Abstract
Multiple common variations discovered via genome-wide association studies (GWASs) were shown to have a minimal association with breast cancer (BC) risk in Vietnamese women. This study analyzed the cumulative effect in predicting BC risk of ten single nucleotide polymorphisms (SNPs) identified by previous GWAS and were common in Vietnamese. In this case-control research, 240 BC patients and 271 healthy controls were recruited to assess candidate SNPs’ association with BC risk. A polygenic risk score (PRS) was then created from SNPs strongly related to the risk of BC among the assessed population. The area under the receiver operating characteristic curve (AUC) was used to assess the effectiveness of the PRS model with BC risk. Logistic regression results showed seven individual SNPs (rs2155209, rs4784227, rs2605039, rs3817198, rs2981582, rs11614913, and rs12325489) were significantly associated with BC risk after multiple testing. These SNPs were then used to create the PRS model. Compared with women in the lowest quartile, women in the highest quartile of PRS had a considerably higher risk (odds ratio 2.65; 95% confidence interval (95% CI) 1.61–4.40) with AUC at 71%. These findings suggest that the 7-SNP PRS would effectively distinguish between women with high and low risk of BC, indicating the genetic marker for BC risk prediction in a Vietnamese population.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 INTRODUCTION
Breast cancer (BC) is one of the most prevalent malignancies among women worldwide, including Vietnamese. It is recognized that BC is a complicated disease that is affected by hereditary and environmental variables. The heredity of BC was estimated in recent times to be 31% and its typical environmental components to just 16% (Möller, 2016), suggesting that a significant concern for recent scientific investigations into BC is their variety in disease-related genes.
Many genome-wide association studies (GWASs) have already found many susceptibility variations related to BC in Caucasian and Asian populations (Hunter, 2007; Gold, 2008; Stacey Simon N, 2008; Cai, 2011; Kim, 2012; Han M.-R., 2016). However, most were variations with modest penetration risk than those in prevalent genes such as BRCA1 or BRCA2 (Bradbury, 2007). The two genes make up 25% of the family risk and around 5% of the incidence of BC through uncommon mutation frequencies (Peto, 1999; Pharoah, 2004). Over 100 single nucleotide polymorphisms (SNPs) have been discovered thus far. Although most of these SNPs were discovered in primarily Caucasian groups (Turnbull, 2010; Fletcher, 2011; Haiman, 2011), a few SNPs were discovered in Asian populations (Kim, 2012; Cai, 2014; Han M.-R., 2016). Several studies have shown that specific SNPs exhibit ethnic-specific characteristics and should not be tested in other populations (Kim, 2012; Chen Yazhen, 2016; Han M.-R., 2016; Xu M., 2016). Fine-scale mapping of GWAS-identified areas (Glubb, 2015; Orr, 2015; Shi, 2016), as well as meta-analysis of previous GWAS (Lindström, 2014; Michailidou, 2015; Couch, 2016), have recently contributed to the enormously increasing number of SNPs related to BC susceptibility in certain ethnic groups (Zheng Y., 2013).
Even if a specific SNP is related to the risk of BC, this single variation conferring risk is minimal. As a result, polygenic risk scores (PRS) have been developed to assess the cumulative effect of specific SNPs related to BC (Mavaddat, 2015). A PRS considers each SNP’s odds ratio (OR) and the overall number of risk alleles that an individual holds. The majority of PRS generated were taken from population data sets of the Caucasian population. Instead, a small amount of research in Asian populations is developing PRS models for the risk of BC using associated SNPs.
This study aimed to examine the relationship of known SNPs with the risk of BC in Vietnamese women. The combinations of relevant SNPs associated with BC were also utilized to generate a PRS that was then assessed to predict Vietnamese BC risk.
2 MATERIALS AND METHODS
2.1 Study Population
This study was a population case-control study of BC. Women aged 28–81 yr with histologically confirmed initial primary in situ or invasive BC were identified as cases at Oncology Hospital in Ho Chi Minh City, Vietnam between 2015 and 2017 (n = 240). Healthy women with no known cancer were recruited at the Oncology Hospital in Ho Chi Minh City to serve as population-based healthy controls. This study included 271 controls without a cancer record, typically matching an approximate cases’s age distribution.
2.2 DNA Extraction, SNP Selection, and Genotyping
The salting-out procedure was used to extract genomic DNA from peripheral blood and kept at –80°C until it was needed for further research. SNP selection was carried out through a review of GWASs or candidate-gene association studies based on the existing genotype analysis literature (Gapska, 2009; Qian, 2011; Li N., 2014; Han M.R., 2015; Qi, 2015; Wu, 2015; Chen Y., 2016; Hein, 2017; Zhang H., 2017; Zhu, 2017). The ten SNPs, which were most significantly associated with BC in other populations and common in Vietnamese population (MAF > 10%), were selected including VDR (rs2228570), IGF-I (rs7965399), miR-146A (rs2910164), MRE11A (rs2155209), TOX3 (rs4784227), HSPD1 (rs2605039), LSP1 (rs3817198), FGFR2 (rs2981582), miR-196A2 (rs11614913), and miR-370 (rs12325489).
SNP genotyping was performed using the High-resolution method (HRM) in a LightCycler 96 System (Roche Diagnostics Penzberg Germany). Comprehensive quality control (QC) procedures were followed to ensure genotyping quality, including duplicate genotype identification using Sanger, a Hardy-Weinberg equilibrium (HWE) test, and a call rate of greater than 99%. A total of 30 (5.9%) quality control samples were successfully genotyped, with a 100% concordance rate.
2.3 Statistical Analysis
The odds ratio (OR) and 95% confidence interval (CI) were computed using a logistic regression model to examine the association between the SNPs and BC risk. HWE was examined among controls using a goodness-of-fit Chi-squared test. Student’s t-test and Chi-squared test were used to compare cases and controls. R version 4.1.0 was used for all statistical analyses.
A PRS was established to estimate the polygenic contribution of BC susceptibility loci using marginally significant SNPs associated with BC risk (p < 0.05) based on any one of the per-allele, codominant, dominant, overdominant, or recessive logistic regression models. For robust linkage disequilibrium SNPs located on the same gene or chromosome (each D' > 0.9), the one variant with the lowest P-value as a candidate was chosen. Then, a weighted PRS was calculated for each individual using the formula: PRS = β1x1 + β1x1 +… βkxk … + βnxn where βk is the per-allele log odds ratio (OR) for BC associated with the minor allele for SNP k, and xk is the number of risk alleles for the same SNP.
Logistic regression analysis was performed to investigate the association between BC and PRS, with PRS being a continuous variable (Mavaddat, 2015). When stratified by menopausal status, a logistic regression model was created to evaluate the association between PRS and BC risk. In addition, ORs based on logistic regression models were estimated for different PRS quartiles, with the first quartile being the reference. The area under the receiver operating characteristic curve (AUC) was applied to evaluate the model’s discriminative ability.
3 RESULTS
Distributions of the characteristic of BC patients and healthy controls are shown in Table 1. The mean age of cases and controls were 51 ± 8.8 and 50 ± 8.9, respectively. The mean age was not significantly different between cases and controls. Due to a lack of comprehensive information, this study relied solely on age as a rough measure for menopausal status (Premenopausal: age ≤50, Postmenopausal: age >50) (Hill, 1996). The percentages of menopause status in patients were 47 and 53% for premenopausal women and postmenopausal women, respectively. These percentages in controls were 60% premenopausal women and 40% postmenopausal women. Interestingly, a significantly more significant proportion of cases were postmenopausal than those of similar ages in controls.
Table 2 indicates the association between the ten candidate SNPs and BC risk in the assessed Vietnamese population. For the per-allele model, two SNPs, rs2605039 and rs2981582, revealed significant associations with the risk of BC. The three SNPs, named rs4784227, rs2605039, and rs11614913, showed significant association with the risk of BC under the dominant model. The four SNPs, rs2155209, rs2605039, rs3817198, and rs2981582, demonstrated significant association with BC risk under the recessive model. CASC22 gene rs12325489 was significantly associated with the risk of BC either in the codominant or under the overdominant model. The remaining three SNPs, VDR rs2228570, IGF-I rs7965399, and miR-146A rs2910164, showed a non-significant association with the risk of BC. Seven SNPs, namely rs2155209, rs4784227, rs2605039, rs3817198, rs2981582, rs11614913, and rs12325489, were selected to create PRS model as their association with BC risk were marginally significant. In addition, since seven SNPs were not in strong linkage disequilibrium (D' < 0.8), all these SNPs were chosen for establishing the PRS model.
According to the quartile distribution, women in the second (from –0.18 to –0.06), third (from –0.06 to 0.07), and fourth (from 0.07 to 0.48) quartiles had 1.79-, 2.03-, and 2.65-fold increased BC risks compared to women in the first quartile (from –0.49 to –0.18), showing a significant increasing trend (p = 0.01). The trend was also significant when the same risk score was separately applied to premenopausal women (p = 0.007). Postmenopausal women in the 4th quartile showed a significant 2.38-fold increase in the risk of BC (p = 0.03) (Table 3).
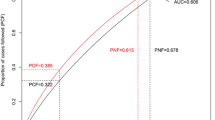
The AUC was then calculated to evaluate the effectiveness of the PRS model (Fig. 1). The AUC was estimated at 71, 75, and 68% for the PRS in all ages, premenopausal and postmenopausal status, respectively (Table 3). The estimated AUC of models was higher than 70%, corresponding to an acceptable discriminative ability to diagnose patients with and without the disease (Safari, 2016).
4 DISCUSSION
The risk associated with high- and moderate-risk breast cancer susceptibility genes is modified by polygenic risk scores (Kuchenbaecker, 2017). When PRS was included in the risk prediction model, Kuchenbaecker et al. found significant differences in the risk for breast cancer (Kuchenbaecker, 2017). Mainly, with a PRS in the top quintile, BRCA1 carriers had a 56% risk of developing breast cancer by the age of 80. Others with a PRS in the 90th percentile, on the other hand, had a 75% risk of breast cancer by 80 yr old. There was also evidence of subtype-specific PRS, with a PRS adjusted for ER-negative risk having the highest correlation with the risk of breast cancer in BRCA1 carriers (Kuchenbaecker, 2017). The highest discrimination was found in the ER-negative PRS model for BRCA1 carriers (AUC = 0.58). However, since the literature remains limited to only a few research findings, more investigation is warranted.
Presently, genetic screening of high- and moderate-penetrance genes obtains non-informative results for most women at high risk of breast cancer. Breast cancer prevention strategies are underutilized within this group of women (Schwartz, 2012). Consequently, novel risk prediction methods are needed to notify risk management decisions for women who have received non-informative outcomes from monogenic testing. According to studies investigating the use of polygenic factors, PRS was predictive of breast cancer risk in women with non-informative BRCA1/2 results (Dite, 2013; Dite, 2016; Li H., 2017; Lakeman, 2019).
There is currently no guideline to help clinical genetic services implement breast cancer polygenic testing. Usually, genetic services primarily concentrate on testing monogenic risk genes (e.g., BRCA1/2) and their familial consequences. Since polygenic screening has become more widely used in clinical practice, a transformation toward personalized care will be necessary. Breast cancer PRS testing is now available in clinics (Hughes, 2017; Black, 2018), and it is aimed at women who have received non-informative results from monogenic testing.
This study evaluated ten BC-related SNPs to determine the possible relationships with BC risk in the Vietnamese population (Table 2). Among them, seven SNPs were significantly associated with BC risk in our Vietnamese population, including MRE11A rs2155209, TOX3 rs4784227, HSPD1 rs2605039, FGFR2 rs2981582, LSP1 rs3817198, miR-196A2 rs11614913, and CASC22 rs12325489 after multiple testing. The findings provide additional evidence for follow-up GWAS studies in BC, particularly those carried out in Asian populations. In addition, the PRSs were constructed using seven selected SNPs to measure the cumulative effect of variants. Furthermore, the PRS models were developed to discriminate against women according to BC risk, which provided adequate power with an AUC of 71%. In addition, this PRS model seems to be most effective in premenopausal women with an AUC of 75%.
In the current association study, of the seven BC‑related SNPs, four SNPs were associated with an increased BC risk in Vietnamese, namely TOX3 rs4784227, FGFR2 rs2981582, LSP1 rs3817198, and CASC22 rs12325489. These four SNPs have been previously reported to be significantly associated with BC risk across different ethnicities with a similar direction of effect to our findings (Long, 2010; Fernandez-Navarro, 2013; Lin, 2014; Na Li, 2014; Zhang Y., 2017; Zuo, 2020). However, several studies were showing that rs3817198 was statistically insignificant as a BC risk factor in some populations, including Turkey (Ozgoz, 2020), China (Chen Y., 2016; Tan, 2017), Brazil (Fernandes, 2016), Tunisia (Shan J., 2012), and Germany (Campa, 2011). The differences between studies indicated that rs3817198 is population-specific SNPs, thus could act as a specific marker for the Vietnamese population. Regarding the molecular mechanism of effect, these SNPs interfere with distinct pathways underlying BC’s growth. By modulating the expression of TOX3, rs4784227 could target and affect BRCA1, which significantly involves controlling genome stability and DNA repairing (Shan Jingxuan, 2013; Tajbakhsh, 2019). SNP rs2981582 and rs3817198, meanwhile, are located in the intron region of FGFR2 (Easton, 2007) and LSP1 (Harrison, 2004), respectively, which have a role in promoting cell proliferation, differentiation (Ricol, 1999; Yu, 2003; Fogarty, 2007) and controlling cell cycle, and apoptosis (MacLachlan, 1995). As for rs12325489, it dictates the transcription of a gene engaging in cancer tumorigenesis and metastasis—lincRNA CASC22 via creating a binding site for miR-370 (Dinger, 2008; Guttman, 2009; Huarte, 2010; Gibb, 2011).
Other significant associations identified in our study included MRE11A rs2155209, HSPD1 rs2605039, and miR-196A2 rs11614913. The current study has demonstrated that these three SNPs showed a significantly decreased risk for BC in a Vietnamese population. The effect trend was likely the same as the previously reported studies for two SNPs, rs2605039 (Zhu, 2017) and rs11614913 (Xu W., 2011; Wang J., 2012; Wang P.Y., 2013; Chen, 2014; Dai Z.J., 2015; Dai Z.M., 2016; Mu, 2017; Zhang H., 2017; Bastami, 2019; Choupani, 2019). In terms of rs2155209, a reverse direction of effect was observed between Vietnamese and Chinese. While rs2155209 showed decreased BC risk in Vietnam, it was associated with an increased BC susceptibility in China (Wu, 2015). This finding is the first study in Vietnam to explore the association between rs2155209 and the risk of BC; thus, further large-scale studies should be carried out to confirm our results. SNP rs2155209 is located in the 3'UTR of MRE11A, a gene responsible for repairing DNA damage when a double-strand break occurs, leading to BC development (Lobrich, 2007). SNP rs2605039 is a genetic variant in the intron region of HSPD1, which encodes for a heat shock protein controlling the expression of anti-apoptotic proteins—BCL-2 and BCL-XL (Ghosh, 2008; Pace, 2013). By dictating the binding region of miR-196A2 and homeobox genes, rs11614913 significantly influences cell proliferation and DNA repair (Easton, 2007; Ma, 2007; Stacey S.N., 2007).
As an individual, the effect of each SNP on the risk of BC is modest. However, it has been demonstrated that their combined impact, as PRS, provides a level of risk discrimination that could be utilized to stratify individuals into distinct disease risk groups. Previous studies that examine the influence of PRS on the risk of BC have consistently shown greater PRS in women who have been diagnosed with BC than in the controls (Sawyer, 2012; Muranen, 2016; Evans, 2017). Overall, European population research shows that the lowest and highest quartile PRS distribution is at least a two-fold difference in the risk of BC (Wacholder, 2010; Darabi, 2012; Allman, 2015; Vachon, 2015). Similar findings have also been reported across other populations, including African American and Asian ancestry (Zheng W., 2010; Allman, 2015; Hsieh, 2017; Chan, 2018; Starlard-Davenport, 2018). Our PRS results also showed a linear association with an increased risk of BC, indicating at least 2.65-fold risk for Vietnamese women in the highest quartile compared to those in the lowest (Table 3). In the combination of PRS and premenopausal status, women in the third and fourth quartiles had 3.65- and 3.32-fold increased BC risks than women in the first quartile (Table 3). These ORs were much higher compared to previous studies. Hsieh et al. created a PRS composed of six SNPs and found that the OR was around 2.26-fold for women in the highest quintile compared to those in the lowest score [36]. Mavaddat et al. constructed a 77-SNP PRS for BC and found a threefold increase in risk when comparing the highest and the middle quintiles (Vachon, 2015).
In addition, this study has assessed the discriminatory accuracy of BC PRS. The discriminatory accuracy of PRS has been most commonly assessed by calculating AUC. The reported AUC for BC PRS has been modest, ranging from 0.59 to 0.69 for European populations and 0.57 to 0.72 for non-European populations (Table 4). The obtained AUCs in our study (>70%) were relatively high compared to previous retrospective and non-familial studies in American, European, and Asian populations (Table 4). A study with a similar AUC result was reported by Shieh et al., 2016 (Shieh, 2016). The study showed an AUC of 0.72 within a sample size of 51 cases and 51 controls in Asian Americans. The difference was that Shieh et al.'s study generated PRS from 76 variants, while this study obtained PRS using only 7 SNPs. There has not been a common consensus on whether fewer or more SNPs would render a better PRS model. In two separate studies conducted in Asians having a similar sample size, one obtained an AUC of 0.60 using only 6 SNPs in their PRS (Shieh, 2017), while the other obtained an AUC of 0.57 using a 46-SNP PRS (Chan, 2018). These findings imply that the choice of ethical SNPs for the populations under study must be tailored. In addition, the sample size seems to have no noticeable impact on the discriminant ability of PRS models. In two separate European studies obtaining a similar AUC of 0.62, one conducted with 1664 cases and 1636 controls (Mealiffe, 2010) while the other conducted with a much larger sample size 33 673 cases and 33 381 controls (Vachon, 2015).
The novelty of this study is that compared to other studies utilizing PRS (Table 4), this study has evaluated four SNPs (miR-196A2 rs11614913, CASC22 rs12325489, MRE11A rs2155209, and HSPD1 rs2605039) that have not been previously included in any other PRS. Of the above four SNPs, there were two SNPs (rs11614913 and rs12325489) on miRNA and long non-coding RNA gene, suggesting the potential use of these non-coding RNAs in further BC studies. Three SNPs (rs11614913, rs2155209, and rs2605039) out of four SNPs above were associated with a reduced risk of BC in Vietnamese. This finding could have contributed to the increased discriminant efficiency of the PRS model in this study.
Nevertheless, some concerns must be addressed in order for this work to be correctly interpreted. First, the power of logistic regression analysis in this study could reach 80% in detecting a log-additive OR of 1.38 with a MAF of 14%. However, other SNPs with lower ORs and MAFs may need a larger sample size to reach this statistical power. Second, our current analysis was limited to 10 common BC-risk variants (>10% in the Vietnamese population) identified by previous association studies with OR values higher than 1.6 or lower than 0.7. Shortly, larger effect sizes of sequence variants are likely to be uncovered. Therefore, our PRS results should be interpreted carefully. In addition, due to a lack of data on clinicopathological characteristics, we could not conduct subgroup analyses in terms of different cancer subtypes. Further studies should take subgroup analyses to differentiate BC risk using PRS.
5 CONCLUSIONS
Our data evaluated and identified the significant association of seven SNPs out of the ten SNPs with BC risk in a Vietnamese population. The PRS model included seven BC-related SNPs that are significantly related to BC risk. The seven-SNP PRS only and menopausal status help discriminate women at high risk of BC from those at low risk. Future comprehensive evaluations of the genetic risk variants in a larger population are warranted.
REFERENCES
Allman, R., Dite, G.S., Hopper, J.L., et al., SNPs and breast cancer risk prediction for African American and Hispanic women, Breast Cancer Res. Treat., 2015, vol. 154, no. 3, pp. 583–589. https://doi.org/10.1007/s10549-015-3641-7
Bastami, M., Choupani, J., Saadatian, Z., et al., Evidences from a systematic review and meta-analysis unveil the role of miRNA polymorphisms in the predisposition to female neoplasms, Int. J. Mol. Sci., 2019, vol. 20, no. 20, art. ID 5088. https://doi.org/10.3390/ijms20205088
Black, M.H., Li, S., LaDuca, H., et al., Polygenic risk score for breast cancer in high-risk women, J. Clin. Oncol., 2018, vol. 36, pp. 1508-1508. https://doi.org/10.1200/JCO.2018.36.15_suppl.1508
Bradbury, A.R. and Olopade, O.I., Genetic susceptibility to breast cancer, Rev. Endocr. Metab. Disord., 2007, vol. 8, no. 3, pp. 255–267. https://doi.org/10.1007/s11154-007-9038-0
Cai, Q., Long, J., Lu, W., et al., Genome-wide association study identifies breast cancer risk variant at 10q21. 2: results from the Asia Breast Cancer Consortium, Hum. Mol. Genet., 2011, vol. 20, no. 24, pp. 4991–4999. https://doi.org/10.1093/hmg/ddr405
Cai, Q., Zhang, B., Sung, H., et al., Genome-wide association analysis in East Asians identifies breast cancer susceptibility loci at 1q32.1, 5q14.3 and 15q26.1, Nature Genetics, 2014, vol. 46, no. 8, pp. 886–890. https://doi.org/10.1038/ng.3041
Campa, D., Kaaks, R., Le Marchand, L., et al., Interactions between genetic variants and breast cancer risk factors in the breast and prostate cancer cohort consortium, J. Natl. Cancer Inst., 2011, vol. 103, no. 16, pp. 1252–1263. https://doi.org/10.1093/jnci/djr265
Chan, C.H.T., Munusamy, P., Loke, S.Y., et al., Evaluation of three polygenic risk score models for the prediction of breast cancer risk in Singapore Chinese, Oncotarget, 2018, vol. 9, no. 16, pp. 12796–12804. https://doi.org/10.18632/oncotarget.24374
Chen, Q.H., Wang, Q.B., and Zhang, B., Ethnicity modifies the association between functional microRNA polymorphisms and breast cancer risk: a HuGE meta-analysis, Tumor Biol., 2014, vol. 35, no. 1, pp. 529–543. https://doi.org/10.1007/s13277-013-1074-7
Chen, Y., Fu, F., Lin, Y., et al., The precision relationships between eight GWAS-identified genetic variants and breast cancer in a Chinese population, Oncotarget, 2016, vol. 7, no. 46, art. ID 75457. https://doi.org/10.18632/oncotarget.12255
Chen, Y., Shi, C., and Guo, Q., TNRC9 rs12443621 and FGFR2 rs2981582 polymorphisms and breast cancer risk, World J. Surg. Oncol., 2016, vol. 14, no. 1, art. ID 50. https://doi.org/10.1186/s12957-016-0795-7
Choupani, J., Nariman-Saleh-Fam, Z., Saadatian, Z., et al., Association of mir-196a-2 rs11614913 and mir-149 rs2292832 polymorphisms with risk of cancer: an updated meta-analysis, Front. Genet., 2019, vol. 10, art. ID 186. https://doi.org/10.3389/fgene.2019.00186
Couch, F.J., Kuchenbaecker, K.B., Michailidou, K., et al., Identification of four novel susceptibility loci for oestrogen receptor negative breast cancer, Nat. Commun., 2016, vol. 7, no. 1, art. ID 11375. https://doi.org/10.1038/ncomms11375
Dai, Z.J., Shao, Y.P., Wang, X.J., et al., Five common functional polymorphisms in microRNAs (rs2910164, rs2292832, rs11614913, rs3746444, rs895819) and the susceptibility to breast cancer: evidence from 8361 cancer cases and 8504 controls, Curr. Pharm. Des., 2015, vol. 21, no. 11, pp. 1455–1463. https://doi.org/10.2174/1381612821666141208143533
Dai, Z.M., Kang, H.F., Zhang, W.G., et al., The Associations of Single Nucleotide Polymorphisms in miR196a2, miR-499, and miR-608 with breast cancer susceptibility: A STROBE-compliant observational study, Medicine (Baltimore), 2016, vol. 95, no. 7, art. ID e2826. https://doi.org/10.1097/MD.0000000000002826
Darabi, H., Czene, K., Zhao, W., et al., Breast cancer risk prediction and individualised screening based on common genetic variation and breast density measurement, Breast Cancer Res., 2012, vol. 14, no. 1, art. ID R25. https://doi.org/10.1186/bcr3110
Dinger, M.E., Amaral, P.P., Mercer, T.R., et al., Long noncoding RNAs in mouse embryonic stem cell pluripotency and differentiation, Genome Res., 2008, vol. 18, no. 9, pp. 1433–1445. https://doi.org/10.1101/gr.078378.108
Dite, G.S., MacInnis, R.J., Bickerstaffe, A., et al., Breast cancer risk prediction using clinical models and 77 independent risk-associated SNPs for women aged under 50 years: australian breast cancer family registry, Cancer Epidemiol. Biomarkers Prev., 2016, vol. 25, no. 2, pp. 359–365. https://doi.org/10.1158/1055-9965.Epi-15-0838
Dite, G.S., Mahmoodi, M., Bickerstaffe, A., et al., Using SNP genotypes to improve the discrimination of a simple breast cancer risk prediction model, Breast Cancer Res. Treat., 2013, vol. 139, no. 3, pp. 887–896. https://doi.org/10.1007/s10549-013-2610-2
Easton, D.F., Pooley, K.A., Dunning, A.M., et al., Genome-wide association study identifies novel breast cancer susceptibility loci, Nature, 2007, vol. 447, no. 7148, pp. 1087–1093. https://doi.org/10.1038/nature05887
Evans, D.G., Brentnall, A., Byers, H., et al., The impact of a panel of 18 SNPs on breast cancer risk in women attending a UK familial screening clinic: a case–control study, J. Med. Genet., 2017, vol. 54, no. 2, art. ID 111113. https://doi.org/10.1136/jmedgenet-2016-104125
Fernandes, G.C., Michelli, R.A., Scapulatempo-Neto, C., et al., Association of polymorphisms with a family history of cancer and the presence of germline mutations in the BRCA1/BRCA2 genes, Hered. Cancer Clin. Pract., 2016, vol. 14, no. 1, art.ID. 2. https://doi.org/10.1186/s13053-015-0042-1
Fernandez-Navarro, P., Pita, G., Santamarina, C., et al., Association analysis between breast cancer genetic variants and mammographic density in a large population-based study (Determinants of Density in Mammographies in Spain) identifies susceptibility loci in TOX3 gene, Eur. J. Cancer, 2013, vol. 49, no. 2, pp. 474–481. https://doi.org/10.1016/j.ejca.2012.08.026
Fletcher, O., Johnson, N., Orr, N., et al., Novel breast cancer susceptibility locus at 9q31.2: results of a genome-wide association study, J. Natl. Cancer Inst., 2011, vol. 103, no. 5, pp. 425–435. https://doi.org/10.1093/jnci/djq563
Fogarty, M.P., Emmenegger, B.A., Grasfeder, L.L., et al., Fibroblast growth factor blocks Sonic hedgehog signaling in neuronal precursors and tumor cells, Proc. Natl. Acad. Sci., 2007, vol. 104, no. 8, pp. 2973–2978. https://doi.org/10.1073/pnas.0605770104
Gapska, P., Scott, R.J., Serrano-Fernandez, P., et al., Vitamin D receptor variants and breast cancer risk in the Polish population, Breast Cancer Res. Treat., 2009, vol. 115, no. 3, pp. 629–633. https://doi.org/10.1007/s10549-008-0107-1
Ghosh, J.C., Dohi, T., Kang, B.H., et al., Hsp60 regulation of tumor cell apoptosis, J. Biol. Chem., 2008, vol. 283, no. 8, pp. 5188–5194. https://doi.org/10.1074/jbc.M705904200
Gibb, E.A., Brown, C.J., and Lam, W.L., The functional role of long non-coding RNA in human carcinomas, Mol. Cancer, 2011, vol. 10, no. 1, art. ID 38. https://doi.org/10.1186/1476-4598-10-38
Glubb, D.M., Maranian, M.J., Michailidou, K., et al., Fine-scale mapping of the 5q11.2 breast cancer locus reveals at least three independent risk variants regulating MAP3K1, Am. J. Hum. Genet., 2015, vol. 96, no. 1, pp. 5–20. https://doi.org/10.1016/j.ajhg.2014.11.009
Gold, B., Kirchhoff, T., Stefanov, S., et al., Genome-wide association study provides evidence for a breast cancer risk locus at 6q22.33, Proc. Natl. Acad. Sci., 2008, vol. 105, no. 11, pp. 4340–4345. https://doi.org/10.1073/pnas.0800441105
Guttman, M., Amit, I., Garber, M., et al., Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals, Nature, 2009, vol. 458, no. 7235, pp. 223–227. https://doi.org/10.1038/nature07672
Haiman, C.A., Chen, G.K., Vachon, C.M., et al., A common variant at the TERT-CLPTM1L locus is associated with estrogen receptor–negative breast cancer, Nat. Genet., 2011, vol. 43, no. 12, pp. 1210–1214. https://doi.org/10.1038/ng.985
Han, M.-R., Long, J., Choi, J.-Y., et al., Genome-wide association study in East Asians identifies two novel breast cancer susceptibility loci, 2016, Hum. Mol. Genet., vol. 25, no. 15, pp. 3361-3371. https://doi.org/10.1093/hmg/ddw164
Han, M.R., Deming-Halverson, S., Cai, Q., et al., Evaluating 17 breast cancer susceptibility loci in the Nashville breast health study, Breast Cancer, 2015, vol. 22, no. 5, pp. 544–551. https://doi.org/10.1007/s12282-014-0518-2
Harrison, R.E., Sikorski, B.A., and Jongstra, J., Leukocyte-specific protein 1 targets the ERK/MAP kinase scaffold protein KSR and MEK1 and ERK2 to the actin cytoskeleton, J. Cell Sci., 2004, vol. 117, no. 10, pp. 2151–2157.https://doi.org/10.1242/jcs.00955
Hein, A., Rack, B., Li, L., et al., Genetic breast cancer susceptibility variants and prognosis in the prospectively randomized SUCCESS a study, Geburtshilfe Frauenheilk., 2017, vol. 77, no. 6, pp. 651–659. https://doi.org/10.1055/s-0042-113189
Hill, K., The demography of menopause, Maturitas, 1996, vol. 23, no. 2, pp. 113–127. https://doi.org/10.1016/0378-5122(95)00968-x
Hsieh, Y.C., Tu, S.H., Su, C.T., et al., A polygenic risk score for breast cancer risk in a Taiwanese population, Breast Cancer Res. Treat., 2017, vol. 163, no. 1, pp. 131–138. https://doi.org/10.1007/s10549-017-4144-5
Huarte, M. and Rinn, J.L., Large non-coding RNAs: missing links in cancer?, Hum. Mol. Genet., 2010, vol. 19, no. R2, pp. R152–R161. https://doi.org/10.1093/hmg/ddq353
Hughes, E., Judkins, T., Wagner, S., et al., Development and validation of a residual risk score to predict breast cancer risk in unaffected women negative for mutations on a multi-gene hereditary cancer panel, J. Clin. Oncol., 2017, vol. 35, pp. 1579–1579. https://doi.org/10.1200/JCO.2017.35.15_suppl.1579
Hunter, D.J., Kraft, P., Jacobs, K.B., et al., A genome-wide association study identifies alleles in FGFR2 associated with risk of sporadic postmenopausal breast cancer, Nat. Genet., 2007, vol. 39, no. 7, pp. 870–874. https://doi.org/10.1038/ng2075
Kim, H.-C., Lee, J.-Y., Sung, H., et al., A genome-wide association study identifies a breast cancer risk variant in ERBB4 at 2q34: results from the Seoul Breast Cancer Study, Breast Cancer Res., 2012, vol. 14, no. 2, art. ID R 56. https://doi.org/10.1186/bcr3158
Kuchenbaecker, K.B., McGuffog, L., Barrowdale, D., et al., Evaluation of polygenic risk scores for breast and ovarian cancer risk prediction in BRCA1 and BRCA2 mutation carriers, J. Natl. Cancer Inst., 2017, vol. 109, no. 7. https://doi.org/10.1093/jnci/djw302
Lakeman, I.M.M., Hilbers, F.S., Rodríguez-Girondo, M., et al., Addition of a 161-SNP polygenic risk score to family history-based risk prediction: impact on clinical management in non-BRCA1/2 breast cancer families, J. Med. Genet., 2019, vol. 56, no. 9, pp. 581–589. https://doi.org/10.1136/jmedgenet-2019-106072
Li, H., Feng, B., Miron, A., et al., Breast cancer risk prediction using a polygenic risk score in the familial setting: a prospective study from the Breast Cancer Family Registry and kConFab, Genet. Med., 2017, vol. 19, no. 1, pp. 30–35. https://doi.org/10.1038/gim.2016.43
Li, N., Zhou, P., Zheng, J., et al., A polymorphism rs12325489C>T in the LincRNA-ENST00000515084 exon was found to modulate breast cancer risk via GWAS-based association analyses, PLoS One, 2014, vol. 9, no. 5, art. ID e98251. https://doi.org/10.1371/journal.pone.0098251
Lin, Y., Fu, F., Chen, M., et al., Associations of two common genetic variants with breast cancer risk in a Chinese population: a stratified interaction analysis, PLoS One, 2014, vol. 9, no. 12, pp. 1–12. https://doi.org/10.1371/journal.pone.0115707
Lindström, S., Thompson, D.J., Paterson, A.D., et al., Genome-wide association study identifies multiple loci associated with both mammographic density and breast cancer risk, Nat. Commun., 2014, vol. 5, no. 1, art. ID 5303. https://doi.org/10.1038/ncomms6303
Lobrich, M. and Jeggo, P., The impact of a negligent G2/M checkpoint on genomic instability cancer induction, Nat. Rev. Cancer, 2007, vol. 7, no. 11, pp. 861–869. https://doi.org/10.1038/nrc2248
Long, J., Shu, X.O., Cai, Q., et al., Evaluation of breast cancer susceptibility loci in Chinese women, Cancer Epidemiol., Biomarkers Prev., 2010, vol. 19, no. 9, pp. 2357–2365. https://doi.org/10.1158/1055-9965.EPI-10-0054
Ma, L., Teruya-Feldstein, J., and Weinberg, R.A., Tumour invasion and metastasis initiated by microRNA-10b in breast cancer, Nature, 2007, vol. 449, no. 7163, pp. 682–688. https://doi.org/10.1038/nature06174
MacLachlan, T.K., Sang, N., and Giordano, A., Cyclins, cyclin-dependent kinases and Cdk inhibitors: implications in cell cycle control and cancer, Crit. Rev. Eukaryotic Gene Expression, 1995, vol. 5, no. 2, pp. 127–156. https://doi.org/10.1615/critreveukargeneexpr.v5.i2.20
Mavaddat, N., Pharoah, P.D., Michailidou, K., et al., Prediction of breast cancer risk based on profiling with common genetic variants, J. Natl. Cancer Inst., 2015, vol. 107, no. 5, art. ID djv036. https://doi.org/10.1093/jnci/djv036
Mealiffe, M.E., Stokowski, R.P., Rhees, B.K., et al., Assessment of clinical validity of a breast cancer risk model combining genetic and clinical information, J. Natl. Cancer Inst., 2010, vol. 102, no. 21, pp. 1618–1627. https://doi.org/10.1093/jnci/djq388
Michailidou, K., Beesley, J., Lindstrom, S., et al., Genome-wide association analysis of more than 120,000 individuals identifies 15 new susceptibility loci for breast cancer, Nat. Genet., 2015, vol. 47, no. 4, pp. 373–380. https://doi.org/10.1038/ng.3242
Möller, S., Mucci, L.A., Harris, J.R., et al., The heritability of breast cancer among women in the nordic twin study of cancer, Cancer Epidemiol. Prev. Biomarkers, 2016, vol. 25, no. 1, pp. 145–150. https://doi.org/10.1158/1055-9965.EPI-15-0913
Mu, K., Wu, Z.Z., Yu, J.P., et al., Meta-analysis of the association between three microRNA polymorphisms and breast cancer susceptibility, Oncotarget, 2017, vol. 8, no. 40, pp. 68809–68824. https://doi.org/10.18632/oncotarget.18516
Muranen, T.A., Mavaddat, N., Khan, S., et al., Polygenic risk score is associated with increased disease risk in 52 Finnish breast cancer families, Breast Cancer Res. Treat., 2016, vol. 158, no. 3, pp. 463–469. https://doi.org/10.1007/s10549-016-3897-6
Na, Li., Ping, Zhou., Jian, Zheng., et al., A polymorphism rs12325489C>T in the LincRNA-ENST00000515084 exon was found to modulate breast cancer risk via GWAS-based association analyses, PLoS One, 2014, vol. 9, no. 5, pp. e98251–e98251. https://doi.org/10.1371/journal.pone.0098251
Orr, N., Dudbridge, F., Dryden, N., et al., Fine-mapping identifies two additional breast cancer susceptibility loci at 9q31.2, Hum. Mol. Genet., 2015, vol. 24, no. 10, pp. 2966–2984. https://doi.org/10.1093/hmg/ddv035
Özgöz, A., İçduygu, F.M., Yükseltürk, A., et al., Low-penetrance susceptibility variants and postmenopausal oestrogen receptor positive breast cancer, J. Genet., 2020, vol. 99, no. 1, art. ID 15. https://doi.org/10.1007/s12041-019-1174-2
Pace, A., Barone, G., Lauria, A., et al., Hsp60, a novel target for antitumor therapy: structure-function features and prospective drugs design, Curr. Pharm. Des., 2013, vol. 19, no. 15, pp. 2757–2764. https://doi.org/10.2174/1381612811319150011
Peto, J., Collins, N., Barfoot, R., et al., Prevalence of BRCA1 and BRCA2 gene mutations in patients with early-onset breast cancer, J. Natl. Cancer Inst., 1999, vol. 91, no. 11, pp. 943–949. https://doi.org/10.1093/jnci/91.11.943
Pharoah, P.D., Dunning, A.M., Ponder, B.A., et al., Association studies for finding cancer-susceptibility genetic variants, Nat. Rev. Cancer, 2004, vol. 4, no. 11, pp. 850–860. https://doi.org/10.1038/nrc1476
Qi, P., Wang, L., Zhou, B., et al., Associations of miRNA polymorphisms and expression levels with breast cancer risk in the Chinese population, Genet. Mol. Res., 2015, vol. 14, no. 2, pp. 6289–6296. https://doi.org/10.4238/2015.June.11.2
Qian, B., Zheng, H., Yu, H., et al., Genotypes and phenotypes of IGF-I and IGFBP-3 in breast tumors among Chinese women, Breast Cancer Res. Treat., 2011, vol. 130, no. 1, pp. 217–226. https://doi.org/10.1007/s10549-011-1552-9
Ricol, D., Cappellen, D., El Marjou, A., et al., Tumour suppressive properties of fibroblast growth factor receptor 2-IIIb in human bladder cancer, Oncogene, 1999, vol. 18, no. 51, pp. 7234–7243. https://doi.org/10.1038/sj.onc.1203186
Safari, S., Baratloo, A., Elfil, M., et al., Evidence based emergency medicine; part 5 receiver operating curve and area under the curve, Emergency (Tehran, Iran), 2016, vol. 4, no. 2, pp. 111–113. https://doi.org/10.22037/aaem.v4i2.232
Sawyer, S., Mitchell, G., McKinley, J., et al., A role for common genomic variants in the assessment of familial breast cancer, J. Clin. Oncol., 2012, vol. 30, no. 35, pp. 4330–4336. https://doi.org/10.1200/JCO.2012.41.7469
Schwartz, M.D., Isaacs, C., Graves, K.D., et al., Long-term outcomes of BRCA1/BRCA2 testing: risk reduction and surveillance, Cancer, 2012, vol. 118, no. 2, pp. 510–517. https://doi.org/10.1002/cncr.26294
Shan, J., Dsouza, S.P., Bakhru, S., et al., TNRC9 downregulates BRCA1 expression and promotes breast cancer aggressiveness, Cancer Res., 2013, vol. 73, no. 9, pp. 2840–2849. https://doi.org/10.1158/0008-5472.CAN-12-4313
Shan, J., Mahfoudh, W., Dsouza, S.P., et al., Genome-Wide Association Studies (GWAS) breast cancer susceptibility loci in Arabs: susceptibility and prognostic implications in Tunisians, Breast Cancer Res. Treat., 2012, vol. 135, no. 3, pp. 715–724. https://doi.org/10.1007/s10549-012-2202-6
Shi, J., Zhang, Y., Zheng, W., et al., Fine-scale mapping of 8q24 locus identifies multiple independent risk variants for breast cancer, Int. J. Cancer, 2016, vol. 139, no. 6, pp. 1303–1317. https://doi.org/10.1002/ijc.30150
Shieh, Y., Hu, D., Ma, L., et al., Breast cancer risk prediction using a clinical risk model and polygenic risk score, Breast Cancer Res. Treat., 2016, vol. 159, no. 3, pp. 513–525. https://doi.org/10.1007/s10549-016-3953-2
Shieh, Y., Hu, D., Ma, L., et al., Joint relative risks for estrogen receptor-positive breast cancer from a clinical model, polygenic risk score, and sex hormones, Breast Cancer Res. Treat., 2017, vol. 166, no. 2, pp. 603–612. https://doi.org/10.1007/s10549-017-4430-2
Stacey, S.N., Manolescu, A., Sulem, P., et al., Common variants on chromosomes 2q35 and 16q12 confer susceptibility to estrogen receptor-positive breast cancer, Nat. Genet., 2007, vol. 39, no. 7, pp. 865–869. https://doi.org/10.1038/ng2064
Stacey, S.N., Manolescu, A., Sulem, P., et al., Common variants on chromosome 5p12 confer susceptibility to estrogen receptor–positive breast cancer, Nat. Genet., 2008, vol. 40, no. 6, pp. 703–706. https://doi.org/10.1038/ng.131
Starlard-Davenport, A., Allman, R., Dite, G.S., et al., Validation of a genetic risk score for Arkansas women of color, PLoS One, 2018, vol. 13, no. 10, art. ID e0204834. https://doi.org/10.1371/journal.pone.0204834
Tajbakhsh, A., Farjami, Z., Darroudi, S., et al., Association of rs4784227-CASC16 (LOC643714 locus) and rs4782447-ACSF3 polymorphisms and their association with breast cancer risk among Iranian population, EXCLI J., 2019, vol. 18, pp. 429–438. https://doi.org/10.17179/excli2019-1374
Tan, T., Zhang, K., Chen, W., Genetic variants of ESR1 and SGSM3 are associated with the susceptibility of breast cancer in the Chinese population, Breast Cancer, 2017, vol. 24, no. 3, pp. 369–374. https://doi.org/10.1007/s12282-016-0712-5
Turnbull, C., Ahmed, S., Morrison, J., et al., Genome-wide association study identifies five new breast cancer susceptibility loci, Nat. Genet., 2010, vol. 42, no. 6, pp. 504–507. https://doi.org/10.1038/ng.586
Vachon, C.M., Pankratz, V.S., Scott, C.G., et al., The contributions of breast density and common genetic variation to breast cancer risk, J. Natl. Cancer Inst., 2015, vol. 107, no. 5, art. ID dju397. https://doi.org/10.1093/jnci/dju397
Wacholder, S., Hartge, P., Prentice, R., et al., Performance of common genetic variants in breast-cancer risk models, N. Engl. J. Med., 2010, vol. 362, no. 11, pp. 986–993. https://doi.org/10.1056/NEJMoa0907727
Wang, J., Wang, Q., Liu, H., et al., The association of miR-146a rs2910164 and miR-196a2 rs11614913 polymorphisms with cancer risk: a meta-analysis of 32 studies, Mutagenesis, 2012, vol. 27, no. 6, pp. 779–788. https://doi.org/10.1093/mutage/ges052
Wang, P.Y., Gao, Z.H., Jiang, Z.H., et al., The associations of single nucleotide polymorphisms in miR-146a, miR-196a and miR-499 with breast cancer susceptibility, PLoS One, 2013, vol. 8, no. 9, art. ID e70656. https://doi.org/10.1371/journal.pone.0070656
Wen, W., Shu, X.O., Guo, X., et al., Prediction of breast cancer risk based on common genetic variants in women of East Asian ancestry, Breast Cancer Res., 2016, vol. 18, no. 1, art. ID 124. https://doi.org/10.1186/s13058-016-0786-1
Wu, Z., Wang, P., Song, C., et al., Evaluation of miRNA-binding-site SNPs of MRE11A, NBS1, RAD51 and RAD52 involved in HRR pathway genes and risk of breast cancer in China, Mol. Genet. Genomics, 2015, vol. 290, no. 3, pp. 1141–1153. https://doi.org/10.1007/s00438-014-0983-5
Xu, M., Xu, Y., Chen, M., et al., Association study confirms two susceptibility loci for breast cancer in Chinese Han women, Breast Cancer Res. Treat., 2016, vol. 159, no. 3, pp. 433–442. https://doi.org/10.1007/s10549-016-3952-3
Xu, W., Xu, J., Liu, S., et al., Effects of common polymorphisms rs11614913 in miR-196a2 and rs2910164 in miR-146a on cancer susceptibility: a meta-analysis, PLoS One, 2011, vol. 6, no. 5, art. ID e20471. https://doi.org/10.1371/journal.pone.0020471
Yu, K., Xu, J., Liu, Z., et al., Conditional inactivation of FGF receptor 2 reveals an essential role for FGF signaling in the regulation of osteoblast function and bone growth, Development, 2003, vol. 130, no. 13, pp. 3063–3074. https://doi.org/10.1242/dev.00491
Zhang, H., Zhang, Y., Yan, W., et al., Association between three functional microRNA polymorphisms (miR-499 rs3746444, miR-196a rs11614913 and miR-146a rs2910164) and breast cancer risk: a meta-analysis, Oncotarget, 2017, vol. 8, no. 1, pp. 393–407. https://doi.org/10.18632/oncotarget.13426
Zhang, Y., Zeng, X., Liu, P., et al., Association between FGFR2 (rs2981582, rs2420946 and rs2981578) polymorphism and breast cancer susceptibility: a meta-analysis, Oncotarget, 2017, vol. 8, no. 2, pp. 3454–3470. https://doi.org/10.18632/oncotarget.13839
Zheng, W., Wen, W., Gao, Y.-T., et al., Genetic and clinical predictors for breast cancer risk assessment and stratification among Chinese women, J. Natl. Cancer Inst., 2010, vol. 102, no. 13, pp. 972–981. https://doi.org/10.1093/jnci/djq170
Zheng, Y., Ogundiran, T.O., Falusi, A.G., et al., Fine mapping of breast cancer genome-wide association studies loci in women of African ancestry identifies novel susceptibility markers, Carcinogenesis, 2013, vol. 34, no. 7, pp. 1520–1528. https://doi.org/10.1093/carcin/bgt090
Zhu, R.M., Lin, W., Zhang, W., et al., Modification effects of genetic polymorphisms in FTO, IL-6, and HSPD1 on the associations of diabetes with breast cancer risk and survival, PLoS One, 2017, vol. 12, no. 6, art. ID e0178850. https://doi.org/10.1371/journal.pone.0178850
Zuo, X., Wang, H., Mi, Y., et al., The association of CASC16 variants with breast Cancer risk in a northwest Chinese female population, Mol. Med., 2020, vol. 26, no. 1, pp. 1–10. https://doi.org/10.1186/s10020-020-0137-7
FUNDING
This research is funded by University of Science, VNU-HCM under grant number T2021-50.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest. The authors declare that they have no conflicts of interest.
Statement of compliance with standards of research involving humans as subjects. During research enrollment, participants signed informed consent. The Ethical Committee approved this study of Oncology Hospital Ho Chi Minh City (no. 177/ÐÐÐ-CÐT November 18th, 2014).
About this article
Cite this article
Thanh Thi Ngoc Nguyen, Nguyen, T.H., Phan, H.N. et al. Seven-Single Nucleotide Polymorphism Polygenic Risk Score for Breast Cancer Risk Prediction in a Vietnamese Population. Cytol. Genet. 56, 379–390 (2022). https://doi.org/10.3103/S0095452722040065
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S0095452722040065