Abstract
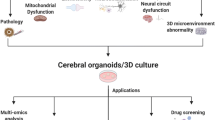
The prevalence of dementia and other neurodegenerative diseases is rapidly increasing in aging nations. These relentless and progressive diseases remain largely without disease-modifying treatments despite decades of research and investments. It is becoming clear that traditional two-dimensional culture and animal model systems, while providing valuable insights on the major pathophysiological pathways associated with these diseases, have not translated well to patients’ bedside. Fortunately, the advent of induced-pluripotent stem cells and three-dimensional cell culture now provide tools that are revolutionizing the study of human diseases by permitting analysis of patient-derived human tissue with non-invasive procedures. Specifically, brain organoids, self-organizing neural structures that can mimic human fetal brain development, have now been harnessed to develop alternative models of Alzheimer’s disease, Parkinson’s disease, motor neuron disease, and Frontotemporal dementia by recapitulating important neuropathological hallmarks found in these disorders. Despite these early breakthroughs, several limitations need to be vetted in brain organoid models in order to more faithfully match human tissue qualities, including relative tissue immaturity, lack of vascularization and incomplete cellular diversity found in this culture system. Here, we review current brain organoid protocols, the pathophysiology of neurodegenerative disorders, and early studies with brain organoid neurodegeneration models. We then discuss the multiple engineering and conceptual challenges surrounding their use and provide possible solutions and exciting avenues to be pursued. Altogether, we believe that brain organoids models, improved with classical and emerging molecular and analytic tools, have the potential to unravel the opaque pathophysiological mechanisms of neurodegeneration and devise novel treatments for an array of neurodegenerative disorders.
Similar content being viewed by others
Introduction
Epidemiological transitions and an increasing life expectancy are projected to lead the rapid expansion in the global prevalence of dementia [1]. For example, Alzheimer’s disease alone currently affects approximately half a million individuals in Canada; a number projected to double by 2031 [2]. Despite this, there are still no disease-modifying drugs available to prevent or reverse these neurodegenerative phenomena. One of the existing barriers to potential therapy development is the remarkably poorly understood nature of even common neurodegenerative diseases. While the late stage hallmarks of Alzheimer’s (AD), Parkinson’s Disease (PD) and motor neuron diseases are better characterized, early, and potentially reversible pathophysiological events remain unresolved. These gaps are in part due to the difficulty in harvesting central nervous system (CNS) tissue in early stages of neurodegenerative diseases and the inability of animal models to accurately recapitulate the neuropathological hallmarks of normal human aging and neurodegenerative disorders. Similarly, scalability and ethical barriers limit the high-throughput drug-testing potential of the established animal models.
Alternative laboratory approaches are now addressing some of these limitations. The discovery of induced pluripotent stem cells (iPSCs) has permitted the engineering of human CNS cells from non-invasive skin biopsies of patient with neurodegenerative diseases. Combined with CRISPR technology, these tools have allowed for scalable testing of neurodegeneration-associated genes and therapeutic agents entirely in human systems. However, while alleviating some concerns regarding the use of animals to study complex human disease, these two-dimensional, often monomorphic cell cultures, have raised alternate criticisms for failing to model the cellular heterogeneity and architectural complexity of human neural tissue. Fortunately, timely emerging tools are now poised to address this particular challenge. Specifically, significant advances in the development of three-dimensional (3D) culture systems allow for the generation of brain-like tissue that self-organize from pluripotent cells. Importantly, these “cerebral organoids” recapitulate many aspects of the early human brain development, notably absent from lower species including the presence of an outer subventicular zone (oSVZ), response to specific trophic factors (e.g. PDGRF), and an ability to undergo cortical folding [3,4,5,6]. Cerebral organoid techniques continue to evolve, allowing derivation from both embryonic or iPSCs, with more precise neural layering and identity [7]. Different culture conditions that aim to mimic developmental signaling patterns also allow tuning and generation of regional brain tissue structures including cerebral [8,9,10,11], optic cup [12, 13], hypothalamus [14], midbrain [15,16,17], cerebellar [18], spinal [19, 20], and hippocampal [21]-specific organoids. Given this ability to recapitulate many aspects of normal brain development, cerebral organoids have been particularly successful at modeling both normal and pathological aspects of early human brain development [22]. For example, two recent studies examined the effects of schizophrenia-associated mutations on brain development [23, 24], one of which correlated patient’s clinical phenotype with findings in their own patient-derived organoids [24]. Other studies have also helped to understand Zika-associated microcephaly [14, 25], autism spectrum disorder [26, 27], and even the effects of psychoactive drugs on human neural tissue development [28]. Despite these diverse successes of brain organoids in modeling neurodevelopmental disorders, the inherent maturational plateau at roughly the 2nd trimester fetal stage and lack of mesodermal elements (e.g. vessels, immune cells) have led to relatively fewer applications to model late-onset neurodegenerative disorders.
Here, we briefly review brain organoid culture protocols, current hypotheses of neurodegenerative mechanisms, and the early literature that has successfully applied organoid models to these adult-onset disorders. We use this as a starting point to discuss how introduction of emerging molecular strategies to promote aging, vascularization, cellular diversity, and connectivity can overcome some existing limitations. While brain organoids have been originally heralded as in vitro models of early human brain development, there is now significant reason for optimism that they can also act as powerful avatars to understand human neurodegenerative conditions.
Brain organoids culture
While brain organoids are relatively new in the scope of cell and tissue culture techniques, the protocol have rapidly evolved in the last decade. In vitro growth of 3D neural tissue (other than brain slices) was first established in 2008 using a technique coined serum-free floating culture of embryoid body-like aggregates with quick reaggregation (SFEBq) (Fig. 1a) [29]. This technique takes advantage of the property that when embryonic stem cells (ESCs) are grown in suspension, in the absence of serum or inhibitors of neural differentiation (BMP, Nodal and Wnts), they have a proclivity to aggregate into spheres and undergo spontaneous neural differentiation [29]. These neural-enriched embryoid bodies show structured apico-basal polarization and develop in cortical tissue with four distinct cell zones (ventricular, early and late cortical-plate, and an outer zone containing Cajal–Retzius cells). If left without inhibitors of neural differentiation, this cortical tissue will eventually adopt a rostral hypothalamic-like fate. A few years later, a breakthrough to further direct embryoid bodies into various brain regions came with the use of Matrigel, a gelatinous laminin-rich extracellular matrix secreted by the Engelbreth–Holm–Swarm mouse sarcoma cell line [30]. Eiraku and colleagues found that embryoid bodies stabilized in Matrigel would spontaneously develop buds of retinal primordial tissue resembling optic vesicles [12] (Fig. 1a). Then in 2013, Lancaster and Knoblich found that embryoid bodies, left in Matrigel without growth factors to promote regional identities, grew larger neuroepithelial buds that spontaneously develop adjacent brain regions to form true cerebral organoids, defined as neural structures showing representation of multiple brain regions [8] (Fig. 1b). This later approach has since become the foundation used by many groups to engineer a variety of different pharmacological cocktails to derive different regional identities including dorsal cortex [11], ventral forebrain [10], midbrain-hindbrain boundary [14,15,16], choroid plexus [21], hypothalamus [14], hippocampus [21], and spinal cord [19, 20] (Table 1, Fig. 1c). While these cerebral organoids models have already proven to be invaluable tools for studying neurodevelopmental perturbations involved in microcephaly [8, 31, 32], autism [26, 27], schizophrenia [23, 24], and Zika virus infection [3, 25], the ability to define recipes of a variety of different brain regions offers precision tools to enable study and elucidation of regional pathological mechanisms specific to more complex neurological disorders.
Basic 3D neural cell culture techniques. a Evolution of techniques to generate three-dimensional neural cultures. Green-blue shades indicate increasing neuronal fates. (i) Spontaneously aggregating neural spheres. Multipotent neural stem cells are grown for four days without retinoic acid then four days with retinoid acid. The cells are then plated for further characterization. (ii) Serum-free floating culture of embryoid body-like aggregates with quick reaggregation (SFEBq). Pluripotent cells are grown in serum-free and retinoic acid-free media (Knockout serum replacement (KSR) media) for 5 days during which they spontaneously aggregate into neural precursors. (iii) SFEBq culture followed by Matrigel addition leads to increased stability and allows for sub-organoid development (here shown forebrain organoid). b Timelines and example brightfield and immunofluorescence images of organoid protocol developed by Lancaster, Knoblich and colleagues. c Regional brain organoids generation. Differentiation of organoids into dorsal fate, ventral fate, midbrain, hypothalamus and hippocampus with the indicated lineage markers (for specific induction protocol, please see Table 1)
Neurodegeneration pathophysiological cascades
Given the limited ability of the CNS to regenerate, neurodegenerative disorders appear to be relentlessly progressive and irreversible. Therefore, much effort by the research community has been geared toward understanding and reversing the pathogenic events upstream of the well-recognized and end-stage neurodegenerative process. Cellular aging and senescence, toxic protein aggregation, reactive oxygen species (ROS), cellular quality control, and neuroinflammation are pathophysiological mechanisms that were recognized in the last few decades as central to the neurodegenerative process. However, many questions regarding the specific sequence of these events during the long evolution of the neurodegenerative process need to be resolved to understand what pathophysiological mechanisms should receive further focus in brain organoid models and eventually on drug-testing platforms (Fig. 2a, b).
Mechanisms of neurodegeneration and their investigation in brain organoids. a Interplay between mechanisms of neurodegeneration. b Dynamics of protective mechanisms in neurodegeneration. c Investigation of pathological cascades with brain organoid models. Cells from patients and controls are rejuvenated by the iPSC process and grown into brain organoids. Treatment with ROS and/or SASP -inducing stimuli, with or without drug treatment can then be applied, with endpoints consisting of neurodegeneration and/or neurodegeneration-associated proteinaceous aggregates accumulation in the brain organoids. The brain organoids can be interrogated with proteomics, epigenomics, metabolomics, architectural and functional methods in time and space to better understand the interplay between neurodegeneration mechanisms
There is perhaps little debate that one of the most common and important risk factors in the vast majority of neurodegenerative diseases is biological aging. Aging is macroscopically associated with gray and white matter atrophy, ventricular enlargement [33, 34] and histologically correlates with reduced dendritic connections and neuronal death [35]. While the pathological process of atrophy has been hypothesized to be mediated by aggregation of toxic proteins such as extracellular amyloid-β (aβ) and intracellular phosphorylated tau (pTau) in AD, these protein aggregates also accumulate in clinically healthy older individuals [36,37,38,39]. Similarly, the decline in human brain function during aging parallels that of other organs, suggesting that pan-cellular mechanisms of aging, rather than exclusive tissue-specific protein dysfunction, also contribute to the molecular underpinnings upstream of cell death and senescence [40]. Of those, the accumulation of DNA damage has emerged as an important pathophysiological process associated with neurodegeneration [41,42,43]. DNA breaks can be normally repaired by at least four distinct cellular pathways: nucleotide-excision repair (NER), base excision repair (BER), homologous recombination (HR), and non-homologous end joining (NHEJ). However, because mature neurons are post-mitotic cells, they preferentially utilize NHEJ, which by definition is error-prone and induces small genomic deletions [44]. This accumulation of such errors promotes activation of senescence and apoptosis in neurons as a hypothesized evolutionary mechanism to prevent malignant transformation [45]. In turn, the establishment of senescence-activated secretory phenotypes (SASP) in astrocytes and microglia, following neuronal DNA damage, promotes chronic neuroinflammation and is increasingly recognized as an important pathogenic mechanism in the neurodegenerative cascade [46]. Indeed, experimental and clinical studies of both progeric syndromes (e.g. Cockayne Syndrome (CS), Xeroderma Pigmentosum (XP), Ataxia-Telengiectasia (ATG), Hutchinson-Gilford progeric syndrome (HGPS)) and neurodegenerative diseases (e.g. AD, PD and Amyotrophic Lateral Sclerosis (ALS)) show increased DNA damage in examined neurons [47,48,49,50,51,52,53,54,55,56,57,58,59]. Encouragingly, increasing activity of the DNA damage checkpoint protein ATM extended the lifespan of mouse models of HGPS, providing optimism that similar approaches can alter the pathogenesis of neurodegenerative disease [60].
Other than defective DNA repair mechanisms and normal exposition to ambient radiation, intracellular ROS has been hypothesized as the most important etiology of DNA damage. Mitochondrial dysfunction has been shown to be the principal intracellular source of ROS [61]. Interestingly, mitochondrial DNA damage accumulates in both PD [54, 55] and ALS [57] and is hypothesized to create a feed-forward loop where mitochondrial DNA defects induce mitochondrial dysfunction, which in turn creates more ROS and DNA damage. In support of this, mitophagy (mitochondrial-specific autophagy) and mitochondria-derived vesicles (MDVs; vesicular mitochondrial quality pathways) are quality-control mechanism that have been shown to be defective in cellular models of PD [62]. Moreover, a recent study showed that a specific MDVs pathway normally repressed by PD-associated proteins contribute to mitochondrial antigen presentation on MHC class I molecules in the CNS, creating a possible link between DNA damage, mitochondrial quality control, and neuroinflammation [63]. In addition, a number of mitochondrial toxins have been implicated in PD pathogenesis [64]. Together, these studies suggest that mitochondria-generated ROS is central to neurodegeneration and that DNA repair and intracellular quality-control mechanisms limiting subsequent cellular oxidative stress and neuroinflammatory phenotypes could be defective in neurodegenerative disorders. Indeed, dysfunction of important cellular quality control mechanisms such as the proteasome, mitochondrial proteases, and general autophagy have also been linked to neurodegenerative disorders [65,66,67,68]. Interestingly, autophagy is activated both by ROS and DNA damage, and its dysregulation impairs DNA repair [69] and degradation of noxious aggregated proteins such as tau, α-synuclein, and amyloid-β. Paradoxically, these aggregates can also overwhelm the cellular machinery leading to autophagic dysfunction and thus creating a threshold point in these pathological cascades of neurodegeneration.
On the other hand, the role of inflammation in the neurodegenerative process appears more complex, with various neuroinflammatory cascades recently suggested to promote both neuronal survival and cell death in neurodegeneration. These dichotomous roles are best exemplified with M1 and M2 types of microglia, which show pro-inflammatory and anti-inflammatory phenotypes, respectively [70]. α-synuclein and amyloid-β protein have both been shown to differentially activate these microglia subtypes via toll-like receptor (TLR) pathways in a context-specific manner based on their conformation and cellular environment [71, 72]. Importantly, M2 microglia has been shown to help clear amyloid-β aggregates in the CNS, opening interesting pharmaceutical possibilities. Of relevance, a rare mutation in TREM2 (triggering receptor expressed on myeloid cells-2), a gene specifically regulating brain microglia, has been shown in GWAS studies to be associated with AD [73]. Conversely, activation of the SASP in astrocytes or microglia secondary to DNA damage leads to secretion of pro-inflammatory molecules that promote M1-type microglial activation to further exacerbate neuronal injury [46]. While the dichotomization of microglia into M1 and M2 types is useful to conduct experiments, it is clear that there is a continuum of functional states in microglia encompassing support, antigen presentation and phagocytosis roles [74]. Thus, a better understanding of the complex dynamics of CNS neuroinflammation cascades appears essential to identify precise pharmacological targets to tackle neurodegenerative disorders.
In light of this knowledge, organoids, by their very nature, appear as an attractive system to understand the dynamics between these various mechanisms leading to neurodegeneration. Reprogramming of adult cells to form iPSC allows engineering of young brain tissue with a lifetime of genetic events. This results in a temporally condensed system in which pathogenic molecular and cellular mechanisms upstream of protein aggregation and cell death can be interrogated, and ultimately where the entire pathological cascades can be scrutinized and characterized at every step by -OMICS technologies to establish causal relationships between genetic changes and neuronal health (Fig. 2c). iPSC-derived cerebral organoids from older patients with “sporadic” neurodegeneration provides an attractive avenue to study these relationships not possible in static pathological tissue from human brain or via inference of pathological findings from animal neurodegenerative models. Below, we discuss the pioneering studies that highlight these exciting avenues and look forward to how many of the potential challenges can be overcome.
Neurodegenerative models in brain organoids
Alzheimer’s disease
Despite the inherent limitations in modeling age-associated disorders in embryonical-derived cells, recent studies have provided significant promise for the use of cerebral organoids to understand neurodegenerative diseases. For example, by infecting human neural stem cells with lentiviral construct expressing β-amyloid precursor protein (APP) or mutated presinilin 1 (PSEN1) (found in familial Alzheimer’s disease (FAD) cases) and growing them in Matrigel, Choi and colleagues developed 3D models to study Alzheimer’s disease phenotypes [75]. While this approach experimentally deviates for the classic organoid workflow, these 3D colonies developed both extracellular β-amyloid plaques and neuronal aggregates of phosphorylated tau, findings that tend not to be simultaneously observed in two-dimensional neuronal cultures or animal models. The authors hypothesized that perhaps animal models might be too genetically removed from humans to recapitulate the neurodegenerative process and that two-dimensional neuronal cultures lack the architectural and density requirements needed to allow focal accumulation and polymerization of misfolded proteins into characteristic plaques and tangles. In addition to providing insight into the dynamics of β-amyloid plaques and neurofibrillary tangles (NFTs) formation, this model provided a novel platform for drug screening. As a proof of principle, they showed that inhibition of glycogen synthase 3 (GSK3) attenuated β-amyloid plaques and NFTs formation, confirming a hypothesis previously only shown in animal models [76, 77]. This is a timely innovation as multiple potential drugs for Alzheimer’s disease developed in animal models have failed late-phase clinical trials [78,79,80]. This 3D culture system could serve as a complementary and scalable tool to test and nominate new therapeutic agents.
Lee and colleagues devised a different approach by generating cortical organoids from sporadic AD patient-derived iPSC. Despite these more physiological conditions, not confounded by overexpression, they were also able to generate β-amyloid plaques [81]. Similarly, they showed that the pathogenic phenotype could be rescued by inhibition of β-secretase (BACE1) or γ-secretase, again demonstrating the drug-testing potential of this system. Interestingly, inhibition of these proteases was less efficient at reducing plaque load in organoids when compared to complementary two-dimensional cultures. This strengthens the hypothesis that differences in diffusion between the two culture systems may account for this potentially more representative modeling of plaque formation and may allow discovery and selection of molecules that have improved bioavailability and pharmacokinetics. Moreover, they demonstrated significant variations in protease efficacy between five patient-derived brain organoids, even with similar initial plaque loads. These findings motivated the authors to further explore which molecular factors can explain this inter-patient variations in proteases efficiency. By performing a proteome analysis of the five patient-derived organoids, they identified variations in specific protein levels (e.g. clathrin), that correlate with protease-induced plaque load reduction. Brain organoids may thus also allow a mechanism for evaluation and selection of drug in a personalized patient-specific manner. Finally, they also validated the efficacy of the BACE1 and γ-secretase inhibitors in the 3D neuronal culture model developed by Choi and colleagues and showed that they inhibited NFT formation as well. This highlights how different 3D neuronal culture models can be used in a complimentary fashion to accelerate drug discovery efforts. Similarly, the ability to form pathologic hallmarks of neurogenerative conditions such as NFT and β-amyloid plaques provides dynamic models to understand the early molecular underpinnings driving their development. Recently, Amiri and colleagues showed that repeated RNA sequencing of cerebral organoids through development provided a dynamic portrait on how the brain is patterned [82]. Similar approaches with organoid models of disease could provide temporal insight into the aberrant pathways and molecular programs operational early before NFT and plaque formation.
A third group also confirmed that brain organoids from AD patient-derived iPSCs form β-amyloid plaques and NFTs that can be reversed using β-secretase and γ-secretase inhibitors [83]. They also show that organoids preserve the abnormal endosome morphology and function previously shown in mouse models and brains of sporadic and familial AD patients [84, 85]. Interestingly, they used brain organoids derived from healthy individuals to demonstrate that they also accumulate small “baseline” levels of β-amyloid deposits and NFTs. This crucially demonstrates that these “precursor” and/or “physiological” levels of β-amyloid deposits and NFTs, commonly found in the brains of clinically healthy individuals [36,37,38,39], can also be modeled and studied. This same group later demonstrated the potential of using iPSC combined with CRISPR technology to study neurodegenerative disease pathogenic cascades by generating multiple isogenic APOE4 or APOE3 brain cell types [86]. The APOE allele (variants either APOE2, APOE3, APOE4) remains the strongest genetic factor for Alzheimer’s disease, where APOE4 is deleterious and APOE2 protective [87, 88]. Through transcriptional profiling of these cells, the authors were able to identify cell type-specific molecular pathways associated with APOE4 and identify causal relationships between AD-associated pathogenic mechanisms in these human cells. In particular, they found that APOE4 increased expression of genes involved in lipid metabolism and immune responses, partly through activation of nuclear transcription factors nuclear factor 1 (NF-1), activator protein 1 (AP-1) and NF-κB. Moreover, APOE4 increased levels of genes involved in synapse formation in neurons. Together, these APOE4-dependent changes in the transcriptome led to decreased cell proliferation, increased synaptic formation, and most importantly increased endosomal trafficking and amyloid-β42 secretion by APOE4 neurons. Moreover, when they added isogenic APOE microglia to their previously established organoid model grown from patient-derived iPSC carrying an APP duplication, they found that the APOE4 microglia displayed altered morphologies and were impaired in their ability to phagocytose amyloid-β aggregates compared to APOE3 microglia. APOE4 microglia also showed increased soluble levels of TREM2, a gene associated with AD that is increased in amyloid-β phagocytosing microglia promoting pathogenic inflammation [73]. These co-culture systems highlight the potential flexibility of organoid cultures, where various cell type with isogenic backgrounds can be combined to study the role of genetic variants in specific cell types in human tissue.
In a similar approach, Park and colleagues used a clever microfluidic system to introduce microglia into mixed 3D astrocytic and neuronal co-cultures and investigate the role of neuroinflammation in AD [89]. By doing so, they where able not only to recapitulate amyloid-β accumulation and phosphorylated tau aggregation, but also elements of neuroinflammation associated with AD. Interestingly, the healthy control-derived brain organoids produced relatively very little amyloid-β, even when cultured for a similar length to Choi and Raja and their colleagues [75, 83]. This might reflect the variability in amyloid-β and tau deposits found in humans, or potentially protective effect mediated by the initial inclusion of astrocytes in the 3D culture. By comparing AD-derived vs control-derived organoids with and without microglia, Park and colleagues were able to identify both AD-associated microglia-attracting chemokines in CCL2, CCL5, and CX3CL1 and microglia-secreted pro-inflammatory cytokines including Il-6, Il-8, TNF-α, MIF, and PAI-1. Moreover, they were able to validate the hypothesis that microglial migration to amyloid-β, secretion of pro-inflammatory molecules and subsequent neuronal loss was dependent of TLR-4 and IFN-γ. This spatially and temporally controlled addition of microglia demonstrates the potential and flexibility of 3D cultures to investigate complex cellular interactions in human tissue in the context of neurodegeneration. One could indeed imagine adding not only various cell types, but therapeutic or noxious molecules to these cultures in such controlled fashion, establishing additional cell-based high-throughput testing platforms to design symbiotic immune-based therapies.
Frontotemporal dementia
While brain organoids have proven useful to study AD, emerging studies are now also demonstrating their potential to study other neurodegenerative disorders. Seo and colleagues generated cerebral organoids from iPSCs derived from patients with Frontotemporal Dementia (FTD) [90]. These patients carried the Tau P301L mutation, and the CRISPR-CAS9 system was used to create a companion isogenic control culture by correcting this defect. Based on their previous finding that a non-cleavable mutant P35 protein attenuated amyloidosis and improved cognitive function in a FAD mouse model, they introduced a mutation in p35 (Δp35KI) in the P301L background. This showed decreased levels of phosphorylated tau and increased the expression of synaptophysin, a marker of functional synapses. Unfortunately, these authors do not compare tau phosphorylation to healthy control-derived organoids in this study. It would have been interesting to see if they could detect some phosphorylated tau in these control organoids as shown by Raja et al. as it was undetectable in wildtype mice. This could provide insight of how brain organoid models, derived from human iPSC, compare to animal models at recapitulating hallmarks of FTD neuropathology.
Parkinson’s disease
A longstanding hypothesis of the pathophysiology of PD, based mostly on neuropathological findings by Braak and Braak, is that toxic aggregates of α-synuclein can propagate themselves from the neurons in the gut, up toward the brainstem, in a prion-like manner [91]. This is also substantiated by genetic evidence, such that LRRK2, the most common genetic variant in PD (variant G2019S, N551K, N2081D), is also a genetic risk factor for Crohn’s disease (variants N551K, N2081D) [92]. However, only recently has this theory been demonstrated in animal models [93], and despite a number of associative studies, has yet to be convincingly shown in human cells or tissue [94]. A study by Son and colleagues demonstrates the power of 3D cell cultures as they interrogate the genetic changes in both intestinal and neural cellular spheres with and without LRRK2 G2019S mutations (the most common PD-associated causal mutation) [95]. While they provide evidence that LRRK2 G2019S induces similar genetic changes in synaptic transmission pathway both in neural and intestinal cells, the strength of this study was to rather demonstrate the potential of simultaneously studying multiple organ systems as organoids. Combined intestinal and brain organoids might eventually become a powerful tool to investigate synuclein transmission in human systems.
More recently, two groups took advantage of the newly developed capacity to produce region-specific brain organoids, in this case midbrain, to develop PD models. Kim and colleagues introduced the LRRK2 G2019S mutation in midbrain organoids that show both markers of dopaminergic neurons and dopamine production [96]. Interestingly, they not only found diminished dopamine transporter (DAT) and tyrosine hydroxylase (TH) markers in the LRRK G2019S organoids, but also increased activated caspase-3 levels in dopaminergic neurons, suggesting that LRRK2 G2019S mutation induces dopaminergic neuron death [96]. Moreover, they found increased phosphorylated S129 synuclein co-localizing with endosomes, reproducing the hallmark of synuclein aggregation and mislocalization in PD. Interestingly, they also compared the transcriptome of LRRK2 G2019S midbrain organoids culture vs sporadic PD-patient-derived brain tissue and found an enrichment of PD-associated genetic changes that were not found in a corresponding 2D LRRK2 G2019S culture. Finally, silencing the gene controlling the expression of TXNIP, a protein found to be central to the LRRK2 G2019S-associated transcriptomic changes seen in both midbrain organoids and sporadic PD-patient-derived brain tissue, led to a decrease in synuclein aggregation. Thus, these authors demonstrated both that midbrain organoids can be a faithful model of PD with neuropathological hallmarks but also a great comparative tool to reveal the most important pathways involved in PD. Similarly, Smith and colleagues also constructed a midbrain organoid model but instead of introducing the LRRK2 G2019S mutation, they used iPSCs generated from patient harboring the mutation, and engineered isogenic controls, to produce the organoids [97]. The midbrain organoids again displayed dopaminergic neuron marker TH and midbrain floorplate markers EN-1 and FOX A1. Interestingly, these organoids were also found to produce dopamine in an order of magnitude greater than those of Kim and colleagues (20ng/ml at day 35 and around 300 ng/ml at day 70 for Smits et al, 2ng/ml at day 45 for Kim et al.), suggesting a much higher percentage of dopaminergic differentiation. Importantly, they were able to show pacemaking activity of the dopaminergic neurons, a feature hypothesized to underly selective neuronal vulnerability to ROS and toxic agents used to model PD such as MPTP [98]. Interestingly, the patient lines, irrespective of the LRRK2 G2019S mutation, both showed approximately 3-fold reduction in the number of dopaminergic neurons growing in the midbrain organoids compared to the control cell lines. This suggests that even rejuvenation by reprogramming, and independently of the LRRK2 G2019S mutation effects, patient cell lines still carried genetic or epigenetic dysfunctions that prevented the midbrain organoids from generating as many dopaminergic neurons as those from control individuals. However, they did find that the presence of LRRK2 G2019S mutation, whether introduced in controls or naturally found in patients, seems to decrease the number of dendrites branching and thus the complexity of the dopaminergic dendritic tree, favoring non-dopaminergic neuronal proliferation.
Motor neuron disease
While there are limited studies to date that use organoids as models for motor neuron diseases, they are already showing great promise. Kawada and colleagues for example used microfluidic technology to develop axon fascicles from neural spheres [19]. Briefly, they custom made a culture microdevice containing a chamber to receive neural spheroids, a microchannel for axon fascicle formation and a target chamber accommodating axon terminals. The authors first differentiated iPSCs into motor neurons precursors by adding retinoic acid, Smoothened agonist and an FGFR antagonist prior to neural sphere formation. These led to neural spheres in which >60% of cells demonstrated HB9 staining, a motor neuron marker. Interestingly, differentiating the iPSCs in a preferred neuronal precursor population prior to organoid growth represents a different strategy than most protocols already discussed and opens additional avenues to design specific subregions in more organized and reliable manners without compromising the cellular diversity of the organoids culture system. Indeed, only a minor fraction of the organoid cells expressed GFAP and almost none expressed O4, an oligodendrocyte marker, or Nestin, a neural stem cell marker. Once the spinal motor neuron precursors-enriched neural spheres were put in the recipient chamber, they spontaneously started to grow axons in the microchannel which formed a single, straight, unidirectional fascicle by 20 days. These fascicles were shown to express both axonal marker Tau1 and presynaptic marker Synapsin1 but no nuclear or dendritic markers. Importantly, the authors observed synchronized electrophysiological activity in the organoids suggesting that the neurons form a functional network. These models were then used to demonstrate possible applications for neurodegenerative studies by using peroxide (H2O2), a ROS-generating molecule, to study its impact on the axonal bundle integrity. Not surprisingly, they found that it altered the structural integrity and coherence of the bundle, demonstrating that this model can be easily used for interrogating axonal degeneration.
Another group used similar differentiation factors (retinoic acid to caudalize, puromorphamine (Sonic Hedgehog agonist) to ventralize and promote spinal motor neuron differentiation) following the establishment of neural spheres [20]. By doing so, they yielded organoids with a homogeneous expression of neural stem cell markers Sox1 and Nestin, which later developed into rosettes with neural stem cell at the apex and a motor neuron marker (ISL1+)-expressing cells at the base. Interestingly, these motor neuron precursors were mostly positive for HOXB4, a cervical level marker, HOXC8, a brachial/thoracic level marker but not HOXC10, a lumbar level marker. Strikingly, the authors were able to show contraction of mouse myofibers co-cultured with the organoids and showed proximity of motor neuron axons (SMI-32 positive) to acetylcholine receptors at the NMJ (labeled with alpha bungarotoxin), suggesting that the spinal cord organoids can form functional synapses. Finally, they also show significant expression of astrocytic marker S100β and V1 inhibitory interneurons marker (Renshaw cells) PAX2 and LHS1 in those organoids, demonstrating more cellular diversity than those of the Kawada group, perhaps as a consequence of introducing the caudal and ventral differentiation factors after the establishment of neural spheres. Using this protocol, Hor and colleagues used iPSCs derived from Spinal Motor Atrophy (SMA) type I and type II patients to generate ventral spinal cord organoids. Interestingly, they observed that while the patient-derived organoids initially formed a similar number of ISL1+ neurons in early culture (day 14-28), they eventually expressed approximately 1.6-3.2 fold of those compared to the control organoids in later culture (day 35), suggesting that in patient-derived organoids ISL1+ neurons degenerate following relatively normal neurogenesis and differentiation. Finally, by performing transcriptome analysis, the authors confirmed previous findings from 2D culture that patient-established or siRNA-induced downregulation of the SMN gene increased CDK and cyclin protein expression. This seems to prevent neuronal precursors from establishing themselves by post-mitotic differentiation and increased apoptosis, which can remarkably be reversed with CDK inhibitors. Altogether, this study demonstrates how that spinal cord organoids can be generated by precisely controlling caudal and ventral signaling and represent a very flexible model to study spinal motor neuron diseases. Indeed, the spinal cord architecture is simple compared to that of the brain and it would be very interesting and technically possible to combine the use of microfluidics to establish gradients of caudal and ventral differentiation factors to generate more diverse and faithful models of the spinal cord.
Psychiatric disorders with neurodegeneration (Schizophrenia)
Multiple decades of research now suggest that some chronic psychiatric disorders are both neurodevelopmental and neurodegenerative processes [99]. However, animal models are poorly representative of the most complex aspects of human cognitive neurobiology and, not surprisingly, have yielded very few major discoveries with therapeutic potential. Brain organoids, by their ability to model both early cerebral development and by recapitulating features of neurodegeneration, are poised to bring a revolution to this field. A few studies over the last few years are beginning to illustrate this potential. Stachowiak and colleagues used iPSCs from three control and four schizophrenia patients to grow cerebral organoids and model the first trimester of development [23]. Both controls and patient-derived organoids formed typical rosettes with subventricular zones, intermediate zones and cortical zones that appeared grossly similar. However, a more detailed cellular analysis revealed that the patient-derived organoids lacked mature neurons (Pan-Neu +) in the cortical zone and instead contained abundant proliferating neuronal precursors in the intermediate zone that extended as far as the cortical zone. Moreover, they went to show that the patient-derived organoids cortical zone neurons exhibit a striking loss of nuclear FGFR1 staining. Together, these findings support a previously established theory that a premature neuronal maturation dependant on integrated nuclear FGFR1 signaling is central to schizophrenia pathophysiology.
Another study, by Johnstone and colleagues, exploited brain organoids to investigate the neurodevelopmental consequences of the 16p13.11 microduplication, a genetic alteration known to be associated with both schizophrenia, developmental delay and seizure disorders [24]. In this article, the authors correlate the clinical and imaging findings of 16p13.11 microduplication schizophrenia patients with their own patient-derived organoid features. Specifically, they found reduced cortical thickness on MRI correlated with a decreased proliferation and organoid size in patients compared to control. Interestingly, an increased ratio of asymmetrical (one neural progenitor producing a neural progenitor and a neuron) to symmetrical divisions (one neural progenitor producing two neural progenitors) seems to underly the defect. An RNA analysis on 2D grown iPSC patient-derived neurons then showed a significant decrease in NFκβ signaling in patient cells, in particular phosphorylation of NFκβ at S536 (also known as p65). To demonstrate the relevance of their findings, they then used small molecules to induce translocation of p65 to the nucleus, which rescued the proliferation deficit in patient-derived neural cell progenitors. However, given that these compounds also increase p65 translocation in control cells, this might only demonstrate that NFκβ activation has a proliferative effect in neural progenitor cells independent of the effects of the 16p13.11 microduplication. Nevertheless, this study offers a tantalizing window into personalized medicine by correlating patient phenotype with their organoid behavior, creating a prognostication and drug-testing platform for this disease.
Together, these early studies show that 3D cell culture models, including brain and spinal cord organoids, can be harnessed to demonstrate defects in human cells that recapitulate pathological cascades found in neurodegenerative diseases. Both patients and control-derived brain organoids demonstrate neuropathological hallmarks of human neurodegeneration that were challenging to model in animal models over the last few decades of research. They appear poised to overcome some of the previous barriers and limitations of two-dimensional cultures and become complementary models and platforms to test novel genetic interactions and therapeutic agents. There are however many outstanding clinical and neuropathologic questions and considerations, that if addressed, we believe could dramatically improve the ability of cerebral organoids to model neurodegenerative conditions.
Exploiting the limitations of current brain organoids system for neurodegeneration studies
Maturity and aging
While there are several neurodegenerative disorders with juvenile or early onset, aging remains the strongest and most common risk factor of neurodegenerative diseases. Brain organoids are currently generated from either ESCs or iPSCs. While the former are by definition young and immature cells, iPSCs have been shown to become rejuvenated during reprogramming, having lengthened telomeres and mitochondrial network with increased fitness [100, 101]. These processes are similar to those happening during fertilization in vivo [101] and mitigate some of the benefits of patient-derived models. Moreover, processes associated with senescence phenotypes, such as abnormal nuclear morphology and DNA damage foci have been shown to be reduced after induced pluripotency [102]. It is therefore not surprising that brain organoids closely match the genetic, epigenetic and transcriptomic signatures of human fetal cortex up to the second trimester of development [9, 14, 103,104,105]. Thus, even with the aforementioned promise, caution should be exercised when brain organoid models are used to used to understand neurodegeneration.
Different strategies, based on the growing molecular understanding driving aging, can perhaps help overcome this limitation. One approach could involve leveraging the biology of known human premature aging disorders and engineering disease-associated mutations or overexpression of progeric genes (Fig. 3). Alternatively, induction of ROS, DNA damage and mitochondrial damage with the use of toxins could be explored [106, 107]. Using the former approach, Miller and colleagues overexpressed progerin, a truncated form of lamin A associated with HGPS to promote aging in iPSC-derived dopaminergic neurons [102]. Using this strategy, they were able to induce aging-related markers and features including neuromelanin accumulation. More interestingly, when progeria was expressed in Parkinson’s disease patient iPSC-derived dopaminergic neurons, disease phenotypes such as Lewy-body precursor inclusions, enlarged mitochondria, loss of tyrosine hydroxylase and dendrite degeneration were produced. This not only illustrates the potential to use aging strategies with iPSCs to better reproduce neurodegenerative pathology but also highlights the feasibility of using a “young” cell system such as iPSCs to study the full spectrum of aging. Other genetic changes associated with progeroid syndromes, such as DC, CS and ATG have yet to be leveraged to induce aging in neurodegenerative iPSC models (Table 2). Interestingly, iPSCs derived from progeroid syndromes patients’ cells were shown to regain aging features after an initial rejuvenation [108,109,110,111], demonstrating that some disease-associated genetic or epigenetic changes (in addition to the causative mutations) associated with age must be conserved even after the iPSC process. This seems to be true of cells from older patients without clinical disease as well [112]. Indeed, Zhang and colleagues demonstrated that, as opposed to young cells, aged-cell derived iPSCs have a decreased expression of cell-specific glucose transporter 3 (GLUT3), which in turn decreases glycoslysis and impairs the ability of these cells to suppress oxidative phosphorylation. In addition to directly causing DNA damage, ROS further decreases an already age-dependent blunting of DNA damage response, accelerating the aging process in those rejuvenated cells [112]. Therefore, progeroid syndromes patients-derived iPSC themselves can serve as valuable tools to study aging and neurodegenerative processes. Finally, bypassing the intermediate pluripotent state by direct reprogramming of differentiated cells into neurons has been shown to better maintain cellular aging hallmarks [113]. Indeed, Tang and colleagues demonstrated that it is possible to directly reprogram ALS donor fibroblasts into motor neurons. Interestingly, contrary to neurons engineered by the iPSC method, the motor neurons maintained the extensive DNA damage, loss of heterochromatin and nuclear organization, and increased SA-β-Gal activity. However, it remains to be seen how this method can be used in conjunction with organoids generation as fully differentiated cells might not be able to undergo spontaneous organization into 3D tissue.
Functional and Architectural analysis of brain organoids for neurodegenerative diseases. a High-throughput platform for drug testing. Brain organoids can be functionally analyzed in vivo with intrinsic (CRISPR-CAS9 integrated probes) or extrinsic (Calcium-imaging, mitochondrial membrane potential imaging, ROS imaging) molecular probes and with electrophysiology techniques (optogenetically based interrogation or spontaneous activity). Once fixed, the organoids can be entirely processed by tissue-clearing methods to visualize the entire neuronal networks. Alternatively, in vivo imaging of cellular activity (microglia, astrocytes) and organellar dynamics (CRISPR-CAS9 integrated specific organellar reporters and fluorescent proteins) can also be achieved. The complex and large amounts of data can then be sub-processed in part by deep-learning systems and help to identify functional and architectural markers and patterns of neurodegeneration. b Brain organoids as avatar of personalized medicine. Brain organoids can be generated directly from patient cells to test various compounds in different concentration to achieve the best possible therapeutic effect
The use of toxins inducing ROS and/or mitochondrial damage might represent a more versatile and complementary approach to promote aging in organoids. Interestingly, the iPSCs process-driven switch from oxidative phosphorylation (OXPHOS) to glycolysis appears to be partially suppressed in older patient-derived iPSCs [112]. Moreover, some neurodegenerative diseases patient-derived iPSCs have been shown to be inherently susceptible to oxidative stress [114, 115]. This suggests that both DNA-damage and ROS strategies could be particularly effective in promoting and modeling aspects of the aging process in iPSCs (Table 2). For example, one group treated human iPSCs with paraquat, a known mitochondrial Complex I toxin that promotes the formation of ROS, to induce chronic stress in retinal pigment epithelium (RPE) and model age-related macular degeneration [115]. Using this approach, they identified micro-RNAs regulating RPE oxidative stress responses and implicating the NRF2-KEAP1 regulatory pathway. Importantly, some doses of paraquat can induced chronic stress without cell death whereas higher concentrations triggered apoptosis. This dose-dependant effect illustrates the experimental flexibility of a toxin-induced aging strategy to tease out early versus late changes in neurodegeneration molecular cascades. However, translating this approach might be a challenge in organoids, as ROS-inducing molecules might not penetrate 3D tissue as efficiently. On the other hand, this can potentially be turned into an advantage to trigger neurodegeneration in a heterogeneous manner, and subsequently simultaneously interrogate the interactions between damaged and healthy tissue. That gradient of toxin concentration in tissue might translate into a gradient of senescence, allowing us to better characterize simultaneously different stages of neurodegeneration.
Cellular and regional diversity, connectivity, and myelination
Neurodegenerative diseases vary in both cellular type and brain region susceptibility over the evolution of the disease. Organoids however are largely exposed to relatively homogeneously dispersed and fairly simple cocktails of differentiation factors that lack the true variety and gradient-effects that surrounding mesenchymal embryonic structures provide during development and limit the full complement of cells found in neural tissue. Multiple groups have now shown that it is possible to develop microglial or astrocytes precursors from embryonic or iPSCs origin and successfully integrate them into growing organoids [86, 89, 116] (Table 2). These non-neuronal precursors can either be added initially with neuronal precursors or once the organoids have started to grow. Conversely, it appears that cells within the organoids, while displaying neuronal fates in early culture, have the potential to differentiate into other cell types. Two groups have now shown that successive cycles of cell dissociation from the organoids can generate mature and relatively pure astrocytic cultures [117, 118]. Moreover, Quadrato and colleagues recently demonstrated that cell diversity in brain organoids, much like in the developing human brain, increase with days in culture. By performing single-cell RNA analysis on more than 80,000 cells from 31 organoids, they found a surprisingly varied pool of cell precursor types, including clusters with increased mesenchymal markers. Other neural cell types such as photoreceptors and astrocytes however did not appear before 6 months of growth. This confirms the potential of organoid cells to self-pattern into more diversified and complex cell fates than originally thought. Once a cell differentiates itself spontaneously into a different fate than the rest of the organoid, whether spontaneously or guided by architectural cues, the differentiation factors gradient likely becomes less homogeneous and triggers the differentiation of more cells, in a feed-forward or cascading fashion, mirroring that of normal development. Therefore, as long as growth does not plateau, varying culture periods can be exploited to tune the diversity of cell types available (Table 2). Similarly, improved culturing techniques including the use of bioengineering filaments provide refined approaches that limit the stochastic production of non-neural elements [119]. Similarly, Ormel and colleagues also showed that brain organoids grown using the Lancaster protocol can intrinsically develop mesodermal-derived elements such as microglia that have transcriptomic signatures closely resembling that of post-mortem human microglia [120]. Moreover, they demonstrate that decreasing neuroectodermal-stimulants such as heparin in culture and delayed Matrigel embedding can also be leveraged to increase the yield of microglial elements in the generated organoid. Therefore, there is optimism that the appropriate balance of mesodermal, endodermal and ectodermal elements could be achieved with slight protocol modifications prior to orienting the growth of the organoid toward more ectodermal fates.
Another solution to generate cellular and regional diversity that has emerged is the fusion of region-specific brain organoids to form “assembloids”. This creates gradients of trophic factors that induce the migration of cells within neural tissue, generating more cellularly diverse and architecturally advanced brain regions. These complex structures also form intermediate regions with a new identities. For example, Xiang and colleagues first developed Median-Ganglionic Eminence-like (MGE) organoids by dual SMAD and canonical WNT inhibition and activation of sonic hedgehog (SHH)-signaling pathway [10]. By mixing ventral fate MGE organoids with organoids showing a more dorsal cortical fate, the authors saw interneuron migration from MGE into the cortical layer. Two other groups used a similar method and further characterized interactions between excitatory and inhibitory neuron types [121, 122].
Another challenge of brain organoids is the prospect of developing functional synapses and neurochemistry that reflects that of the human brain. This would afford researchers the ability to study finer pathophysiological changes and test neuromodulatory agents for potential application to psychiatric and neurodegenerative diseases. The optimism is so great in this domain that special committees have formed to discuss the ethical implications with developing potentially pain-sensing “mini-brains” [123, 124]. However, several challenges must be addressed before brain organoids can display such organized signal processing. Brain organoids culture typically take a few months to reach maximal size of about 4 mm and about 6 months to display mature connections [7, 9, 125, 126]. In addition to necessitating a relatively long period to develop functional synapses, it is likely that brain organoids of this size will not be able to model the connectivity of long white matter tracts found in the human brain that are essential for high-order neural processing. A possible solution to partially model these connections however is again to fuse region-specific organoids (Table 2). As previously mentioned, doing so not only promotes the migration of specific neuronal types but also increases local cellular type diversity [10, 121, 122]. This approach might not be suitable for the longest CNS tracts but has the advantage of isolating the contribution of one region to the development of another, something which otherwise would be difficult to unravel in the complexity of the developing brain. Interestingly, a recent study by the Lancaster group has shown that growing slices of mature cortical organoids at the liquid-air interface promotes the growth of longer tracts, enhances connectivity between cells and furthers cortical layers differentiation [127]. Furthermore, the emergent axonal bundles exhibited neuronal guidance behaviors similarly to in vivo models and could form functional synapses with isolated mouse spinal cord. Finally, possibly by having a constant and homogeneous supply of oxygen and a reduced thickness, these organoids cortical slices can survive much longer (up to 1 year) in culture, overcoming another limiting aspect of organoid culture. This innovative approach however comes at a cost of reducing the complexity of the 3D architecture that appears to be a strong asset to modeling neurodegenerative inclusions.
Another key component to connectivity is the myelination of axons (Table 2). While some groups have managed to integrate oligodendrocytes to brain organoids [7], mature myelination of axons in these has yet to be achieved. In a recent study, Matsui and colleagues shows that brain organoids generated from H9 human ESCs for 6 months harbour oligodendrocyte with simple myelin sheaths around neurons, similar to what can be observed in prenatal human brains [7]. Consequently, the maturity of the organoid itself appears to be the limiting factor for the myelination process. Accordingly, the same aging-promoting strategies mentioned in a previous section might be instrumental in achieving proper myelination. Finally, another strategy would be to add separately grown iPSC-derived oligodendrocytes at various stages of organoid growth as it might better reflect normal developmental phases and result in more robust myelination.
Growth, reproducibility, and blood supply
Due to biophysical constraints, brain organoids currently reach a maximal growth of about 4 mm after multiple months of culture [7, 128]. The limited diffusion of oxygen and nutrients stunts growth and induces necrosis at the core of the organoids. Moreover, while self-organizing cultures have proven to be cheaper and more accurate methods of growing brain organoids, they can lack homogeneity between batches given their initial spatial organization on plates and their development in heterogeneous extracellular matrices (Matrigel). Growth of organoids in smaller miniaturized spinning bioreactors has been shown to accelerate growth and to provide more consistency between batches. Combining 3D printing technology and biophysical analysis, Qian and colleagues developed SpinΩ, a bioreactor that can fit into a conventional 12 well that can be stacked to further increase throughput [14]. Other groups demonstrated that scalability and automation can be improved by starting cultures with neurospheres instead of adherent colonies [129]. This method also seems to greatly decrease variation in those 3D cultures, and in doing so removing an important barrier to achieve a functional 3D cell culture platform for drug testing. Quadrato and colleagues used similar bioreactors but also found simple adjustments in the culture protocol that promote organoids growth and sustainability. They added brain-derived neurotrophic factor (BDNF) to the final differentiation medium, and used a reduced number of initial pluripotent stem cells, which led to reduced programmed cell death and necrosis and viable cultures up to 9 months [126]. Nonetheless, the growth of brain organoids eventually plateaus and due to the limited access to nutrients and oxygen. As the field is still very much in its infancy, it is expected that groups will continue to propose individualized modifications that aim to improve consistency of various organoids culture system [130]. It is important to highlight that these rapidly evolving protocol could have modifications (e.g. more directed differentiation by use of additional growth factors) that come at the potential expense of other important features to improve consistency (e.g. autonomous self-organizing cellular programs that inherently increase cell diversity). Diligences in the annotation of culture conditions is thus required to ensure reproducibility in modeling subtle aspects of neurodegenerative disorders in this culture system as the field develops.
Vascular pathology is central to the two most common neurodegenerative diseases worldwide, AD and vascular dementia. In addition, vascularization has been shown to be important for the later stages of neocortex development, including inducing neurogenesis and serving as guides to neuronal migration [131, 132]. The development of blood vessels in organoids was first realized by adding mesenchymal stem cells to tissue-specific cells by two groups in 2015 [133]. With this technique, Takebe and colleagues generated vascularized organ buds that acquired functional blood vessels after transplant in severe combined immunodeficient mouse models. Importantly, they found that the ability of the tissue-specific cells to condensate and form vascularized organ buds upon the addition of mesenchymal stem cell was dependent of the density and biochemical properties of the substrate (Matrigel) in which they grow in. Schwartz and colleagues on the other hand added endothelial cells and mesenchymal stem cells to neural progenitors already organized into multilayers with radially oriented neural and glial populations [116]. These constructs expressed an extensive capillary network by three weeks of growth and were used to validate the drug-testing potential of this system. Notably, they also added microglial precursors and showed that these can adapt into ramified and ameboid morphologies that associate with the endothelial tubules (Table 2). Finally, Mansour and colleagues bypassed the need for mesenchymal or microglial progenitors by grafting brain organoids onto adult mouse brain [134]. Strikingly, the organoids demonstrated vascularized and functional neuronal networks with integration of microglia. Together, these studies show that growing organoids possess the inherent ability and trophic factors needed to promote angiogenesis when cultured in a favorable environment, whether constituted from mesenchymal and/or endothelial stem cells or by a living organism. It would be interesting to compare this ability in young versus old, immature versus mature brain organoids to explore the impact of age and maturity on the brain parenchyma capacity to support strong vasculature (Table 2). In addition, models of ischemia and vascular-dementia could be devised simply by providing nutrients and oxygen via the developing vasculature and later occluding it (Table 2). Finally, vascular disease-associated patient-derived iPSCs differentiated into mesenchymal or endothelial stem cells can be used to study angiogenic mechanisms implicated in neurodegenerative diseases (Table 2).
Brain organoids as functional avatars for precision diagnostics and personalized prognostication in neurodegenerative diseases
Neurodegenerative diseases, by their progressive and irreversible nature, have been challenging for clinicians to diagnose and prognosticate. Reliable markers of early disease have yet to be identified and brain tissue biopsies rarely provide enough benefits to outweigh the risks of the procedure. iPSC and brain organoid culture therefore present a great opportunity to reliably identify individuals at the risk of dementia or movement disorders, and even possibly predict the course of their disease. This would rely on first identifying robust differences between brain organoids grown from patient- versus control-derived iPSCs. Given the limitations of brain organoids culture mentioned earlier in the text, and their intrinsic complexity, finding robust and reliable differences might appear at first a formidable task. However, the advent of emerging downstream molecular readouts (e.g. proteomics), and automated and objective pattern recognition tools (e.g. deep-learning networks) for quantitative histological analysis could help overcome these barriers (Fig. 3) [135,136,137,138]. Mass spectrometry-based proteomics is now increasingly used as a global profiling tool to probe clinically relevant biological differences is human disease [135, 137, 139,140,141,142]. Proteomics offers the advantage over genomic technologies of not only detecting protein-level differences, which do not always correlate with mRNA levels, but it also allows detection of post-transcriptional modifications (e.g. phosphorylation) that have particular relevance for aggregates in neurodegenerative disease (e.g. hyperphosphorylated tau) and potential upstream pathogenic pathways. Coupled with dynamic tracking of the formation of protein aggregates in organoids, this technology could offer new insights into disease pathogenesis previously not possible. Similarly, the transition from 2D to 3D tissue systems increases the importance of tools that allow accurate quantification of complex histological patterns of disease. Technological improvements in both computational software and hardware tools now allow for true automation in image analysis that has recently matched, and in cases surpassed, the capabilities of pathologists and other subspecialized physician experts alike [136, 138, 143, 144]). Our own group has focused on developing “hypothesis free” unsupervised deep-learning approaches capable of defining “anomalous” outlier patterns in histological data [145]. Thus, while it may not be feasible to carry out molecular profiling on organoids interrogated by large drug libraries/experimental conditions, automated analysis of H&E and immunofluorescence images of organoids could allow detection of even subtle and unexpected structural changes in various organoids models of neurodegeneration.
Two-photon microscopy offers a mechanism to bypass tissue sectioning but has required organisms with transparent tissue such as zebrafish or tadpoles to overcome the heterogeneous refractive indexes of different cellular elements. Modern tissue-clearing methods have therefore first relied on dehydration followed by immersion in a solvent to homogenize these refractive indexes, a process named 3DISCO (3D imaging of solvent-cleared organs) [146]. Dodt and colleagues for example developed a solvent-based tissue-clearing technique in 2007 that allows imaging of artificially expressed fluorescent proteins and some auto-fluorescent proteins in whole organs, but lost most other epitopes in doing so [147]. In 2013, the Deisseroth group devised a tissue-clearing method (CLARITY) based on cross-linking a hydrogel composed of acrylamide to the tissue followed by electrochemical ionic detergent extraction of lipid molecules [148]. This method allowed to clear the tissue to near-transparency while maintaining structural integrity and most epitopes available to immunolabelling. Consequently, imaging multiple proteins of interest in integral tissue with light sheet fluorescence microscopy (LSFM) is now feasible. A number of methods were then derived from this breakthrough, including PACT (passive clarity technique), which utilizes ionic detergent extraction without an electrochemical gradient. This provides greater epitope stability and creates less damage due to heating [149]. Another method called CUBIC (clear, unobstructed brain imaging cocktails and computational analysis), used cheaper and more widely available aminoalcohols to clear the lipids, and was later shown in de-paraffinized paraffin-embedded tissue [150], opening the technique to pathological diagnostic applications [151]. Finally, ScaleS, a tissue-clearing method using sorbitol as a solvent was shown to be particularly good at preserving structural details, a feature that is very useful to image the architecture and disposition of complex protein aggregates such as amyloid plaques [152]. Thus, multiple tissue-clearing techniques with different advantages and disadvantages are now available [153], and it will be important to determine which one is more amenable to image neurodegenerative features in brain organoids (Fig. 3).
With the help of CRISPR-CAS9 genome-editing techniques, fluorescent labels and functional reporter systems can be integrated in pluripotent cells prior to organoid growth. These can be used to interrogate regional connectivity, cellular migration and sub-cellular mechanisms. This will be particularly useful to establish optogenetic systems in 3D cultures. Interestingly, recent studies showed that light can induce neuronal activity in organoids that have developed photoreceptor cells [126], suggesting that brain organoids are sufficiently translucid to allow probing with light. In this manner, specific neurotransmitter pathways can be interrogated in vivo in human tissue and analyzed in various genetic background. Extrinsic reporters of ROS, mitochondria membrane potential and oxygen consumption can also be used to monitor mitochondrial function in vivo. Finally, electrophysiological interrogation of brain organoids in conjunction with calcium-imaging can be used to monitor the neuronal networks fitness.
Once the organoid is architecturally and functionally imaged, the data obtained will be many levels of complexity greater than what obtained with two-dimensional tissue. By feeding the data into emerging deep-learning pattern recognition systems discussed above, we might be able to discover novel histological and cellular patterns of early neurodegenerative process that will be difficult for humans to detect and analyze using the traditional qualitative microscopy examination (Fig. 3). These changes and readouts promise to improve diagnostics and predictive abilities in the development of neurodegenerative disorders in pre-symptomatic patients and expedite early initiation of therapy.
Conclusion
Despite multiple decades of research and the continued refinement of neurodegenerative hypotheses and models, we have yet to nominate successful disease-modifying drug for these debilitating diseases. This is in part because animal models and relatively simple two-dimensional cell culture models have failed to replicate and effectively dissect the human-specific pathogenic cascades leading to neuronal death. While brain organoids are perhaps best known and used to study early neural development, there is now increasing interests to translate them to other neurological disorders. The flexibility that combines the benefits of human-specific processes with an experimentally malleable model to explore spatiotemporal molecular mechanisms of disease has profound implication for the neurodegenerative community. This promise and accumulating advances that allow integration of additional variables and risk factors (toxic agents, vasculature) makes brain organoids a formidable and scalable system to improve our understanding, provide precision to diagnostic and prognostic predictions and personalize drug discovery efforts for neurodegenerative diseases. Overcoming the existing challenges through the adoption of various emerging technologies and approaches discussed in this review aims to help accelerate progress toward this important goal.
References
Fiest KM, Jette N, Roberts JI, Maxwell CJ, Smith EE, Black SE, et al. The prevalence and incidence of dementia: a systematic review and meta-analysis. Can J Neurol Sci Le J Can Sci Neurol. 2016;43(Suppl 1):S3–S50.
Wong SL, Gilmour H, Ramage-Morin PL. Alzheimer's disease and other dementias in Canada. Health Rep. 2016;27:11–16.
Li Y, Muffat J, Omer A, Bosch I, Lancaster MA, Sur M, et al. Induction of expansion and folding in human cerebral organoids. Cell Stem Cell. 2017;20:385–96 e383.
Lancaster MA, Knoblich JA. Generation of cerebral organoids from human pluripotent stem cells. Nat Protoc. 2014;9:2329–40.
Hansen DV, Lui JH, Parker PR, Kriegstein AR. Neurogenic radial glia in the outer subventricular zone of human neocortex. Nature. 2010;464:554–61.
Lui JH, Nowakowski TJ, Pollen AA, Javaherian A, Kriegstein AR, Oldham MC. Radial glia require PDGFD-PDGFRbeta signalling in human but not mouse neocortex. Nature. 2014;515:264–8.
Matsui TK, Matsubayashi M, Sakaguchi YM, Hayashi RK, Zheng C, Sugie K, et al. Six-month cultured cerebral organoids from human ES cells contain matured neural cells. Neurosci Lett. 2018;670:75–82.
Lancaster MA, Renner M, Martin CA, Wenzel D, Bicknell LS, Hurles ME, et al. Cerebral organoids model human brain development and microcephaly. Nature. 2013;501:373–9.
Pasca AM, Sloan SA, Clarke LE, Tian Y, Makinson CD, Huber N, et al. Functional cortical neurons and astrocytes from human pluripotent stem cells in 3D culture. Nat Methods. 2015;12:671–8.
Xiang Y, Tanaka Y, Patterson B, Kang YJ, Govindaiah G, Roselaar N, et al. Fusion of regionally specified hPSC-derived organoids models human brain development and interneuron migration. Cell Stem Cell. 2017;21:383–98 e387.
Krefft O, Jabali A, Iefremova V, Koch P, Ladewig J. Generation of standardized and reproducible forebrain-type cerebral organoids from human induced pluripotent stem cells. J Vis Exp: JoVE. 2018;131:56768.
Eiraku M, Takata N, Ishibashi H, Kawada M, Sakakura E, Okuda S, et al. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature. 2011;472:51–56.
Nakano T, Ando S, Takata N, Kawada M, Muguruma K, Sekiguchi K, et al. Self-formation of optic cups and storable stratified neural retina from human ESCs. Cell Stem Cell. 2012;10:771–85.
Qian X, Nguyen HN, Song MM, Hadiono C, Ogden SC, Hammack C, et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell. 2016;165:1238–54.
Jo J, Xiao Y, Sun AX, Cukuroglu E, Tran HD, Goke J, et al. Midbrain-like organoids from human pluripotent stem cells contain functional dopaminergic and neuromelanin-producing neurons. cell stem cell. 2016;19:248–57.
Monzel AS, Smits LM, Hemmer K, Hachi S, Moreno EL, van Wuellen T, et al. Derivation of human midbrain-specific organoids from neuroepithelial stem cells. Stem Cell Rep. 2017;8:1144–54.
Qian X, Jacob F, Song MM, Nguyen HN, Song H, Ming GL. Generation of human brain region-specific organoids using a miniaturized spinning bioreactor. Nat Protoc. 2018;13:565–80.
Muguruma K, Nishiyama A, Ono Y, Miyawaki H, Mizuhara E, Hori S, et al. Ontogeny-recapitulating generation and tissue integration of ES cell-derived Purkinje cells. Nat Neurosci. 2010;13:1171–80.
Kawada J, Kaneda S, Kirihara T, Maroof A, Levi T, Eggan K, et al. Generation of a motor nerve organoid with human stem cell-derived neurons. Stem Cell Rep. 2017;9:1441–9.
Hor JH, Soh ES, Tan LY, Lim VJW, Santosa MM, Winanto, et al. Cell cycle inhibitors protect motor neurons in an organoid model of Spinal Muscular Atrophy. Cell Death Dis. 2018;9:1100.
Sakaguchi H, Kadoshima T, Soen M, Narii N, Ishida Y, Ohgushi M, et al. Generation of functional hippocampal neurons from self-organizing human embryonic stem cell-derived dorsomedial telencephalic tissue. Nat Commun. 2015;6:8896.
Di Lullo E, Kriegstein AR. The use of brain organoids to investigate neural development and disease. Nat Rev Neurosci. 2017;18:573–84.
Stachowiak EK, Benson CA, Narla ST, Dimitri A, Chuye LEB, Dhiman S, et al. Cerebral organoids reveal early cortical maldevelopment in schizophrenia-computational anatomy and genomics, role of FGFR1. Transl Psychiatry. 2017;7:6.
Johnstone M, Vasistha NA, Barbu MC, Dando O, Burr K, Christopher E, et al. Reversal of proliferation deficits caused by chromosome 16p13.11 microduplication through targeting NFkappaB signaling: an integrated study of patient-derived neuronal precursor cells, cerebral organoids and in vivo brain imaging. Mol psychiatry. 2019;24:294–311.
Qian X, Nguyen HN, Jacob F, Song H, Ming GL. Using brain organoids to understand Zika virus-induced microcephaly. Development. 2017;144:952–7.
Mellios N, Feldman DA, Sheridan SD, Ip JPK, Kwok S, Amoah SK, et al. MeCP2-regulated miRNAs control early human neurogenesis through differential effects on ERK and AKT signaling. Mol psychiatry. 2018;23:1051–65.
Mellios N, Feldman DA, Sheridan SD, Ip JPK, Kwok S, Amoah SK, et al. Human cerebral organoids reveal deficits in neurogenesis and neuronal migration in MeCP2-deficient neural progenitors. Mol psychiatry. 2018;23:791.
Dakic V, Minardi Nascimento J, Costa Sartore R, Maciel RM, de Araujo DB, Ribeiro S, et al. Short term changes in the proteome of human cerebral organoids induced by 5-MeO-DMT. Sci Rep. 2017;7:12863.
Eiraku M, Watanabe K, Matsuo-Takasaki M, Kawada M, Yonemura S, Matsumura M, et al. Self-organized formation of polarized cortical tissues from ESCs and its active manipulation by extrinsic signals. Cell Stem Cell. 2008;3:519–32.
Kleinman HK, McGarvey ML, Hassell JR, Star VL, Cannon FB, Laurie GW, et al. Basement membrane complexes with biological activity. Biochemistry. 1986;25:312–8.
Li R, Sun L, Fang A, Li P, Wu Q, Wang X. Recapitulating cortical development with organoid culture in vitro and modeling abnormal spindle-like (ASPM related primary) microcephaly disease. Protein Cell. 2017;8:823–33.
Bershteyn M, Nowakowski TJ, Pollen AA, Di Lullo E, Nene A, Wynshaw-Boris A, et al. Human iPSC-derived cerebral organoids model cellular features of lissencephaly and reveal prolonged mitosis of outer radial glia. Cell Stem Cell. 2017;20:435–49 e434.
Double KL, Halliday GM, Kril JJ, Harasty JA, Cullen K, Brooks WS, et al. Topography of brain atrophy during normal aging and Alzheimer's disease. Neurobiol aging. 1996;17:513–21.
Pini L, Pievani M, Bocchetta M, Altomare D, Bosco P, Cavedo E, et al. Brain atrophy in Alzheimer's disease and aging. Ageing Res Rev. 2016;30:25–48.
Dumitriu D, Hao J, Hara Y, Kaufmann J, Janssen WG, Lou W, et al. Selective changes in thin spine density and morphology in monkey prefrontal cortex correlate with aging-related cognitive impairment. J Neurosci : Off J Soc Neurosci. 2010;30:7507–15.
Xekardaki A, Kovari E, Gold G, Papadimitropoulou A, Giacobini E, Herrmann F, et al. Neuropathological changes in aging brain. Adv Exp Med Biol. 2015;821:11–17.
Kovacs GG, Lee VM, Trojanowski JQ. Protein astrogliopathies in human neurodegenerative diseases and aging. Brain Pathol. 2017;27:675–90.
Kovacs GG, Milenkovic I, Wohrer A, Hoftberger R, Gelpi E, Haberler C, et al. Non-Alzheimer neurodegenerative pathologies and their combinations are more frequent than commonly believed in the elderly brain: a community-based autopsy series. Acta Neuropathol. 2013;126:365–84.
Bennett DA, Schneider JA, Arvanitakis Z, Kelly JF, Aggarwal NT, Shah RC, et al. Neuropathology of older persons without cognitive impairment from two community-based studies. Neurology. 2006;66:1837–44.
Mattson MP, Arumugam TV. Hallmarks of brain aging: adaptive and pathological modification by metabolic states. Cell Metab. 2018;27:1176–99.
Ioannidou A, Goulielmaki E, Garinis GA. DNA damage: from chronic inflammation to age-related deterioration. Front Genet. 2016;7:187.
Chow HM, Herrup K. Genomic integrity and the ageing brain. Nat Rev Neurosci. 2015;16:672–84.
Madabhushi R, Pan L, Tsai LH. DNA damage and its links to neurodegeneration. Neuron. 2014;83:266–82.
McKinnon PJ. Maintaining genome stability in the nervous system. Nat Neurosci. 2013;16:1523–9.
d'Adda di Fagagna F. Living on a break: cellular senescence as a DNA-damage response. Nat Rev Cancer. 2008;8:512–22.
Chinta SJ, Woods G, Rane A, Demaria M, Campisi J, Andersen JK. Cellular senescence and the aging brain. Exp Gerontol. 2015;68:3–7.
van der Horst GT, van Steeg H, Berg RJ, van Gool AJ, de Wit J, Weeda G, et al. Defective transcription-coupled repair in Cockayne syndrome B mice is associated with skin cancer predisposition. Cell. 1997;89:425–35.
Andressoo JO, Weeda G, de Wit J, Mitchell JR, Beems RB, van Steeg H, et al. An Xpb mouse model for combined xeroderma pigmentosum and cockayne syndrome reveals progeroid features upon further attenuation of DNA repair. Mol Cell Biol. 2009;29:1276–90.
Laposa RR, Huang EJ, Cleaver JE. Increased apoptosis, p53 up-regulation, and cerebellar neuronal degeneration in repair-deficient Cockayne syndrome mice. Proc Natl Acad Sci USA. 2007;104:1389–94.
Kuljis RO, Xu Y, Aguila MC, Baltimore D. Degeneration of neurons, synapses, and neuropil and glial activation in a murine Atm knockout model of ataxia-telangiectasia. Proc Natl Acad Sci USA. 1997;94:12688–93.
Adamec E, Vonsattel JP, Nixon RA. DNA strand breaks in Alzheimer's disease. Brain Res. 1999;849:67–77.
Jacobsen E, Beach T, Shen Y, Li R, Chang Y. Deficiency of the Mre11 DNA repair complex in Alzheimer's disease brains. Brain Res Mol Brain Res. 2004;128:1–7.
Shackelford DA. DNA end joining activity is reduced in Alzheimer's disease. Neurobiol aging. 2006;27:596–605.
Bender A, Krishnan KJ, Morris CM, Taylor GA, Reeve AK, Perry RH, et al. High levels of mitochondrial DNA deletions in substantia nigra neurons in aging and Parkinson disease. Nat Genet. 2006;38:515–7.
Kraytsberg Y, Kudryavtseva E, McKee AC, Geula C, Kowall NW, Khrapko K. Mitochondrial DNA deletions are abundant and cause functional impairment in aged human substantia nigra neurons. Nat Genet. 2006;38:518–20.
Kisby GE, Milne J, Sweatt C. Evidence of reduced DNA repair in amyotrophic lateral sclerosis brain tissue. Neuroreport. 1997;8:1337–40.
Kikuchi H, Furuta A, Nishioka K, Suzuki SO, Nakabeppu Y, Iwaki T. Impairment of mitochondrial DNA repair enzymes against accumulation of 8-oxo-guanine in the spinal motor neurons of amyotrophic lateral sclerosis. Acta Neuropathol. 2002;103:408–14.
Ferrante RJ, Browne SE, Shinobu LA, Bowling AC, Baik MJ, MacGarvey U, et al. Evidence of increased oxidative damage in both sporadic and familial amyotrophic lateral sclerosis. J Neurochem. 1997;69:2064–74.
Sau D, De Biasi S, Vitellaro-Zuccarello L, Riso P, Guarnieri S, Porrini M, et al. Mutation of SOD1 in ALS: a gain of a loss of function. Hum Mol Genet. 2007;16:1604–18.
Qian M, Liu Z, Peng L, Tang X, Meng F, Ao Y, et al. Boosting ATM activity alleviates aging and extends lifespan in a mouse model of progeria. Elife. 2018;7:e34836.
Balaban RS, Nemoto S, Finkel T. Mitochondria, oxidants, and aging. Cell. 2005;120:483–95.
Roberts RF, Tang MY, Fon EA, Durcan TM. Defending the mitochondria: the pathways of mitophagy and mitochondrial-derived vesicles. Int J Biochem cell Biol. 2016;79:427–36.
Matheoud D, Sugiura A, Bellemare-Pelletier A, Laplante A, Rondeau C, Chemali M, et al. Parkinson's disease-related proteins PINK1 and parkin repress mitochondrial antigen presentation. Cell. 2016;166:314–27.
Helley MP, Pinnell J, Sportelli C, Tieu K. Mitochondria: a common target for genetic mutations and environmental toxicants in Parkinson's disease. Front Genet. 2017;8:177.
Deger JM, Gerson JE, Kayed R. The interrelationship of proteasome impairment and oligomeric intermediates in neurodegeneration. Aging cell. 2015;14:715–24.
Baker MJ, Palmer CS, Stojanovski D. Mitochondrial protein quality control in health and disease. Br J Pharm. 2014;171:1870–89.
Misgeld T, Schwarz TL. Mitostasis in neurons: maintaining mitochondria in an extended cellular architecture. Neuron. 2017;96:651–66.
Scrivo A, Bourdenx M, Pampliega O, Cuervo AM. Selective autophagy as a potential therapeutic target for neurodegenerative disorders. Lancet Neurol. 2018;17:802–15.
Hewitt G, Korolchuk VI. Repair, reuse, recycle: the expanding role of autophagy in genome maintenance. Trends Cell Biol. 2017;27:340–51.
Shapouri-Moghaddam A, Mohammadian S, Vazini H, Taghadosi M, Esmaeili SA, Mardani F, et al. Macrophage plasticity, polarization, and function in health and disease. J Cell Physiol. 2018;233:6425–40.
Caplan IF, Maguire-Zeiss KA. Toll-like receptor 2 signaling and current approaches for therapeutic modulation in synucleinopathies. Front Pharm. 2018;9:417.
Gambuzza ME, Sofo V, Salmeri FM, Soraci L, Marino S, Bramanti P. Toll-like receptors in Alzheimer's disease: a therapeutic perspective. CNS Neurol Disord Drug Targets. 2014;13:1542–58.
ElAli A, Rivest S. Microglia in Alzheimer's disease: a multifaceted relationship. Brain, Behav, Immun. 2016;55:138–50.
Hickman S, Izzy S, Sen P, Morsett L, El Khoury J. Microglia in neurodegeneration. Nat Neurosci. 2018;21:1359–69.
Choi SH, Kim YH, Hebisch M, Sliwinski C, Lee S, D'Avanzo C, et al. A three-dimensional human neural cell culture model of Alzheimer's disease. Nature. 2014;515:274–8.
Jaworski T, Dewachter I, Lechat B, Gees M, Kremer A, Demedts D, et al. GSK-3alpha/beta kinases and amyloid production in vivo. Nature. 2011;480:E4–5. discussion E6
Phiel CJ, Wilson CA, Lee VM, Klein PS. GSK-3alpha regulates production of Alzheimer's disease amyloid-beta peptides. Nature. 2003;423:435–9.
Salloway S, Sperling R, Fox NC, Blennow K, Klunk W, Raskind M, et al. Two phase 3 trials of bapineuzumab in mild-to-moderate Alzheimer's disease. New Engl J Med. 2014;370:322–33.
Doody RS, Raman R, Farlow M, Iwatsubo T, Vellas B, Joffe S, et al. A phase 3 trial of semagacestat for treatment of Alzheimer's disease. New Engl J Med. 2013;369:341–50.
Doody RS, Thomas RG, Farlow M, Iwatsubo T, Vellas B, Joffe S, et al. Phase 3 trials of solanezumab for mild-to-moderate Alzheimer's disease. New Engl J Med. 2014;370:311–21.
Lee HK, Velazquez Sanchez C, Chen M, Morin PJ, Wells JM, Hanlon EB, et al. Three dimensional human neuro-spheroid model of Alzheimer's disease based on differentiated induced pluripotent stem cells. PloS ONE. 2016;11:e0163072.
Amiri A, Coppola G, Scuderi S, Wu F, Roychowdhury T, Liu F, et al. Transcriptome and epigenome landscape of human cortical development modeled in organoids. Science. 2018;362:eaat6720.
Raja WK, Mungenast AE, Lin YT, Ko T, Abdurrob F, Seo J, et al. Self-organizing 3D human neural tissue derived from induced pluripotent stem cells recapitulate Alzheimer's disease phenotypes. PloS ONE. 2016;11:e0161969.
Cataldo AM, Peterhoff CM, Troncoso JC, Gomez-Isla T, Hyman BT, Nixon RA. Endocytic pathway abnormalities precede amyloid beta deposition in sporadic Alzheimer's disease and Down syndrome: differential effects of APOE genotype and presenilin mutations. Am J Pathol. 2000;157:277–86.
Cataldo A, Rebeck GW, Ghetri B, Hulette C, Lippa C, Van Broeckhoven C, et al. Endocytic disturbances distinguish among subtypes of Alzheimer's disease and related disorders. Ann Neurol. 2001;50:661–5.
Lin YT, Seo J, Gao F, Feldman HM, Wen HL, Penney J, et al. APOE4 causes widespread molecular and cellular alterations associated with Alzheimer's disease phenotypes in human iPSC-derived brain cell types. Neuron. 2018;98:1294.
Corder EH, Saunders AM, Strittmatter WJ, Schmechel DE, Gaskell PC, Small GW, et al. Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer's disease in late onset families. Science. 1993;261:921–3.
Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer's disease. Nat Genet. 2013;45:1452–8.
Park J, Wetzel I, Marriott I, Dreau D, D'Avanzo C, Kim DY, et al. A 3D human triculture system modeling neurodegeneration and neuroinflammation in Alzheimer's disease. Nat Neurosci. 2018;21:941–51.
Seo J, Kritskiy O, Watson LA, Barker SJ, Dey D, Raja WK, et al. Inhibition of p25/Cdk5 attenuates tauopathy in mouse and iPSC models of frontotemporal dementia. J Neurosci : Off J Soc Neurosci. 2017;37:9917–24.
Braak H, Del Tredici K, Rub U, de Vos RA, Jansen Steur EN, Braak E. Staging of brain pathology related to sporadic Parkinson's disease. Neurobiol aging. 2003;24:197–211.
Barrett JC, Hansoul S, Nicolae DL, Cho JH, Duerr RH, Rioux JD, et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn's disease. Nat Genet. 2008;40:955–62.
Holmqvist S, Chutna O, Bousset L, Aldrin-Kirk P, Li W, Bjorklund T, et al. Direct evidence of Parkinson pathology spread from the gastrointestinal tract to the brain in rats. Acta Neuropathol. 2014;128:805–20.
Lionnet A, Leclair-Visonneau L, Neunlist M, Murayama S, Takao M, Adler CH, et al. Does Parkinson's disease start in the gut? Acta Neuropathol. 2018;135:1–12.
Son MY, Sim H, Son YS, Jung KB, Lee MO, Oh JH, et al. Distinctive genomic signature of neural and intestinal organoids from familial Parkinson's disease patient-derived induced pluripotent stem cells. Neuropathol Appl Neurobiol. 2017;43:584–603.
Kim H, Park HJ, Choi H, Chang Y, Park H, Shin J, et al. Modeling G2019S-LRRK2 sporadic Parkinson's disease in 3D midbrain organoids. Stem Cell Rep. 2019;12:518–31.
Smits LM, Reinhardt L, Reinhardt P, Glatza M, Monzel AS, Stanslowsky N, et al. Modeling Parkinson's disease in midbrain-like organoids. NPJ Park's Dis. 2019;5:5.
Surmeier DJ. Determinants of dopaminergic neuron loss in Parkinson's disease. FEBS J. 2018;285:3657–68.
Rund BR. Is there a degenerative process going on in the brain of people with Schizophrenia? Front Hum Neurosci. 2009;3:36.
Marion RM, Strati K, Li H, Tejera A, Schoeftner S, Ortega S, et al. Telomeres acquire embryonic stem cell characteristics in induced pluripotent stem cells. Cell Stem Cell. 2009;4:141–54.
Suhr ST, Chang EA, Tjong J, Alcasid N, Perkins GA, Goissis MD, et al. Mitochondrial rejuvenation after induced pluripotency. PloS ONE. 2010;5:e14095.
Miller JD, Ganat YM, Kishinevsky S, Bowman RL, Liu B, Tu EY, et al. Human iPSC-based modeling of late-onset disease via progerin-induced aging. cell stem cell. 2013;13:691–705.
Camp JG, Badsha F, Florio M, Kanton S, Gerber T, Wilsch-Brauninger M, et al. Human cerebral organoids recapitulate gene expression programs of fetal neocortex development. Proc Natl Acad Sci USA. 2015;112:15672–7.
Luo C, Lancaster MA, Castanon R, Nery JR, Knoblich JA, Ecker JR. Cerebral organoids recapitulate epigenomic signatures of the human fetal brain. Cell Rep. 2016;17:3369–84.
Yoon KJ, Ringeling FR, Vissers C, Jacob F, Pokrass M, Jimenez-Cyrus D, et al. Temporal control of mammalian cortical neurogenesis by m(6)A methylation. Cell. 2017;171:877–89 e817.
Jin K. Modern biological theories of aging. Aging Dis. 2010;1:72–74.
Guillaumet-Adkins A, Yanez Y, Peris-Diaz MD, Calabria I, Palanca-Ballester C, Sandoval J. Epigenetics and oxidative stress in aging. Oxid Med Cell Longev. 2017;2017:9175806.
Agarwal S, Loh YH, McLoughlin EM, Huang J, Park IH, Miller JD, et al. Telomere elongation in induced pluripotent stem cells from dyskeratosis congenita patients. Nature. 2010;464:292–6.
Batista LF, Pech MF, Zhong FL, Nguyen HN, Xie KT, Zaug AJ, et al. Telomere shortening and loss of self-renewal in dyskeratosis congenita induced pluripotent stem cells. Nature. 2011;474:399–402.
Andrade LN, Nathanson JL, Yeo GW, Menck CF, Muotri AR. Evidence for premature aging due to oxidative stress in iPSCs from Cockayne syndrome. Hum Mol Genet. 2012;21:3825–34.
Mertens J, Paquola ACM, Ku M, Hatch E, Bohnke L, Ladjevardi S, et al. Directly reprogrammed human neurons retain aging-associated transcriptomic signatures and reveal age-related nucleocytoplasmic defects. Cell Stem Cell. 2015;17:705–18.
Zhang C, Skamagki M, Liu Z, Ananthanarayanan A, Zhao R, Li H, et al. Biological significance of the suppression of oxidative phosphorylation in induced pluripotent stem cells. Cell Rep. 2017;21:2058–65.
Tang Y, Liu ML, Zang T, Zhang CL. Direct reprogramming rather than iPSC-based reprogramming maintains aging hallmarks in human motor neurons. Front Mol Neurosci. 2017;10:359.
Lin L, Goke J, Cukuroglu E, Dranias MR, VanDongen AM, Stanton LW. Molecular Features underlying neurodegeneration identified through in vitro modeling of genetically diverse Parkinson's disease patients. Cell Rep. 2016;15:2411–26.
Garcia TY, Gutierrez M, Reynolds J, Lamba DA. Modeling the dynamic AMD-associated chronic oxidative stress changes in human ESC and iPSC-derived RPE cells. Invest Ophthalmol Vis Sci. 2015;56:7480–8.
Schwartz MP, Hou Z, Propson NE, Zhang J, Engstrom CJ, Santos Costa V, et al. Human pluripotent stem cell-derived neural constructs for predicting neural toxicity. Proc Natl Acad Sci USA. 2015;112:12516–21.
Dezonne RS, Sartore RC, Nascimento JM, Saia-Cereda VM, Romao LF, Alves-Leon SV, et al. Derivation of functional human astrocytes from cerebral organoids. Sci Rep. 2017;7:45091.
Sloan SA, Darmanis S, Huber N, Khan TA, Birey F, Caneda C, et al. Human astrocyte maturation captured in 3D cerebral cortical spheroids derived from pluripotent stem cells. Neuron. 2017;95:779–90 e776.
Lancaster MA, Corsini NS, Wolfinger S, Gustafson EH, Phillips AW, Burkard TR, et al. Guided self-organization and cortical plate formation in human brain organoids. Nat Biotechnol. 2017;35:659–66.
Ormel PR, Vieira de Sa R, van Bodegraven EJ, Karst H, Harschnitz O, Sneeboer MAM, et al. Microglia innately develop within cerebral organoids. Nat Commun. 2018;9:4167.
Bagley JA, Reumann D, Bian S, Levi-Strauss J, Knoblich JA. Fused cerebral organoids model interactions between brain regions. Nat methods. 2017;14:743–51.
Birey F, Andersen J, Makinson CD, Islam S, Wei W, Huber N, et al. Assembly of functionally integrated human forebrain spheroids. Nature. 2017;545:54–59.
Aach J, Lunshof J, Iyer E, Church GM Addressing the ethical issues raised by synthetic human entities with embryo-like features. Elife 2017;6:e20674.
Munsie M, Hyun I, Sugarman J. Ethical issues in human organoid and gastruloid research. Development. 2017;144:942–5.
Vera E, Studer L. When rejuvenation is a problem: challenges of modeling late-onset neurodegenerative disease. Development. 2015;142:3085–9.
Quadrato G, Nguyen T, Macosko EZ, Sherwood JL, Min Yang S, Berger DR, et al. Cell diversity and network dynamics in photosensitive human brain organoids. Nature. 2017;545:48–53.
Giandomenico SL, Mierau SB, Gibbons GM, Wenger LMD, Masullo L, Sit T, et al. Cerebral organoids at the air-liquid interface generate diverse nerve tracts with functional output. Nat Neurosci. 2019;22:669–79.
Hartley BJ, Brennand KJ. Neural organoids for disease phenotyping, drug screening and developmental biology studies. Neurochem Int. 2017;106:85–93.
Rigamonti A, Repetti GG, Sun C, Price FD, Reny DC, Rapino F, et al. Large-scale production of mature neurons from human pluripotent stem cells in a three-dimensional suspension culture system. Stem Cell Rep. 2016;6:993–1008.
Yoon SJ, Elahi LS, Pasca AM, Marton RM, Gordon A, Revah O, et al. Reliability of human cortical organoid generation. Nat methods. 2019;16:75–78.
Lange C, Turrero Garcia M, Decimo I, Bifari F, Eelen G, Quaegebeur A, et al. Relief of hypoxia by angiogenesis promotes neural stem cell differentiation by targeting glycolysis. EMBO J. 2016;35:924–41.
Vasudevan A, Long JE, Crandall JE, Rubenstein JL, Bhide PG. Compartment-specific transcription factors orchestrate angiogenesis gradients in the embryonic brain. Nat Neurosci. 2008;11:429–39.
Takebe T, Enomura M, Yoshizawa E, Kimura M, Koike H, Ueno Y, et al. Vascularized and complex organ buds from diverse tissues via mesenchymal cell-driven condensation. Cell Stem Cell. 2015;16:556–65.
Mansour AA, Goncalves JT, Bloyd CW, Li H, Fernandes S, Quang D, et al. An in vivo model of functional and vascularized human brain organoids. Nat Biotechnol. 2018;36:432–41.
Archer TC, Ehrenberger T, Mundt F, Gold MP, Krug K, Mah CK, et al. Proteomics, post-translational modifications, and integrative analyses reveal molecular heterogeneity within medulloblastoma subgroups. Cancer Cell. 2018;34:396–410 e398.
Djuric U, Zadeh G, Aldape K, Diamandis P. Precision histology: how deep learning is poised to revitalize histomorphology for personalized cancer care. NPJ Precis Oncol. 2017;1:22.
Djuric U, Rodrigues DC, Batruch I, Ellis J, Shannon P, Diamandis P. Spatiotemporal proteomic profiling of human cerebral development. Mol Cell Proteom : MCP. 2017;16:1548–62.
Coudray N, Ocampo PS, Sakellaropoulos T, Narula N, Snuderl M, Fenyo D, et al. Classification and mutation prediction from non-small cell lung cancer histopathology images using deep learning. Nat Med. 2018;24:1559–67.
Papaioannou MD, Djuric U, Kao J, Karimi S, Zadeh G, Aldape K et al. Proteomic analysis of meningiomas reveals clinically-distinct molecular patterns. Neuro Oncol. 2019;21:1028–38.
Vasaikar S, Huang C, Wang X, Petyuk VA, Savage SR, Wen B, et al. Proteogenomic analysis of human colon cancer reveals new therapeutic opportunities. Cell. 2019;177:1035–49 e1019.
Zhang H, Liu T, Zhang Z, Payne SH, Zhang B, McDermott JE, et al. Integrated proteogenomic characterization of human high-grade serous ovarian cancer. Cell. 2016;166:755–65.
Mertins P, Mani DR, Ruggles KV, Gillette MA, Clauser KR, Wang P, et al. Proteogenomics connects somatic mutations to signalling in breast cancer. Nature. 2016;534:55–62.
Xie Q, Faust K, Van Ommeren R, Sheikh A, Djuric U, Diamandis P. Deep learning for image analysis: personalizing medicine closer to the point of care. Crit Rev Clin Lab Sci. 2019;56:61–73.
Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM, et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017;542:115–8.
Faust K, Xie Q, Han D, Goyle K, Volynskaya Z, Djuric U, et al. Visualizing histopathologic deep learning classification and anomaly detection using nonlinear feature space dimensionality reduction. BMC Bioinforma. 2018;19:173.
Erturk A, Bradke F. High-resolution imaging of entire organs by 3-dimensional imaging of solvent cleared organs (3DISCO). Exp Neurol. 2013;242:57–64.
Dodt HU, Leischner U, Schierloh A, Jahrling N, Mauch CP, Deininger K, et al. Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain. Nat methods. 2007;4:331–6.
Chung K, Wallace J, Kim SY, Kalyanasundaram S, Andalman AS, Davidson TJ, et al. Structural and molecular interrogation of intact biological systems. Nature. 2013;497:332–7.
Yang B, Treweek JB, Kulkarni RP, Deverman BE, Chen CK, Lubeck E, et al. Single-cell phenotyping within transparent intact tissue through whole-body clearing. Cell. 2014;158:945–58.
Susaki EA, Tainaka K, Perrin D, Kishino F, Tawara T, Watanabe TM, et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell. 2014;157:726–39.
Nojima S, Susaki EA, Yoshida K, Takemoto H, Tsujimura N, Iijima S, et al. CUBIC pathology: three-dimensional imaging for pathological diagnosis. Sci Rep. 2017;7:9269.
Hama H, Hioki H, Namiki K, Hoshida T, Kurokawa H, Ishidate F, et al. ScaleS: an optical clearing palette for biological imaging. Nat Neurosci. 2015;18:1518–29.
Yu T, Qi Y, Gong H, Luo Q, Zhu D Optical clearing for multiscale biological tissues. J Biophotonics. 2018;11: e201700187.
Acknowledgements
J.K. is supported by CIHR CGSM studentship. Research time and space for P.D. is provided by the Princess Margaret Cancer Centre and University Health Network Department of Pathology.
Author information
Authors and Affiliations
Contributions
All Authors contributed to the preparation of the manuscript equally.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Grenier, K., Kao, J. & Diamandis, P. Three-dimensional modeling of human neurodegeneration: brain organoids coming of age. Mol Psychiatry 25, 254–274 (2020). https://doi.org/10.1038/s41380-019-0500-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-019-0500-7
- Springer Nature Limited
This article is cited by
-
Generation of ‘semi-guided’ cortical organoids with complex neural oscillations
Nature Protocols (2024)
-
Harnessing the potential of machine learning and artificial intelligence for dementia research
Brain Informatics (2023)
-
Organoids for modeling prion diseases
Cell and Tissue Research (2023)
-
Esophageal organoids: applications and future prospects
Journal of Molecular Medicine (2023)
-
Effects of Sevoflurane Exposure on Fetal Brain Development Using Cerebral Organoids
Journal of Molecular Neuroscience (2022)