Summary
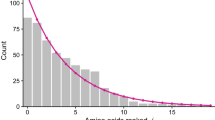
In a study of 25 human variants and 23 “evolutionary alleles” of hemoglobin we show that intraspecific and interspecific patterns of electrophoretic variability are not comparable. Significant deviation from the predicted electrophoretic differentiation between evolutionary alleles is normally found only when amino acid sequence divergence exceeds 10%. When two sequences had diverged at less than 30 out of 287 amino acid residues sites, only 7% of comparisons showed significant deviations from the expected difference of electrophoretic mobility, while significant deviation was shown by 57% of comparisons involving 30–40 residue differences, by 79% in the case of 51–60 differences and by all of the comparisons involving more than 60 differences. In contrast, human variants, which differ by only one or two amino acid residues (less than 1% difference), had significant deviations in 58% of comparisons. Those mutations that appear as fixed differences in the evolutionary material probably represent only a subset of the mutations which can appear within the species. The results suggest that statistical comparisons such as genetic distance may not measure the same process within a species as between species. This is due not to inherent problems with the statistic, but rather to inherent differences in the nature of molecular changes that are detectable by electrophoresis at different stages of population divergence.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Ayala FJ (1975) Genetic differentiation during the speciation process. Evol Biol 8:1–78
Bunn HF, Forget BG, Ranney HM (1977) Human Hemoglobins. W.B. Saunders, Philadelphia
Dayhoff MO (1972) Atlas of Protein Sequence and Structure vol 5, National Biomedical Research Foundation, Washington, DC
Fitch WM (1974) A comparison between evolutionary substitutions and variants in human hemoglobins. Ann NY Acad Sci 241:439–448
Fuerst PA, Ferrell RE (1980) The stepwise mutation model: An experimental evaluation utilizing hemoglobin variants. Genetics 94:185–201
Gorman GC, Kim YJ, Rubinoff R (1976) Genetic relationships of three species ofBathyogobius from the Atlantic and Pacific sides of Panama. Copeia 1976:361–364
Heindell HC, Liu A, Paddock GV, Studnicka GM, Salser WA (1978) The primary sequence of the rabbit α-globin mRNA. Cell 15:43–54
Kirk RL (1976) Genetic distance — A pragmatic approach. Proceedings V. International Congress Human Genetics, Mexico City
Langley CH, Fitch WM (1974) An examination of the constancy of the rate of molecular evolution. J Mol Evol 3:161–177
Nei M (1972) Genetic distance between populations. Am Nat 106:283–292
Nei M, Chakraborty R (1973) Genetic distance and the electrophoretic identity of protein between taxa. J Mol Evol 2:323–328
Ohta T, Kimura M (1971) On the constancy of the evolutionary rate of cistrons. J Mol Evol 1:18–25
Ohta T, Kimura M (1973) A model of mutation appropriate to estimate the number of electrophoretically detectable alleles in a finite population. Genet Res 22:201–204
Ramshaw JAM, Coyne JA, Lewontin RC (1979) The sensitivity of gel electrophoresis as a detector of genetic variation. Genetics 93:1019–1037
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Fuerst, P.A., Ferrell, R.E. The analysis of hidden electrophoretic variation: Interspecific electrophoretic differentiation and amino acid divergence. J Mol Evol 19, 449–454 (1983). https://doi.org/10.1007/BF02102320
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF02102320