Abstract
Membrane fusion is an essential process for the survival of eukaryotes and entry of the enveloped viruses. Fusion machines (proteins) and membrane lipids play their intricate role in a concerted way to complete the process with precision and speed. The in vitro study of fusion does not match the kinetics of biological fusion, nonetheless, it provides molecular details of the process. The fusion proteins are more celebrated due to their obvious role in catalyzing the process, but the lipids are also important as they provide the platform for the process. Lipid composition provides the membrane organization, dynamics, and overall physical properties to the membrane, which are crucial for the lamellar to nonlamellar to lamellar reorganization during fusion. The current review discusses the prospective roles of the players in fusion with a special emphasis on lipidic shape, chain unsaturation, membrane physical properties, water percolation at the acyl chain region, and membrane domain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
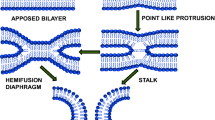
Several fundamental biological processes in eukaryotes are underpinned by a complex choreography of lipids and proteins through which two membranes fuse (fusion) and divide (fission). Fusion is essential for the homeostatic functions of eukaryotes such as exocytosis, endocytosis, intracellular trafficking, muscle development, wound repair, and fertilization. Interestingly, enveloped viruses enter the host cells utilizing the same process. The process of membrane fusion entails the close contact of two lipid bilayers, localized distortion of the distinct bilayers, and reformation into a single, fused membrane [1]. Fusion events are characterized by several intermediary states, referred to as fusion intermediates, distinguishable by their stabilization energies. According to the lipid stalk hypothesis, the stalk, hemifusion diaphragm, and fusion pores are considered as three distinct states en route to the protein-free fusion of lipid membranes (Fig. 1) [1, 2]. The properties of the hemifusion intermediate are acknowledged as its structure determines the fate of the fusion process. In this state, the exterior leaflets of apposed membranes are connected by hemifusion structures, while the inner leaflets remain intact. In addition to the spectroscopic results, the electrophysiological measurements have also bolstered hemifusion [3, 4]. The hemifusion state has typically been defined through lipid mixing without content mixing or as the merging of the lipids of the two contacting bilayers (outer leaflets) without affecting the inner leaflets. The integral membrane proteins are crucial components of numerous biological membranes and are enmeshed within the lipid bilayer, spanning across its width. Fusion proteins are believed to localize at the fusion site and cover the early fusion site with a protein ring. This ring either catalyzes the formation of early fusion intermediates or readies the membrane bilayer for fusion [2, 5]. However, integral proteins do not appear at the hemifusion diaphragm, leaving it as a lipid assembly [6]. In general, the hemifusion diaphragm is a temporary structure that elongates longitudinally to open fusion pores [7].
Diagrammatic representation of different fusion intermediates according to lipid stalk hypothesis. The figure has been adapted from Ref. [30] with permission
The pore opening connects the volumes of two initially distinct membrane interiors. The pore opening has been studied utilizing electrophysiological methods and fluorescence assays. These results have demonstrated that fusion pores can flicker [4] and that the polar heads of the lipids cover the fusion pore edge.
2 Proteins as machines in membrane fusion
In somatic membrane fusion, multiple proteins work in total orchestration to catalyze the fusion process. The transmembrane domain (TMD) of the fusion protein anchors the cell and the N-terminal fusion peptide sticks to the target membrane followed by the docking and fusion. Therefore, fusion proteins play a central role in bringing two membranes into close contact [8,9,10]. In contrast to somatic fusion, viral fusion is majorly carried out by a single fusion protein [11]. Our understanding of fusion protein machines is mostly based on the examination of two biological processes, i.e., viral entry into a host cell and cellular secretory release. Viral entry machines are the simplest fusion tools, and the hemagglutinin (HA) of the influenza virus is the most well-studied fusion protein. The fusion of the influenza virus with the endosomal membrane is driven by HA protein [7]. The potential involvement of several HA regions in the fusion process has drawn attention in attempts to develop a thorough molecular perspective of the function of HA. Epand group has demonstrated that a 127-residue N-terminal construct of HA2 (FHA2-127, comparable to the TBHA2 fragment) generates aggregation, lipid mixing, and content leakage of model membrane vesicles, but could not induce mixing of trapped compartments [12]. This behavior also displayed a native-HA2-like dependence on an intact fusion peptide region and a pH response that was almost identical to that of HA-mediated fusion [13]. It is interesting to note that a shorter construct (FHA2-90) was unable to exhibit the same pH-sensitive membrane-disrupting, and ANS-binding behavior as FHA2-127, indicating that the interhelical loop (residues 105–113) and a brief helical segment (residues 114–127) may be significant in membrane interactions necessary for fusion.
In a reconstructed HA fusion protein glycophosphatidylinositol (GPI) was engineered in place of the TMD demonstrating lipid mixing but incapable of inducing content mixing [14]. The TM section of HA is an α-helix that aggregates in SDS micelles with two to five TM units, angles within 37 of the bilayer normal, and receives exposure to water in bilayers [15]. The formation of fusion pores has also been demonstrated to be hindered or blocked by the deletion or extension of the HA cytoplasmic tail (CT) [16]. The ability of chimeras with the HA TM domain and cytoplasmic tail regions replaced with those of the Sendai virus fusion protein to produce fusion without significantly changing the amount or kinetics of membrane mixing [16], suggests the interchangeability of these components. Interestingly, a single glycine mutation in the TM region eliminated the mixing of contents between vesicular stomatitis-virus-infected cells [17]. The structural makeup of the CT and/or TM regions may have a role in the stabilization or destabilization of an intermediate as well as the induction of lipid rearrangement that results in the development of a stable fusion pore.
Viral fusion proteins initiate the fusion of the viral membrane with the host membrane. A six-helix bundle (coiled-coil motif) development between the heptad repeats 1 and 2 (HR1 and HR2) is essential for the type-I fusion protein. The energy needed for fusion is provided by the development of a six-helix bundle by the HR1 and HR2 domains. However, the six-helix bundle formation is not spontaneous on the viral surface, it occurs when the N-terminal fusion peptide binds with the host cell. A stretch of 15–30 amino acids at the N-terminal of the fusion protein is known as fusion peptide because of their significance in membrane fusion. Studies further suggest that the fusion peptide may alter the membrane interface and trigger the fusion via non-bilayer structures [18]. Therefore, it is presumed that the type-I viral fusion is catalyzed by both the six-helix bundle formation and the destabilization of the host cell membrane by the fusion peptide [18]. We have tabulated the names of fusion proteins and their important domains for somatic and viral fusion in Table 1 [19].
Viral fusion machines are relatively simpler compared to somatic fusion. To understand the functioning of secretory systems, efforts have been made on the protein machines of synaptic vesicle fusion [20]. The process is highly complex and involves several proteins, perhaps because of its fast kinetics (tens of milliseconds as compared to tens to hundreds of seconds for viral fusion). There are at least five phases of synaptic fusion such as targeting, fusion machine assembly or priming, triggering of fusion, fusion and release, disassembly, and machine recycling. The synaptic fusion is accomplished through a class of proteins called Soluble N-ethylmaleimide-sensitive factor Activating protein REceptor (SNAREs), of which SNAP-25 and syntaxin are linked to the target membrane (plasma membrane of neurons) and vesicle-associated membrane protein (VAMP) is associated with the membrane of synaptic vesicles [20]. Another vesicle-associated protein synaptotagmin, is a potential candidate for regulating Ca2+-triggered synaptic vesicle release [21].
Homotypic fusion between endoplasmic reticulum membranes is catalyzed by atlastins and dynamin-like fusogenic GTPases, whereas mitofusins (MFNs) and optic atrophy-1 (OPA1) are responsible for homotypic fusion of mitochondrial outer membrane [22]. Interestingly, a highly stable four-helix bundle formation is the hallmark of SNAREs instead of a six-helix bundle formation in viral fusion. Figure 2 demonstrates the overall mechanism of synaptic, viral, and homotypic fusion. Mounting data backs the idea that both the viral and secretory fusion takes place via molecular rearrangements that are influenced by the presence of proteins, can be investigated, and described at the rudimentary level by using suitable model membrane systems. The functions of a fusion machine, when viewed in the context of the mechanism described here, are to juxtapose lipid bilayers, form highly curved bilayers, and dimpled structures (increasing the free energy of the initial state), stabilize membrane voids (i.e., lowering the free energy barrier to the committed intermediate), and expand the stalk radius, destabilizing the committed intermediate and causing pore formation. Proteins provide the necessary energy and direction, but the materials they must work with are the lipid assemblies, whose physical characteristics ultimately determine and constrain both the process and the final output. Therefore, the role of lipids cannot be ignored though fusion proteins orchestrate the process.
A SNARE forms a four-helix bundle to promote fusion, B six-helix bundle formation by the Type-I viral fusion proteins (FP: fusion peptide, RBD: Receptor Binding Domain), and C GTPase activity and helix bundle formation by mitofusin during fusion of mitochondrial outer membranes. The figure has been adapted from Ref. [19] with permission
3 Lipids as the materials in membrane fusion
Membrane lipids can influence fusion in two distinct ways. The first pathway is referred to as intrinsic because the energetics of the intermediates in lipidic hemifusion are influenced by the head-group or tail-structure of the lipid. The second pathway is referred to as extrinsic, where the energetics of fusion are modified by lipid-fusion protein interactions. Pioneering studies on the modulation of membrane curvature [23], thickness [24], hydration [25,26,27], and dipole moment were described for the intrinsic route. The biological membrane is not uniform, and the lipid composition of various membranes is tailored for a desired biological function. The biological membrane is mostly composed of phospholipids, cholesterol, sphingolipids, and diacylglycerol. The main phospholipids found in biological membranes are phosphatidylcholine (PC), phosphatidylethanolamine (PE), phosphatidic acid (PA), phosphatidylserine (PS), phosphatidylinositol (PI), phosphatidylglycerol (PG), and phosphatidylinositol phosphate (PIP) [19].
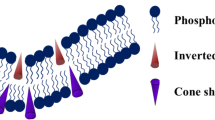
Phospholipids assume different structures based on head-group chemistry and its hydration (Fig. 3). Among them, PC has a cylindrical shape and forms a planar bilayer (Fig. 3) whereas PE and cholesterol have an inverted cone-shaped molecular structure that induces intrinsic negative curvature to the membrane [18, 28,29,30]. On the other hand, lysophosphatidylcholine and oleic acid assume intrinsic positive curvature at physiological pH due to their larger hydrated headgroup compared to the tail thickness [31]. Positive curvature in the bilayer’s outer leaflet typically raises the activation barrier for the formation of the hemifusion intermediate, which inhibits fusion. Negatively charged phospholipids like PS, PA, PG, PI, and PIP bind to the protein’s polybasic regions [32, 33]. However, due to charge-charge repulsion, negatively charged lipids like PS and PG prevent the fusion of model membranes [34]. The presence of Ca2+, which brings PS-doped membranes close to one another, may promote the fusion [34]. Moreover, cholesterol is also one of the important lipid components of the cellular and sub-cellular membranes [35]. It induces a negative curvature in lipid monolayers and modulates the membrane fusion by altering the membrane thickness and fluidity [36]. Membrane fluidity and negative curvature both are responsible for promoting fusion. In contrast, sphingomyelin (SM), is a prime sphingolipid that mostly stabilizes the membrane and inhibits membrane fusion [37]. The unsaturation in the acyl chain further affects the organization and dynamics of membranes, thereby, impacting the fusion process [38, 39].
Different shapes of lipids and their impact on the membrane curvature. The figure has been adapted from Ref. [30] with permission
Joardar et al. have reported that the presence of 30 mol% of PE in PC membranes drives the membrane fusion reaction by circumventing the classical stalk model of fusion [40]. This is solely due to the intrinsic negative curvature of PE that aids the fusion in such a way that the pore opens much earlier in the fusion process. The experimental results support the prediction of the Kasson group from molecular dynamics simulation [41]. It was also shown that the presence of 5–10 mol% oleic acid inhibits fusion due to its intrinsic positive curvature, enhanced hydrophobic tail ordering, and reduction of water penetration into the hydrophobic region [40]. It has been established that water percolation at the acyl chain region of the membrane reduces the activation barrier for fusion [42, 43].
Membrane lipids further modulate the lipid–protein interaction (extrinsic pathway) and alter the fusion process. PS and PIP facilitate the reorganization of fusion proteins and their domains by recruiting the synaptic fusion machinery components [32, 44]. Membrane cholesterol further affects SNARE-mediated membrane fusion by reducing exocytosis in the presence of a lower concentration of membrane cholesterol [45, 46]. Cholesterol is also known to modify the physical characteristics of the membrane and their interaction with fusion proteins (SNAREs) that control the development of cholesterol-dependent domains (raft-like domains) at the fusion sites [47, 48]. Not only in SNARE-mediated fusion, membrane cholesterol also regulates viral infections by impacting the fusion between virus and host cell membranes. It has been demonstrated that higher cholesterol content facilitates entry of influenza by reducing the energy required to generate the hemifusion stalk [49, 50]. Increasing the amount of cholesterol in the membrane increases the likelihood of complete fusion events, for example, virus-like particles with low cholesterol levels exhibit limited cell entry [51]. Cholesterol is further implicated in inducing oligomerization of fusion peptides to assume the fusogenic conformation for the N-terminal fusion peptide of SARS-CoV [52, 53]. Lower membrane cholesterol reduces the pathological syncytia formation for SARS-CoV-2 [54]. Interestingly, an internal fusion peptide of S-glycoprotein of SARS-CoV-2 promotes greater hemifusion formation in cholesterol-rich membranes [55].
Thus, lipid morphologies are significant, because fusion is typically aided by lipids that generate intrinsic negative curvature in the membrane. The fusion proteins often have specific interactions with the lipids that aid to its fusion-triggering abilities, therefore specific lipid composition plays an important role in membrane fusion. Taken together, the role of lipid composition in membrane fusion requires detailed exploration.
4 Lipidic domain in membrane fusion
Lipidic domains were considered the detrimental patches for membrane fusion because of their ordered and rigid organization and relatively slower lipid dynamics. However, recent reports highlight the importance of lipidic domains in the entry of enveloped viruses. HIV fusion peptide, which induces fusion between two sets of membranes, binds at the boundaries of liquid-ordered (Lo) and liquid-disordered (Ld) domains and reduces the line tension [45, 56]. The released energy from the reduction of line tension assists in crossing the activation barrier for the fusion reaction. Results suggest that the fusion peptide promotes more fusion in a membrane where both Lo and Ld phases coexist (heterogeneous) rather than simple Lo or Ld membranes (homogeneous) [11]. Another study showed that HIV-1 particle production is supported by a viral protein (Gag), which binds at the cholesterol-rich domains of the plasma membrane, whereas, depletion of cholesterol leads to a reduction in HIV-1 particle production. Therefore, it is conjectured that the binding of Gag at the cholesterol-rich domain is associated with the production of HIV-1 particles and hence higher infectivity [57]. Moreover, the T cell activation is linked to the rearrangement of Lo and Ld phases, which makes T cells susceptible to infection [58,59,60]. Therefore, modification of membrane organization could be exploited to inhibit the entry of enveloped viruses. It is known that polyunsaturated fatty acids (PUFAs) modify the membrane organization and dynamics by redistributing cholesterol from Lo to Ld phases, [61, 62] thereby, modulating the lipid–protein interaction. Eicosapentaenoic acid (EPA) and docosahexaenoic acid (DHA) are the two most extensively studied PUFAs. It was observed from the DPH anisotropy data that DHA has a higher membrane fluidizing ability than EPA, and further addition of cholesterol does not alter the membrane fluidity in the presence of DHA [63,64,65]. DHA was found to improve the fluidity of the acyl chain area in heterogeneous model membranes (POPC/Cholesterol, 50 mol%/50 mol%), however, there are no reports of EPA exhibiting any such impact [62]. Joardar et al. demonstrated that the DHA and EPA have relatively lower fusion inhibitory ability in Lo–Ld coexisting membranes (DOPC/SM/CH, 40/40/20 mol%) than in Ld membranes [66]. Membrane order even generates differential mechanical properties and is responsible for the permeability of small molecules [67, 68].
5 Effect of fusion peptide on membrane organization and dynamics
It is well known that fusion peptides are essential for membrane fusion, however, it is still unclear how the fusion peptide induces membrane fusion [42, 69,70,71,72]. Measurement of membrane organization and dynamics could be a better way to understand the role of fusion peptides in membrane fusion. Surprisingly, other than the conserved location of glycine and alanine at the N-terminus, there is no structural and sequence homology among the fusion peptides of different viral fusion proteins. In addition, the fusion peptide exhibits structural variation in response to lipid composition [18]. The process gets further complicated by the impact of lipid composition on the effectiveness of fusion peptide [18]. In terms of polarity, stiffness (order), segmental motion, and capacity to establish hydrogen bonds, biological membranes are innately anisotropic along the bilayer normal [73,74,75]. A bilayer is formed by two layers of lipid molecules with their phosphate groups oriented away from each other, with one layer (leaflet) mirroring the other leading to similar physical properties of two leaflets. As a result, we detect a substantial fluctuation in stiffness, polarity (dielectric constant varies from about 70 to 5), and segmental motion in only 20 Å length when going from the membrane surface to the center of the bilayer, which is unrivaled by any synthetic nanomaterial. A gradient of solvent mobility exists along the bilayer normal, having a significant influence on the lipid–protein interaction [75, 76]. Besides stiffness and polarity, environmental heterogeneity varies along the bilayer normally in a depth-dependent manner (Fig. 4) [77]. Fluorescence spectroscopy is a powerful tool for studying the fluidity, polarity, and heterogeneity of lipid bilayers. The fluorescence anisotropy of a probe reflects the rotational constraint in its near surroundings. A series of depth-dependent probes have been generated by adding anthroyloxy at various carbons of stearic acid. The n-(9-anthroyloxy) stearic acid (n-AS) in tandem with 1, 6-diphyenyl-1,3,5-hexatriene (DPH) and its trimethylammonium derivative (TMA-DPH) have been utilized effectively to determine the graded rigidity, polarity, and heterogeneity of the membrane [42, 70, 77,78,79,80,81,82]. Because of the hydrogen bonding and electrostatic interactions between the lipid headgroup and surface waters, the head-group area of the membrane is highly organized and descends exponentially along the bilayer normal. The depth-dependent polarity of the membrane is provided by the fluorescence lifetime observations of these probes. The lifetime distribution analysis offers a unique method to determine the environmental heterogeneity from its width of distribution [77].
Diagrammatic representation of the gradient of the membrane’s stiffness, polarity, and heterogeneity along the bilayer normal. The figure has been adapted from Ref. [76] with permission
Fusion peptides are known to reduce the bending energy of the bilayer, filling void space at intermediate structures, and changing the membrane curvature to trigger the peptide-induced membrane fusion [72]. Furthermore, fusion peptide enhances membrane fusion by altering the depth-dependent membrane ordering. The wild-type fusion peptide from influenza HA fosters rigidity at the head-group and interfacial region, but has little effect on the acyl chain order close to the center of the dimyristoylphosphatidylcholine (DMPC) and 1-palmitoyl-2-oleoylphosphatidylcholine (POPC) bilayers, according to the study conducted by Freed and coworkers [83]. The effect is more prominent at pH 5 compared to pH 7. Interestingly, the influenza virus fuses with the endosome at an acidic pH (~ pH 5). The impact of a fusion-inefficient G1 mutant on the depth-dependent ordering of the lipid bilayer has also been examined. Surprisingly, in contrast to the wild-type HA fusion peptide, the G1 mutant did not cause any modification in the membrane order at the head-group and interfacial region [83]. In a related investigation, it was shown that the wild-type gp41 fusion peptide from the HIV promotes order at the head-group and interfacial area but has no influence on the acyl chain packing further into the bilayer of POPC/POPG membranes [84]. Interestingly, the gp41 fusion peptide displays a better membrane penetration in the presence of 30 mol% of cholesterol. Moreover, the gp41 fusion peptide has been shown to undergo a conformational shift from helix to sheet as a function of increasing concentration of membrane cholesterol [85]. The cholesterol-dependent enhancement of the fusogenicity of gp41 could be a collective effect of deeper penetration and peptide conformation. However, it is significant to note that the circular dichroism measurements for secondary structure prediction were carried out at a lipid-to-peptide ratio of 100:1, whereas the fusion tests were conducted at a lipid-to-peptide ratio of 40:1. This knowledge is crucial since the conformational change of the gp41 fusion peptide is concentration dependent [86].
Membrane cholesterol imparts a unique effect on the depth-dependent organization, dynamics, and mechanical stability of the membrane [87]. Therefore, the secondary structure and fusogenic property of the gp41 fusion peptide are highly connected with the cholesterol-induced ordering of the membrane. It is interesting to note that the V2E mutant of gp41 barely affects the membrane order and leaves the peptide entirely ineffective for fusion [88, 89]. Furthermore, according to Dimitrov et al. the full-length gp160 with the V2E mutation did not exhibit membrane destabilization and fusion [90]. This supports the idea that peptide-induced bilayer organization is crucial for catalyzing membrane fusion. In a different investigation, the fluorescence anisotropies of DPH and TMA-DPH were examined to track the order of the membrane at the acyl chain and interfacial region, respectively. The results demonstrate that the interfacial region is disturbed by the gp41 fusion peptide at all concentrations. Moreover, the acyl chain order in POPC membranes is disrupted at lower peptide concentrations in the deeper area of the bilayer but is restored at higher peptide concentrations [52]. The findings from the fluorescence measurements contradict the observation from ESR measurements [84]. A detailed molecular dynamics simulation study might be important to bridge the gap between two widely used experimental methodologies. Structural flexibility of the fusion peptide [91] might be important in promoting fusion as the peptide is required to accommodate itself in the highly curved nonlamellar intermediates.
6 Water percolation at the acyl chain region in promoting fusion
Studies have shown that water forms hydrogen bonds with phospholipids and takes part in the formation of the lipid bilayer [92,93,94,95,96,97]. The “hydration layer” is a network of hydrogen-bonded water molecules that extends between the lipids and includes the water molecules in the head-group area [98, 99]. A “bound” hydration shell of 9–20 fluids per phospholipid gives rise to this layer of hydration. Water is present not only in the lipid head-group region but also between the fatty-acyl chains, as “interstitial hydration”.
Fluorescence lifetime can be used as an authentic marker to map the water penetration at the various depths of the membrane using depth-dependent fluorophores. Measurement of fluorescence lifetime is simple yet robust as it averages many experiments. The effect of cholesterol and phosphatidylcholine on water penetration into the hydrophobic region of the membrane bilayer was carried out by using DPH and TMA-DPH [100, 101]. The changing concentration of interstitial or interchain water is mostly to blame for the decrease in fluorescence lifetime of DPH with greater unsaturation and in the presence of cholesterol [102, 103]. The Stubbs group demonstrated the variation of DPH-PC lifetime in various phospholipid bilayers [75]. In the case of DPH-PC, the DPH is tied to the sn-2 position of the PC so that its specific location can be determined. The lifetime reduces with higher unsaturation levels or protein addition. Contrarily, cholesterol lengthens the lifetime of DPH-PC, consistent with its known ability to dehydrate the bilayer acyl chains. For example, in dansyl-PE, where the probe is at the head-group region, the effect of increasing unsaturation again decreases the fluorescence lifetime. Properties such as polarity, fluidity, segmental motion, ability to form hydrogen bonds, and extent of water penetration vary in a depth-dependent manner in the membrane. Chakraborty et al. have utilized red edge excitation shift (REES) to monitor water penetration and its dynamics at different depths in 1,2-diheptanoyl-sn-glycero-3-phosphocholine (DHPC) and polar lipid fraction E (PLFE) from archaeal membranes [80]. From REES data it was concluded that the anthroyloxy moiety in 2-AS is localized in a motionally restricted region of the DHPC membrane that offers considerable resistance to solvent reorientation in the excited state while the motional restriction is completely absent in the location of deeper n-AS probes (or lack of polar solvent in the deeper regions of the membrane). This indicates differential extents of motional restriction along the bilayer normal in DHPC membranes. From the intensity-averaged fluorescence lifetime observations of n-AS probes in PLFE and DHPC membranes, it was observed that each n-AS probe displays a higher lifetime in PLFE membranes indicating that the PLFE membranes are less polar than DHPC membranes.
Water penetration at the acyl chain region is often correlated with the fusogenicity of the membrane. The presence of water at the acyl chain region increases the energy of the bilayer that partially compensates the activation energy for the intermediate and pore formation, thereby, facilitating the fusion process [42]. Our group has demonstrated that fusion-inducing peptides enhance water penetration whereas fusion-inhibiting peptides block water penetration at the acyl chain region [43, 104, 105]. The presence of unsaturation in the added fatty acids promotes water penetration at the acyl chain region of DOPC/DOPE (70/30 mol%) membranes resulting in a decrease in DPH fluorescence lifetime [39]. Moreover, the fluorescence lifetime of DPH (an indicator of water penetration at the acyl chain region) is directly correlated to the unsaturation present in the fatty acids (Fig. 5).
Plot correlating fluorescence lifetime of DPH with the rate of pore formation (k3) in the presence of 5 mol% OA, LA, and ALA in DOPC/DOPE (70/30 mol%) membranes at fixed lipid concentration of 200 µM. All experiments were performed in a pH 7.4 buffer containing 10 mM TES, 100 mM NaCl, 1 mM CaCl2, and 1 mM EDTA. The figure has been adapted from Ref. [39] with permission
The results with unsaturated fatty acids demonstrate the significance of water percolation at the acyl chain region of the membrane in predicting the fusogenic ability of added perturbant. Experiments with fusion inhibitory peptides vouch for the same observation with the unsaturated fatty acids. A coronin 1-derived peptide, TG-23, inhibits the fusion of DOPC/DOPE/DOPG membranes by around 40% significantly blocking water percolation at the acyl chain region of the membrane [43].
7 Membrane voids in fusion
The stalk hypothesis proposed the formation of non-bilayer intermediates during the fusion pore opening. These intermediates are characterized by the formation of symmetric and interstice voids (evident in Fig. 1). Generated void spaces between the bilayer leaflets in the hemifusion intermediate make the intermediate formation energetically unfavorable [106]. The significance of void spaces in membrane fusion has been justified by the fusion experiments carried out in the presence of hexadecane and tetradecane. Being long-chain alkane, hexadecane and tetradecane can fill void space formed at the non-bilayer intermediate states, and reduce the overall energy of the intermediates, thereby, promoting vesicular fusion by lowering the energy of the hydrophobic interstices [42, 107,108,109,110,111]. Many fusion peptides are known to promote the hemifusion intermediate by filling the void space at the intermediate states [42, 81]. The partitioning of C6-NBD-PC in the membrane is utilized to measure the void space in the bilayer. The externally added C6-NBD-PC in the membrane demonstrates two different fluorescence lifetime values corresponding to the aqueous and membrane environments [81]. Therefore, by measuring the fluorescence lifetime of C6-NBD-PC, one can estimate the available void space in the membrane. The same experiment is used to determine the void-filling ability of peptides and other small molecules. The gp41 and HA fusion peptides reduce membrane partitioning of C6-NBD-PC, indicating their void-filling ability, and promote lipid mixing in polyethylene glycol-induced small unilamellar vesicles fusion assays [81]. Figure 6 demonstrates a schematic illustration of void space and how fusion peptide can fill that void space.
Schematic representation of A void at the fusion intermediate and B being filled by a peptide and partitioning into the membrane thus stabilizing the fusion intermediate. The figure has been adapted from Ref. [72] with permission
The TMD of vesicular stomatitis virus (VSV) also promotes fusion by occupying the void space of the hemifusion intermediate to facilitate fusion [112]. The fact that the TMD of VSV competes with hexadecane to promote the rate of intermediate formation further supports this assertion. It is important to optimize the stabilization of the intermediate structure such that it promotes the formation of hemifusion intermediates without impeding the later steps of the process. Chemistry of the lipidic tail further contributes to the void volume (or defects) in the membrane. Lipids with identical fatty acyl tails generate mismatch in the tail length and it has been observed that dioleoyl membranes are more fusogenic than the palmitoyl-oleoyl lipidic membranes. Similarly, membranes with unsaturated lipids are more fusogenic than their saturated counterpart. The saturated lipids form a rigid bilayer due to its compact packing whereas unsaturated lipids reduce rigidity because of kink formation at the unsaturated site [30].
8 Lipid dynamics and fusion
Though the overall physical properties of the membrane have been correlated to fusion in various aspects, there is not much information on the impact of lipid dynamics on membrane fusion. In membrane fusion two distinct bilayers convert into a single bilayer through the reorganization of lipids and it depends on the dynamical nature of the lipids in the leaflets. It has been shown that lipids in Lo phase becomes more dynamic in presence of gp41 fusion peptide resulting in increase in diffusion coefficient value, and eventually the lipidic nanodomain merge with each other to form a larger domain [113]. This microscopic change in lipid dynamics has further been correlated to the macroscopic fusogenic properties of the same peptide. This result implicates the significance of lipid dynamics in membrane fusion. The wide variety of lipid molecules and their unique mixing characteristics enable their complex behavior [114,115,116]. It is well recognized that the resulting structural characteristics of the membranes, such as bilayer thickness and lipid packing, have an impact on membrane permeability, elasticity, and protein-membrane interactions [113,114,115,116,117]. Lipid molecules also experience continuous thermal motions in the form of segmental dynamics and orientational changes. Solid-state NMR is one of the few experimental methods that can identify and measure these thermal movements at the molecular level [118,119,120,121,122,123]. The carbon–hydrogen (CH or deuterium) bonds are sensitive to alterations in the lipid acyl chains. The combination of NMR line shape and relaxation time measurements allows us to determine the bilayer fluctuations. The NMR line shape quantifies the order parameters of the carbons throughout the lipid chains and displays the levels of molecular motion [124,125,126]. At the same time, data on nuclear spin relaxation rates for each carbon reveal correlations in CH bond fluctuations over various time scales [127]. To have detailed information on the membrane order and lipid dynamics, the molecular dynamics simulation results should be correlated with the experimental results. In doing so, Doktorova et al. developed a methodology to relate membrane elastic properties and collected molecular dynamics by determining mean-square amplitude (representing order parameter) and relaxation rate (representing correlation time). Therefore, all-atom molecular dynamics simulations of the lipid bilayer can be directly compared to the CH bond relaxation results obtained from NMR spectroscopy [128]. Results highlight the critical role of interleaflet coupling in the mechanical properties of the membrane.
The correlated lipid fluctuation in two opposing inner leaflets is considered to be essential for the pore to open from the extended TM contact intermediate [129]. However, it is experimentally challenging to monitor the fluctuation at the intermediate state because of its transient nature. With the technological development of single molecular fluorescence and super-resolution microscopy efforts may be given to decipher the complex nature of lipid dynamics and membrane fusion. Furthermore, the measurement of lipid dynamics is important to understand membrane mechanical properties at the molecular level, and correlate membrane mechanical properties with the abilities of membrane fusion.
9 Observables in membrane fusion
The mechanism of the membrane fusion can be studied by measuring three observables such as lipid mixing, content mixing and content leakage. The lipid mixing and content mixing describe the mixing of leaflet lipids and inner contents between two apposed bilayers, respectively. Instead, content leakage provides us information regarding membrane integrity during the fusion. Generally, fluorescence resonance energy transfer (FRET) is used to monitor the kinetics of these three observables [133]. Lentz group has developed an analytical model of membrane fusion based on the lipid stalk hypothesis where they have utilized kinetic data of lipid mixing, content mixing and content leakage to decipher the rate constant of different steps. The model assumes that these observables have finite probability at each state of the fusion intermediates [133, 134]. As the stalk hypothesis assumes the mixing of inner leaflets (which contains about 60% lipids in a small unilamellar vesicle) to form the stalk and minimum probability of content mixing at the stalk state. As the outer leaflet comprises around 60% lipids in small unilamellar vesicles, the probability of lipid mixing at the stalk state is generally expected to be close to 0.6 [40]. Therefore, probability of lipid mixing at the stalk intermediate should always be higher than the probability of content mixing if the fusion follows classical stalk model. On contrary, if the probability of lipid mixing is substantially lower than 0.6 with a concomitant increase in probability of content mixing at stalk state the fusion mechanism is considered to proceed via non-classical stalk model [40]. Therefore, analyzing the kinetic data of fusion observables in an analytical model provides in-depth information on fusion mechanism.
10 Conclusion
Membrane fusion is an essential biological process in the existence of eukaryotic cells, which occurs through the formation of multiple non-bilayer hemifusion intermediates. There is a long-standing conflict on the significance of proteins and lipids in modulating membrane fusion. It is noteworthy to mention that both proteins and lipids play their parts in a successful fusion event. Fusion protein not only supplies activation energy for the fusion process through conformational changes but also alters the lipid organization and dynamics of the membrane. The extent and location (depth-dependency) of organizational and dynamical deformity in the bilayer affect the fusion process. Therefore, proteins and lipids work in complete coherence for a successful and efficient fusion pore opening. Our review discusses several aspects of the fusion proteins, peptides, and lipids in the fusion process keeping the lipid stalk model as the frame of reference. It is obvious that both the shape and chain unsaturation of the lipids have a major impact on the fusion process. Fluorescence lifetime measurements of DPH demonstrate that the water percolation at the acyl chain region of the membrane is correlated to the degree of unsaturation of the added fatty acids. This emphasizes the importance of looking at the fusion events from the extent of water percolation at the acyl chain region. Experimental results with fusion peptides and TMDs of different viral fusion peptides elucidate their ability to induce fusion by filling up the void space generated at the non-bilayer intermediates. Finally, we aimed to correlate membrane dynamics with its mechanical properties and fusion. Overall, our review presents a concise yet critical discussion of the different physical and biological aspects that govern membrane fusion.
Data availability
No data associated in the manuscript.
References
L.V. Chernomordik, M.M. Kozlov, Mechanics of membrane fusion. Nat. Struc. Mol. Biol. 15, 675–683 (2008)
L.V. Chernomordik, M.M. Kozlov, Protein-lipid interplay in fusion and fission of biological membranes. Annu. Rev. Biochem. 72, 175–207 (2003)
L.V. Chernomordik, G.B. Melikyan, Y.A. Chizmadzhev, Biomembrane fusion: a new concept derived from model studies using two interacting planar lipid bilayers. Biochim. Biophys. Acta 906, 309–352 (1987)
A. Chanturiya, L.V. Chernomordik, J. Zimmerberg, Flickering fusion pores comparable with initial exocytotic pores occur in protein-free phospholipid bilayers. Proc. Natl. Acad. Sci. U.S.A. 94, 14423–14428 (1997)
A. Sapir, O. Avinoam, B. Podbilewicz, L.V. Chernomordik, Viral and developmental cell fusion mechanisms: conservation and divergence. Dev. Cell 14, 11–21 (2008)
F.S. Cohen, G.B. Melikyan, The Energetics of Membrane Fusion from Binding, through Hemifusion, Pore Formation, and Pore Enlargement. J. Membr. Biol. 199, 1–14 (2004)
B.R. Lentz, V. Malinin, M.E. Haque, K. Evans, Protein machines and lipid assemblies: current views of cell membrane fusion. Curr. Opin. Struct. Biol. 10, 607–615 (2000)
C.M. Carr, P.S. Kim, A spring-loaded mechanism for the conformational change of influenza hemagglutinin. Cell 73, 823–832 (1993)
J.J. Skehel, D.C. Wiley, Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin. Annu. Rev. Biochem. 69, 531–569 (2000)
T.C. Südhof, J.E. Rothman, Membrane fusion: grappling with SNARE and SM proteins. N.Y. Sci. J. 323, 474–477 (2009)
E. London, Membrane fusion: A new role for lipid domains? Nat. Chem. Biol. 11, 383–384 (2015)
R.F. Epand, J.C. Macosko, C.J. Russell, Y.K. Shin, R.M. Epand, The ectodomain of HA2 of influenza virus promotes rapid pH dependent membrane fusion. Mol. Biol. 286, 489–503 (1999)
D.L. LeDuc, Y.K. Shin, R.F. Epand, R.M. Epand, Factors determining vesicular lipid mixing induced by shortened constructs of influenza hemagglutinin. Biochemistry 39, 2733–2739 (2000)
G.W. Kemble, T. Danieli, J.M. White, Lipid-anchored influenza hemagglutinin promotes hemifusion, not complete fusion. Cell 76, 383–391 (1994)
S.A. Tatulian, L.K. Tamm, Secondary structure, orientation, oligomerization, and lipid interactions of the transmembrane domain of influenza hemagglutinin. Biochemistry 39, 496–507 (2000)
B. Schroth-Diez, E. Ponimaskin, H. Reverey, M.F. Schmidt, A. Herrmann, Fusion activity of transmembrane and cytoplasmic domain chimeras of the influenza virus glycoprotein hemagglutinin. Virol. J. 72, 133–214 (1998)
D.Z. Cleverley, J. Lenard, The transmembrane domain in viral fusion: essential role for a conserved glycine residue in vesicular stomatitis virus G protein. Proc. Natl. Acad. Sci. U.S.A. 95, 3425–3430 (1998)
G. Meher, H. Chakraborty, Membrane composition modulates fusion by altering membrane properties and fusion peptide structure. J. Membr. Biol. 252, 261–272 (2019)
A. Sardar, N. Dewangan, B. Panda, D. Bhowmick, P.K. Tarafdar, Lipid and lipidation in membrane fusion. J. Membr. Biol. 255, 691–703 (2022)
J.B. Bock, R.H. Scheller, SNARE proteins mediate lipid bilayer fusion. Proc. Natl. Acad. Sci. U.S.A. 96, 12227–12229 (1999)
R.B. Sutton, J.A. Ernst, A.T. Brunger, Crystal structure of the cytosolic C2A–C2B domains of synaptotagmin III. Implications for Ca(+2)-independent snare complex interaction. J. Cell Biol. 147, 589–598 (1999)
Y. Moon, Y. Jun, The effects of regulatory lipids on intracellular membrane fusion mediated by dynamin-like GTPases. Front. Cell Dev. Biol. 8, 518 (2020)
S.A. Akimov, R.J. Molotkovsky, P.I. Kuzmin, T.R. Galimzyanov, O.V. Batishchev, Continuum models of membrane fusion: evolution of the theory. Int. J. Mol. Sci. 21, 3875 (2020)
K. Lähdesmäki, O.H.S. Ollila, A. Koivuniemi, P.T. Kovanen, M.T. Hyvönen, Membrane simulations mimicking acidic pH reveal increased thickness and negative curvature in a bilayer consisting of lysophosphatidylcholines and free fatty acids. Biochim. Biophys. Acta - Biomembr. 1798, 938–946 (2010)
H. Chakraborty, T. Sengupta, B.R. Lentz, pH Alters PEG-mediated fusion of phosphatidylethanolamine-containing vesicles. Biophys. J. 107, 1327–1338 (2014)
A. Efrat, L.V. Chernomordik, M.M. Kozlov, Point-like protrusion as a prestalk intermediate in membrane fusion pathway. Biophys. J. 92, L61–L63 (2007)
R.P. Rand, V.A. Parsegian, Hydration forces between phospholipid bilayers. Biochim. Biophys. Acta - Biomembr. 988, 351–376 (1989)
N. Fuller, R.P. Rand, The influence of lysolipids on the spontaneous curvature and bending elasticity of phospholipid membranes. Biophys. J. 81, 243–254 (2001)
H.T. McMahon, E. Boucrot, Membrane curvature at a glance. J. Cell Sci. 128, 1065–1070 (2015)
A. Joardar, G.P. Pattnaik, H. Chakraborty, Mechanism of membrane fusion: interplay of lipid and peptide. J. Membr. Biol. 255, 211–224 (2022)
K. Lähdesmäki, O.H. Ollila, A. Koivuniemi, P.T. Kovanen, M.T. Hyvönen, Membrane simulations mimicking acidic pH reveal increased thickness and negative curvature in a bilayer consisting of lysophosphatidylcholines and free fatty acids. Biochim. Biophys. Acta 1798, 938–946 (2010)
G. van den Bogaart, K. Meyenberg, H.J. Risselada, H. Amin, K.I. Willig, B.E. Hubrich, M. Dier, S.W. Hell, H. Grubmüller, U. Diederichsen, R. Jahn, Membrane protein sequestering by ionic protein-lipid interactions. Nature 479, 552–555 (2011)
P.K. Tarafdar, H. Chakraborty, M.J. Bruno, B.R. Lentz, Phosphatidylserine-dependent catalysis of stalk and pore formation by synaptobrevin JMR-TMD peptide. Biophys. J.. J. 109, 1863–1872 (2015)
P.K. Tarafdar, H. Chakraborty, S.M. Dennison, B.R. Lentz, Phosphatidylserine inhibits and calcium promotes model membrane fusion. Biophys. J. 103, 1880–1889 (2012)
H.I. Ingólfsson, M.N. Melo, F.J. van Eerden, C. Arnarez, C.A. Lopez, T.A. Wassenaar, X. Periole, A.H. de Vries, D.P. Tieleman, S.J. Marrink, Lipid organization of the plasma membrane. J. Am. Chem. Soc. 136, 14554–14559 (2014)
W. Wang, L. Yang, H.W. Huang, Evidence of cholesterol accumulated in high curvature regions: implication to the curvature elastic energy for lipid mixtures. Biophys. J. 92, 2819–2830 (2007)
M.E. Haque, T.J. McIntosh, B.R. Lentz, Influence of lipid composition on physical properties and peg-mediated fusion of curved and uncurved model membrane vesicles: “nature’s own” fusogenic lipid bilayer. Biochem. 40, 4340–4348 (2001)
M. Pinot, S. Vanni, S. Pagnotta, S. Lacas-Gervais, L.A. Payet, T. Ferreira, R. Gautier, B. Goud, B. Antonny, H. Barelli, Lipid cell biology. Polyunsaturated phospholipids facilitate membrane deformation and fission by endocytic proteins. N. Y. Sci. J. 345, 693–697 (2014)
A. Joardar, S. Pandia, H. Chakraborty, Effect of polyunsaturated free fatty acids on the membrane fusion mechanism. Soft Matter 19, 733–742 (2023)
A. Joardar, G.P. Pattnaik, H. Chakraborty, Effect of phosphatidylethanolamine and oleic acid on membrane fusion: phosphatidylethanolamine circumvents the classical stalk model. J. Phys. Chem. B 125, 13192–13202 (2021)
P.M. Kasson, V.S. Pande, Control of membrane fusion mechanism by lipid composition: predictions from ensemble molecular dynamics. PLoS Comput. Biol. 3, e220 (2007)
H. Chakraborty, P.K. Tarafdar, D.G. Klapper, B.R. Lentz, Wild-type and mutant hemagglutinin fusion peptides alter bilayer structure as well as kinetics and activation thermodynamics of stalk and pore formation differently: mechanistic implications. Biophys. J. 105, 2495–2506 (2013)
G.P. Pattnaik, H. Chakraborty, Coronin 1 derived tryptophan-aspartic acid containing peptides inhibit membrane fusion. Chem. Phys. Lipids 217, 35–42 (2018)
Á. Pérez-Lara, A. Thapa, S.B. Nyenhuis, D.A. Nyenhuis, P. Halder, M. Tietzel, K. Tittmann, D.S. Cafiso, R. Jahn, PtdInsP(2) and PtdSer cooperate to trap synaptotagmin-1 to the plasma membrane in the presence of calcium, Elife 5, (2016).
S.T. Yang, V. Kiessling, L.K. Tamm, Line tension at lipid phase boundaries as driving force for HIV fusion peptide-mediated fusion. Nat. Commun. 7, 11401 (2016)
J. Han, K. Pluhackova, R.A. Böckmann, The Multifaceted Role of SNARE Proteins in Membrane Fusion. Front. physiol. 8, 5 (2017)
D.H. Murray, L.K. Tamm, Clustering of syntaxin-1A in model membranes is modulated by phosphatidylinositol 4,5-bisphosphate and cholesterol. Biochem. 48, 4617–4625 (2009)
D.H. Murray, L.K. Tamm, Molecular mechanism of cholesterol- and polyphosphoinositide-mediated syntaxin clustering. Biochem. 50, 9014–9022 (2011)
P. Chlanda, E. Mekhedov, H. Waters, C.L. Schwartz, E.R. Fischer, R.J. Ryham, F.S. Cohen, P.S. Blank, J. Zimmerberg, The hemifusion structure induced by influenza virus haemagglutinin is determined by physical properties of the target membranes. Nat. Microbiol. 1, 16050 (2016)
K.N. Liu, S.G. Boxer, Target Membrane Cholesterol Modulates Single Influenza Virus Membrane Fusion Efficiency but Not Rate. Biophys. J. 118, 2426–2433 (2020)
J. Lee, A.J.B. Kreutzberger, L. Odongo, E.A. Nelson, D.A. Nyenhuis, V. Kiessling, B. Liang, D.S. Cafiso, J.M. White, L.K. Tamm, Ebola virus glycoprotein interacts with cholesterol to enhance membrane fusion and cell entry. Nat. Struct. Mol. Biol. 28, 181–189 (2021)
G. Meher, S. Bhattacharjya, H. Chakraborty, Membrane Cholesterol Modulates Oligomeric Status and Peptide-Membrane Interaction of Severe Acute Respiratory Syndrome Coronavirus Fusion Peptide. J. Phys. Chem. B 123, 10654–10662 (2019)
G. Meher, S. Bhattacharjya, H. Chakraborty, Membrane cholesterol regulates the oligomerization and fusogenicity of SARS-CoV fusion peptide: implications in viral entry. Phys. Chem. Chem. Phys. 25, 7815–7824 (2023)
D.W. Sanders, C.C. Jumper, P.J. Ackerman, D. Bracha, A. Donlic, H. Kim, D. Kenney, I. Castello-Serrano, S. Suzuki, T. Tamura, A.H. Tavares, M. Saeed, A.S. Holehouse, A. Ploss, I. Levental, F. Douam, R.F. Padera, B.D. Levy, C.P. Brangwynne, SARS-CoV-2 requires cholesterol for viral entry and pathological syncytia formation. Elife (2021). https://doi.org/10.7554/eLife.65962
G.P. Pattnaik, S. Bhattacharjya, H. Chakraborty, Enhanced cholesterol-dependent hemifusion by internal fusion peptide 1 of SARS coronavirus-2 compared to its N-Terminal counterpart. Biochem. 60, 559–562 (2021)
S.T. Yang, V. Kiessling, J.A. Simmons, J.M. White, L.K. Tamm, HIV gp41-mediated membrane fusion occurs at edges of cholesterol-rich lipid domains. Nat. Chem. Biol. 11, 424–431 (2015)
A. Ono, E.O. Freed, Plasma membrane rafts play a critical role in HIV-1 assembly and release. Proc. Natl. Acad. Sci. U.S.A. 98, 13925–13930 (2001)
S. Mahammad, J. Dinic, J. Adler, I. Parmryd, Limited cholesterol depletion causes aggregation of plasma membrane lipid rafts inducing T cell activation. Biochim. Biophys. Acta 180, 625–634 (2010)
T. Zech, C.S. Ejsing, K. Gaus, B. de Wet, A. Shevchenko, K. Simons, T. Harder, Accumulation of raft lipids in T-cell plasma membrane domains engaged in TCR signalling. EMBO J. 28, 466–476 (2009)
T. Harder, C. Rentero, T. Zech, K. Gaus, Plasma membrane segregation during T cell activation: probing the order of domains. Curr. Opin. Immunol. 19, 470–475 (2007)
M.L. Jacobs, H.A. Faizi, J.A. Peruzzi, P.M. Vlahovska, N.P. Kamat, EPA and DHA differentially modulate membrane elasticity in the presence of cholesterol. Biophys. J. 120, 2317–2329 (2021)
R.P. Mason, R.F. Jacob, S. Shrivastava, S.C.R. Sherratt, A. Chattopadhyay, Eicosapentaenoic acid reduces membrane fluidity, inhibits cholesterol domain formation, and normalizes bilayer width in atherosclerotic-like model membranes. Biochim. Biophys. Acta 1858, 3131–3140 (2016)
M. Hashimoto, S. Hossain, H. Yamasaki, K. Yazawa, S. Masumura, Effects of eicosapentaenoic acid and docosahexaenoic acid on plasma membrane fluidity of aortic endothelial cells. Lipids 34, 1297–1304 (1999)
M. Hashimoto, M. Katakura, S. Hossain, A. Rahman, T. Shimada, O. Shido, Docosahexaenoic acid withstands the Aβ(25–35)-induced neurotoxicity in SH-SY5Y cells. J. Nutr. Biochem. 22, 22–29 (2011)
D.C. Mitchell, B.J. Litman, Effect of cholesterol on molecular order and dynamics in highly polyunsaturated phospholipid bilayers. Biophys. J. 75, 896–908 (1998)
A. Joardar, H. Chakraborty, Differential behavior of eicosapentaenoic and docosahexaenoic acids on the organization, dynamics, and fusion of homogeneous and heterogeneous membranes. Langmuir 39, 4439–4449 (2023)
C. Hannesschlaeger, A. Horner, P. Pohl, Intrinsic membrane permeability to small molecules. Chem. Rev. 119, 5922–5953 (2019)
M.B. Sankaram, T.E. Thompson, Modulation of phospholipid acyl chain order by cholesterol. A solid-state 2H nuclear magnetic resonance study. Biochemistry 29, 10676–10684 (1990)
R.F. Epand, I. Martin, J.M. Ruysschaert, R.M. Epand, Membrane orientation of the SIV fusion peptide determines its effect on bilayer stability and ability to promote membrane fusion. Biochem. Biophys. Res. Commun. 205, 1938–1943 (1994)
M.E. Haque, H. Chakraborty, T. Koklic, H. Komatsu, P.H. Axelsen, B.R. Lentz, Hemagglutinin fusion peptide mutants in model membranes: structural properties, membrane physical properties, and PEG-mediated fusion. Biophys. J. 101, 1095–1104 (2011)
Y. Kozlovsky, M.M. Kozlov, Stalk model of membrane fusion: solution of energy crisis. Biophys. J. 82, 882–895 (2002)
G.P. Pattnaik, G. Meher, H. Chakraborty, Exploring the mechanism of viral peptide-induced membrane fusion. Adv. Exp. Med. Biol. 1112, 69–78 (2018)
A. Chattopadhyay, Exploring membrane organization and dynamics by the wavelength-selective fluorescence approach. Chem. Phys. Lipids 122, 3–17 (2003)
S. Haldar, A. Chaudhuri, A. Chattopadhyay, Organization and dynamics of membrane probes and proteins utilizing the red edge excitation shift. J. Phys. Chem. B 115, 5693–5706 (2011)
C.D. Stubbs, C. Ho, S.J. Slater, Fluorescence techniques for probing water penetration into lipid bilayers. J. Fluoresc. 5, 19–28 (1995)
G. Meher, H. Chakraborty, The role of fusion peptides in depth-dependent membrane organization and dynamics in promoting membrane fusion. Chem. Phys. Lipids 234, 105025 (2021)
S. Haldar, M. Kombrabail, G. Krishnamoorthy, A. Chattopadhyay, Depth-dependent heterogeneity in membranes by fluorescence lifetime distribution analysis. J. Phys. Chem. Lett. 3, 2676–2681 (2012)
F.S. Abrams, A. Chattopadhyay, E. London, Determination of the location of fluorescent probes attached to fatty acids using parallax analysis of fluorescence quenching: effect of carboxyl ionization state and environment on depth. Biochem. 31, 5322–5327 (1992)
F.S. Abrams, E. London, Extension of the parallax analysis of membrane penetration depth to the polar region of model membranes: use of fluorescence quenching by a spin-label attached to the phospholipid polar headgroup. Biochem. 32, 10826–10831 (1993)
H. Chakraborty, S. Haldar, P.L. Chong, M. Kombrabail, G. Krishnamoorthy, A. Chattopadhyay, Depth-dependent organization and dynamics of archaeal and eukaryotic membranes: development of membrane anisotropy gradient with natural evolution. Langmuir 31, 11591–11597 (2015)
M.E. Haque, V. Koppaka, P.H. Axelsen, B.R. Lentz, Properties and structures of the influenza and HIV fusion peptides on lipid membranes: implications for a role in fusion. Biophys. J. 89, 3183–3194 (2005)
G. Meher, S. Sinha, G.P. Pattnaik, S. Ghosh Dastidar, H. Chakraborty, Cholesterol modulates membrane properties and the interaction of gp41 fusion peptide to promote membrane fusion. J. Phys. Chem. B 123, 7113–7122 (2019)
M. Ge, J.H. Freed, Fusion peptide from influenza hemagglutinin increases membrane surface order: an electron-spin resonance study. Biophys. J. 96, 4925–4934 (2009)
A.L. Lai, J.H. Freed, HIV gp41 fusion peptide increases membrane ordering in a cholesterol-dependent fashion. Biophys. J. 106, 172–181 (2014)
A.L. Lai, A.E. Moorthy, Y. Li, L.K. Tamm, Fusion activity of HIV gp41 fusion domain is related to its secondary structure and depth of membrane insertion in a cholesterol-dependent fashion. J. Mol. Biol. 418, 3–15 (2012)
V. Volkov, M. Bonn, Structural properties of gp41 fusion peptide at a model membrane interface. J. Phys. Chem. B 117, 15527–15535 (2013)
R.M. Epand, Proteins and cholesterol-rich domains. Biochim. Biophys. Acta 1778, 1576–1582 (2008)
M.D. Delahunty, I. Rhee, E.O. Freed, J.S. Bonifacino, Mutational analysis of the fusion peptide of the human immunodeficiency virus type 1: identification of critical glycine residues. Virol. J. 218, 94–102 (1996)
C.M. Gabrys, W. Qiang, Y. Sun, L. Xie, S.D. Schmick, D.P. Weliky, Solid-state nuclear magnetic resonance measurements of HIV fusion peptide 13CO to lipid 31P proximities support similar partially inserted membrane locations of the α helical and β sheet peptide structures. J. Phys. Chem. A 117, 9848–9859 (2013)
A.S. Dimitrov, S.S. Rawat, S. Jiang, R. Blumenthal, Role of the fusion peptide and membrane-proximal domain in HIV-1 envelope glycoprotein-mediated membrane fusion. Biochem. 42, 14150–14158 (2003)
D.K. Chang, S.F. Cheng, V. Deo Trivedi, S.H. Yang, The amino-terminal region of the fusion peptide of influenza virus hemagglutinin HA2 inserts into sodium dodecyl sulfate micelle with residues 16–18 at the aqueous boundary at acidic pH. Oligomerization and the conformational flexibility. J. Biol. Chem. 275, 19150–19158 (2000)
E.G. Finer, A. Darke, Phospholipid hydration studied by deuteron magnetic resonace spectroscopy. Chem. Phys. Lipids 12, 1–16 (1974)
G.L. Jendrasiak, J.H. Hasty, The hydration of phospholipids. Biochim. Biophys. Acta 337, 79–91 (1974)
B.D. Ladbrooke, D. Chapman, Thermal analysis of lipids, proteins and biological membranes. A review and summary of some recent studies. Chem. Phys. Lipids 3, 304–356 (1969)
G.C. Newman, C. Huang, Structural studies on phophatidylcholine-cholesterol mixed vesicles. Biochemistry 14, 3363–3370 (1975)
D.M. Small, Phase equilibria and structure of dry and hydrated egg lecithin. J. Lipid Res. 8, 551–557 (1967)
I. Ueda, H.S. Tseng, Y. Kaminoh, S.M. Ma, H. Kamaya, S.H. Lin, Anesthetics release unfreezable and bound water in partially hydrated phospholipid lamellar systems and elevate phase transition temperature. Mol. Pharmacol. 29, 582–588 (1986)
M. Prats, J.F. Tocanne, J. Teissie, Lateral proton conduction at a lipid/water interface. Effect of lipid nature and ionic content of the aqueous phase. Eur. J. Biochem.Biochem. 162, 379–385 (1987)
J. Teissié, M. Prats, A. LeMassu, L.C. Stewart, M. Kates, Lateral proton conduction in monolayers of phospholipids from extreme halophiles. Biochemistry 29, 59–65 (1990)
M. Straume, B.J. Litman, Equilibrium and dynamic structure of large, unilamellar, unsaturated acyl chain phosphatidylcholine vesicles. Higher order analysis of 1,6-diphenyl-1,3,5-hexatriene and 1-[4-(trimethylammonio)phenyl]- 6-phenyl-1,3,5-hexatriene anisotropy decay. Biochemistry 26, 5113–5120 (1987)
M. Straume, B.J. Litman, Influence of cholesterol on equilibrium and dynamic bilayer structure of unsaturated acyl chain phosphatidylcholine vesicles as determined from higher order analysis of fluorescence anisotropy decay. Biochemistry 26, 5121–5126 (1987)
C. Ho, C.D. Stubbs, Hydration at the membrane protein-lipid interface. Biophys. J. 63, 897–902 (1992)
K. Kinosita, A. Ikegami, Reevaluation of the wobbling dynamics of diphenylhexatriene in phosphatidylcholine and cholesterol/phosphatidylcholine membranes. Biochim. Biophys. Acta 769, 523–527 (1984)
G.P. Pattnaik, H. Chakraborty, Fusogenic effect of cholesterol prevails over the inhibitory effect of a peptide-based membrane fusion inhibitor. Langmuir 37, 3477–3489 (2021)
G.P. Pattnaik, H. Chakraborty, Cholesterol alters the inhibitory efficiency of peptide-based membrane fusion inhibitor. Biochim. Biophys. Acta 1861, 183056 (2019)
D.P. Siegel, The modified stalk mechanism of lamellar/inverted phase transitions and its implications for membrane fusion. Biophys. J. 76, 291–313 (1999)
G. Basáñez, F.M. Goñi, A. Alonso, Effect of single chain lipids on phospholipase C-promoted vesicle fusion. A test for the stalk hypothesis of membrane fusion. Biochemistry 37, 3901–3908 (1998)
A. Chanturiya, E. Leikina, J. Zimmerberg, L.V. Chernomordik, Short-chain alcohols promote an early stage of membrane hemifusion. Biophys. J. 77, 2035–2045 (1999)
Z. Chen, R.P. Rand, Comparative study of the effects of several n-alkanes on phospholipid hexagonal phases. Biophys. J. 74, 944–952 (1998)
R.P. Rand, N.L. Fuller, S.M. Gruner, V.A. Parsegian, Membrane curvature, lipid segregation, and structural transitions for phospholipids under dual-solvent stress. Biochemistry 29, 76–87 (1990)
A. Walter, P.L. Yeagle, D.P. Siegel, Diacylglycerol and hexadecane increase divalent cation-induced lipid mixing rates between phosphatidylserine large unilamellar vesicles. Biophys. J. 66, 366–376 (1994)
T. Sengupta, H. Chakraborty, B.R. Lentz, The transmembrane domain peptide of vesicular stomatitis virus promotes both intermediate and pore formation during PEG-mediated vesicle fusion. Biophys. J. 107, 1318–1326 (2014)
A. Chaudhury, S. Swarnakar, G.P. Pattnaik, G. Varshney, H. Chakraborty, J. Basu, Peptide induced fusion of dynamic membrane nanodomains: Implications in viral entry. Langmuir 39, 17713–17722 (2023)
A.J. Sodt, M.L. Sandar, K. Gawrisch, R.W. Pastor, E. Lyman, The molecular structure of the liquid-ordered phase of lipid bilayers. J. Am. Chem. Soc. 136, 725–732 (2014)
J.H. Lorent, K.R. Levental, L. Ganesan, G. Rivera-Longsworth, E. Sezgin, M. Doktorova, E. Lyman, I. Levental, Plasma membranes are asymmetric in lipid unsaturation, packing and protein shape. Nat. Chem. Biol. 16, 644–652 (2020)
I. Levental, K.R. Levental, F.A. Heberle, Lipid rafts: controversies resolved, Mysteries Remain. Trends Cell Biol. 30, 341–353 (2020)
J.H. Lorent, B. Diaz-Rohrer, X. Lin, K. Spring, A.A. Gorfe, K.R. Levental, I. Levental, Structural determinants and functional consequences of protein affinity for membrane rafts. Nat. Commun. 8, 1219 (2017)
M. Doktorova, F.A. Heberle, R.L. Kingston, G. Khelashvili, M.A. Cuendet, Y. Wen, J. Katsaras, G.W. Feigenson, V.M. Vogt, R.A. Dick, Cholesterol promotes protein binding by affecting membrane electrostatics and solvation properties. Biophys. J.. J. 113, 2004–2015 (2017)
M. Doktorova, F.A. Heberle, D. Marquardt, R. Rusinova, R.L. Sanford, T.A. Peyear, J. Katsaras, G.W. Feigenson, H. Weinstein, O.S. Andersen, Gramicidin increases lipid flip-flop in symmetric and asymmetric lipid vesicles. Biophys. J.. J. 116, 860–873 (2019)
A. Ghysels, A. Krämer, R.M. Venable, W.E. Teague Jr., E. Lyman, K. Gawrisch, R.W. Pastor, Permeability of membranes in the liquid ordered and liquid disordered phases. Nat. Commun. 10, 5616 (2019)
E.G. Kelley, P.D. Butler, R. Ashkar, R. Bradbury, M. Nagao, Scaling relationships for the elastic moduli and viscosity of mixed lipid membranes. Proc. Natl. Acad. Sci. U.S.A. 117, 23365–23373 (2020)
E.J. Dufourc, C. Mayer, J. Stohrer, G. Althoff, G. Kothe, Dynamics of phosphate head groups in biomembranes. Comprehensive analysis using phosphorus-31 nuclear magnetic resonance lineshape and relaxation time measurements. Biophys. J.. J. 61, 42–57 (1992)
K. Weisz, G. Gröbner, C. Mayer, J. Stohrer, G. Kothe, Deuteron nuclear magnetic resonance study of the dynamic organization of phospholipid/cholesterol bilayer membranes: molecular properties and viscoelastic behavior. Biochemistry 31, 1100–1112 (1992)
A. Nowacka, P.C. Mohr, J. Norrman, R.W. Martin, D. Topgaard, Polarization transfer solid-state NMR for studying surfactant phase behavior. Langmuir 26, 16848–16856 (2010)
A. Leftin, C. Job, K. Beyer, M.F. Brown, Solid-state 13C NMR reveals annealing of raft-like membranes containing cholesterol by the intrinsically disordered protein α-Synuclein. J. Mol. Biol. 425, 2973–2987 (2013)
A. Leftin, T.R. Molugu, C. Job, K. Beyer, M.F. Brown, Area per lipid and cholesterol interactions in membranes from separated local-field (13)C NMR spectroscopy. Biophys. J. 107, 2274–2286 (2014)
M.F. Brown, Collective dynamics in lipid membranes, characterization of biological membranes: structure and dynamics (De Gruyter, Berlin, 2019), pp.231–268
J.P. Douliez, A. Léonard, E.J. Dufourc, Restatement of order parameters in biomembranes: calculation of C-C bond order parameters from C-D quadrupolar splittings. Biophys. J. 68, 1727–1739 (1995)
J.-P. Douliez, A. Ferrarini, E.-J. Dufourc, On the relationship between C-C and C-D order parameters and its use for studying the conformation of lipid acyl chains in biomembranes. J. Chem. Phys. 109, 2513–2518 (1998)
M.F. Brown, S.I. Chan, Bilayer Membranes: Deuterium and Carbon‐13 NMR, eMagRes (2007).
M.F. Brown, R.L. Thurmond, S.W. Dodd, D. Otten, K. Beyer, Elastic deformation of membrane bilayers probed by deuterium NMR relaxation. J. Am. Chem. Soc. 124, 8471–8484 (2002)
M. Doktorova, G. Khelashvili, R. Ashkar, M.F. Brown, Molecular simulations and NMR reveal how lipid fluctuations affect membrane mechanics. Biophys. J. 122, 984–1002 (2023)
H. Chakraborty, P.K. Tarafdar, M.J. Bruno, T. Sengupta, B.R. Lentz, Activation thermodynamics of poly(ethylene glycol)-mediated model membrane fusion support mechanistic models of stalk and pore formation. Biophys. J. 102, 2751–2760 (2012)
G. Weinreb, B.R. Lentz, Analysis of membrane fusion as a two-state sequential process: evaluation of the stalk model. Biophys. J.. J. 92, 4012–4029 (2007)
Acknowledgements
This work was supported by the Core Research Grant (CRG/2021/001515) of the Science and Engineering Research Board (SERB), and the SERB-Science and Technology Award (STR/2021/000029) for Research. H.C. and S. P. thank the University Grants Commission and SERB, New Delhi for the UGC-Assistant Professor position and research fellowship, respectively. We thank members of the Chakraborty laboratory for their comments and discussions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pandia, S., Chakraborty, H. Navigating the mechanistic pathways of membrane fusion: the lipid perspective. Eur. Phys. J. Spec. Top. (2024). https://doi.org/10.1140/epjs/s11734-024-01106-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1140/epjs/s11734-024-01106-5