Abstract
Neurological and psychiatric disorders account for an increasing proportion of the global disease burden. Correspondingly the neuropharmaceutical industry has experienced a significant contraction in recent years resulting in a poor variety of therapies available to treat an expanding range of conditions. Perhaps the greatest contributor to this failure in drug-discovery is the lack of understanding of the underlying biology of the nervous system and how molecular scale events translate into macroscale pathologies. Due to the unique nature of the human nervous system commonly used model organisms are often poorly representative of human pathologies resulting in a need for the development of advanced in vitro models that are capable of faithfully modeling complex structures within the brain. In this prospective, strategies for the generation of neuronal circuits and cultivation of complex three-dimensional (3D) cultures are explored. Frequently these constructs provide valuable insights into systems and processes that are difficult to explore in vivo due to the isolated and delicate nature of neuronal tissues. New developments are required to assess the physiological functions of 3D tissues in vitro.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Neurodegenerative diseases and mental disorders currently present a huge burden to national healthcare systems, a burden that is predicted to become increasingly overwhelming as the Western population ages. As of 2004 mental and neurological disorders represented the greatest burden of disease in Europe[1] with up to 38.2% of Europeans suffering from some form of disorder in 2010, up from 27. 4% in 2005.[2] As of 2010 the total cost of this burden was an estimated €798 billion with 60% of this figure resulting from direct costs of treatment and the remaining 40% due to lost productivity.[3] The approach to this daunting challenge has been somewhat variable, with international efforts to map the human brain and connectome underway in the US, EU, and China, along with attempts to raise awareness and diagnosis of mental disorders, both coinciding with a 70% contraction in neuropharmaceutical drug research programs over the 10-year period between 2002 and 2012.[4] The reasons for this contraction are varied, however major contributors include extremely low rates of drug candidates reaching phase I clinical trials, with just 8% of compounds reaching such a stage between 1993 and 2004,[5] along with poor assignment of manpower and capital. Primarily this lack of success is the result of a poor understanding of the underlying biology of neurodegenerative and mental health disorders, and a paucity of effective models for the translation of in vitro assays to in vivo therapies.[6] There are numerous examples of clinical trial failures that have been based wholly on animal model results. The usefulness of animal models has been questioned for many years, with some suggesting that results from these models are not representative of likely effects found in humans due to interspecies variance. Certainly the gap between in vitro methodologies and human clinical trials has not been reduced substantially for neurodegenerative diseases, with translation of therapies into either clinical trial or therapeutics taking a considerable amount of time. With trials failing there is now a growing concern over how well-adapted future methodologies must be to bridge this gap and better inform the development of clinical trial design. In vitro models which better reflect the natural tissue are difficult to develop, yet such a tool would address these difficulties and aid the development of treatments and their translation. In this prospective, we highlight several such ambitious methodologies.
The human brain is one of the most structurally complex organs known to science and is arguably the defining feature of the “homo” genus from which we derive the intelligence and adaptability to create suitable ecological niches in almost every environment. Historically functions of the brain, such as voluntary movement or language expression, have been correlated to specific anatomical structures within the tissue with distinct neuronal populations. While this regional specialization has been validated to some degree by functional magnetic resonance imaging (fMRΓ) and studies of dysfunction,[7] it cannot be ignored that cerebral and nervous function stems from the molecular activity of neuronal populations and the synaptic circuits formed between various populations, and that as such anatomical distinctions alone are of insufficient resolution to define function. That is to say, analyses of anatomical regions of the brain alone cannot provide sufficient mechanistic explanations of a pathophysiology if the pathology arises from a breakdown of multiple circuits and inter-regional connections. More recently efforts have focused primarily on unraveling the complex network of synaptic connections formed by neurons, known as the connectome. In the human brain this connectome may amount to upwards of 1015 individual synapses,[8] and up to 1012 in mice.[9] As such tracing these networks on a cellular scale in vivo will likely remain an extreme technical challenge for the foreseeable future.
Molecular analysis of gene expression suggests that the primate central nervous system (CNS) is produced by a series of evolutionarily adaptive gene networks.[10] Genes relating to the development of the nervous system including “AHI1”, which aids axonal path finding, and “FOXP2”, which is required for the development of vocalization, are differentially expressed in primate development compared with common mammalian model organisms and often contain mutations unique to Homo sapiens.[11] Although the use of standardized Carnegie stages has been extrapolated to neurogenesis during development, the underlying genetic and molecular differences between humans and model organisms result in significantly different end-points of development While the initiation of developmental landmarks may be comparable, the progression of this development clearly results in vastly different cytoarchitecture as demonstrated by the greatly increased encephalization quotient and cerebral volume found in H. sapiens. Early in the development of the primate brain unique patterns are evident, such as the differentiation of distinct inner and outer radial glia (a supporting cell type specifically involved in establishing neuronal architecture during neurogenesis) separated by an inner fiber layer of the sub-ventricular zone that is not present in rodent neurogenesis.[12] Considering these translational difficulties, it is perhaps unsurprising that existing in vivo models are often unfit for proving efficacy in the treatment of neurological and psychiatric disorders, and that in vitro models of isolated neuronal circuits using human-derived cell cultures could potentially provide a platform from which neuropharmaceutical drug discovery could be revolutionized. Such brain-on-a-chip devices offer great potential, with advancing technologies emerging and combining to enable highly precise neuronal circuits as excellent functional mimics of discrete neural tissue.
Biochemical niche of neurogenesis
Novel in vitro cell culture methodologies, particularly within three-dimensional (3D) culture, have looked toward mimicking environmental cues found in the normal cellular niche. These have included chemical and biochemical markers (often added as supplements to media), micro and nanotopographical cues (to mimic extracellular structures) and more recently mechanical properties, such as stiffness and elasticity. The latter has been a focal point of 3D culture systems due to the use of hydrogels to support cell culture. Here we review some of the main drivers and highlight research in this area.
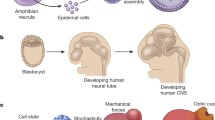
The mechanical properties of nervous tissue are in large part dictated by the make-up of the extracellular matrix (ECM), which changes dramatically according to a predictable spatio-temporal pattern over the course of development and maturation.[13] During development, neuronal outgrowth and circuit formation is guided by a series of chemotrophic factors and biophysical cues (Fig. 1). Radial glia migrate from the basal lamina of the cerebral sub-ventricular zone and lay down the developmental neural matrix along which neural progenitor cells migrate. Multiple distinct populations of radial glia have been identified[14] with sub-populations providing progenitor cells for the rapidly expanding glial and neuronal populations.[15] In the developing cortex, distinct anatomical regions are formed by varying composition of the ECM and associated growth factors.[16] The cell bodies of neural progenitors remain in a laminin-rich region of the ventricular zone and migrate along the processes of radial glia extending to the basal lamina and pia mater. These processes pass through a region rich in chondroitin sulfate proteogylcans (CSPGs) in the cortical intermediate zone before making contact with the laminin-rich basal lamina,[17] thus as progenitor cell migrate along extended processes of radial glia, they encounter a variety of distinct mechanical and biochemical environments due to differential mechanical properties, exact matrix composition, and retention of growth factors, which serve to guide differentiation and axon outgrowth.[18,19] Ventral-dorsal and rostral-caudal patterning are achieved via gradients of Wnt/bone morphogenic protein, and retinoic acid/fibroblast growth factor, respectively[20] from which neural progenitors differentiate to form the spinal cord, hindbrain, midbrain, and forebrain.
Neurogenesis in the mammalian brain occurs over the course of a series of well-defined processes. Briefly, neuroepithelial cells and radial glial adhere to the laminin-rich meninges and ventricular zone. Neural and glia progenitor cells migrate along these processes to the relevant region of the brain passing through a variety of biochemical niches. This highly complex process is reviewed extensively.[21] Copied with permission from Paridaen and Huttner.[21]
Mature neuronal ECM
The decellularized ECM of nervous tissue can promote varied cellular responses depending upon its anatomical source. ECM derived from the spinal cord has been demonstrated to produce significantly reduced neurite outgrowth in vitro in 3D hydrogel-based cultures of N1E-115 cells, compared with cortical ECM derivatives.[22] Cortical ECM contains a significantly higher proportion of sulfated glycosaminoglycans (GAGs) than that derived from the spinal cord, the latter showing a significantly greater proportion of collagen within its composition.[23] These differences may influence cell behavior by varying mechanical properties or due to differential binding of integrin proteins on the neuronal membrane.[24] Work by Saha et al.[25] highlighted that neuronal stem cell proliferation and differentiation fate can be guided by the stiffness of the substrate material. On materials with a Young’s modulus ≤10 Pa both proliferation and differentiation capacity are inhibited; proliferation and neuronal differentiation are promoted in the range modulus 100–500 Pa, whereas glial differentiation is greatly favored on materials with a modulus of 1000–10,000 Pa.
The ECM of the CNS (Fig. 2) is comprised a dense network of α-laminin and collagen IV patterned with a variety of linker proteins which serves as a barrier to guide patterning of radial glia migration and subsequent cellular architecture of neuronal progenitor cells during development.[26] The remaining neuronal ECM can be divided into the diffuse interstitial matrix and the much denser perineuronal nets (PNNs) that surround neuronal soma, nodes of Ranvier, and unmyelinated neurites.[27] In both matrices, the bulk material is comprised of hyaluronan and CSPGs onto which covalently bound peptide motifs and proteoglycans are presented, creating more complex regional cellular patterning. The PNN forms as a neuron becomes mature and typically exhibits inhibitory activity with regards to plasticity and neurite outgrowth.[28] Beyond the GAG backbone, the PNN is comprised primarily of the CSPG aggrecan, hyaluronan, proteoglycan link protein 1 (HAPLN1), revican, and tenascin. Removal of various components of the PNN, including aggrecan,[29] and tenascin-R,[30] has been demonstrated to recover rates of neurite outgrowth however whether this is due to changes in the mechanical properties of the perineuronal space, removal of an inhibitory chemical signal, or some combination of both factors is not yet fully understood.
The neuronal ECM forms a collagen and laminin-rich basal lamina surrounding cerebral blood vessels. A diffuse ECM consisting primarily of hyaluronan and CSPGs such as aggrecan forms the bulk of the tissue; however, this matrix condenses around neuronal cell bodies and synapses to form a dense perineural net that inhibits plasticity and cell migration.[27] Coped with permission from Bradbury and Burnside.[27]
In vitro tissue slice culture models
Organotypic tissue-slice cultures are a commonly used method of assessing the physiological properties of anatomical structures in their native conformation. While these methods provide a means to investigate/interrogate living tissue, there are disadvantages over in vivo studies surrounding the damage to outer cell layers when cutting, as well as limitations on functional tissue being dissected due to loss of region connectivity along an axis orthogonal to the slide direction. Notwithstanding, tissue slice models offer major benefits over single-cell populations studies in conventional well plates, not least due to the complexity of tissue obtained and plethora of interconnecting neural cell types.
The first description of nervous tissue in slice culture appears in the early 1960s following the work of Bousquet and Meunier.[31] Organotypic cultures allow for the investigation of cell function and response to stimuli, while the cells retain the native ECM and phenotypic distribution, and may be cultured for several weeks or months depending on the age of the donor tissue and substrate. Modern neuronal slice culture protocols call for the use of a supporting permeable membrane with the slice culture on one side and a supply of media on the other to ensure sufficient nutrient provision as well as aeration. Earlier methods of providing trophic support include in oculo cultures in which the tissue slice is grown in the anterior chamber of a rodent eye and visualized through the cornea.[32] Significant differences have been observed in mature, postnatal, and embryonic organisms in the outcomes of tissue-slice preparation. Provided sufficient trophic support is made available to the culture, a process that often requires significant optimization,[33,34] cell outgrowth and a degree of tissue remodeling can occur over the course of a slice culture’s lifetime. In slice cultures of embryonic nervous tissue, this has amounted to as much as an eightfold increase in size from E14 rat spinal cord slices.[35] Preparation of slice cultures from postnatal donors may result in similar, if reduced, rates of expansion; however, cellular phenotypes are considered to be more mature and cultures may survive for several months in optimal conditions.[36] Slice cultures prepared from adult or even post-mortem donors are likely to be more representative as models of a mature nervous system; however, there are currently few optimized techniques for their long-term culture and significant issues with neuronal cell death and gliosis remain.[36] While slice cultures preserve the native cytoarchitechture with a high degree of fidelity, their use for studying neuronal circuits as they exist in vivo is limited by the severance of extrinsic neurite projections. Although synaptic connections and neural circuits that exist within the slice are likely to be preserved, a significant proportion of the neuronal population undergoes axotomy (axonal damage) during slice preparation. In embryonic and postnatal cultures axotomy results in significant rates of neuronal cell death, which may be attenuated via treatment with neural or glial-derived neurotrophic factors via agonism of tyrosine receptor kinases.[37,38] In contrast, slice cultures from adult donors experience much higher rates of neuronal survival and reorganization of the tissue cytoarchitechture, likely due to the presence of locally produced and secreted growth factors within the slice culture.[36]
Elasticity of neuronal tissue
By their nature tissue slice cultures allow for comparatively trivial manipulation of cells in their native conformation due to removal of obstructing protective features found in vivo, such as the skull and blood-brain barrier. While representation of a healthy physiological state is restricted in neuronal slice cultures by the effects of axotomy and circuit-breaking as described above, the electrophysiological and mechanical properties of tissue samples can be readily assessed on a single-cell scale using extremely mature assays such as patch-clamping and atomic-force microscope (AFM) indentation. AFM probing of neuronal organotypic organ slice cultures has been used to demonstrate that the structural complexity of the tissue is reflected in widespread heterogeneity of mechanical properties, both within anatomical structures and between individual cells. Elkin et al.[39] demonstrated via AFM indentation on 1 mm-thick slices of rat hippocampi that the elastic modulus varies significantly between the various cell layers that make up the tissue, ranging from 308 Pa in response to a 3 µm indentation of the striatum radiatum, to 137 Pa in the CA1 pyramidal cell layer. Fine-detail mapping of the pituitary gland via AFM has also demonstrated significant variations in tissue elasticity of the order of 12 kPa/µm.[40] More recently Moeendarbary et al.[41] have demonstrated using acute slice cultures of rat tissues that stab wounds to the hippocampus and crush wounds to the spine result in significant reduction in the elasticity and stiffness of both tissues up to 400 µm from the site of injury, with a drop of 80% in elastic modulus noticed within the scar tissue itself. Significant loss of tissue stiffness and elasticity of spinal cord gray matter was observed outside the wound lesion in spinal crush models. The observed reduction in stiffness and elasticity was persistent for a 3-week period, trending toward a return to the original mechanical properties of the tissue. Upregulation of glial fibrillary acidic protein, an indication of increased glial infiltration and the formation of a glial scar was observed up to 1000 µm from the wound lesion. Together these observations oppose the long-held understanding that material stiffness is able to some degree to direct neural stem cell fate, with glial differentiation occurring preferentially on stiffer substrates.[25,42,43] However, this reduction in stiffness may also simply be due to temporary changes in the ECM generated by increased rates of cell migration and recruitment to the wound site.[43]
Organoid fabrication
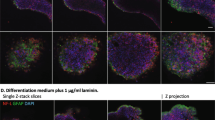
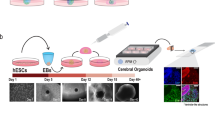
Isolated pluripotent or embryonic stem cells provide an alternative approach to modeling in vivo cellular environment. These cells can be cultured to produce non-adhered spheroids, also termed “neurospheres” or more recently “organoids”. These can be guided to present different differentiated cell types through the application of specific differentiation factors doped into the cell culture media in vitro. These 3D biologic constructs present organized cytoarchitechture without the restrictions that two-dimensional (2D) (largely monolayer) cell culture offers, which can be used to understand migration, growth factor distribution and focal adhesion (FA) clustering.[44] Organoids can be constructed either through aggregation of non-adherent cells within a supporting hydrogel matrix such as Matrigel, or through the seeding of cells onto a scaffold substrate. As the focus of organoid culture is to mimic the native conformation of a given tissue in vitro, decellularized samples of ECM present an ideal substrate to recreate a biomimetic niche and have been used to generate renal organoids,[45] cochlear tissue,[46] and nervous tissue.[47,48] While appealing, the use of decellularized matrices to generate neural organoids presents serious technical challenges due to the delicate and cell-dense nature of the neural tissue. The unique composition of neuronal ECM plays an essential role in guiding neuroblast and glial progenitor differentiation that would likely be absent in non-neuronal ECM as determined by comparisons of neuronal differentiation and rates of neurite outgrowth on cortical, spinal, and bladder-derived ECM.[27,49] Native decellularized ECM presents an ideal replica of a mature tissue, although it suffers from distinct limitations such as an inability to recreate developmental guidance cues due to the temporal changes that occur both in the cellular population, and distribution of growth factors and ECM components.[50] As such it has become preferable to fabricate organoid cultures using embryonic or pluripotent stem cells to generate an in vitro neuroepithelium that can then be directed via the addition of small molecules and growth factors to undergo neurogenesis.[51–53] The formation of neural and cortical organoid constructs from induced human pluripotent stem cells has gained increasing attention as a tool from which the developmental processes of the human brain [54–56] and its response to stimuli and treatments can be studied.[56] Certainly, these model systems offer potential for investigative approaches of multiplexed organoid cell structures [Fig. 3(a)], although discrete neuronal connectivity is difficult to interrogate.
(a) Tissue slice of a cerebral organoid generated using human-induced pluripotent stem cells.[54] Scale bar represents 1000 µm. The pluripotency marker SOX2 is seen to be localized around pseudo-ventricles as found in vivo. The TUJ1 neuronal filament marker is seen localized primarily to exterior cortical layers, again mimicking anatomical features of the tissue found in vivo. While a useful tool for developmental studies, such constructs suffer from many of the issues found in living neuronal tissue, notably a complexity of neural circuitry that is difficult to trace and untangle. (b, c) Schematic representation of permissive and promoting hydrogels, respectively. While permissive hydrogels readily support the growth of floating cellular aggregates in three dimensions, promoting hydrogels allow for integrin-binding and cell-stretching, which play a role in organization of nuclear actin and thus gene expression. Cells that are bound to a 3D matrix more accurately represent cells in their native niche in vivo. Copied with permission from Lancaster et al.[55] and Tibbitt and Anseth.[80]
From constructs to circuits
The use of contact guidance assays to manipulate and guide neurite outgrowth has been a major tool in developing our understanding of how neurons align and form organized networks in vivo using in vitro culture techniques. Chemical patterning and physical guidance using micropatterning fabrication techniques allows for the control of cell alignment and serves to direct neurite outgrowth such that neural circuits can be created in a controlled and defined manner, allowing for the relative comparison of different material properties and features on neuronal functions. It is worth noting that on the scale with which cells interact with their substrate, micropatterning techniques often generate physical structures that may themselves serve to guide FA formation. Investigations into the molecular process by which cells adhere to surface motifs demonstrate that cells are sensitive to topographical cues on the nanometer scale; collagen I, II, III, and V fibrils in vivo all present a banding pattern known as the D-period with a regular interval of 67 nm.[57] Removal of this D-period in collagen I has been shown to completely erase the ability of aligned fibrils on a 2D substrate to produce cultures of aligned fibroblasts in vitro suggesting a key role played by this pattern in guiding cell alignment.[58] Indeed, molecular analysis of the internal molecular architecture of FAs within the cell membrane suggests a minimum interval of ~20–40 nm between adjacent FAs,[59] while in vitro studies have demonstrated that FA formation is inhibited if integrin-binding features large enough to support a single integrin dimer are separated by distances >73 nm.[60] As the size of an individual integrin dimer is in the region of 8–12 nm[61] these findings suggest that integrin clustering during FA formation plays an essential role in mechanotransduction and in helping the cell sense the physical dimensions of its environment. Due to these realities, it should be noted that methods of chemical surface patterning that do not maintain a feature height below this 40 nm range should not truly be considered 2D from a cellular perspective. Certain commonly used fabrication methods, such as wet lithography nanopatterning,[62] should be approached with caution as the resulting patterns are often treated as 2D despite producing features that range up to a micrometer above the substrate surface—this is commonly termed 2.5D.[63] When reviewing work using structured surfaces it is essential to unpick the effects of surface features “visible” to cells, be these either chemical modifications of the surface or structures presenting defined edges with which cells can interact. Structures in the nanometer range are often considered too small for cells to directly interact, yet the proteins adsorbing on them are known to be altered by features in this size range and subsequently impact on cell-surface interaction. The competitive adsorption of serum proteins to a substrate, known as the Vroman effect[64] suggests that in serumcontaining cultures cell-substrate interactions will be mediated by an intervening layer of adsorbed serum proteins. Thus, in standard culture protocols great care should be taken to identify whether cell responses to chemical patterning are truly the result ofintegrin-substrate binding, due to the physical structure of the pattern itself, or due to the differential protein adsorption and conformation of serum components bound to varying substrate components.[65]
2D neuronal cell circuit formation
The 2D model circuits have been generated in vitro using polydimethylsiloxane (PDMS) stamps and devices generated via photolithographic fabrication methods to align segregated populations. Small diameter microfluidic channels, typically in the range of 5–50 µm, are used to guide neurite outgrowth such that these segregated populations form connected circuits that can model circuits identified in vivo.[66–68] Such devices can be readily adhered to microelectrode arrays (MEAs), generating a controlled neural circuit that can be assayed to record synaptic activity and action potential spike patterns in long-term cultures, providing valuable functional information regarding the effects of downstream signaling within a circuit following insult or treatment of one of the defined regions with a chemical agent.[69] Such MEA-linked devices provide quantitative information regarding the signaling activity within a circuit; however, are limited by the need for cell-electrode contact to be made for consistent measurements. In contrast, more mature techniques for measuring neuronal activity, such as imaging of calcium fluxes, provide more qualitative and temporally limited data, but are more directly applicable to in vivo analysis[70] without the problems electrode probes tend to encounter when used in vivo such as reactive gliosis producing signal impedance.[71] Although these 2D platforms give extraordinary insight into functional activity of connected neuronal networks, they do not present “normal” 3D native tissue structure, which is considered to play a dominant role in cellular function.
3D models of neuronal cell circuits
Biologic and synthetic polymers have been utilized in hydrogel-based 3D cell culture.[72,73] These are comprised a network of hydrophilic polymer chains that sequester copious quantities of water, up to hundreds of times the dry weight of the polymer content of the gel.[74] The polymers used, and network morphology fabricated, can be altered to change the chemical and physical characteristics of these materials. A variety of techniques have been established to provide some degree of control over these properties. Briefly, these techniques include freeze-thawing of gels on aligned surfaces,[75] modifying the cross-linking agent and target sites on the polymer chains,[76] modifying the relative strength of the cross-linking agent,[77] or by controlling the exposure to light in photosensitive polymers.[78]
The polymer network of a hydrogel may be fixed or dynamic depending on whether the constituent polymer chains are held together by direct covalent bonding or a more dynamic attraction such as hydrogen bonding, ionic bonding, or hydrophobic forces. Distinctions should be made between water molecules interacting directly with the polymer chains, known as bound water, and free water that fills the bulk space between polymer chains. It has been demonstrated that changes in the relative quantities of bound and free water, such as when shear stress is applied to the polymer network causing changes in polymer chain alignment, can produce significant and irreversible changes in the mechanical properties of polysaccharide-based hydrogels, which demonstrates implications in the synthesis and handling protocols when using hydrogels in cell culture.[79]
Hydrogels can further be categorized as “permissive” or “promoting” (Fig. 3). Promoting hydrogels consist of biologically derived or otherwise modified polymers that present chemical motifs that are recognized by integrin proteins on the cellular membrane, allowing the cell to adhere to the polymer chain and spread, whereas permissive hydrogels support the proliferation of an encapsulated non-adherent cell population.[80] While both hydrogels support the formation of an endogenous ECM that is laid down by the cells in culture, it has been demonstrated that cell spreading plays a significant role in mechanotransduction and results in significant reorganization of nuclear actin,[81] suggesting cellular adhesion to, as opposed to simple encapsulation within, a supporting matrix may play a role in mediating gene expression. Similarly encapsulation within a hydrogel that does not itself support cellular adhesion is associated with significantly reduced neurite outgrowth and as such does not support functionality in neuronal culture.[82,83] Furthermore, hydrogel random interconnected pore structure does not enable control of neurite outgrowth direction. Formation of designer neuronal connectivity, i.e., artificially designed living cell circuits, requires the use of material cues such as those topographic or chemical patterning methods described previously. While these have been employed within 2D (or sometimes termed 2.5D) materials allowing this directional control within 3D are highly sought. More recently numerous hydrogel materials have been used to specifically mimic the native properties of neural tissue and to support 3D neuronal cell cultures. Materials used for such cultures include gellan gum,[84] functionalized chitosan and hyaluronan,[85] and methacrylate-modified hyaluronic acid.[86]
Neuronal cellular guidance on 3D fibers
The use of aligned electrospun fibers to guide neurite outgrowth was first reported by Yang et al. in 2005[87] and has since achieved widespread use in neural tissue engineering and research.[88] Aligned electrospun fibers on a flat surface present a topographical cue along which neurites can extend from the neuronal soma; however, nanoscale features such as kinks that are formed when fibers overlap may interfere with this alignment.[89] The alignment of neurite outgrowth to micro- and nanoscale fibers is not entirely dogmatic, indeed it has been demonstrated that high fiber density and surface treatment with biologically relevant peptide motifs such as laminin can overcome physical alignment and produce neurite outgrowth perpendicular to the physical guidance cue.[90] Analysis of physical guidance cues and surface wettability in tandem suggest that there is no clear defining signal and that instead surface patterning of deposited serum-proteins determined by these substrate characteristics likely plays a prominent role.[91] As such it might be useful to consider physical and chemical modification of surfaces as guides for protein deposition rather than directly controlling any cellular response, although cellular adhesion can be controlled relatively precisely via pre-treatment of a surface with adhered peptide motifs or proteins that are recognized by cellular integrins.
The choice of fiber diameter has also been demonstrated to impact the physiological response of cultured cells. In vivo the diameter of neurites ranges from 0.08 to 20 µm depending on cell type, location, and presence of a myelin sheath.92 The filopodia extended by neurites to probe their surroundings and establish new FAs are typically in the range of 100–300 nm in diameter.[93] On large diameter electrospun fibers of polycaprolactone ranging from 1 to 8 µm in diameter, neurite length increases with fiber diameter, while neurite diameter decreases as fiber diameter increases.[94] Similarly, the length of the primary dendrite of rat subventricular neurons was found to increase with fiber diameter from 400-1200 nm on electrospun silk fibroin fibers, while total dendrite number was found to decrease.[95] Chick dorsal root ganglion explants cultured on electrospun fibers of poly-L-lactic acid (PLLA) with a fiber diameter ranging from 1700 to 230 nm have been similarly demonstrated to show significantly greater neurite length on wider fibers, while growth perpendicular to fiber alignment is greatest on small diameter fibers.[96] In contrast to these findings, assessment of neuronal response to aligned and randomly deposited electrospun fibers of PLLA of varying diameters conducted by He et al.[97] demonstrated a range of responses, with maximum neurite outgrowth occurring on aligned fibers with a diameter of 500 nm, which decreased at lower and higher fiber diameters. Similarly, neural progenitors derived from human embryonic stem cells cultured on silk fibroin fibers have been demonstrated by Wang et al.[98] to present significantly reduced neurite outgrowth on fiber diameters of 800 nm compared with 400 nm diameter fibers. 400 nm diameter fibers also significantly upregulated markers of neuronal differentiation, which contrasts with findings from Christopherson et al.[99] in which neural stem cells cultured on fibers of polyethersulfone showed an upregulation of markers of neuronal differentiation on fiber length of 749 nm when compared with similar fibers with a diameter of 283 nm. In conclusion, FA formation is inhibited on fibers below 100 nm in diameter, whereas fibers with a diameter of the order of a micrometer or above struggle to produce alignment of neurite outgrowth due to there being sufficient space for FA formation to develop across a much wider path.
There is clearly intense research activity in the area of novel (bio)materials and assessment of how biologic fluids, proteins, and cells are affected during interactions with these. The ambition for many of these explorations is the development of materials with advanced properties, enabling the fabrication of complex neural architecture. Nano and microfibers offer a means to rapidly produce mesh networks, onto which neural cells can adhere and be guided in the direction of the fiber axis. Further, due to the nature of the fabrication methods, these can also be layered to form 3D networks, with control over cell alignment within a 3D stack of aligned fiber meshes.[87]
Summary and conclusions
Recreating the neuronal niche in three dimensions in vitro is likely to provide significant challenges for the foreseeable future. In vivo neurons and glia interact to produce a matrix that is patterned on macro-, micro-, and nanoscales each of which provide physical guidance and chemical cues to create genetically defined functional circuits. While increased substrate stiffness can guide neural stem cells down a glial differentiation pathway in vitro, it has been demonstrated that developmental brain tissue is stiffer than that of mature brain tissue, and that glial scar tissue formation (due to trauma damage) results in softening of the surrounding matrix. Considering this uncertainty, care should be taken in the tuning of substrate mechanical properties; however, soft materials with elastic moduli in the range of 1-10 kPa are suitably biomimetic to support proliferation and differentiation of neuronal progenitors. The use of aligned fibers capable of supporting integrinmediated adhesion to align neurite outgrowth, both on flat surfaces and as a promoting hydrogel matrix, and the use of microfluidic systems to provide physical guidance are clearly viable tools with which defined neuronal networks can be established in vitro. While the specific response of a given neuron to its physical environment is still indeterminate, initial analysis from data generated thus far suggests that neurite extension and alignment is likely ideal when the width of the cell-substrate interface is in the region of 400–1000 nm; beyond this range the chance of random neurite outgrowth and loss of alignment increases. The combination of these patterning and guidance techniques, using the physical barrier created by microfluidic channels to maintain segregation of neuronal populations while permitting neurite outgrowth, along with a dynamic hydrogel matrix to replicate the 3D ECM of the native cellular environment, and micro- and nanoscale fibers to guide neurite extension, can readily allow for the fabrication of complex 3D neuronal circuits. There is clear need for advanced neural 3D in vitro models to bridge the gap between the limitations of cellularly complex yet impaired functional tissue slices and that of very basic single cell-type populations. We have highlighted here that many methods are emerging, and being synergistically combined to enable such advances, although the importance of determining the physiological parameters presented by these models must be kept in mind when comparing with the normal in vitro tissue architecture and its function.
References
C. Mathers, D.M. Fat, and J.T. Boerma: The Global Burden of Disease: 2004 Update (World Health Organization, Switzerland, 2008).
H.U. Wittchen, F. Jacobi, J. Rehm, A. Gustavsson, M. Svensson, and B. Jönsson: The size and burden of mental disorders and other disorders of the brain in Europe 2010. Eur. Neuropsychopharmacol. 21(9), 655–679 (2010).
J. Olesen, A. Gustavsson, M. Svensson, H.U. Wittchen, and B. Jönsson: The economic cost of brain disorders in Europe. Eur. J. Neurol. 19, 155–162 (2012).
inVentive Health PR Group. An Advocacy Rx for Progress in Mental Health, InVentiv Health Blog (2017). Available at: http://eddas.eu (accessed March 20, 2017).
S.E. Hyman: Revolution stalled. Sci. Transl. Med. 4(155), 155–159 (2012).
J. Skripka-Serry: The great neuro-pipeline brain drain. Drug Discov. World. Blog post (2013). Available at http://www.ddw-online.com (accessed March 22, 2017).
N. Kanwisher: Functional specificity in the human brain: a window into the functional architecture of the mind. Proc. Natl. Acad. Sci. USA 107, 11163–11170 (2010).
O. Sporns, G. Tononi, and R. Kötter: The human connectome: a structural description of the human brain. PLoS Comput. Biol. 1, e42 (2005).
S. Herculano-Houzel, B. Mota, and R. Lent: Cellular scaling rules for rodent brains. Proc. Natl. Acad. Sci. USA 103, 12138–12143 (2006).
S. Dorus, E.J. Vallender, P.D. Evans, J.R. Anderson, S.L. Gilbert, M. Mahowald, G.J. Wyckoff, C.M. Malcom, and B.T. Lahn: Accelerated evolution of nervous system genes in the origin of Homo sapiens. Cell 119, 1027–1040 (2004).
C.C. Sherwood, F. Subiaul, and T.W. Zawidzki: A natural history of the human mind: tracing evolutionary changes in brain and cognition. J. Anat. 212, 426–454 (2008).
I.H. Smart, C. Dehay, P. Giroud, M. Berland, and H. Kennedy: Unique morphological features of the proliferative zones and postmitotic compartments of the neural epithelium giving rise to striate and extrastriate cortex in the monkey. Cereb. Cortex 12, 37–53 (2002).
B. Sadaghiani, B.J. Crawford, and J.R. Vielkind: Changes in the distribution of extracellular matrix components during neural crest development in Xiphophorus spp. embryos. Can. J. Zool. 72, 1340–1353 (1994).
E. Hartfuss, R. Galli, N. Heins, and M. Götz: Characterization of CNS precursor subtypes and radial glia. Dev. Biol. 229, 15–30 (2001).
T.E. Anthony, C. Klein, G. Fishell, and N. Heintz: Radial glia serve as neuronal progenitors in all regions of the central nervous system. Neuron 41, 881–890 (2004).
R.T. Ferri and P. Levitt: Regulation of regional differences in the differentiation of cerebral cortical neurons by EGF family-matrix interactions. Development 121, 1151–1160 (1995).
C.S. Barros, S.J. Franco, and U. Müller: Extracellular matrix: functions in the nervous system. Cold Spring Harb. Perspect. Biol. 3, a005108 (2011).
J.M. Barnes, L. Przybyla, and V.M. Weaver: Tissue mechanics regulate brain development, homeostasis and disease. J. Cell Sci. 130, 71–82 (2017).
D.E. Koser, A.J. Thompson, S.K. Foster, A. Dwivedy, E.K. Pillai, G.K. Sheridan, H. Svoboda, M. Viana, L. da F Costa, J. Guck, and C.E. Holt: Mechanosensing is critical for axon growth in the developing brain. Nat. Neurosci. 19, 1592–1598 (2016).
T.J. Petros, J.A. Tyson, and S.A. Anderson: Pluripotent stem cells for the study of CNS development. Front. Mol. Neurosci. 4, 30–41 (2011).
J.T. Paridaen and W.B. Huttner: Neurogenesis during development of the vertebrate central nervous system. EMBO Rep. 15, 351–364 (2014).
C.J. Medberry, P.M. Crapo, B.F. Siu, C.A. Carruthers, M.T. Wolf, S.P. Nagarkar, V. Agrawal, K.E. Jones, J. Kelly, S.A. Johnson, and S.S. Velankar: Hydrogels derived from central nervous system extracellular matrix. Biomaterials 34, 1033–1040 (2013).
J.T. Rutka, G. Apodaca, R. Stern, and M. Rosenblum: The extracellular matrix of the central and peripheral nervous systems: structure and function. J. Neurosurg. 69, 155–170 (1988).
R. Eva and J. Fawcett: Integrin signalling and traffic during axon growth and regeneration. Curr. Opin. Neurobiol. 27, 179–185 (2014).
K. Saha, A.J. Keung, E.F. Irwin, Y. Li, L. Little, D.V. Schaffer, and K.E. Healy: Substrate modulus directs neural stem cell behavior. Biophys. J. 95, 4426–4438 (2008).
P.K. Grant and C.B. Moens: The neuroepithelial basement membrane serves as a boundary and a substrate for neuron migration in the zebrafish hindbrain. Neural Dev. 5, 9 (2010).
E.R. Burnside and E.J. Bradbury: Review: manipulating the extracellular matrix and its role in brain and spinal cord plasticity and repair. Neuropathol. Appl. Neurobiol. 40, 26–59 (2014).
A. Dityatev, G. Brückner, G. Dityateva, J. Grosche, R. Kleene, and M. Schachner: Activity-dependent formation and functions of chondroitin sulfate-rich extracellular matrix of perineuronal nets. Dev. Neurobiol. 67, 570–588 (2007).
K.A. Giamanco, M. Morawski, and R.T. Matthews: Perineuronal net formation and structure in aggrecan knockout mice. Neuroscience 170, 1314–1327 (2010).
P. Weber, U. Bartsch, M.N. Rasband, R. Czaniera, Y. Lang, H. Bluethmann, R.U. Margolis, S.R. Levinson, P. Shrager, D. Montag, and M. Schachner: Mice deficient for tenascin-R display alterations of the extracellular matrix and decreased axonal conduction velocities in the CNS. J. Neurosci. 19, 4245–4262 (1999).
J. Bousquet and J.M. Meunier: Organotypic culture, on natural and artificial media, of fragments of the adult rat hypophysis. C. R. Seances Soc. Biol. Fil. 156, 65–67 (1962).
A.G. Bragin and O.S. Vinogradova: Comparison of neuronal activity in septal and hippocampal grafts developing in the anterior eye chamber of the rat. Dev. Brain Res. 10, 279–286 (1983).
R.L. Fisher and A.E. Vickers: Preparation and culture of precision-cut organ slices from human and animal. Xenobiotica 43, 8–14 (2013).
D. Pozzi, J. Ban, F. Iseppon, and V. Torre: An improved method for growing neurons: comparison with standard protocols. J. Neurosci. Methods 280, 1–10 (2017).
A. Henschen, B. Hoffer, and L. Olson: Spinal cord grafts in oculo: survival, growth, histological organization and electrophysiological characteristics. Exp. Brain Res. 60, 38–47 (1985).
J. Marksteiner and C. Humpel: Beta-amyloid expression, release and extracellular deposition in aged rat brain slices. Mol. Psychiatry 13, 939–952 (2008).
S. Alcántara, J. Frisén, J.A. del Río, E. Soriano, M. Barbacid, and I. Silos-Santiago: TrkB signaling is required for postnatal survival of CNS neurons and protects hippocampal and motor neurons from axotomy-induced cell death. J. Neurosci. 17, 3623–3633 (1997).
C. Weis, J. Marksteiner, and C. Humpel: Nerve growth factor and glial cell line-derived neurotrophic factor restore the cholinergic neuronal phenotype in organotypic brain slices of the basal nucleus of Meynert. Neuroscience 102, 129–138 (2001).
B.S. Elkin, E.U. Azeloglu, K.D. Costa, and B. Morrison III: Mechanical heterogeneity of the rat hippocampus measured by atomic force microscope indentation. J. Neurotrauma 24, 812–822 (2007).
N. Bouchonville, M. Meyer, C. Gaude, E. Gay, D. Ratel, and A. Nicolas: AFM mapping of the elastic properties of brain tissue reveals kPa mm(−1) gradients of rigidity. Soft Mat. 12, 6232–6239 (2016).
E. Moeendarbary, I.P. Weber, G.K. Sheridan, D.E. Koser, S. Soleman, B. Haenzi, E.J. Bradbury, J. Fawcett, and K. Franze: The soft mechanical signature of glial scars in the central nervous system. Nat. Commun. 8, 14787 (2017).
N.D. Leipzig and M.S. Shoichet. The effect of substrate stiffness on adult neural stem cell behavior. Biomaterials 30, 6867–6878 (2009).
P. Moshayedi, G. Ng, J.C. Kwok, G.S. Yeo, C.E. Bryant, J.W. Fawcett, K. Franze, and J. Guck: The relationship between glial cell mechanosensitivity and foreign body reactions in the central nervous system. Biomaterials 35, 3919–3925 (2014).
B.M. Baker and C.S. Chen: Deconstructing the third dimension—how 3D culture microenvironments alter cellular cues. J. Cell Sci. 125, 3015–3024 (2012).
M. Caralt, J.S. Uzarski, S. Iacob, K.P. Obergfell, N. Berg, B.M. Bijonowski, K.M. Kiefer, H.H. Ward, A. Wandinger-Ness, W.M. Miller, and Z.J. Zhang: Optimization and critical evaluation of decellularization strategies to develop renal extracellular matrix scaffolds as biological templates for organ engineering and transplantation. Am. J. Transpl. 15, 64–75 (2015).
A.J. Mellott, H.E. Shinogle, J.G. Nelson-Brantley, M.S. Detamore, and H. Staecker: Exploiting decellularized cochleae as scaffolds for inner ear tissue engineering. Stem Cell Res. Ther. 8, 41 (2017).
J. De Waele, K. Reekmans, J. Daans, H. Goossens, Z. Berneman, and P. Ponsaerts: 3D culture of murine neural stem cells on decellularized mouse brain sections. Biomaterials 41, 122–131 (2015).
D. Sood, K. Chwalek, E. Stuntz, D. Pouli, C. Du, M. Tang-Schomer, I. Georgakoudi, L.D. Black III, and D.L. Kaplan: Fetal brain extracellular matrix boosts neuronal network formation in 3d bioengineered model of cortical brain tissue. ACS Biomater. Sci. Eng. 2, 131–140 (2015).
A. Faissner and J. Reinhard: The extracellular matrix compartment of neural stem and glial progenitor cells. Glia 63, 1330–1349 (2015).
Temple S.: The development of neural stem cells. Nature 414, 112–117 (2001).
K. Watanabe, D. Kamiya, A. Nishiyama, T. Katayama, S. Nozaki, H. Kawasaki, Y. Watanabe, K. Mizuseki, and Y. Sasai: Directed differentiation of telencephalic precursors from embryonic stem cells. Nat. Neurosci. 8, 288–296 (2005).
M. Eiraku, K. Watanabe, M. Matsuo-Takasaki, M. Kawada, S. Yonemura, M. Matsumura, T. Wataya, A. Nishiyama, K. Muguruma, and Y. Sasai: Self-organized formation of polarized cortical tissues from ESCs and its active manipulation by extrinsic signals. Cell Stem Cell 3, 519–532 (2008).
A.M. Maroof, S. Keros, J.A. Tyson, S.W. Ying, Y.M. Ganat, F.T. Merkle, B. Liu, A. Goulburn, E.G. Stanley, A.G. Elefanty, and H.R. Widmer: Directed differentiation and functional maturation of cortical interneurons from human embryonic stem cells. Cell Stem Cell 12, 559–572 (2013).
J. Mariani, M.V. Simonini, D. Palejev, L. Tomasini, G. Coppola, A.M. Szekely, T.L. Horvath, and F.M. Vaccarino: Modeling human cortical development in vitro using induced pluripotent stem cells. Proc. Natl. Acad. Sci. USA 109, 12770–12775 (2012).
M.A. Lancaster, M. Renner, C.A. Martin, D. Wenzel, L.S. Bicknell, M.E. Hurles, T. Homfray, J.M. Penninger, A.P. Jackson, and J.A. Knoblich: Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
A.M. Paşca, S.A. Sloan, L.E. Clarke, Y. Tian, C.D. Makinson, N. Huber, C.H. Kim, J.Y. Park, N.A. O’rourke, K.D. Nguyen, and S.J. Smith: Functional cortical neurons and astrocytes from human pluripotent stem cells in 3D culture. Nat. Methods 12, 671–678 (2015).
R. Mallinger, W. Kulnig, and P. Böck: Symmetrically banded collagen fibrils: observations on a new cross striation pattern in vivo. Anat. Rec. 232, 45–51 (1992).
K. Poole, K. Khairy, J. Friedrichs, C. Franz, D.A. Cisneros, J. Howard, and D. Mueller: Molecular-scale topographic cues induce the orientation and directional movement of fibroblasts on two-dimensional collagen surfaces. J. Mol. Biol. 349, 380–386 (2005).
I. Patla, T. Volberg, N. Elad, V. Hirschfeld-Warneken, C. Grashoff, R. Fässler, J.P. Spatz, B. Geiger, and O. Medalia: Dissecting the molecular architecture of integrin adhesion sites by cryo-electron tomography. Nat. Cell Biol. 12, 909–915 (2010).
M. Arnold, E.A. Cavalcanti-Adam, R. Glass, J. Blümmel, W. Eck, M. Kantlehner, H. Kessler, and J.P. Spatz: Activation of integrin function by nanopatterned adhesive interfaces. Chemphyschem 5, 383–388 (2004).
X. Wang, K. Ye, Z. Li, C. Yan, and J. Ding: Adhesion, proliferation, and differentiation of mesenchymal stem cells on RGD nanopatterns of varied nanospacings. Organogenesis 9, 280–286 (2013).
M. Cavallini, C. Albonetti, and F. Biscarini: Nanopatterning soluble multifunctional materials by unconventional wet lithography. Adv. Mater. 21, 1043–1053 (2009).
B. Chelli, M. Barbalinardo, F. Valle, P. Greco, E. Bystrenova, M. Bianchi, and F. Biscarini: Neural cell alignment by patterning gradients of the extracellular matrix protein laminin. Interface Focus 4, 20130041 (2014).
S.L. Hirsh, D.R. McKenzie, N.J. Nosworthy, J.A. Denman, O.U. Sezerman, and M.M. Bilek: The Vroman effect: competitive protein exchange with dynamic multilayer protein aggregates. Colloids Surf. B: Biointerfaces 103, 395–404 (2013).
P. Roach, D. Farrar, and C.C. Perry: Interpretation of protein adsorption: surface-induced conformational changes. J. Am. Chem. Soc. 127, 8168–8173 (2005).
F. Morin, N. Nishimura, L. Griscom, B. LePioufle, H. Fujita, Y. Takamura, and E. Tamiya: Constraining the connectivity of neuronal networks cultured on microelectrode arrays with microfluidic techniques: a step towards neuron-based functional chips. Biosens. Bioelectron. 21, 1093–1100 (2006).
R. Renault, N. Sukenik, S. Descroix, L. Malaquin, J.L. Viovy, J.M. Peyrin, S. Bottani, P. Monceau, E. Moses, and M. Vignes: Combining microfluidics, optogenetics and calcium imaging to study neuronal communication in vitro. PLoS ONE 10, e0120680 (2015).
L. Pan, S. Alagapan, E. Franca, S.S. Leondopulos, T.B. DeMarse, G.J. Brewer, and B.C. Wheeler: An in vitro method to manipulate the direction and functional strength between neural populations. Front. Neural Circuits 9, 32 (2015).
M.E.J. Obien, K. Deligkaris, T. Bullmann, D.J. Bakkum, and U. Frey: Revealing neuronal function through microelectrode array recordings. Front. Neurosci. 8, 423 (2015).
C. Grienberger and A. Konnerth: Imaging calcium in neurons. Neuron 73, 862–885 (2012).
J. Groothuis, N.F. Ramsey, G.M. Ramakers, and G. van der Plasse: Physiological challenges for intracortical electrodes. Brain Stimul. 7, 1–6 (2014).
S. Van Vlierberghe, P. Dubruel, and E. Schacht: Biopolymer-based hydrogels as scaffolds for tissue engineering applications: a review. Biomacromolecules 12, 1387–1408 (2011).
J. Thiele, Y. Ma, S. Bruekers, S. Ma, and W.T. Huck: Designer hydrogels for cell cultures: a materials selection guide. Adv. Mater. 26, 125–148 (2014).
A.S. Hoffman: Hydrogels for biomedical applications. Adv. Drug Deliv. Rev. 64, 18–23 (2012).
M.K. Lee, M.H. Rich, K. Baek, J. Lee, and H. Kong: Bioinspired tuning of hydrogel permeability-rigidity dependency for 3D cell culture. Sci. Rep. 5, 8948 (2015).
S. Khetan, C. Chung, and J.A. Burdick: Tuning hydrogel properties for applications in tissue engineering. In Engineering in Medicine and Biology Society, 2009. EMBC 2009. In Annu. Int. Conf. IEEE (2094–2096). IEEE (2009).
L.S. Wang, J.E. Chung, P.P.Y. Chan, and M. Kurisawa: Injectable biodegradable hydrogels with tunable mechanical properties for the stimulation of neurogenesic differentiation of human mesenchymal stem cells in 3D culture. Biomaterials 31, 1148–1157 (2009).
A.M. Kloxin, A.M. Kasko, C.N. Salinas, and K.S. Anseth: Photodegradable hydrogels for dynamic tuning of physical and chemical properties. Science 324, 59–63 (2009).
D. Pasqui, M. De Cagna, and R. Barbucci: Polysaccharide-based hydrogels: the key role of water in affecting mechanical properties. Polymers 4, 1517–1534 (2012).
M.W. Tibbitt and K.S. Anseth: Hydrogels as extracellular matrix mimics for 3D cell culture. Biotechnol. Bioeng. 103, 655–663 (2009).
M. Plessner, M. Melak, P. Chinchilla, C. Baarlink, and R. Grosse: Nuclear F-actin formation and reorganization upon cell spreading. J. Biol. Chem. 290, 11209–11216 (2015).
K.J. Lampe, A.L. Antaris, and S.C. Heilshorn: Design of three-dimensional engineered protein hydrogels for tailored control of neurite growth. Acta Biomater. 9, 5590–5599 (2013).
E.J. Berns, Z. Álvarez, J.E. Goldberger, J. Boekhoven, J.A. Kessler, H.G. Kuhn, and S.I. Stu: A tenascin-C mimetic peptide amphiphile nanofiber gel promotes neurite outgrowth and cell migration of neurosphere-derived cells. Acta Biomater. 37, 50–58 (2016).
J.T. Koivist, T. Joki, J.E. Parraga, R. Pääkkönen, L. Ylä-Outinen, L. Salonen, I. Jönkkäri, M. Peltola, T.O. Ihalainen, and S. Narkilahti: Bioamine-crosslinked gellan gum hydrogel for neural tissue engineering. Biomed. Mater. 12(2), 025014 (2017).
T.R. Ham, M. Farrag, and N.D. Leipzig: Covalent growth factor tethering to direct neural stem cell differentiation and self-organization. Acta Biomater. 53, 140–151 (2017).
Z. Zhang, B.C. Freitas, H. Qian, J. Lux, A. Acab, C.A. Trujillo, R.H. Herai, V.A.N. Huu, J.H. Wen, S. Joshi-Barr, J.V. Karpiak, A.J. Engler, X. Fu, A.R. Muotri, and A. Almutairi: Layered hydrogels accelerate iPSC-derived neuronal maturation and reveal migration defects caused by MeCP2 dysfunction. Proc. Natl. Acad. Sci. USA 113, 3185–3190 (2016).
F. Yang, R. Murugan, S. Wang, and S. Ramakrishna: Electrospinning of nano/micro scale poly (L-lactic acid) aligned fibers and their potential in neural tissue engineering. Biomaterials 26, 2603–2610 (2005).
N.J. Schaub, C.D. Johnson, B. Cooper, and R.J. Gilbert: Electrospun fibers for spinal cord injury research and regeneration. J. Neurotrauma 33, 1405–1415 (2016).
V. Cirillo, V. Guarino, M.A. Alvarez-Perez, M. Marrese, and L. Ambrosio: Optimization of fully aligned bioactive electrospun fibers for “in vitro” nerve guidance. J. Mater. Sci., Mater. Med. 25(10), 2323–2332 (2014).
J. Xie, W. Liu, M.R. MacEwan, P.C. Bridgman, and Y. Xia: Neurite outgrowth on electrospun nanofibers with uniaxial alignment: the effects of fiber density, surface coating, and supporting substrate. ACS Nano 8(2), 1878–1885 (2014).
P. Roach, T. Parker, N. Gadegaard, and M.R. Alexander: A bio-inspired neural environment to control neurons comprising radial glia, substrate chemistry and topography. Biomater. Sci. 1, 83–93 (2013).
D. Debanne, E. Campanac, A. Bialowas, E. Carlier, and G. Alcaraz: Axon physiology. Physiol. Rev. 91, 555–602 (2011).
P.K. Mattila and P. Lappalainen: Filopodia: molecular architecture and cellular functions. Nat. Rev. Mol. Cell Biol. 9, 446–454 (2008).
M.F. Daud, K.C. Pawar, F. Claeyssens, A.J. Ryan, and J.W. Haycock: An aligned 3D neuronal-glial co-culture model for peripheral nerve studies. Biomaterials 33, 5901–5913 (2012).
J. Qu, D. Wang, H. Wang, Y. Dong, F. Zhang, B. Zuo, and H. Zhang: Electrospun silk fibroin nanofibers in different diameters support neurite outgrowth and promote astrocyte migration. J. Biomed. Mater. Res. A 101, 2667–2678 (2013).
H.B. Wang, M.E. Mullins, J.M. Cregg, C.W. McCarthy, and R.J. Gilbert: Varying the diameter of aligned electrospun fibers alters neurite outgrowth and Schwann cell migration. Acta Biomater. 6, 2970–2978 (2010).
L. He, S. Liao, D. Quan, K. Ma, C. Chan, S. Ramakrishna, and J. Lu: Synergistic effects of electrospun PLLA fiber dimension and pattern on neonatal mouse cerebellum C17. 2 stem cells. Acta Biomater. 6, 2960–2969 (2010).
J. Wang, R. Ye, Y. Wei, H. Wang, X. Xu, F. Zhang, J. Qu, B. Zuo, and H. Zhang: The effects of electrospun TSF nanofiber diameter and alignment on neuronal differentiation of human embryonic stem cells. J. Biomed. Mater. Res. A 100, 632–645 (2011).
G.T. Christopherson, H. Song, and H.Q. Mao: The influence of fiber diameter of electrospun substrates on neural stem cell differentiation and proliferation. Biomaterials 30, 556–564 (2009).
Acknowledgment
Authors would like to thank the UK EPRSC for support with funding via the centre for doctoral training in regenerative medicine (EP/L015072/1).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Merryweather, D., Roach, P. The need for advanced three-dimensional neural models and developing enabling technologies. MRS Communications 7, 309–319 (2017). https://doi.org/10.1557/mrc.2017.50
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1557/mrc.2017.50