined the production of the M1-related Th1 cell-attracting chemokine CXCL10 in supernatants of macrophages which were differentiated with GM-CSF or with GM-CSF supplemented with histamine, 2-pyridylethylamine (H1R agonist), amthamine (H2R agonist) or with ST-1006. Supernatants were taken from cultured M1 macrophages at day 10. We observed a significantly reduced CXCL10 production when histamine or ST-1006 had been presented during the differentiation process when compared to controls (Fig. 2e). The histamine or ST-1006 mediated down-regulation of CXCL10 occurs in a dose dependent manner and was most pronounced in the concentration of 10 µM (data not shown). Treatment with 2-pyridylethylamine (H1R agonist), amthamine (H2R agonist) had no effect CXCL10 production (data not shown).
The typical M1 macrophage morphology changes when the H4R agonist ST-1006 is presented during the differentiation process of M1 macrophages
It has been shown in previous studies that macrophages adopt different geometries in vivo in response to physical cues, e.g., specific tissue architecture or stimuli such as cytokines or microbial products which are present in the extracellular environment. In vitro differentiated macrophages display a typical morphology with a central nucleus, flatly outspread cytoplasm and multiple pseudopodia [23]. To investigate if histamine or ST-1006 is able to modulate macrophages morphology and cell shape, we differentiated monocytes to M1 macrophages with GM-CSF or with GM-CSF in the presence of histamine or ST-1006. Interestingly, the morphology of the M1 macrophages changed vigorously in the presence of ST-1006. Macrophages generated in medium supplemented with GM-CSF only or, in particular, in the presence of GM-CSF together with histamine exhibited the typical morphology with an elongated cell shape and multiple pseudopodia (Fig. 4a, b). In contrast, macrophages generated in the presence of GM-CSF and ST-1006 exhibited a round fried egg-like cell shape and obvious fewer pseudopodia (Fig. 4c).
Cell shape and organisation of actin filaments in monocyte-derived macrophages are influenced by histamine or the H4R agonist ST-1006 during the differentiation process. Primary monocytes were differentiated into M1 macrophages with GM-CSF (10 ng/ml) in the presence or absence of histamine (10 µM) or ST-1006 (10 µM). a M1 macrophages, generated in medium supplemented with GM-CSF or b in medium supplemented with GM-CSF and histamine, show an elongated cell shape. c M1 macrophages, generated in medium supplemented with GM-CSF and ST-1006, show round cell shape. Representative photomicrographs out of 4 experiments are shown. Scale bars denote 200 µm. d Induction of the formation of filamentous F-actin in response to stimulation with the anaphylatoxin C5a. Formation of filamentous F-actin as an indicator of changes in the cell shape to stimulation with 1 µg/ml C5a was determined in fully differentiated M1 macrophages as well as in histamine or in ST-1006 treated M1 macrophages. Time course of actin polymerisation to F-actin in response to the anaphylatoxin C5a is depicted. The median and ±SEM out of n = 9 independent experiments are shown
The H4R agonist ST-1006 prevents filamentous F-actin formation in M1 macrophages in response to C5a
We studied if the atypical round cell shape of M1 macrophages differentiated in the presence of ST-1006 goes along with the absence of reorganisation of the cytoskeleton. Since the formation of filamentous F-actin through polymerisation of A-actin determines cell morphology and motility, we applied an actin polymerisation assay as readout to assess changes in the cytoskeleton reorganisation. We previously reported that DCs when exposed to the anaphylatoxin C5a showed a robust actin polymerisation to F-actin [16]. Therefore, we stimulated the M1 macrophages with C5a for short periods of time and assessed formation of F-actin by flow cytometry. We detected that M1 macrophages differentiated with histamine showed a clear actin polymerisation to F-actin in response to C5a over time. Non-stimulated M1 macrophages showed a weak actin polymerisation to F-actin in response to C5a over time in some patients and, in particular, macrophages differentiated in the presence of the H4R agonist ST-1006 failed to do (Fig. 4d). This observation was related to the atypical round cell shape of the ST-1006 treated cells. During the process of F-actin formation, macrophages form filopodia which are rich in filamentous actin. The absence of F-actin formation seems to contribute to the round pan cake like cell shape of ST-1006 treated M1 macrophages.
Histamine and the H4R agonist ST-1006 dose-dependently down-regulate the expression of CCL4 in IFN-Ƴ and LPS-activated human M1 macrophages
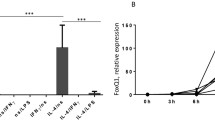
Th1-related pro-inflammatory cytokines such as IFN-Ƴ as well as microbial products like LPS polarize macrophages to an activated M1 phenotype. The influence of histamine or ST-1006 on the production of M1 related Th1 cell-attracting chemokines CXCL10, CCL3, and CCL4 and the cytokine IL-23 was investigated. We observed that stimulation of M1 macrophages with histamine or ST-1006 for 24 h before activation with IFN-Ƴ and LPS led to a down-regulation of CCL4 mRNA- and protein production (Fig. 5a, b). Pre-incubation with the selective H4R antagonist JNJ7777120 reversed the H4R-induced down-regulation of CCL4 production, showing that the effect was specific for the H4R (Fig. 5c). The down-regulation of CCL4 by histamine and ST-1006 occured dose-dependently and was most pronounced in the concentration of 10 µM (Fig. 5d, e). Pre-incubation with different concentrations of JNJ7777120 showed also an optimal blocking effect in the concentration of 10 µM (Fig. 5f). In contrast to the histamine or ST-1006 stimulated cells 2-pyridylethylamine (H1R agonist) and amthamine (H2R agonist) had no effect on the CCL4 protein production (Fig. 5g).
CCL4 mRNA expression and protein secretion are down-regulated dose-dependently by histamine or the H4R agonist ST-1006 in activated M1 macrophages. M1 macrophages were stimulated for 24 h with histamine (10 µM) or ST-1006 (10 µM) or in different concentrations as indicated and with 2-pyridylethylamine (H1R agonist) (10 µM) and amthamine (H2R agonist) (10 µM) followed by activation with IFN-Ƴ (200 ng/ml) and LPS (50 ng/ml) for 24 h. a CCL4 mRNA levels were quantified by real-time PCR calculated by the [delta] [delta] Ct method and normalized to the non-stimulated samples. b CCL4 protein production was analyzed by ELISA. c Pre-incubation with the H4R antagonist JNJ7777120 (10 µM) blocked the H4R-induced down-regulation of CCL4 production. d M1 macrophages were stimulated for 24 h with histamine in different concentrations as indicated. e M1 macrophages were stimulated for 24 h with ST-1006 in different concentrations as indicated. f Blocking experiments by pre-incubation with the H4R antagonist JNJ7777120 in different concentrations as indicated. Only experiments where a down-regulation of CCL4 via H4R was observed were included in the blocking experiments. g M1 macrophages were stimulated for 24 h with 2-pyridylethylamine (H1R agonist) (10 µM) and amthamine (H2R agonist) (10 µM). The CCL4 protein production was analyzed by ELISA. Only experiments where the non-stimulated samples had a protein secretion > 500 pg/ml were included in ELISA analysis. (a n = 7 experiments; b n = 18 experiments; c n = 6 experiments, d n = 8 experiments; e n = 9 experiments; f n = 6 experiments; g n = 4 experiments). Each dot of the NS samples represents one donor with paired observations in the stimulated groups. Significant differences as determined by the Wilcoxon signed rank test are indicated as follows: *p < 0.05; **p < 0.01; ***p < 0.001; medians are shown in the graphs. NS non-stimulated, JNJ JNJ7777120
Opposed to our observations in macrophages during the differentiation process, no consistent effect on the regulation of CXCL10 by histamine receptor agonists was detected in activated M1 macrophages (data not shown). Histamine and ST-1006 did not regulate the production of CCL3 or IL-23 in activated M1 macrophages (Suppl. Figure 1, a and b).
Discussion
Human macrophages can be found in low numbers in the circulation and are integrated or recruited into specific tissues. In different compartments of the tissues macrophages encounter diverse signals and respond with high adaptivity and functional polarization of their phenotype. Inflammatory stimuli such as the TLR4 ligand LPS and IFN-Ƴ induce an activated inflammatory phenotype in macrophages that arise from the differentiation of monocytes [2, 4].The induction of more than 400 genes and the repression of more than 700 genes in response to LPS, as described by Lang et al. [24], point to a regulatory role of macrophages in orchestrating the cellular network by differential expression of cytokines or chemokines. We hypothesized that histamine, known as an endogenous mediator tightly connected to the effector phase of allergic reactions in the skin, provides a local signal to influence macrophage differentiation and functional capacity in their niche.
In the present study, we show that H1R-, H2R-, and H4R mRNA were expressed in untouched monocytes obtained from human PBMCs by adherence. During in vitro differentiation of monocytes in the presence of GM-CSF to macrophages exhibiting a M1 phenotype, the H1R- and H2R mRNA expression were significantly up-regulated. The H4R mRNA was lower expressed in monocytes, but showed also a trend to increase the expression levels on fully differentiated macrophages.
Opposed to our results Capelo et al. [25] showed that the H2R mRNA is highly expressed on human monocytes, whereas the H1R- and H4R mRNA are mostly absent. In their study, human monocytes were isolated by CD14 immunomagnetic positive selection [25] which might influence histamine receptor expression. However, H4R mRNA was also detected in human monocytes isolated from healthy donors using CD14 positive selection in another study. Calculating the delta Ct values (normalized to two different housekeeping genes) of the H4R mRNA expression in H4R-transfected HEK293 cells versus human monocytes, the H4R mRNA was apparently also expressed in monocytes in these investigations [26]. Triggiani et al. [27] detected H1R- and H2R mRNA expression in freshly isolated monocytes obtained by negative immunomagnetic selection and showed an up-regulation of the H1R mRNA after differentiation to macrophages. The mRNA levels of the H2R were markedly reduced during differentiation of monocyte-derived marcophages. The expression profile of the H4R was not investigated in their study.
Consistent with the findings of Capelo et al. [25] we detected high donor dependent variability of the histamine receptor mRNA expression levels in monocytes and macrophages in all experiments in our study.
Furthermore, we observed that activation of fully differentiated macrophages with IFN-Ƴ and LPS led to down-regulation of mRNA expression levels of the H1R and H4R by trend. Interestingly, we could show that the mRNA expression of the H2R was significantly down-regulated by IFN-Ƴ and LPS. These findings are in part consistent with data from Capelo et al. [25]. They observed in M1 macrophages stimulated with IFN-Ƴ a down-regulation of the H1R- and H2R mRNA expression whereas the H4R mRNA was not regulated by IFN-Ƴ and expressed at very low levels. The treatment of M1 macrophages with TLR4 ligand LPS did not affect the H4R mRNA expression levels in their study [25]. Previous investigations of our research group showed in one study that IFN-Ƴ up-regulates H4R mRNA expression in a subgroup of inflammatory DCs in AD patients [28].
Our new findings in regard to the published data provide additional evidence for high variability of the mRNA expression levels of the histamine receptors: (1) in various primary immune cells which might be in part influenced by the different methods to isolate primary cells; (2) during the process of differentiation maturation of immune cells; and (3) in response to inflammatory cytokines/stimuli or in different pathological conditions such as in AD or psoriasis.
The endosomal expressed CD68 represents a common marker to monitor the differentiation process of monocytes to macrophages [29,30,31]. We observed that the expression of CD68 which characterizes fully differentiated macrophages was further up-regulated in macrophages differentiated in the presence of the H4R agonist and in some cases in the presence of histamine. The endosomal expression of CD68 is associated with a protective role of lysosomal membranes against lysosomal hydrolases [31]. The up-regulation of CD68 via H4R suggests a function of the receptor in promoting the lysosomal integrity. Down-regulation of CXCL10, a chemokine serving as biomarker for severity of various inflammatory diseases, was observed by histamine and by ST-1006 when presented during the differentiation process.
In addition, we detected that the presence of the H4R agonist during differentiation process changed the morphology of macrophages. Macrophages treated with ST-1006 developed a round cell shape when compared to the more elongated cell shape of untreated or histamine treated cells. It has been shown in former reports that macrophages display markedly different cell morphologies dependent on the activation factors, e.g., cytokines or inflammatory stimuli present in culture medium. Moreover, it was demonstrated that changes in the cell shape are caused by actin reorganisation creating F-actin filaments [23, 32]. Therefore, it was not surprising that we found formation of F-actin in response to the anaphylatoxin C5a in untreated macrophages of some patients and particularly in histamine treated macrophages, whereas we failed to detect changes in the cytoskeleton by actin polymerisation in macrophages treated with the H4R agonist during the period of differentiation. The inhibition of cytoskeleton reorganisation via H4R may reflect the lower motility of these cells which relates the morphological change in response to the H4R agonist to physiological meaning.
Since it is supposed in general that an increase in the number of the histamine receptors expressed on human immune cells might be a means to modulate the cell responsiveness to histamine, we stimulated the monocyte-derived M1 macrophages with histamine or ST-1006 at the end of the differentiation process, when the cells displayed the highest expression levels of H1R, H2R, or H4R. To investigate functional effects of histamine in an in vitro model resembling inflammatory situations, we activated the macrophages thereafter with the TLR4 ligand LPS that mimics bacterial infection and IFN-y which is secreted in high amounts from Th1 cells and NK cells in chronic inflammatory diseases.
We demonstrated that the IFN-Ƴ- and LPS-induced production of the Th1 cell-attracting chemokine CCL4 was down-regulated in fully differentiated M1 macrophages when the cells were pre-treated with histamine or ST-1006. The effects of histamine or the H4R agonist were selective as pre-incubation with the specific H4R antagonist reversed the H4R-mediated effect. However, the expression levels of CXCL10, IL-23, and CCL3 were not regulated under the same conditions.
As opposed to our findings on human macrophages, Czerner et al. [33] observed in murine bone marrow cell derived macrophages that histamine did not affect the LPS-induced up-regulation of inflammatory mediators.
Among other chemokines, CCL4 which is up-regulated in the upper compartment in skin lesions of acute AD has been linked to recruitment of inflammatory cell types into the skin. In parallel frequency of CCR5 (CCL4 receptor) bearing DCs increased after allergen application in the skin and expression of CCR5 on DCs is higher in acute than in chronic AD lesions [34]. Besides DCs, the chemokine receptor CCR5 is selectively expressed on Th1 lymphocytes [35]. The altered expression profile of CCL4 in response to histamine in M1 macrophages may limit migration of CCR5-expressing inflammatory cells, in particular of Th1 effector cells, and may present an important mechanism shifting to a more homeostatic Th2-dominated situation which could be crucial for the course of atopic diseases [36].
In this respect, our novel scientific data provide further indication that histamine, in particular via the H4R, plays a potent role in the pathophysiology of allergic diseases through tight control of Th1–Th2 mediator production favouring Th2 lymphocyte accumulation in inflamed tissues. Lastly, our results fit to a growing evidence of data [37, 38] suggesting that the H4R may serve as a therapeutic target in allergic diseases.
Abbreviations
- AD:
-
Atopic dermatitis
- DCs:
-
Dendritic cells
- H4R:
-
Histamine H4 receptor
References
Hume DA, Summers KM, Rehli M. Transcriptional regulation and macrophage differentiation. Microbiol Spectr. 2016;MCHD-0024-2015.
Mosser DM. The many faces of macrophage activation. J Leukoc Biol. 2003;73:209–12.
Stout RD, Jiang C, Matta B, Tietzel I, Watkins SK, Suttles J. Macrophages sequentially change their functional phenotype in response to changes in microenvironmental influences. J Immunol. 2005;175:342–9.
Martinez FO, Gordon S, Locati M, Mantovani A. Transcriptional profiling of the human monocyte-to-macrophage differentiation and polarization: new molecules and patterns of gene expression. J Immunol. 2006;177(10):7303–11.
Anthony RM, Urban JF Jr, Alem F, Hamed HA, Rozo CT, Boucher JL, et al. Memory T(H)2 cells induce alternatively activated macrophages to mediate protection against nematode parasites. Nat Med. 2006;12:955–60.
Biswas SK, Mantovani A. Macrophage plasticity and interaction with lymphocyte subsets: cancer as a paradigm. Nat Immunol. 2010;11:889–96.
Ginhoux F, Schultze JL, Murray PJ, Ochando J, Biswas SK. New insights into the multidimensional concept of macrophage ontogeny, activation and function. Nat Immunol. 2015;17:34–40.
Kasraie S, Werfel T. Role of macrophages in the pathogenesis of atopic dermatitis. Mediators Inflamm. 2013;2013:942375.
Mantovani A, Sica A, Sozzani S, Allavena P, Vecchi A, Locati M. The chemokine system in diverse forms of macrophage activation and polarization. Trends Immunol. 2004;25:677–86.
Shaik-Dasthagirisaheb YB, Conti P. Letter to editor: chemokine network involved in inflammatory skin diseases. Ann Clin Lab Sci. 2015;45:452–7.
Ruzicka T, Glück S. Cutaneous histamine levels and histamine releasability from the skin in atopic dermatitis and hyper-IgE-syndrome. Arch Dermatol Res. 1983;275:41–4.
Krogstad AL, Lonnroth P, Larson G, Wallin BG. Increased interstitial histamine concentration in the psoriatic plaque. J Invest Dermatol. 1997;109:632–5.
Gschwandtner M, Bunk H, Kother B, Thurmond RL, Kietzmann M, Werfel T, et al. Histamine down-regulates IL-27 production in antigen-presenting cells. J Leukoc Biol. 2012;92:21–9.
Glatzer F, Mommert S, Kother B, Gschwandtner M, Stark H, Werfel T, et al. Histamine downregulates the Th1-associated chemokine IP-10 in monocytes and myeloid dendritic cells. Int Arch Allergy Immunol. 2014;163:11–9.
Gutzmer R, Diestel C, Mommert S, Kother B, Stark H, Wittmann M, et al. Histamine H4 receptor stimulation suppresses IL-12p70 production and mediates chemotaxis in human monocyte-derived dendritic cells. J Immunol. 2005;174:5224–32.
Gschwandtner M, Mommert S, Kother B, Werfel T, Gutzmer R. The histamine H4 receptor is highly expressed on plasmacytoid dendritic cells in psoriasis and histamine regulates their cytokine production and migration. J Invest Dermatol. 2011;131:1668–76.
Mommert S, Dittrich-Breiholz O, Stark H, Gutzmer R, Werfel T. The histamine H4 receptor regulates chemokine production in human natural killer cells. Int Arch Allergy Immunol. 2015;166:225–30.
Gutzmer R, Mommert S, Gschwandtner M, Zwingmann K, Stark H, Werfel T. The histamine H4 receptor is functionally expressed on T(H)2 cells. J Allergy Clin Immunol. 2009;123:619–25.
Mommert S, Gschwandtner M, Koether B, Gutzmer R, Werfel T. Human memory Th17 cells express a functional histamine H4 receptor. Am J Pathol. 2012;180:177–85.
Sander K, Kottke T, Tanrikulu Y, Proschak E, Weizel L, Schneider EH, et al. 2,4-diaminopyrimidines as histamine H4 receptor ligands–scaffold optimization and pharmacological characterization. Bioorg Med Chem. 2009;17:7186–96.
Gschwandtner M, Koether B, Werfel T, Stark H, Gutzmer R. Profiling of histamine H4 receptor agonists in native human monocytes. Br J Pharmacol. 2013;170:136–43.
Weinmann O, Gutzmer R, Zwirner J, Wittmann M, Langer K, Lisewski M, et al. Up-regulation of C5a receptor expression and function on human monocyte derived dendritic cells by prostaglandin E2. Immunology. 2003;110:458–65.
McWhorter FY, Wang T, Nguyen P, Chung T, Liu WF. Modulation of macrophage phenotype by cell shape. Proc Natl Acad Sci USA. 2013;110:17253–8.
Lang R, Patel D, Morris JJ, Rutschman RL, Murray PJ. Shaping gene expression in activated and resting primary macrophages by IL-10. J Immunol. 2002;169:2253–63.
Capelo R, Lehmann C, Ahmad K, Snodgrass R, Diehl O, Ringleb J, et al. Cellular analysis of the histamine H4 receptor in human myeloid cells. Biochem Pharmacol. 2016;103:74–84.
Werner K, Neumann D, Buschauer A, Seifert R. No evidence for histamine H4 receptor in human monocytes. J Pharmacol Exp Ther. 2014;351:519–26.
Triggiani M, Petraroli A, Loffredo S, Frattini A, Granata F, Morabito P, et al. Differentiation of monocytes into macrophages induces the upregulation of histamine H1 receptor. J Allergy Clin Immunol. 2007;119:472–81.
Gschwandtner M, Schakel K, Werfel T, Gutzmer R. Histamine. H(4) receptor activation on human slan-dendritic cells down-regulates their pro-inflammatory capacity. Immunology. 2011;132:49–56.
Holness CL, Simmons DL. Molecular cloning of CD68, a human macrophage marker related to lysosomal glycoproteins. Blood. 1993;81:1607–13.
O’Reilly D, Greaves DR. Cell-type-specific expression of the human CD68 gene is associated with changes in pol II phosphorylation and short-range intrachromosomal gene looping. Genomics. 2007;90:407–15.
Gottfried E, Kunz-Schughart LA, Weber A, Rehli M, Peuker A, Muller A, et al. Expression of CD68 in non-myeloid cell types. Scand J Immunol. 2008;67:453–63.
Rey-Giraud F, Hafner M, Ries CH. In vitro generation of monocyte-derived macrophages under serum-free conditions improves their tumor promoting functions. PLoS One. 2012;7:e42656.
Czerner CP, Klos A, Seifert R, Neumann D. Histamine induces chemotaxis and phagocytosis in murine bone marrow-derived macrophages and RAW 264.7 macrophage-like cells via histamine H4-receptor. Inflamm Res. 2014;63:239–47.
Gros E, Bussmann C, Bieber T, Förster I, Novak N. Expression of chemokines and chemokine receptors in lesional and nonlesional upper skin of patients withatopic dermatitis. J Allergy Clin Immunol. 2009;124:753–60.
Kutukculer N, Azarsiz E, Aksu G, Karaca NE. CD4 + CD25 + Foxp3 + T regulatory cells, Th1 (CCR5, IL-2, IFN-γ) and Th2 (CCR4, IL-4, Il-13) type chemokine receptors and intracellular cytokines in children with common variableimmunodeficiency. Int J Immunopathol Pharmacol. 2016;29:241–51.
Werfel T, Allam JP, Biedermann T, Eyerich K, Gilles S, Guttman-Yassky E, et al. Cellular and molecular immunologic mechanisms in patients with atopic dermatitis. J Allergy Clin Immunol. 2016;138:336–49.
Kollmeier A, Francke K, Chen B, Dunford PJ, Greenspan AJ, Xia Y, et al. The histamine H(4) receptor antagonist, JNJ 39758979, is effective in reducing histamine-induced pruritus in a randomized clinical study in healthy subjects. J Pharmacol Exp Ther. 2014;350:181–7.
Werfel T, Lynch V, Asher A, Tsianakas A, Gupta B, Sarmiento R, et al. Allergy. 2016;71(Suppl.102):95.
Acknowledgements
The authors would like to thank Brigitta Koether and Kira Herwig for excellent technical assistance.
Funding
This study was supported by Grants from the Deutsche Forschungsgemeinschaft DFG: Gu434/6-1. Funding for this research was provided by Janssen Research & Development, LLC.
Author information
Authors and Affiliations
Contributions
SM is the primary author and analyzed the histamine receptor expression on human macrophages, conducted data collection, analysis, interpretation of the data and writing the first draft of the manuscript. LR performed most of the experiments, generating and stimulating the macrophages and performing ELISA and qPCR. HS provided the H4R-agonist ST-1006 and made contributions to the conception, design and interpretation of the data. RG and TW made significant and substantial contributions to the conception, design and interpretation of the data. All authors reviewed, revised, and approved the manuscript for publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare.
Additional information
Responsible Editor: Bernhard Gibbs.
Electronic supplementary material
Below is the link to the electronic supplementary material.


Rights and permissions
About this article
Cite this article
Mommert, S., Ratz, L., Stark, H. et al. The histamine H4 receptor modulates the differentiation process of human monocyte-derived M1 macrophages and the release of CCL4/MIP-1β from fully differentiated M1 macrophages. Inflamm. Res. 67, 503–513 (2018). https://doi.org/10.1007/s00011-018-1140-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00011-018-1140-0