Summary
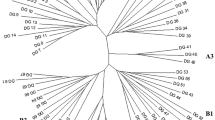
RFLP variability was studied in eight U.S. peanut cultivars, representing the four market types, and in 14 wild Arachis species accessions, using random genomic clones from a PstI library. Very low levels of RFLP variability were found among the allotetraploids, which included the U.S. cultivars and Arachis monticola, a wild species. The diploid wild species were very diverse, however. RFLP patterns of the allotetraploids were more complex than the diploids, and the two constituent genomes could usually be distinguished. On the basis of RFLP band sharing, A. ipaensis, A. duranensis, and A. spegazzinii appeared most closely related to the diploid progenitor species of the allotetraploids. A dendrogram of relationships among the diploid wild species was constructed based on band sharing.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bernatzky R, Tanksley SD (1986) Genetics of actin-related sequences in tomato. Theor Appl Genet 72:314–321
Burr B, Burr RA, Thompson KH, Albertsen MC, Stuber CW (1988) Gene mapping with recombinant inbreds in maize. Genetics 118:519–526
Feinberg AP, Vogelstein B (1984) A technique for radiolabeling DNA restriction fragments to a high specific activity. Anal Biochem 132:6–13
Gregory WC, Gregory MP, Krapovickas A, Smith BW, Yarbrough A (1973) Structures and genetic resources of peanuts. In: Peanuts — culture and uses. American Peanut Research and Education Society, Yoakum/TX, pp 47–135
Grieshammer U (1989) Isozymes in peanuts: variability among U.S. cultivars and Mendelian and non-Mendelian inheritance. MS thesis, North Carolina State University, Raleigh/NC
Hammons RO (1982) Origin and early history of the peanut. In: Pattee HE, Young CT (eds) Peanut science and technology. American Peanut Research and Education Society, Yoakum/TX, pp 1–20
Kirti PB, Murty UR, Bharathi M, Rao NGP (1982) Chromosome pairing in F1 hybrid Arachis hypogaea L. x A. monticola Krap. et Rig. Theor Appl Genet 62:139–144
Klozová E, Turková V, Smartt J, Pitterová K, Svachulová J (1983) Immunochemical characterization of seed proteins of some species of the genus Arachis L. Biol Plant 25:201–208
Knauft DA, Gorbet DW (1989) Genetic diversity among peanut cultivars. Crop Sci 29:1417–1422
Krapovickas A, Fernandez A, Seeligmann P (1974) Recuperacion de la fertilidad en un hiberido interspecifico esteril de Arachis (Leguminosae). Bonplandia Corrientes 3:129–142
Krishna TG, Mitra R (1988) The probable genome donors to Arachis hypogaea L. based on arachin seed storage protein. Euphytica 37:47–52
Murray M, Thompson WF (1980) Rapid isolation of highmolecular-weight plant DNA. Nucleic Acids Res 8:4321–4325
Rigby P, Dieckmann M, Rhodes C, Berg P (1977) Labeling deoxyribonucleic acid to high specific activity in vitro by Nick-translation with DNA polymerase I. J Mol Biol 113:237–251
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor/NY
Singh AK (1986) Utilization of wild relatives in the genetic improvement of Arachis hypogaea L. 8. Synthetic amphidiploids and their importance in interspecific breeding. Theor Appl Genet 72:433–439
Singh AK (1988) Putative genome donors of Arachis hypogaea (Fabaceae), evidence from crosses with synthetic amphidiploids. Plant Syst Evol 160:143–151
Singh AK, Moss JP (1982) Utilization of wild relatives in genetic improvement of Arachis hypogaea L. 2. Chromosome complements of species in section Arachis. Theor Appl Genet 61:305–314
Singh AK, Moss JP (1984) Utilization of wild relatives in the genetic improvement of Arachis hypogaea L. 5. Genome analysis in section Arachis and its implications in gene transfer. Theor Appl Genet 68:355–364
Smartt J, Stalker HT (1982) Speciation and cytogenetics in Arachis. In: Pattee HE, Young CT (eds) Peanut science and technology. American Peanut Research and Education Society, Yoakum/TX, pp 21–49
Smartt J, Gregory WC, Phluge M (1978) The genomes of Arachis hypogaea. 1. Cytogenetic studies of putative genome donors. Euphytica 27:665–675
Song KM, Osborn TC, Williams PH (1988) Brassica taxonomy based on nuclear restriction fragment length polymorphisms (RFLPs). 1. Genome evolution of diploid and amphidiploid species. Theor Appl Genet 75:784–794
Song KM, Osborn TC, Williams PH (1990) Brassica taxonomy based on nuclear restriction fragment length polymorphisms (RFLPs). 3. Genome relationships in Brassica and related genera and the origin of B. oleracea and B. rapa (syn. campestris). Theor Appl Genet 79:497–506
Southern EM (1975) Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol 98:503–517
Stalker HT, Moss JP (1987) Speciation, cytogenetics, and utilization of Arachis species. Adv Agron 41:1–39
Tanksley SD, Miller J, Paterson A, Bernatzky R (1987) Molecular mapping of plant chromosomes. In: Gustafson JP, Appels RA (eds) Chromosome structure and function. Plenum Press, New York, pp 157–173
Wilimzig R (1985) LiCl boiling method for plasmid minipreps. Trends Genet 1:158
Wynne JC, Coffelt TA (1982) Genetics of Arachis hypogaea L. In: Pattee HE, Young CT (eds) Peanut science and technology, American Peanut Research and Education Society, Yoakum/TX, pp 50–94
Author information
Authors and Affiliations
Additional information
Communicated by P. L. Pfahler
Rights and permissions
About this article
Cite this article
Kochert, G., Halward, T., Branch, W.D. et al. RFLP variability in peanut (Arachis hypogaea L.) cultivars and wild species. Theoret. Appl. Genetics 81, 565–570 (1991). https://doi.org/10.1007/BF00226719
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00226719