Abstract
Objective
The aim of this study was to determine whether tumor necrosis factor alpha inducible protein 3 (TNFAIP3) polymorphisms confer susceptibility to rheumatoid arthritis (RA) in ethnically different populations.
Methods
The authors conducted meta-analyses on associations between the TNFAIP3 polymorphisms and RA susceptibility.
Result
A total of ten comparative studies were included in this meta-analysis, which showed an association between the two allele of rs6920220 and RA in all study subjects [odds ratio (OR) 1.216, 95% confidence interval (CI) 1.166–1.269, p < 1.0 × 10−9]. The two allele of rs6920220 was also significantly associated with RA in Europeans only (OR 1.227, 95% CI 1.175–1.282, p < 1.0 × 10−9). Meta-analysis revealed no association between the two allele of the rs10499194 polymorphism and RA in Europeans, but a significant association was found in Asians (OR 1.254, 95% CI 1.101–1.429, p = 6.7 × 10−4). Furthermore, an association was found between the two allele of rs2230926 and RA in all study subjects (OR 1.390, 95% CI 1.214–2.331, p = 1.9 × 10−6).
Conclusions
This meta-analysis confirms that the TNFAIP3 polymorphisms are associated with RA susceptibility in different ethnic groups, namely, in Europeans for rs6920220 and in Asians for rs10499194.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rheumatoid arthritis (RA) is a chronic inflammatory disease that predominantly involves synovial joints and affects up to 1% of adults worldwide. Although the etiology of RA remains unknown, a genetic component has been established by twin and family studies, in which it was estimated to contribute as much as 60% to RA liability. Human leukocyte antigen (HLA) class II molecules have been shown to be strongly associated with RA, but family studies suggest that this association accounts for only one-third of genetic susceptibility and that non-HLA genes are also involved [1].
The tumor necrosis factor alpha inducible protein 3 gene (TNFAIP3) also encodes ubiquitin-editing protein A20, which is an inhibitor of nuclear factor-κB (NF-κB) activation in several signaling pathways, including those of TNF and Toll-like receptors [2]. Furthermore, A20-deficient mice have been reported to exhibit systematic inflammation and damage to joints, and to develop autoimmunity [3]. TNFAIP3 protein participates in the negative regulation of inflammatory responses, and alterations in the activity or expression of TNFAIP3-encoded A20 may influence the pathogenesis of RA [4]. TNFAIP3 is located at 6q23, and is known to be associated with susceptibility to multiple autoimmune diseases [5]. In a genome-wide association study, it was shown that a strong association exists between the polymorphisms rs6920220 and rs10499194 and RA [6]. These polymorphisms lie between OLG3 and TNFAIP3. Rs2230926 is located in the coding region of TNFAIP3, and an amino acid substitution of Phe to Cys at position 127 in the ovarian tumor domain of TNPAIP3 has been suggested to play a role in the inhibitory function of A20 [7]. Furthermore, it has been reported that the Cys127 allele product is slightly less effective at inhibiting NF-κB activation than the Phe127 allele product [8].
Associations between TNFAIP3 polymorphisms and RA have been reported in different ethnic groups, but reported results are contradictory [9–18]. Typically, the allelic frequencies of genes often differ substantially in different ethnic groups, and thus ethnicity specific association studies are required to determine genetic associations in different populations [19–21]. In the present study, using a meta-analysis approach we investigated whether the TNFAIP3 polymorphisms, rs6920220, rs10499194, and rs2230926, contribute to RA susceptibility in different populations.
Materials and methods
Identification of eligible studies and data extraction
A literature search was conducted for studies that examined associations between the TNFAIP3 polymorphisms and RA. The MEDLINE citation index was utilized to identify articles in which the TNFAIP3 polymorphism was determined in RA patients and controls (until September 2011). In addition, all references mentioned in identified articles were reviewed to identify additional studies not indexed by MEDLINE. The following key words and subject terms were searched: “tumor necrosis factor alpha inducible protein 3,” “TNFAIP3,” “rheumatoid arthritis” and “RA”. The following information was extracted from each study: author, year of publication, ethnicity of the study population, demographics, numbers of cases and controls, and the allele frequencies of the TNFAIP3 polymorphisms.
Evaluation of publication bias
Funnel plots are used to detect publication bias, but require studies with different sizes that involve subjective judgments. Thus, we evaluated publication bias using Egger’s linear regression test [22], which measures funnel plot asymmetry on a natural logarithm scale of odds ratios (ORs).
Evaluation of statistical associations
Allele frequencies of the TNFAIP3 polymorphisms in each of the studies were determined using the allele counting method. Allelic effect contrast for the minor alleles (the two allele) versus the common alleles (the one allele) was examined, and point estimates of risk, ORs, and 95% confidence intervals (CIs) were determined for each study. Cochran’s Q-statistic was used to assess within- and between-study variations and heterogeneities [23]. Heterogeneity testing was performed to assess the probability of the null hypothesis, namely, that all studies evaluated the same effect. When a significant Q-statistic (p < 0.10) indicated heterogeneity across studies, the random effects model was used for meta-analysis, and when heterogeneity was not indicated across studies, the fixed effects model was used. The fixed effects model assumes that genetic factors have similar effects on RA susceptibility across all studies, and that observed variations between studies are caused by chance alone [24]. On the other hand, the random effects model assumes that different studies show substantial diversity, and assesses both within-study sampling errors and between-study variances [25]. The random effects model is used in the presence of significant between-study heterogeneity. We quantified the effect of heterogeneity by using a recently developed measure, namely \( I^{ 2} { = 100}\% \times (Q - {\text{df)/}}Q \) [26]. I 2 ranges between 0 and 100% and represents the proportion of inter-study variability attributable to heterogeneity rather than chance. I 2 values of 25, 50, and 75% were defined as low, moderate, and high estimates, respectively. Statistical manipulations were undertaken using the Comprehensive meta-analysis computer program (Biosta, Englewood, NJ, USA).
Results
Studies included in the meta-analysis
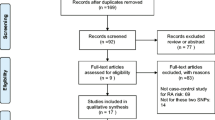
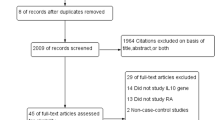
Electronic and manual searches resulted in the identification of 30 studies, and 11 were selected for full-text review based on their titles and abstracts [9–18, 27]. One study was excluded because it contained no extractable data [27]. Thus, ten studies were considered in this meta-analysis [9–18], and comprised six European, two Asian, one African–American, and one Tunisian population (Table 1). Meta-analyses were conducted separately on the rs6920220, rs10499194, and rs2230926 polymorphisms. The selected characteristics of the ten studies included are summarized in Table 1.
Frequency of the two allele of the TNFAIP3 rs6920220 polymorphism in control groups
The mean frequency of the two allele was found to be 15.9% among all controls. Asian controls had the lowest two-allele prevalence (0.067%). Frequencies ranged from 0.067 to 22.4% in the other ethnic groups, and Tunisians had the highest two-allele frequency (22.4%) (Table 2).
Meta-analysis of the associations between the TNFAIP3 polymorphisms and RA
Meta-analysis was performed on all RA patients and on RA patients in each ethnic group. A summary of the meta-analysis findings of the relations between the TNFAIP3 polymorphisms and RA is provided in Table 3. Meta-analysis showed an association between the two allele of rs6920220 and RA in all study subjects (OR 1.216, 95% CI 1.166–1.269, p < 1.0 × 10−9) (Table 3; Fig. 1). Analysis after stratification by ethnicity indicated that the two allele of rs6920220 was significantly associated with RA in Europeans (OR 1.227, 95% CI 1.175–1.282, p < 1.0 × 10−9) (Table 3). The analyses performed in each Asian, African–American, and Tunisian study showed no significant association between the two allele of rs6920220 and RA (Table 3).
Meta-analysis revealed no association between the two allele of the rs10499194 polymorphism and the risk of developing RA in all study subjects, in Europeans, or African–Americans (Table 3; Fig. 2). However, meta-analysis showed a significant association between the two allele of rs10499194 and RA in Asians (OR 1.254, 95% CI 1.101–1.429, p = 6.7 × 10−4) (Table 3; Fig. 3).
Meta-analysis revealed an association between the two allele of rs2230926 and RA in all study subjects (OR 1.390, 95% CI 1.214–2.331, p = 1.9 × 10−6) (Table 3; Fig. 3). The analysis performed in a single Asian population showed a significant association between the two allele of rs2230926 and RA (OR 1.366, 95% CI 1.188–1.571, p = 1.2 × 10−5), and the analysis in the African–American study revealed a significant association between the two allele of rs2230926 and RA (OR 1.847, 95% CI 1.053–3.241, p = 0.032) (Table 3; Fig. 3).
Heterogeneity and publication bias
Some between-study heterogeneity was found during the meta-analyses, but no evidence of heterogeneity was found for rs6920220 in all study subjects or Europeans, for rs10499194 in Asians, or for rs2230926 in all study subjects (Table 3). Funnel plots, which are usually used to detect publication bias, were difficult to correlate, presumably because of the small number of studies included. Egger’s regression test showed no evidence of publication bias in this meta-analysis of TNFAIP3 polymorphisms in any group analyzed (Egger’s regression test p > 0.1).
Discussion
In this meta-analysis, we examined evidence of associations between the rs6920220, rs10499194, and rs2230926 polymorphisms of TNFAIP3 and RA susceptibility. Our results provide evidence of strong associations between these polymorphisms and RA. Although our findings do not support associations between the TNFAIP3 polymorphisms and RA susceptibility in all ethnic groups, our analysis does reveal a significant association between the rs6920220 polymorphism and the risk of developing RA in Europeans. Furthermore, our study shows an association between RA and rs10499194 in Asians and between rs2230926 and RA in all subjects. These findings suggest that the TNFAIP3 polymorphisms are associated with the development of RA in Europeans and Asians.
The disease-associated variant of rs2230926 is a non-synonymous variant that results in a phenylalanine-to-cysteine change at residue 127 of A20, a key player in anti-inflammatory reactions. The risk allele (Cys127) leads to reduced inhibition of NF-κB activation or reduced TNFAIP3 mRNA levels [8]. These findings suggest that reduced A20 negative regulatory activity leads to excessive immune activity, and thus enhanced autoreactivity, and that rs2230926 plays a functional role in the development of RA.
The prevalence of the two allele of the rs6920220 polymorphism was calculated in different populations, and was found to vary between 0.067 and 22.4%. Its mean frequency in all controls was 15.9%, and its frequency was lowest among Asian controls and highest among Tunisian controls. Furthermore, our ethnicity-specific meta-analysis showed that an association exists between the rs6920220 polymorphism and RA in Europeans. Unfortunately, meta-analysis of the rs6920220 polymorphism was not possible in Asians, African Americans, or Tunisians due to limited data.
Several limitations of the present study require consideration. First, publication bias, heterogeneity, and confounding factors may have distorted the meta-analysis. However, most of the studies analyzed demonstrated the same directionality with respect to associations between the TNFAIP3 polymorphisms and RA. Second, only data from European and Asian patients was included in the ethnicity-specific analysis, and thus our results are applicable to only these ethnic groups. Third, in the case of rs6920220, meta-analyses of minor populations like Asians, African–Americans and Tunisians were impossible because of lack of useful data concerning the minor populations. In the meta-analysis of rs2230926, we could include only two studies (African–Americans and Asians). Concerning the allele of rs10499194, it seems to be associated with RA in Asian populations and African–Americans but not so among Europeans. However, the current evidence is too limited to draw a definitive conclusion because the number of studies included in the meta-analysis was small. Thus, this meta-analysis may have not enough power to explore the association between the TNFAIP3 polymorphisms in RA. Fourth, most of the studies included were performed in populations of European and Asian descent––African–Americans and Tunisians were the subjects of only one study apiece. Furthermore, the allelic frequencies of genes often differ substantially between populations, and thus further ethnicity-specific association studies are required to confirm genetic associations with RA susceptibility in different populations. In the present study, we performed meta-analyses on the rs6920220, rs10499194, and rs2230926 polymorphisms, and additional studies are needed to explore the roles of other TNFAIP3 polymorphisms in RA. Fifth, the rs6920220, rs10499194, and rs2230926 polymorphisms have also been reported to be associated with the severity of autoimmune diseases [28], but limited data prevented our examining associations between these polymorphisms and the clinical manifestations of RA.
In conclusion, this meta-analysis of published data confirms that the rs6920220, rs10499194, and rs2230926 polymorphisms are associated with RA susceptibility in Europeans and Asians. Furthermore, it shows that the prevalence of the two allele of rs6920220 in the TNFAIP3 gene is ethnicity-dependent. Further studies are required to determine whether TNFAIP3 polymorphisms contribute to RA susceptibility in different ethnic groups.
References
Choi SJ, Rho YH, Ji JD, Song GG, Lee YH. Genome scan meta-analysis of rheumatoid arthritis. Rheumatology (Oxford). 2006;45:166–70.
Boone DL, Turer EE, Lee EG, Ahmad RC, Wheeler MT, Tsui C, et al. The ubiquitin-modifying enzyme A20 is required for termination of toll-like receptor responses. Nat Immunol. 2004;5:1052–60.
Lee EG, Boone DL, Chai S, Libby SL, Chien M, Lodolce JP, et al. Failure to regulate TNF-induced NF-kappaB and cell death responses in A20-deficient mice. Science. 2000;289:2350–4.
Tavares RM, Turer EE, Liu CL, Advincula R, Scapini P, Rhee L, et al. The ubiquitin modifying enzyme A20 restricts B cell survival and prevents autoimmunity. Immunity. 2010;33:181–91.
Dieude P, Guedj M, Wipff J, Ruiz B, Riemekasten G, Matucci-Cerinic M, et al. Association of the TNFAIP3 rs5029939 variant with systemic sclerosis in the European caucasian population. Ann Rheum Dis. 2010;69:1958–64.
Graham RR, Cotsapas C, Davies L, Hackett R, Lessard CJ, Leon JM, et al. Genetic variants near TNFAIP3 on 6q23 are associated with systemic lupus erythematosus. Nat Genet. 2008;40:1059–61.
Coornaert B, Carpentier I, Beyaert R. A20: central gatekeeper in inflammation and immunity. J Biol Chem. 2009;284:8217–21.
Musone SL, Taylor KE, Lu TT, Nititham J, Ferreira RC, Ortmann W, et al. Multiple polymorphisms in the TNFAIP3 region are independently associated with systemic lupus erythematosus. Nat Genet. 2008;40:1062–4.
Ben Hamad M, Cornelis F, Maalej A, Petit-Teixeira E. A Tunisian case–control association study of a 6q polymorphism in rheumatoid arthritis. Rheumatol Int. 2011 (in press). doi: 10.1007/s00296-011-1996-6.
Musone SL, Taylor KE, Nititham J, Chu C, Poon A, Liao W, et al. Sequencing of TNFAIP3 and association of variants with multiple autoimmune diseases. Genes Immun. 2011;12:176–82.
Hughes LB, Reynolds RJ, Brown EE, Kelley JM, Thomson B, Conn DL, et al. Most common single-nucleotide polymorphisms associated with rheumatoid arthritis in persons of European ancestry confer risk of rheumatoid arthritis in African Americans. Arthritis Rheum. 2010;62:3547–53.
Shimane K, Kochi Y, Horita T, Ikari K, Amano H, Hirakata M, et al. The association of a nonsynonymous single-nucleotide polymorphism in TNFAIP3 with systemic lupus erythematosus and rheumatoid arthritis in the Japanese population. Arthritis Rheum. 2010;62:574–9.
Han TU, Bang SY, Kang C, Bae SC. TRAF1 polymorphisms associated with rheumatoid arthritis susceptibility in Asians and in caucasians. Arthritis Rheum. 2009;60:2577–84.
Coenen MJ, Trynka G, Heskamp S, Franke B, van Diemen CC, Smolonska J, et al. Common and different genetic background for rheumatoid arthritis and coeliac disease. Hum Mol Genet. 2009;18:4195–203.
Stark K, Rovensky J, Blazickova S, Grosse-Wilde H, Ferencik S, Hengstenberg C, et al. Association of common polymorphisms in known susceptibility genes with rheumatoid arthritis in a Slovak population using osteoarthritis patients as controls. Arthritis Res Ther. 2009;11:R70.
Dieguez-Gonzalez R, Calaza M, Perez-Pampin E, Balsa A, Blanco FJ, Canete JD, et al. Analysis of TNFAIP3, a feedback inhibitor of nuclear factor-kappaB and the neighbor intergenic 6q23 region in rheumatoid arthritis susceptibility. Arthritis Res Ther. 2009;11:R42.
Raychaudhuri S, Remmers EF, Lee AT, Hackett R, Guiducci C, Burtt NP, et al. Common variants at CD40 and other loci confer risk of rheumatoid arthritis. Nat Genet. 2008;40:1216–23.
Thomson W, Barton A, Ke X, Eyre S, Hinks A, Bowes J, et al. Rheumatoid arthritis association at 6q23. Nat Genet. 2007;39:1431–3.
Lee YH, Rho YH, Choi SJ, Ji JD, Song GG, Nath SK, et al. The PTPN22 C1858T functional polymorphism and autoimmune diseases—a meta-analysis. Rheumatology (Oxford). 2007;46:49–56.
Lee YH, Witte T, Momot T, Schmidt RE, Kaufman KM, Harley JB, et al. The mannose-binding lectin gene polymorphisms and systemic lupus erythematosus: two case–control studies and a meta-analysis. Arthritis Rheum. 2005;52:3966–74.
Lee YH, Harley JB, Nath SK. Meta-analysis of TNF-alpha promoter -308 A/G polymorphism and SLE susceptibility. Eur J Hum Genet. 2006;14:364–71.
Egger M. Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997;315:629–34.
Davey Smith G, Egger M. Meta-analyses of randomised controlled trials. Lancet. 1997;350:1182.
Egger M, Smith GD, Phillips AN. Meta-analysis: principles and procedures. BMJ. 1997;315:1533–7.
DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986;7:177–88.
Higgins JP, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med. 2002;21:1539–58.
Bowes J, Lawrence R, Eyre S, Panoutsopoulou K, Orozco G, Elliott KS, et al. Rare variation at the TNFAIP3 locus and susceptibility to rheumatoid arthritis. Hum Genet. 2010;128:627–33.
Bates JS, Lessard CJ, Leon JM, Nguyen T, Battiest LJ, Rodgers J, et al. Meta-analysis and imputation identifies a 109 kb risk haplotype spanning TNFAIP3 associated with lupus nephritis and hematologic manifestations. Genes Immun. 2009;10:470–7.
Acknowledgments
This study is supported by a grant from the Korea Healthcare Technology R&D Project, Ministry of Health and Welfare, Republic of Korea (A102065).
Conflict of interest
We have no financial and non-financial conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Andras Falus.
Rights and permissions
About this article
Cite this article
Lee, Y.H., Bae, SC., Choi, S.J. et al. Associations between TNFAIP3 gene polymorphisms and rheumatoid arthritis: a meta-analysis. Inflamm. Res. 61, 635–641 (2012). https://doi.org/10.1007/s00011-012-0455-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00011-012-0455-5