Summary
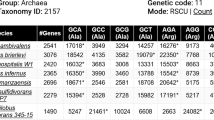
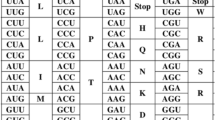
We construct a “codon space” in which a given DNA sequence can be plotted as a function of its base composition in each of the three codon positions. We demonstrate that the base composition is very highly nonrandom, with sequences from more primitive organisms having the least random compositions. By using cluster analysis on the points plotted in codon space we show that there is a strong correlation between base composition and type of organism, with the most primitive organisms having the highest A or T content in the second and third codon positions. A smooth transition toward lower A+T and higher G+C content is observed in the second and third codon positions as the evolutionary complexity of the organism increases. Besides this general trend, more detailed structure can be observed in the clustering that will become clearer as the data base is increased.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Anderberg MR (1973) Cluster analysis for applications. Academic Press, New York
Gillespie DT (1975) The Monte Carlo method for evaluating integrals. Naval Weapons Center, China Lake, California
Grantham R (1980) Workings of the genetic code. Trends Biochem Sci 5:327–331
Grantham R, Gautier C, Gouy M (1980) Codon frequences in 119 individual genes confirm consistent choices of degenerate bases according to genome type. Nucleic Acids Res 8:1893–1912
Grantham R, Gautier C, Gouy M, Jacobzone M, Mercier R (1981) Codon catalog usage as a genome strategy modulated for gene expressivity. Nucleic Acids Res 9:r43-r74
Rowe GW, Trainor LEH (1983a) On the informational content of viral DNA. J Theor Biol 101:151–170
Rowe GW, Trainor LEH (1983b) A thermodynamic theory of codon bias in viral genes. J Theor Biol 101:171–203
Shepherd JCW (1981) Periodic correlations in DNA sequences and evidence suggesting their evolutionary origin in a commaless genetic code. J Mol Evol 17:94–102
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Rowe, G.W., Szabo, V.L. & Trainor, L.E.H. Cluster analysis of genes in codon space. J Mol Evol 20, 167–174 (1984). https://doi.org/10.1007/BF02257377
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF02257377