Abstract
Cytotoxicity, speedy degradation, and limited cellular absorption are the foremost features influencing the successful delivery of RNAs. Chitosan (Cs) is a polymer that offers an advantage due to its bio-compatibility and biodegradable nature, making it an ideal polycationic vector for delivering siRNA. In this study, chitosan has been modified with arginine in order to increase its encapsulation of siRNA and improve cellular absorption. It was discovered that arginine and guanidino moieties could transport through membranes of cells and play an important part in membrane permeability. FTIR and 13C NMR were used to characterize the complex. These chitosan-arginine (CsAr) siRNA complexes are further encapsulated in anionic DPPC/cholesterol liposomes to combine the effects of liposome-chitosan-arginine complexes called lipopolyplexes (LCAr). Formed LCAr were investigated for their lipid/CsAr-siRNA ratios, size, zeta-potential, heparin, and serum RNase stability by agarose gel retardation, and cell uptake efficiency compared to their “parent” polyplexes. Results revealed complete lipidation of CsAr-siRNA polyplexes at lipid mass ratio 10 resulting in lipopolyplexes in the 120 to 230nm range. Polyplex entrapped ~70% of siRNA, whereas lipidation increases siRNA encapsulation to ~95%. Developed LCAr showed ~4 times less hemolytic potential as compared to the parent polyplexes at the highest siRNA dose. The CsAr-siRNA and its lipid-coated form showed enhanced cellular association as compared to the marketed Lipofectamine 2000 proving its effectiveness in siRNA delivery. CsAr-liposome conjugation is simple and safe, and serves as a robust carrier for gene transport in physiological situations without compromising transfection efficacy.
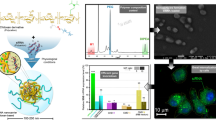
Graphical Abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Gene therapy involves the introduction of a corrected gene or the inhibition of the gene accountable for the excessive expression of a specific disease-causing protein [1]. The transport of small-interfering RNA (siRNA) into cells is one way of gene silencing. Site-specific selectivity and adaptability lead to siRNA, a potential candidate for gene therapy agents in a range of disorders [2]. siRNA molecules are inherently unstable, particularly at very low concentrations following administration, and one of the main challenges is protecting them from degradation by RNases in the in-vivo environment [3]. Furthermore, considering their substantial molecular weight and phosphodiester backbone, siRNA molecules cannot effectively travel through cell membranes [4,5,6]. Cs, a positively charged polysaccharide, is an extensively employed polymer in non-viral gene therapy and formulations for ophthalmic drug delivery [7,8,9,10,11]. Nonetheless, when native Cs were being used, complex durability and transfection effectiveness were often low at physiological pH [12, 13]. Cs-built nucleic acid delivery systems are most effective within a pH varying between 6.8 and 7.0. However, this pH range is not suitable for in vivo drug delivery applications [14]. Several chemical changes have been reported to enhance Cs’ transgenic potency [15, 16]. Various ligands have been linked with Cs for enhancing its complexation with siRNA like branched polyethylenimine, poly-l-arginine, polyguluronate, cyclodextrin, and histidine [17, 18]. Arginine residues have been identified in numerous cell-penetrating peptides (CPPs), playing a critical role in mediating the interaction between CPPs and various receptors [19].
In this study, we have prepared the conjugate of Cs and arginine (Ar) to enrich the complexation capability of Cs alongside anionic siRNA, thereby improving cellular interaction and gene silencing proficiency of CsAr-siRNA complexes. However, high cationic charges may lead to toxicity; hence, the lipidation of these CsAr-siRNA complexes by phospholipids has been explored. Phospholipids play an essential role in forming cell membranes, demonstrating favorable biocompatible nature, and have obtained clinical validation for drug delivery applications [20, 21]. When compared to the unmodified complex, phospholipid-containing lipopolyplexes had much lower surface charges, higher transfection efficiency, lower cytotoxicity, and higher colloidal stability [22, 23]. In vivo research revealed blood longevity without any side effects [24, 25]. With a proper amalgamation of the properties of liposomes and Cs, it is possible to create liposomal formulations that exhibit specific, sustained, and meticulous release characteristics. Quite a few studies have been reported on using chitosan and liposome together to improve drug release stability [8, 26,27,28]. Thus, the physicochemical properties of CsAr-siRNA complexes and their lipid-coated form LCAr were compared for their physiochemical properties, transfection efficiency, and toxicity (in-vitro as well as ex-vivo).
Materials and Methods
Materials
MISSION® siRNA Fluorescent Universal Negative Control #1, 6-FAM (FAM-siRNA), MISSION® siRNA Universal Negative Control #1 (Sc-siRNA), chitosan (medium molecular weight), 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide (EDC), and N-hydroxysuccinimide (NHS) were bought from Merck, India. The 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC) and cholesterol were acquired as generous gifts from Lipoid Ltd., Ludwigshafen, Germany. The Narayana Nethralaya Foundation in Bangalore, India, kindly donated the ARPE19 retinal cells for our research purposes. Cell media and other reagents like DMEM F12 (Dulbecco’s Modification of Eagle’s Medium), antibiotic solution (penicillin G, streptomycin, and amphotericin B solution), 3-(4,5-dimethylthiazol-2-ly)-2,5-diphenyl-tetrazolium bromide (MTT), Dulbecco’s phosphate-buffered saline (DPBS), l-glutamine, and fetal bovine serum (FBS) were bought from Himedia (Mumbai, India). Heparin injection IP (FLAGORIN-5000™) was used as a source of heparin. The rest of the remaining chemicals employed in the research are of laboratory grade.
Cs-Arginine (Ar) Synthesis
Purified Cs and arginine conjugation were carried out by employing EDC and NHS in the role of linking reagents [29, 30]. In short, purified Cs (1 g) plus arginine (0.3g) were solubilized in TEMED/HCl buffer solution (50 ml) (pH 4.7). In this experiment, a solution containing 0.14 g of NHS and 0.6 g of EDC was added to the mixture. The suspension was then subjected to magnetic stirring for a duration of 24 h at room temperature. This allowed for the conjugation reaction between chitosan and arginine to take place, forming stable amide bonds (Fig. 1). The resulting solution was dialyzed against distilled water with the help of a 2000Da MWCO dialysis membrane for 5 days. After dialysis, the purified material was lyophilized to obtain a dry, solid product. The final product was characterized by nuclear magnetic resonance spectroscopy (NMR) and FTIR for their chemical properties.
Characterization of Cs Conjugated with Arginine
Total Reflectance-Fourier Transform Infrared Spectroscopy
The chemical conjugation between arginine and Cs was analyzed using total reflectance-fourier transform infrared spectroscopy (ATR-FTIR). The Cs and arginine, as well as CsAr (conjugated product) specimens, were scanned in the spectral range of 800–4000 cm−1 at a resolution of 4 cm−1. ATR-FTIR spectroscopy allows for the identification and characterization of chemical bonds and functional groups present in the samples, providing valuable information about the conjugation reaction and structural changes that may have occurred.
Nuclear Magnetic Resonance Spectroscopy
For NMR spectroscopy, freeze-dried CsAr was suspended within D2O/H2O (1ml) (1:1 v/v) solution. NMR spectroscopy was performed for 13C NMR on a 600 MHz, NMR spectrometer, Japan. Similarly, the NMR spectra for arginine and purified Cs were recorded.
Preparation of CsAr-siRNA Complexes (polyplex) and Their Lipidation (LPP)
To avoid RNase contamination, polymer solutions, as well as liposomes, were prepared in diethyl pyrocarbonate (DEPC)–treated water/buffer and purified via membrane filters (0.22 μm, Merck, India). During each investigation, RNase-free products along with environments were utilized. To prepare CsAr-siRNA complexes, the freeze-dried CsAr was dispersed in a sodium acetate buffer (0.1 M, pH 5.5). Increasing amounts of CsAr were added to a fixed quantity of siRNA (20 pmol). The complexation between CsAr and siRNA was analyzed by gel electrophoresis and EtBr intercalation method, associating the results with those obtained with naked siRNA.
Thin-film hydration was implemented to create DPPC/Cholesterol liposomes (10:26 mol/mol). In this process, DPPC and cholesterol were dissolved in chloroform and subsequently evaporated below 600 mmHg vacuum and 40°C temperature at 100 rpm speed. A rotating flask evaporator was used to create the transparent lipid film. The generated lipid film was then hydrated for an hour in 1 ml of DEPC sterile water at 55°C. Following hydration, the mixture underwent sonication with a bath sonicator over 15 min. Unilamellar liposomes were generated by extruding the suspension through a polycarbonate membrane (200 nm) for eleven counts employing a Mini-Extruder (Avanti Polar Lipids), at a controlled temperature of 45°C. The lipidation of the CsAr-siRNA polyplex was obtained by incubating DPPC liposomes with CsAr-siRNA polyplex in the different mass ratios (i.e., 1–12) at 37°C for half an hour. Formed LPP was then investigated for its heparin compatibility, serum, and RNase assay [25].
Characterization of CsAr-siRNA Complexes (polyplex) and Their Lipidation (LPP)
Ethidium Bromide (EtBr) Intercalation Assays
Complexation of CsAr-siRNA and Lipidation in a Different Mass Ratio
When EtBr intercalates with siRNA, it exhibits fluorescence. Therefore, a boost in fluorescence specifies an existence of uncomplexed, loose, or freed siRNA. A creation of complexes including siRNA and CsAr or lipid complexes of CsAr may lower fluorescence due to decreased intercalating properties. The reduced fluorescence was utilized to quantify siRNA condensation in polyplexes or lipopolyplexes. In the positive control well, 50 μl of siRNA (20 pmol) was added. The remaining wells, also containing 20 pmol of siRNA, were supplemented with varying concentrations of CsAr (ranging from 5 to 150 μg/ml). Ethidium bromide (40 μl, 0.25 μg/ml) was applied right away into the wells. Negative controls consisted of wells devoid of RNA as well as polyplex. The fluorescence intensity was measured after mixing 45 min of incubation at 37℃ by using a microplate reader. A fluorescence was quantified on excitation and emission wavelengths set at 510 and 590 nm, respectively. The background fluorescence of siRNA/ethidium bromide solution without polymers was set to 100, whereas that of ethidium bromide without siRNA was set to 0 [31, 32]. The relative fluorescence was determined by subtracting the fluorescence of EtBr from the fluorescence of the complexed siRNA and EtBr. This value was divided by the difference between the fluorescence of uncomplexed siRNA and EtBr and the fluorescence of EtBr alone. The resulting quotient was multiplied by 100 to obtain the relative fluorescence as a percentage.
Competitive ligand assay
The optimized complexes (CsAr and LCAr) were evaluated for their siRNA holding capacity in the presence of strong polyanionic compounds like heparin. Optimized CsAr-siRNA (20pmol) and its lipid-coated complex (lipidated CsAr-siRNA (20pmol) were exposed to the different concentrations of heparin from 0.25 to 2 IU, for half an hour. Following a half-hour incubation, the samples were transferred to a 96-well plate (Nunc, USA). The quantification of siRNA release, measured as % relative fluorescence, was performed via EtBr intercalation assay at various concentrations of heparin.
Agarose Gel Electrophoresis
Confirmation of siRNA and CsAr complexation, as well as the formation of LPPs, was achieved through a gel electrophoresis assay. Concisely, a mixture of 20 ul of the complexes comprising siRNA (20pmol) plus 2 µl loading buffer (6×) (Thermofisher Scientific) was loaded in the wells upon 2.7%w/v agarose gel comprising 0.5ug/ml EtBr. The gel electrophoresis was conducted at 70V for 45 min in 1× TAE Buffer (Tris-acetate ethylenediamine tetraacetic acid buffer (50X), Thermofisher Scientific) (pH 8.4). The band integrity of complexes was then examined in gels using the Alpha imager (Cell Biosciences, CA).
RNase Assay
To assess RNase stability, complexes containing siRNA (20pmol) received treatment by RNase A (10 U/ml) on a ratio of 1 U/μg of siRNA. Incubation was conducted for half an hour on 37°C. To stop the reaction, 0.5 μL of RNase OUT (100 mM) (ThermoFisher Scientific, USA) was added. The complexes were then dissociated by employing triton X 100 plus 50 IU of heparin. Subsequently, the mixture was incubated for 15 min at ambient temperature. Gel electrophoresis was used for examining the samples under the conditions described above.
Serum Stability Assay
Similarly, complexes comprising 1.3µg of siRNA was incubated alongside 10% FBS (final volume of the reaction mixture was 100µl) followed by their incubation at 37°C for 4 h. Following exposure, 50 µl Trizol was mixed into each specimen, and the samples then centrifuged at 12,000×g over 15 min at a temperature of 4. The foremost aquatic layer (20 µl) was isolated from all samples and mixed with 3µl of a loading buffer. All samples were analyzed by gel electrophoresis for the siRNA band integrity as described above. The samples with (+ve) and without (−ve) RNase/serum treatment were compared to assess the RNA integrity [33, 34].
Measurement of Complex Sizes and Zeta Potentials
The formation of nanoscale complexes was monitored using dynamic light scattering (DLS). The size and zeta potentials (ZP) of CsAr/siRNA complexes and LCAr were determined utilizing the Zetasizer Nano ZS-90 (Malvern, UK) at ambient temperature (25°C). All complexes were reconstituted 100-fold by 1% DEPC-treated filtered H2O prior to testing, and analyses were done in triple.
Transmission Electron Microscopy
To confirm the size of CsAr and LCAr, transmission electron microscopy (TEM) was performed. Briefly, the suspensions of CsAr and LCAr were placed on the Formvar/carbon grid followed by the draining of surplus fluid, and the grid was air-dried before being stained negatively with 2% of the phosphotungstic acid hydrate at neutral pH. Imaging was performed by employing the TEM (TECNAI 12 BT/FEI) outfitted with a great-resolution slow-scan CCD camera (GATAN Inc., USA), operating at 300 kV.
In-Vitro Cellular Toxicity Assays
Hemolysis Study
To evaluate the hemocompatibility of the optimized formulations, hemolysis assay was performed. Blood samples were obtained from Sprague-Dawley rats through a retro-orbital puncture using heparin tubes. These tubes were then gently overturned five to six times. Rat red blood cells (RBCs) underwent a PBS (0.144 g/l KH2PO4, 9.0g/l NaCl, 0.726g/l Na2HPO4 pH7.4) wash 3–4 times and suspended in HEPES-buffered glucose (HBG). A suitable dilution was performed on the cell solutions to reach a level of 108 cells/ml and mixed with different amounts of siRNA carriers at final siRNA concentrations of 0, 10, 20, 40, 70, and 100 pmol. Negative and positive controls were prepared using RBCs in pure HBG and 1% Triton X-100 in HBG, respectively. Following a 4-h incubation at 37℃, the samples were spun up at 3000 rpm for 10 min at 4°C. A UV-visible spectrophotometer was used to measure heme release in the supernatant at 540 nm [35]. The percentage of hemolysis was determined using the following equation:
The absorbance measurements at 540 nm of the sample and positive control are represented by A. sample and A. PC, accordingly.
Cell Viability Assay
Human ARPE-19 cells were cultured and kept in an incubator at 37°C with 5% CO2 in a humidified atmosphere. These cells were cultured in a 1:1 mixture of DMEM/F12 medium with 10% FBS and antibiotics. The MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide, Merck) assay was employed to assess the cytotoxicity of CsAr and LCAr in the ARPE-19 cell line in-vitro. Cells were added at 7000 cells/well in a 96-well plate and incubated for 24 h at 37°C with 5% CO2. Formed complexes at different siRNA concentrations, i.e., 1–10pmol of siRNA, were incubated with cells over 4 and 24 h. Following the removal of the culture medium, 200µl of 0.2 mg/ml MTT solution was introduced to every well of the plate. The plate had been incubated again for 4 h. MTT solution was withdrawn following exposure, and 100µl of DMSO was incorporated for dissolving the formazon particles. Utilizing an ELISA plate reader (Bio-Rad, USA), an absorbance of the dissolved solution has been determined between 570 and 650 nm (Bio-Rad, USA). The cells treated with PBS were used as a negative control. All formulation’s cell viability was normalized to the value derived from the negative control. The following formula was used to compute the percentage of cell viability.
Cellular Internalization Study In-Vitro
Flow Cytometry
Flow cytometry was performed to quantify the siRNA uptake after CsAr and LCAr transfection. ARPE 19 cells were seeded in a six-well plate at a density of 5 ×105 cells/well and permitted to grow for 24 h until reaching 75% confluence. The cells were then explored with complexes containing FAM-siRNA (100nM), in 1 mL of serum-free culture medium. The cells were incubated with the complexes for a duration of 4 h. Following the incubation period, the cells were subjected to three washes with DPBS, detached using 0.05% trypsin and 0.02% EDTA, and then washed again with DPBS solution. The cells were subsequently resuspended in 0.3 mL of PBS for flow cytometric analysis using a fluorescence-activated cell sorter (FACS-BD-Aria III, BD, USA) [36, 37]. The fluorescence activity was measured to assess cell integrity. The obtained values were compared to those observed with naked Fam-siRNA and FAM-siRNA complexed with Lipofectamine 2000.
Confocal Laser Scanning Microscopy
Cellular internalization of CsAr and LCAr in ARPE 19 cells was studied by confocal microscopy. A total of 10,000 cells were seeded per well in six-well plates containing glass coverslips (0.17 mm2) at the bottom. After 24 h, the cells were transfected with CsAr and LCAr complexes carrying scrambled FAM-siRNA at a dose of 100 nM siRNA. Following a 4-h incubation, the cells were promptly washed with cold DPBS and then fixed with an ice-cold solution of 4% paraformaldehyde for 10 min. Subsequently, they were washed three times with PBS. For nuclear staining, DAPI (4′,6-diamidino-2-phenylindole) at a concentration of 1 µg/mL was applied for 10 min. After another round of washing, the coverslips were mounted on slides for visualization using a confocal laser scanning microscope (CLSM 710, Carl-Zeiss Inc., USA). The negative and positive controls were naked FAM-siRNA and Lipofectamine2000 complexed FAM-siRNA, respectively.
Statistical Analysis
Unless specified otherwise, the values are presented as the mean ± standard deviation (SD) and are calculated based on a least 3 observations. The statistical analysis for flow cytometry involved the use of the one-way ANOVA test, considering a threshold of P<0.05 for establishing statistical significance.
Results and Discussion
Characterization of Cs Conjugated with Arginine
Chitosan stands out as a promising non-viral vector material for delivering nucleic acids; yet, there are hardly any chitosan-constructed nucleic acid delivery systems that have been employed commercially. This might be due to the numerous shortcomings associated with chitosan like its poor water solubility, meager targeting competencies, and depletion of the charge at biological pH. In the present investigation, we have anticipated simple conjugation of the Cs with arginine followed by their lipidation to promote the binding efficiency of Cs and stability of the siRNA in the presence of serum proteins. Using EDC/NHS as coupling agents, amidation of the main amine groups found in Cs glucosamine (GlcN) units, l-arginine grafted chitosan (Cs-Arg/CsAr) was produced. The covalent conjugation of Cs and arginine was established by FTIR-ATR and NMR. FTIR identified the effective grafting of Ar onto Cs via EDC/NHS coupling chemistry. For Cs, the absorption band for distinguishing NH2 scissoring was obtained at 1639 cm−1, C–O stretching vibrations of the pyranose ring at 1031–1072 cm−1, and carbonyl asymmetric stretching vibration at 1564 cm−1. The arginine used for conjugation showed an absorption band at 1657 cm−1 for the guanido group present in it and the COO− symmetric bending absorption band at 1439 cm−1 has been observed. In addition to this, C–C–N asymmetric bending and COO− scissoring modes were observed at 1136 cm−1 and 768 cm−1 respectively. A characteristic peak of arginine 1639 cm−1 and 1443 cm−1 in addition to ether bond stretching vibrations of the pyranose ring of Cs in conjugate CsAr has been observed as shown in Fig. 2. Followed by FTIR, 13C NMR spectroscopy has been performed to confirm the conjugation of Cs and arginine. It has been observed that many characteristic peaks of Cs and arginine have overlapped in the CsAr as shown in Fig. 3. Amide group peak at 1532 cm−1 was observed in the CsAr which is due to the establishment of a covalent linkage between the carboxylate of arginine and the amino groups of Cs ensuring the successful conjugation of Cs and arginine as shown in Fig. 2 (Liu et al., 2004) (Park et al., 2013).
Formation of CsAr-siRNA Polyplex and their Lipidation (LCAr)
The key significance of arginine is related to the guanidino moiety, which is cationic in nature and has a unique bidentate structure, which may point to the reaction between the vector and the incorporated gene [38, 39]. Arginine’s guanidino group can establish hydrogen bonds with the phosphate backbones of DNA/RNA [40, 41].
The efficiency of CsAr to form complexes with siRNA was analyzed by EtBr dye displacement assay. EtBr can intercalate among base pairs in siRNA because it is double-stranded, and enough fluorescence was observed to investigate polymer-siRNA interactions. The CsAr–siRNA complexation prevents EtBr from binding to siRNA, resulting in a decrease in fluorescence intensity. CsAr was added in increasing concentration (5µg/ml to 150 µg/ml) to 20 pmol of siRNA. The % relative fluorescence intensity is designed as a function of polymer addition in Fig. 4a. The CsAr showed a maximal quenching (approx. 70% siRNA complexation) at 20µg/ml concentration. Further, the fluorescence plateau was reached after 40µg/ml concentration. Hence, CsAr at a concentration of 20µg/ml was further used for the preparation of lipopolyplexes. Additionally, the agarose gel retardation experiment was also used to validate the production of CsAr-siRNA complexes at a concentration of 20µg/ml, which shows maximum complexation of siRNA according to the EtBr exclusion assay. Gel electrophoresis assay was used for authorization of complex integrity among CsAr and siRNA. Freeze-dried CsAr in concentrations ranging from 20 to 120ug/ml was utilized for the synthesis of CsAr-siRNA complexes. It has been observed that the addition of CsAr leads to the complete complexation of siRNA as there is no siRNA release (Fig. 6a). This can be equated to the electrophoretic mobility of free siRNA, which was utilized as a positive control in the study. Arginine’s guanidino moiety might establish hydrogen bonds with the phosphate backbones of RNA thereby increasing their complexation efficiency with Cs [42]. In order to use a minimal quantity of CsAr, 20µg/ml of CsAr was further used for the lipidation.
Ethidium bromide exclusion assay. a CsAr-siRNA condensation assay. siRNA condensation was analyzed as a reduction in fluorescence of EtBr added to polyplexes. b Heparin competition assay. The release profile of siRNA-loaded CsAr polyplex and lipopolyplex (LCAr) at different amounts of heparin, n = 3
To evaluate the effect of lipidation, the ternary complexes were formed mainly by means of electrostatic interaction using DPPC/cholesterol liposomes. As the lipid mass ratio was increased, there was a further decrease in the EtBr fluorescence as shown in Fig. 5b. It has been observed that after lipid mass ratio 10, there is no further decrease in the EtBr fluorescence. At lipid mass ratio 10, the % relative fluorescence was found to be 3.7% ensuring substantial encapsulation of siRNA (precisely around 97.3%). Hence, lipid mass ratio 10 was further used for evaluation of their safety and stability in the existence of serum, RNase, and heparin.
Size and Surface Charge of CsAr-siRNA Polyplex and Their Lipidated Form (LCAr)
When it comes to particle size of this lipid-coated CsAr-siRNA complex, it has been observed that the CsAr-siRNA polyplex showed an average particle diameter of 108.70± 6.81nm with PDI 0.19± 0.03 which was amplified with a rise in the lipid mass ratio. Lipidation of the CsAr-siRNA polyplex leads to an increase in the particle size of the original CsAr-siRNA polyplex as shown in Fig. 5a. The rise was most likely due to chitosan-liposome bridging [43]. The mean diameter of lipopolyplexes of CsAr-siRNA complexes ranges 120–230nm, centered around the mass ratio of liposomes: CsAr-siRNA polyplex.
Zeta potential (ζ potential) is a degree of particle surface electrical charge that is frequently explored to characterize colloidal drug delivery systems. Changes in ZP values provided more evidence of complex formation via electrostatic interactions. The potential of the negatively charged liposomes to conceal the net positive charge of the CsAr-siRNA polyplex resulted in a shift towards less positive values. The influence of lipid coating on the surface charge of the CsAr-siRNA polyplex is shown in Fig. 5a. It was found that with an increase in the lipid mass ratio, the ZP of CsAr-siRNA polyplex drops from 14.73±0.40 to 11.14±0.04mV at lipid mass ratio 12. The zeta potential at lipid mass ratios 10 and 12 did not show any significant difference. There was no further decrease in the ZP observed after lipid mass ratio 10 (11.21±0.24mV). The zeta potential was shown to drop with growing lipid carrier abundance and to stabilize after accomplishment of the complexation point, implying charge neutralization at the complexation point followed by an excess of carriers resulting in positive zeta potential. These findings support the anionic nanocarriers’ ability to neutralize the cationic charge of CsAr-siRNA and form ternary complexes via electrostatic contact. The particle size, PDI, and ZP of the final optimized products were mentioned in the Table I.
Agarose Gel Electrophoresis
The Integrity of siRNA in the Presence of RNase
The capability of CsAr and LCAr to protect siRNA against nuclease enzyme was investigated by gel electrophoresis assay. RNase A destroyed free siRNA in 30 min, whereas siRNA complexed with CsAr and LCAr was seen completely protected from RNase after 30 min. As shown in Fig. 6b, siRNA band intensity of +ve (i.e., RNase treated) and −ve (i.e., RNase non-treated) showed the same band intensity proving no degradation of siRNA molecule in the presence of RNase enzymes.
The siRNA Integrity in the Presence of Serum
Due to the rapid siRNA disintegration in blood by serum proteins, the designed siRNA carrier system must stand the necessary characteristics for preventing encapsulated siRNA from such degradation. Gel electrophoresis assay has shown that siRNA without a complexing agent was destroyed following 4 h of incubation. The free siRNA in CsAr-siRNA also gets degraded to some extent after 4 h of incubation which leads to the low siRNA band intensity of +ve (i.e., serum treated sample) as compared to the −ve (non-serum treated sample) as shown in Fig. 6b. Lipidation of CsAr leads to the complete complexation of siRNA as shown in Fig. 4b which protects the degradation of siRNA from serum proteins.
siRNA Release in the Presence of Competitive Ligand (Heparin)
Heparin, as a polyanion typically assigned to cell surface proteoglycans, strives with cationic complexes and shifts condensed DNA/siRNA, which results in premature siRNA release following contact with cell surfaces [44, 45]. As therefore, we chose to investigate the RNA-protective properties of CsAr polyplex and LCAr in the presence of heparin. The EtBr intercalation experiment verified siRNA release in the presence of heparin. In Fig. 4b, at 2IU of heparin, it was revealed that more than 70% of the siRNA was released from the CsAr-siRNA polyplex, whereas the lipid-coated CsAr-siRNA polyplex LCAr leads to a release of around 50% of siRNA at 2 IU of heparin. This shows better stability. The liposome coating successfully prevents the siRNA release in the presence of polyanions [46].
Morphology of CsAr-siRNA and Its Lipidated Form
TEM imaging of the CsAr-siRNA complexes and lipopolyplexes (LCAr) revealed more information about their structure (Fig. 7). The lipid coat around the CsAr-siRNA complex can be seen with some partial encapsulation (breaks in the lipid coat) (Fig. 7b), which is consistent with earlier results [43, 47]. The size of both complexes matches that of the DLS analysis.
In-Vitro Toxicity Assay
Hemolysis Study
A hemolysis assay was executed to examine the complexes’ suitability for an in-vivo application. Erythrocytes have an anionic charge on their surface and have been shown the interact with cationic-charged polyplexes, resulting in the development of aggregates and systemic toxicity [48]. The amount of hemoglobin released was measured to determine the aptitude of different complexes to impair the erythrocyte membrane. siRNA in varying doses ranging from 10 to 100pmol (respective concentration CsAr and liposomes were also increased) were treated with rat RBCs in HBG buffer for 4 h at 37℃. Liposome-coated CsAr-siRNA complexes, i.e., LCAr, showed higher hemocompatibility as compared to the CsAr-siRNA polyplexes as seen in Fig. 8a. At the highest siRNA concentration, i.e.,100pmol, polyplex CsAr-siRNA complexes showed % hemolysis of 5.69±0.06%, which was successfully lowered by lipidated CsAr-siRNA complexes by 1.98±0.02%. This proves that lipidation improves the hemolytic potential of CsAr-siRNA polyplexes thereby improving their hemocompatibility and making them suitable for systemic administration, which was also supported by the findings of Baghdan and colleagues [43].
In-vitro toxicity assay. a The extent of hemolysis was evaluated by measuring absorbance at 550 nm. The degree of damage caused to blood erythrocytes by different concentrations of siRNA 10–100pmol was calculated as the percentage lysis of the total red blood cells added. The results are averages and standard deviations of three distinct sets of experiments. b, c Cell viability assay of CsAr-siRNA and LCAr complexes by using ARPE-19 cells. The siRNA concentration was taken in the range of 1–10 pmol. The results were displayed as mean ± SD (n = 3)
Cell Viability Assay
An MTT test has been implemented to assess the survival of cells that had been treated with CsAr-siRNA complexes and their lipidated form (LCAr), which was standardized to non-treated cells as a standard. The cells were incubated for 4 and 24 h after being treated with different concentrations of siRNA (1–10pmol). It has been observed that even at a higher dose, i.e.,10 pmol LCAr, showed good cell viability as compared to the CsAr-siRNA complexes (Fig. 8b, c). Lipidation of CsAr-siRNA complexes reduced the toxicity associated with CsAr’s high cationic charge. This is in line with past research indicating that it emphasizes the role of liposomes in diminishing the cationic charge of complexes, consequently improving their biocompatibility. Prior studies have consistently indicated that the heightened surface charge density of polycationic polymers is a key contributor to cytotoxicity [49, 50].
Flow Cytometry
A critical factor that affects the transgenic efficiency of the gene delivery system is the degree to which siRNA complexes are taken up by the cells. As a result, the main emphasis was on investigating whether conjugating arginine moieties onto Cs molecules might improve siRNA cellular uptake. Lipofectamine 2000 which is a traditional transfection gold standard showed an MFI of 79.35±8.84%, whereas CsAr-FAM siRNA complexes showed 89.40±0.85%MFI and its lipidated form LCAr showed an MFI of 88.05±0.21% as shown in Fig. 9. Both forms of CsAr showed higher cellular uptake as compared to the Lipofectamine 2000. The increased positive surface charge and stability of complexes caused by the addition of arginine to chitosan may have led to the enhanced cellular association of the siRNA loaded in CsAr. Naked siRNA is often not efficiently taken up by cells because it faces electrostatic repulsion when interacting with the negatively charged cell membrane [51].
Quantification of cellular uptake as mean fluorescence intensity (MFI) of CsAr and LCAr/siRNA. Flow cytometry data for cellular uptake of CsAr and LCAr/siRNA. Comparative data (n=3) of intracellular uptake of FAM-labeled siRNA in ARPE-19. (**p<0.001 vs. placebo (Control) cells, comparisons with lipofectamine 2000 did not show any significant differences (ns))
Confocal Microscopy
The subcellular location of FAM-labeled siRNA complexed with CsAr and its lipidated form LCAr in ARPE-19 cells was investigated using confocal microscopy. When excited at 493 nm, the fluorophore FAM exhibits green fluorescence. After 4 h of incubation, uniform distribution of FAM-siRNA was observed mainly in the cytoplasm as shown in Fig. 10. Confocal microscopy also revealed that arginine aids in intracellular localization. Zhang and co-workers studied the internalization pathway involved in arginine-conjugated polymers, i.e., chitosan [29]. They demonstrated that conjugating arginine to Cs improves the cellular intake of polymer/DNA particles and causes alterations in endocytic pathways, resulting in the preference for internalization of arginine-conjugated Cs-DNA via caveolin-mediated endocytosis. Interestingly, whereas the majority of other pathways (like the clathrin-dependent pathway, clathrin- and caveolin-independent endocytosis, and phagocytosis) transfer the delivered cargo to lysosomes, the cell’s breakdown compartment, the caveolin-arbitrated pathway appears to avoid this destination [52, 53]. This may be reinforced by the recent findings by Kim et al., which showed that the caveolin-facilitated endocytosis inhibitor reduced gene expression interceded by arginine peptide to a superior level than the clathrin-dependent endocytosis inhibitor [54].
Conclusion
We established in this work that arginine-modified chitosan-siRNA complexes, as well as their lipidated form, can be used to transport siRNA in vitro. Arginine-modified chitosan was created by chemically conjugating arginine to the chitosan backbone and then utilized to prepare polyplexes via charge complex formation with siRNA. Conjugation of arginine with chitosan remarkably improved the cellular uptake and internalization of siRNA. Moreover, the technique of lipidation of CsAr-siRNA polyplexes with phospholipids and cholesterol provides a great advantage in terms of reducing toxicity and improving the stability of the CsAr polyplexes. The combination of liposomes with cationic polymer siRNA complexes assures that the positive charge of the cationic nanoparticles is lowered, making them more biocompatible with an enhanced safety profile, as demonstrated by hemolysis assays. Based on these discoveries, chitosan-arginine-based complexes may serve as a potent carrier for gene delivery under physiological settings without reducing transfection effectiveness.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Shahryari A, Saghaeian Jazi M, Mohammadi S, Razavi Nikoo H, Nazari Z, Hosseini ES, Burtscher I, Mowla SJ, Lickert H. Development and clinical translation of approved gene therapy products for genetic disorders. Front Genet. 2019;10:868. https://doi.org/10.3389/fgene.2019.00868.
Hu B, Zhong L, Weng Y, Peng L, Huang Y, Zhao Y, Liang X-J. Therapeutic siRNA: state of the art. Signal Transduct Target Ther. 2020;5:101. https://doi.org/10.1038/s41392-020-0207-x.
Feng L, Li SK, Liu H, Liu CY, Lasance K, Haque F, Shu D, Guo P. Ocular delivery of pRNA nanoparticles: distribution and clearance after subconjunctival injection. Pharm Res. 2014;31:1046–58. https://doi.org/10.1007/s11095-013-1226-x.
Hamoudi MC, Henry E, Zerrouk N, Scherman D, Arnaud P, Deprez E, Escriou V. Enhancement of siRNA lipid-based vector stability and siRNA integrity in human serum with addition of anionic polymer adjuvant. J Drug Deliv Sci Technol. 2015;26:1–9. https://doi.org/10.1016/j.jddst.2015.01.001.
Arruda DC, Gonzalez IJ, Finet S, Cordova L, Trichet V, Andrade GF, Hoffmann C, Bigey P, de Almeida Macedo WA, da Silva Cunha A, Malachias de Souza A, Escriou V. Modifying internal organization and surface morphology of siRNA lipoplexes by sodium alginate addition for efficient siRNA delivery. J Colloid Interface Sci. 2019;540:342–53. https://doi.org/10.1016/j.jcis.2019.01.043.
Ribeiro MCS, de Miranda MC, Cunha PS, Andrade GF, Fulgêncio GO, Gomes DA, Fialho SL, Pittella F, Charrueau C, Escriou V, Silva-Cunha A. Neuroprotective effect of siRNA entrapped in hyaluronic acid-coated lipoplexes by intravitreal administration. Pharmaceutics. 2021;13:845. https://doi.org/10.3390/pharmaceutics13060845.
Chaharband F, Daftarian N, Kanavi MR, Varshochian R, Hajiramezanali M, Norouzi P, Arefian E, Atyabi F, Dinarvand R. Trimethyl chitosan-hyaluronic acid nano-polyplexes for intravitreal VEGFR-2 siRNA delivery: formulation and in vivo efficacy evaluation. Nanomedicine. 2020;102181. https://doi.org/10.1016/j.nano.2020.102181.
Yao Y, Su Z, Liang Y, Zhang N. pH-Sensitive carboxymethyl chitosan-modified cationic liposomes for sorafenib and siRNA co-delivery. Int J Nanomedicine. 2015;10:6185–97. https://doi.org/10.2147/IJN.S90524.
Ragelle H, Riva R, Vandermeulen G, Naeye B, Pourcelle V, Le Duff CS, D’Haese C, Nysten B, Braeckmans K, De Smedt SC, Jérôme C, Préat V. Chitosan nanoparticles for siRNA delivery: optimizing formulation to increase stability and efficiency. J Control Release. 2014;176:54–63. https://doi.org/10.1016/j.jconrel.2013.12.026.
Jean M, Alameh M, De Jesus D, Thibault M, Lavertu M, Darras V, Nelea M, Buschmann MD, Merzouki A. Chitosan-based therapeutic nanoparticles for combination gene therapy and gene silencing of in vitro cell lines relevant to type 2 diabetes. Eur J Pharm Sci. 2012;45:138–49. https://doi.org/10.1016/j.ejps.2011.10.029.
Howard KA, Rahbek UL, Liu X, Damgaard CK, Glud SZ, Andersen MØ, Hovgaard MB, Schmitz A, Nyengaard JR, Besenbacher F, Kjems J. RNA interference in vitro and in vivo using a novel chitosan/siRNA nanoparticle system. Mol Ther. 2006;14:476–84. https://doi.org/10.1016/j.ymthe.2006.04.010.
Mao S, Sun W, Kissel T. Chitosan-based formulations for delivery of DNA and siRNA. Adv Drug Deliv Rev. 2010;62:12–27. https://doi.org/10.1016/j.addr.2009.08.004.
Xu S, Dong M, Liu X, Howard KA, Kjems J, Besenbacher F. Direct force measurements between siRNA and chitosan molecules using force spectroscopy. Biophys J. 2007;93:952–9. https://doi.org/10.1529/biophysj.106.093229.
Sato T, Ishii T, Okahata Y. In vitro gene delivery mediated by chitosan. Effect of pH, serum, and molecular mass of chitosan on the transfection efficiency. Biomaterials. 2001;22:2075–80. https://doi.org/10.1016/S0142-9612(00)00385-9.
Singh B, Choi YJ, Park IK, Akaike T, Cho CS. Chemical modification of chitosan with ph-sensitive molecules and specific ligands for efficient DNA transfection and siRNA silencing. J Nanosci Nanotechnol. 2014;14:564–76. https://doi.org/10.1166/jnn.2014.9079.
Heidari R, Khosravian P, Mirzaei SA, Elahian F. siRNA delivery using intelligent chitosan-capped mesoporous silica nanoparticles for overcoming multidrug resistance in malignant carcinoma cells. Sci Rep. 2021;11:20531. https://doi.org/10.1038/s41598-021-00085-0.
Sun P, Huang W, Kang L, Jin M, Fan B, Jin H, Wang QM, Gao Z. siRNA-loaded poly(histidine-arginine)6-modified chitosan nanoparticle with enhanced cell-penetrating and endosomal escape capacities for suppressing breast tumor metastasis. Int J Nanomedicine. 2017;12:3221. https://doi.org/10.2147/IJN.S129436.
Ragelle H, Vandermeulen G, Préat V. Chitosan-based siRNA delivery systems. J Control Release. 2013;172:207–18. https://doi.org/10.1016/j.jconrel.2013.08.005.
Zhou Y, Han S, Liang Z, Zhao M, Liu G, Wu J. Progress in arginine-based gene delivery systems. J Mater Chem B. 2020;8:5564–77. https://doi.org/10.1039/d0tb00498g.
Torchilin VP. Recent advances with liposomes as pharmaceutical carriers. Nat Rev Drug Discov. 2005;4:145–60. https://doi.org/10.1038/nrd1632.
Beltrán-Gracia E, López-Camacho A, Higuera-Ciapara I, Velázquez-Fernández JB, Vallejo-Cardona AA. Nanomedicine review: clinical developments in liposomal applications. Cancer Nanotechnol. 2019;10:11. https://doi.org/10.1186/s12645-019-0055-y.
Rezaee M, Oskuee RK, Nassirli H, Malaekeh-Nikouei B. Progress in the development of lipopolyplexes as efficient non-viral gene delivery systems. J Control Release. 2016;236:1–14. https://doi.org/10.1016/j.jconrel.2016.06.023.
Pinnapireddy SR, Duse L, Strehlow B, Schäfer J, Bakowsky U. Composite liposome-PEI/nucleic acid lipopolyplexes for safe and efficient gene delivery and gene knockdown. Colloids Surf B Biointerfaces. 2017;158:93–101. https://doi.org/10.1016/j.colsurfb.2017.06.022.
Ko YT, Kale A, Hartner WC, Papahadjopoulos-Sternberg B, Torchilin VP. Self-assembling micelle-like nanoparticles based on phospholipid–polyethyleneimine conjugates for systemic gene delivery. J Control Release. 2009;133:132–8. https://doi.org/10.1016/j.jconrel.2008.09.079.
Ewe A, Panchal O, Pinnapireddy SR, Bakowsky U, Przybylski S, Temme A, Aigner A. Liposome-polyethylenimine complexes (DPPC-PEI lipopolyplexes) for therapeutic siRNA delivery in vivo. Nanomedicine. 2017;13:209–18. https://doi.org/10.1016/j.nano.2016.08.005.
Alamelu S, Rao KP. Studies on the carboxymethyl chitosan-containing liposomes for their stability and controlled release of dapsone. J Microencapsul. 1991;8:505–19. https://doi.org/10.3109/02652049109021874.
Guo J, Ping Q, Jiang G, Huang L, Tong Y. Chitosan-coated liposomes: characterization and interaction with leuprolide. Int J Pharm. 2003;260:167–73. https://doi.org/10.1016/s0378-5173(03)00254-0.
Şalva E, Turan SÖ, Eren F, Akbuğa J. The enhancement of gene silencing efficiency with chitosan-coated liposome formulations of siRNAs targeting HIF-1α and VEGF. Int J Pharm. 2015;478:147–54. https://doi.org/10.1016/j.ijpharm.2014.10.065.
Zhang H, Zhu D, Song L, Liu L, Dong X, Liu Z, Leng X. Arginine conjugation affects the endocytic pathways of chitosan/DNA nanoparticles. J Biomed Mater Res A. 2011;98:296–302. https://doi.org/10.1002/jbm.a.33115.
Wang K, Qi Z, Pan S, Zheng S, Wang H, Chang Y, Li H, Xue P, Yang X, Fu C. Preparation, characterization and evaluation of a new film based on chitosan, arginine and gold nanoparticle derivatives for wound-healing efficacy. RSC Adv. 2020;10:20886–99. https://doi.org/10.1039/D0RA03704D.
Sutton D, Kim S, Shuai X, Leskov K, Marques JT, Williams BRG, Boothman DA, Gao J. Efficient suppression of secretory clusterin levels by polymer-siRNA nanocomplexes enhances ionizing radiation lethality in human MCF-7 breast cancer cells in vitro. Int J Nanomedicine. 2006;1:155–62. https://doi.org/10.2147/nano.2006.1.2.155.
Safari F, Tamaddon AM, Zarghami N, Abolmali S, Akbarzadeh A. Polyelectrolyte complexes of hTERT siRNA and polyethyleneimine: effect of degree of PEG grafting on biological and cellular activity. Artif Cells Nanomed Biotechnol. 2016;44:1561–8. https://doi.org/10.3109/21691401.2015.1064936.
Konishi M, Kawamoto K, Izumikawa M, Kuriyama H, Yamashita T. Gene transfer into guinea pig cochlea using adeno-associated virus vectors. J Gene Med. 2008;10:610–8. https://doi.org/10.1002/jgm.1189.
Khatri N, Baradia D, Vhora I, Rathi M, Misra A. cRGD grafted liposomes containing inorganic nano-precipitate complexed siRNA for intracellular delivery in cancer cells. J Control Release. 2014;182:45–57. https://doi.org/10.1016/j.jconrel.2014.03.003.
Upadhya A, Sangave PC. Hydrophobic and electrostatic interactions between cell penetrating peptides and plasmid DNA are important for stable non-covalent complexation and intracellular delivery. J Pept Sci. 2016;22:647–59. https://doi.org/10.1002/psc.2927.
Hu C, Chiang C-H, Hong P, Yeh M-K. Influence of charge on FITC-BSA-loaded chondroitin sulfate-chitosan nanoparticles upon cell uptake in human Caco-2 cell monolayers. Int J Nanomedicine. 2012;7:4861–72. https://doi.org/10.2147/IJN.S34770.
Chen C-W, Yeh M-K, Shiau C-Y, Chiang C-H, Lu D-W. Efficient downregulation of VEGF in retinal pigment epithelial cells by integrin ligand-labeled liposome-mediated siRNA delivery. Int J Nanomedicine. 2013;8:2613–27. https://doi.org/10.2147/IJN.S39622.
Boisguérin P, Deshayes S, Gait MJ, O’Donovan L, Godfrey C, Betts CA, Wood MJA, Lebleu B. Delivery of therapeutic oligonucleotides with cell penetrating peptides. Adv Drug Deliv Rev. 2015;87:52–67. https://doi.org/10.1016/j.addr.2015.02.008.
Zou L, Peng Q, Wang P, Zhou B. Progress in research and application of HIV-1 TAT-derived cell-penetrating peptide. J Membr Biol. 2017;250:115–22. https://doi.org/10.1007/s00232-016-9940-z.
Calnan BJ, Tidor B, Biancalana S, Hudson D, Frankel AD. Arginine-mediated RNA recognition: the arginine fork. Science. 1991;252:1167–71. https://doi.org/10.1126/science.252.5009.1167.
Liu L, Bai Y, Song C, Zhu D, Song L, Zhang H, Dong X, Leng X. The impact of arginine-modified chitosan–DNA nanoparticles on the function of macrophages. J Nanoparticle Res. 2010;12:1637–44. https://doi.org/10.1007/s11051-009-9722-y.
Sakai N, Takeuchi T, Futaki S, Matile S. Direct observation of anion-mediated translocation of fluorescent oligoarginine carriers into and across bulk liquid and anionic bilayer membranes. Chembiochem. 2005;6:114–22. https://doi.org/10.1002/cbic.200400256.
Baghdan E, Pinnapireddy SR, Strehlow B, Engelhardt KH, Schäfer J, Bakowsky U. Lipid coated chitosan-DNA nanoparticles for enhanced gene delivery. Int J Pharm. 2018;535:473–9. https://doi.org/10.1016/j.ijpharm.2017.11.045.
Bertschinger M, Backliwal G, Schertenleib A, Jordan M, Hacker DL, Wurm FM. Disassembly of polyethylenimine-DNA particles in vitro: implications for polyethylenimine-mediated DNA delivery. J Control Release. 2006;116:96–104. https://doi.org/10.1016/J.JCONREL.2006.09.006.
Danielsen S, Maurstad G, Stokke BT. DNA-polycation complexation polyplex stability in the presence of competing polyanions. Biopolymers. 2005;77:86–97. https://doi.org/10.1002/bip.20170.
Supe S, Upadhya A, Tripathi S, Dighe V, Singh K. Liposome-polyethylenimine complexes for the effective delivery of HuR siRNA in the treatment of diabetic retinopathy. Drug Deliv Transl Res. 2023. https://doi.org/10.1007/S13346-022-01281-9.
Schäfer J, Sitterberg J, Ehrhardt C, Kumar MNVR, Bakowsky U. A new drug vehicle - lipid coated biodegradable nanoparticles, in: CIMTEC 2008 - Proceedings of the 3rd International Conference on Smart Materials, Structures and Systems - Biomedical Applications of Smart Materials, Nanotechnology and Micro/Nano Engineering, 2008: pp. 148–153. https://doi.org/10.4028/www.scientific.net/AST.57.148.
Kircheis R, Wightman L, Schreiber A, Robitza B, Rössler V, Kursa M, Wagner E. Polyethylenimine/DNA complexes shielded by transferrin target gene expression to tumors after systemic application. Gene Ther. 2001;8:28–40. https://doi.org/10.1038/sj.gt.3301351.
Ewe A, Schaper A, Barnert S, Schubert R, Temme A, Bakowsky U, Aigner A. Storage stability of optimal liposome-polyethylenimine complexes (lipopolyplexes) for DNA or siRNA delivery. Acta Biomater. 2014;10:2663–73. https://doi.org/10.1016/j.actbio.2014.02.037.
Nafee N, Schneider M, Schaefer UF, Lehr C-M. Relevance of the colloidal stability of chitosan/PLGA nanoparticles on their cytotoxicity profile. Int J Pharm. 2009;381:130–9. https://doi.org/10.1016/j.ijpharm.2009.04.049.
Babu A, Muralidharan R, Amreddy N, Mehta M, Munshi A, Ramesh R. Nanoparticles for siRNA-based gene silencing in tumor therapy. IEEE Trans Nanobioscience. 2016;15:849. https://doi.org/10.1109/TNB.2016.2621730.
Shin JS, Abraham SN. Cell biology. Caveolae–not just craters in the cellular landscape. Science. 2001;293:1447–8. https://doi.org/10.1126/science.1061079.
Pelkmans L, Helenius A. Endocytosis via caveolae. Traffic. 2002;3:311–20. https://doi.org/10.1034/j.1600-0854.2002.30501.x.
Kim T, Baek J, Yoon JK, Choi JS, Kim K, Park J. Synthesis and characterization of a novel arginine-grafted dendritic block copolymer for gene delivery and study of its cellular uptake pathway leading to transfection. Bioconjug Chem. 2007;18:309–17. https://doi.org/10.1021/bc0601525.
Acknowledgements
The authors are grateful to Science and Engineering Board for funding this research. We thank SVKM’S NMIMS SPPSPTM (Shobhaben Pratapbhai Patel School of Pharmacy & Technology Management) and NIRRH (National Institute for Research in Reproductive and Child Health) for supporting this research work by providing their facilities. The authors are thankful to Dr. Debashish Das from Narayana Nethralaya Foundation in Bangalore, India, for providing ARPE-19 cell lines for this research.
Funding
This research project is a result of funding received through the Early Career Award granted to Dr. Kavita Singh by the Science and Engineering Research Board, a division of the Department of Science and Technology, Government of India.
Author information
Authors and Affiliations
Contributions
Shibani Supe: examination, tactic, authentication, writing—first draft. Archana Upadhya: conceptualization, examination, methodology, direction. Vikas Dighe: resources, supervision. Kavita Singh: conceptualization, direction, resources, funding acquisition, writing—proofreading and editing.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Supe, S., Upadhya, A., Dighe, V. et al. Development and Characterization of Modified Chitosan Lipopolyplex for an Effective siRNA Delivery. AAPS PharmSciTech 25, 13 (2024). https://doi.org/10.1208/s12249-023-02728-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12249-023-02728-z