Abstract—Aryl-hydrocarbon receptor (Aryl Hydrocarbon Receptor, AHR) is a ligand-dependent transcription factor; its functions are related to xenobiotic detoxification, response to inflammation, and the maintenance of tissue homeostasis. Results of recent studies suggest that AHR also plays an important role in carcinogenesis. Increased expression of AHR is observed in several types of tumors and tumor derived cell lines. In addition, many AHR ligands are included in compositions of pharmaceutical drugs used in oncotherapy. These facts provide some ground to consider AHR as a potential target for anticancer therapy, especially for treatment of severe cancers which have very limited (if any) treatment options. In this review we have considered the examples of the effects of AHR ligands on tumor derived cell cultures and on model mice lines with analysis of the AHR-dependent response.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 GENERAL INFORMATION ON ARYL-HYDROCARBON RECEPTOR AND ITS LIGANDS
Aryl-hydrocarbon receptor (AHR) is a ligand-dependent cytosolic transcription factor, which belongs to the family of heterodimeric transcriptional regulators containing bHLH/PAS motifs (basic-Helix-Loop-Helix/Period [Per]-Aryl hydrocarbon receptor nuclear translocator [ARNT]—single minded [SIM]) [1].
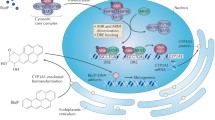
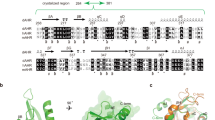
AHR has the N-terminal bHLH motif, which includes two functionally different and highly conservative domains, located at the distance of 60 amino acid residues from each other. At the N-terminal part of this motif there is the main domain responsible for AHR binding with its consensus sequence on DNA (5'-T/GCGTG-3'). This consensus is known in the literature as AHREs (Aryl Hydrocarbon Response Elements) or XREs (Xenobiotic Response Elements), or DREs (Dioxin Response Elements). It is usually located in the promoter zone of the AHR target genes. At the C-terminus of the bHLH motif there is the HLH domain responsible for the protein-protein interaction necessary for heterodimer formation with ARNT (Aryl Hydrocarbon Receptor Nuclear Translocator). The PAS-A and PAS-B domains participate in secondary interactions with ARNT, maintaining specificity of this protein complex. The AHR ligand binding site is located within the PAS-B domain. This site contains several conservative amino acid residues essential for ligand binding. Finally, the C-terminal region of the AHR protein contains a glutamine-rich (Q) domain, which is necessary for the coactivator binding and participation in the activation of transcription (Fig. 1) [1, 2].
The domain structure of the AHR protein and the scheme of its activation by ligands. (a) Domain protein structure. Numbers and lines indicate amino acid sequences corresponding to the functional domains of the AHR protein. (b) The scheme of AHR activation (modified from [2]). Explanations are given in the text.
According to the Human Protein Atlas project [3], human AHR gene mRNA is present in many organs and tissues, with predominance (>10 RPKM Reads per kilo base per million) in the bladder, lungs, liver, stomach, gall bladder , adrenal gland, appendix, intestine, placenta, skin, spleen, thyroid gland and bone marrow. To a lesser extent (<10 RPKM), AHR transcripts are present in the brain, heart, kidneys, pancreas, salivary glands, and testicles. Human tissue-specific distribution of the AHR protein is different: the highest AHR content was found in the brain, lungs, endocrine glands (thyroid, adrenal glands), testicles, muscles, some organs of the urogenital system (kidney, bladder, fallopian tubes), skin, some organs of the gastrointestinal tract. Expression of the AHR protein was not detected in the pancreas and prostate glands, ovaries, and low expression levels were detected in the gallbladder, stomach, lungs, and rectum [3].
Inactive ligand-free AHR is localized in the cytoplasm as part of a multipeptide complex containing two molecules of the HSP90 (Heat Shock Protein 90), co-chaperone p23, and one molecule of immunophilin-like protein XAP2 (also known as AIP, AHR Interacting Protein) (Fig. 1). HSP90 interacts with AHR through the bHLH domains and the domain containing the PAS-B ligand-binding site. When ligand binding to AHR causes conformational changes of this receptor and its N-terminal nuclear localization signal (NLS) becomes active due to its release from XAP2. AHR is then transported to the nucleus, released from the HSP90 chaperone, and dimerized with its partner ARNT. The AHR/ARNT heterodimer interacts with several histone acetyltransferases, chromatin-remodeling factors and a number of coactivators and/or corepressors. The resultant multiprotein complex binds to XRE in the region of enhancers and TATA box, recruits RNA polymerase II and induces transcription of target genes. At the final stage of the transcriptional regulation, AHR is rapidly exported to the cytoplasm by means of the CRM1 protein (Chromosome Region Maintenance 1), where its ubiquitin-dependent degradation occurs in the 26S proteasome [4] (Fig. 1).
The AHR/ARNT complex can influence transcription through binding to consensus sequences on DNA, thus limiting access of other transcription factors to the promoter [5]. In addition, AHR activity in the cell is negatively regulated by the AHRR repressor protein (AHR Repressor protein), and the expression of this protein is controlled by AHR itself. AHRR, as well as AHR, is a bHLH/PAS transcription factor that can dimerize with ARNT and compete with it for XREs binding. This triggers the negative feedback mechanism resulted in decreased activity of AHR target genes decreases [6].
Several other nuclear receptor coactivators also interact with AHR; these include ERAP140, RIP140, BRG1, Rb, PML, NEDD8, SUMO1, and three members of the p160 coactivator family: NCOA1, NCOA2 and NCOA3. Sequential and cyclic association of AHR and coactivators results in acetylation of histones, activation of PolII (RNA polymerase II), and the start of gene transcription. In other cases, AHR activation leads to inhibition of transcriptionally active genes, including genes encoding immunoglobulin heavy chain, estrogen-inducible p27, cathepsin D, and PS2 [7, 8].
Activation of AHR induces transcription of many genes involved in sequential detoxification processes. These include genes encoding phase I enzymes (xenobiotic metabolism phases), for example, CYP1A1, CYP1A2, CYP1B1, and CYP2S1. Other AHR target genes encode phase II enzymes: UDP-glucuronosyltransferase (UGT1A6), NAD(P)H-quinone oxidoreductase 1 (NQO1), aldehyde dehydrogenase (ALDH3A1), and several glutathione-S-transferase. Finally, the third group of AHR target genes encodes phase III xenobiotic transporters (xenobiotic utilization phase). These include genes encoding P-glycoprotein (P-gp), proteins associated with multidrug-resistant (MRP), and organic anion transporter proteins type 2 (OATP2). All these genes are expressed in many tissues and organs (liver, intestines, kidneys, brain) and play an important role in the absorption, distribution, and elimination of drugs from the body. This enzyme system plays a central role in the metabolism, elimination and detoxification (or activation) of xenobiotics, as well as drugs administered into the human body [9]. There are also many other genes whose functions depend on AHR activity. Basically, these genes are involved in control of homeostasis and detoxification processes, as well as division [9, 10], differentiation, polarization, and apoptosis of cells. Some of these genes are responsible for formation of organ-tissue structures of the nervous, immune, cardiovascular, endocrine, generative and excretory systems in higher multicellular organisms. The most studied of them are: Myc, Rbf1, NFKB1, JUN, CDC42, p23, RELA, p53 (and many others) [10, 11].
Recently, the effect of ectopic expression of human AHR on the activity of its target genes has been studied using transgenic lines of Drosophila melanogaster [12]. It has been shown that exogenous AHR agonists can both increase and decrease the transcription level of target genes. It should be noted that the effect of ligands on the expression of AHR target genes is tissue-specific and depends on the stage of development. Some evidence has also been obtained that the activation of targeted AHR genes, including many oncogenes and genes involved in the regulation of homeostasis and “developmental” functions, depends on their epigenetic status. It is possible that epigenetic repressive labels in the promoter region of target genes limit AHR availability to their binding sites [12].
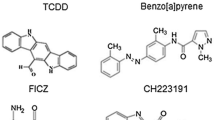
For many years, the major attention has been paid to identification of chemical compounds exhibiting potent agonistic (stimulating) or antagonistic (inhibitory) activity towards AHR. Table 1 lists such compounds of both endogenous and exogenous origin. Despite great diversity of ligands, there are very few studies on kinetics of their interaction with AHR; most of them were focused on 2,3,7,8-tetrachloro-dibenzo-p-dioxin and its derivatives. Table 2 summarizes results of studies on kinetics of ligand/receptor complex formation and their dissociation constants.
Despite enormous studies on analysis of the action of AHR ligands performed during several decades, new substances and pharmacological agents exhibiting affinity for AHR still appear. However, it is almost impossible to find common features in all ligands. Usually, in order to understand whether the effects of a given ligand depend on the action of AHR, additional experiments are performed using known AHR inhibitors or receptor knockdown. Disappearance of the effect strongly suggests its AHR-dependence.
2 AHR AND ITS LIGANDS IN TUMORIGENESIS AND CHEMOTHERAPY OF CANCER
Results of experimental studies provide increasing evidence for the important role of AHR in carcinogenesis and the search for selective AHR modulators is becoming a new promising area of pharmacological research aimed at developing drugs for chemotherapy of certain types of cancer [19].
Standard first-line chemotherapy for most types of cancer involves the use of cytotoxic drugs that selectively affect rapidly dividing cells of malignant tumors and do not damage healthy cells of the body. Activation of several target genes is used in the treatment of cancer. These include genes encoding membrane receptors with tyrosine kinase activity and their ligands, transcription factors and nuclear receptors. More than 80 pharmacological agents designed to activate 18 different nuclear receptors have been approved for use in oncological therapy [20]. However, compounds capable of activating AHR, which is also a ligand-dependent nuclear receptor, have not been approved for pharmacological applications. Only a few AHR ligands, such as aminoflavone and laquinimide, have been used in clinical trials to treat breast cancer and multiple sclerosis, respectively [21, 22].
Most of the initial studies of AHR and its ligands were devoted to the effect of TCDD (2,3,7,8-tetrachlorodibenzodioxin or the trivial name dioxin) on tumor formation after prolonged feeding of rodents. In most cases, TCDD acted as a hepatocarcinogen [23, 24]. TCDD-induced tumors have also been observed in many other organs; however, during the entire feeding time of Sprague-Dawley rats, there was a decrease in the occurrence of spontaneous tumors of the mammary gland and uterus [25]. AHR activity has been studied in many human cell lines and tumors [19]. The development of selective AHR modulators (SAhRM), including AHR-activating pharmacological agents, there is a clear need to use AHR as a potential target for the treatment of cancer and other diseases. Below we consider the role of AHR in carcinogenesis, studied in cultures of tumor cells of different origin and in model lines of mice.
2.1 Malignant Tumors of the Urogenital System
Table 3 shows examples of the effect of several AHR ligands on various malignant tumors of the urogenital system and in provoking prostate cancer in the TRAMP mouse model line [26]. TCDD and other dioxin compounds inhibit proliferation of prostate cancer cells, but their mechanisms of action differ in different cell types [27, 28]. The role of AHR and its ligands in prostate cancer cells depend on the androgen receptor (AR). On the one hand, there is evidence that AHR ligands are anti-androgenic in prostate cancer cells expressing AR, and AHR itself is an inhibitor of tumor growth [29]. According to other data, AHR knockdown using small RNA (siAHR) decreased proliferation of androgen-independent prostate adenocarcinoma cells [30]. Most AHR ligands induce activity of matrix metalloproteinases (MMP), which are able to degrade all types of extracellular matrix proteins [28], while the antagonist CH223191 inhibits cell division [31]. In TRAMP mice with normal AR expression tumor growth was inhibited by ligands, but these data were inconsistent with the contradictory effects of TCDD [26, 32, 33].
Interesting data were obtained during the study of the AHR effect on the expression of miRNA on prostate cancer cell lines. AHR activation by TCDD (10 nM) or DIM (25 nM) resulted in increased expression of miR-150-5p, which had a negative impact on proliferation and invasion of prostate cancer cells [34].
The results of studies of urinary tract tumors suggest that AHR and its ligands increase cancer cell invasion [35], whereas the results obtained on kidney cancer cell lines are contradictory and most likely depend on the line of cultured cells [36, 37] (Table 3).
2.2 Malignant Tumors of the Central Nervous System
Glioblastoma is the most aggressive form of the malignant brain tumors with a poor prognosis. There are limited variants for its treatment and they are not very effective. Initial studies have shown increased expression of AHR in both malignant tumors of the human CNS and in glioblastoma cell cultures with activation of the TGFβ signaling pathway involved in the pro-oncogenic activity of AHR [38]. Knockdown of AHR (by siAHR) or the inhibition by the antagonist CH223191 (10 μM) reduced viability and migration of glioblastoma cells [38]. Subsequent studies by this research group showed that tryptophan-2,3-dioxygenase-mediated metabolism of tryptophan, resulted in its conversion into kynurenine, was a key procarcinogenic event, as kynurenine provoked AHR-dependent survival and mobility of tumor cells [39]. Recent study revealed existence of relationships between AHR, integrin and TGFβ in glioblastoma [40]. It is clear that these studies demonstrate the potential clinical role of AHR antagonists in the treatment of glioblastoma. Studies on other types of central nervous system tumors, including medulloblastoma and pituitary adenomas, also show that AHR is a pro-oncogene [41, 42], while in neuroblastoma cells AHR enhances differentiation [43] and TCDD induces apoptosis of pheochromocytoma (PC12) cultured cells [44]. In the biopsy material, taken from patients with meningiomas different degrees of malignancy, a direct relationship was found between the expression level of the AHR protein and the degree of tumor malignancy [45]. These studies show multipolar opinions on the role of AHR and its ligands in brain carcinogenesis (Table 3).
2.3 Lung and Esophageal Cancer. Melanoma, Leukemia and Lymphoma
Lung cancer is the most common type of cancer among men. Worldwide, more than 1 million new cases of this disease are diagnosed annualy, with about 60 000 of them in Russia. According to World Health Organization (2017), the mortality rate for lung cancer remains high for many years. Patients with diagnosed lung carcinoma demonstrate increased levels of AHR expression. Most studies performed on lung cancer cell cultures show that various AHR agonists, such as tobacco smoke extracts, PAHs (polycyclic aromatic hydrocarbons), β-naphthoflavone, TCDD, induce cancer cell growth by activating gene expression of growth factors and genes that activate cell division [46, 47]. It was also noted that in lung carcinoma cells, AHR expression positively correlated with expression of adrenomedullins and osteopontin, contributing to tumor growth and progression [48, 49] (Table 3).
Using lines of mice with null mutation of AHR (AHR–/–), it was shown that in lung fibroblasts AHR regulated the expression of miRNA (particularly miR-196a) involved in control of cell proliferation and apoptosis. Interestingly, this regulation was independent of the action of xenobiotics [50].
In many cases, AHR overexpression was found in esophageal cancer, leukemia, and lymphoma. It was shown that β-naphthoflavone significantly inhibited invasion of esophageal cancer cells [51]. The action of TCDD on human lymphoma cells and mice with lymphoma increased activity of cyclooxygenase (COX2) and increased resistance to apoptosis [52]. Indole-3-carbinol inhibited cell proliferation, induced apoptosis and cell cycle arrest of THP-1 acute myeloid leukemia cells [53]. The study of primary cultures of NK cells, (natural killer cells), lymphocytes obtained from patients with diagnosed acute myeloid leukemia has shown that AHR activity leads to the induction of miR-29b, thus impairing the development and division of NK cells. The antagonist CH223191 increases apoptotic parameters of blast cells and reduces their resistance to the NK cell cytotoxicity. The authors propose the use of AHR antagonists as therapeutic agents for the treatment of leukemia [54].
Conflicting data have been reported on melanoma. AHR knockdown (by using siAHR) increased tumor formation in vivo, while leflunomide inhibited proliferation of melanoma cells [55, 56]; however, it was also reported that AHR knockdown reduced cell proliferation [57] and TCDD increased tumor invasion and MMP metalloproteinase expression [58] (Table 3). Differences in these data may be attributed to different cell cultures and model mouse strains used in these experiments. Obviously, results of these studies need additional verification.
2.4 Colon and Stomach Cancers
Functions of AHR ligands in colon cancer cells depend on the cell type and ligands (Table 3). Several different ligands, for example, 3-methylcholanthrene (MC) (in studies on Caco-2 and LS174T cells) and TCDD (in studies H508, SN7-C4 cells) show a pro-oncogenic potential, by inducing cell division and expression of genes related to extracellular matrix remodeling (MMP9) and transport of xenobiotics (ABCG2) [58–60]. However, in studies performed on other cell lines of colon tumors, AHR ligands inhibited cell division: FICZ (on the LoVo line) and chrysin (on HCT116, DLD-1 and SW837 lines) [61, 62]. On the contrary, several in vivo studies have shown that in mice with null mutation of AHR (AHR–/–) and in mice with multiple intestinal neoplasia of the APCmin/+ line (with a point mutation in the APC gene) carcinogenesis of the colon and cecum increased, while the effect of I3C ligands and DIM inhibited carcinogenesis [4, 63, 64]. Thus, the positive tumor-suppressor activity of AHR in the development of colon/cecum cancer and the positive role of specific AHR ligands in inhibiting carcinogenesis were clearly shown using the in vivo mouse model.
Studies performed on the culture of gastric carcinoma cells MNK5 in vitro and in vivo (xenotransplantation) have shown that AHR promotes division, migration, and survival of tumor cells [65, 66]. TCDD induced proliferation and invasion of AGS gastric carcinoma cells [67], while DIM reduced division of SGC-7901 cell [68, 69]. However, it remains unclear whether this inhibitory effect of DIM depends on AHR. Expression of constitutively active AHR (CA-AHR) in mice results in formation of a gastric tumor, which indicates the proto-oncogenic role of this receptor [70, 71].
2.5 Liver Cancer
Liver cancer is the second after lung cancer in terms of the highest rates of mortality from malignant tumors worldwide, accounting for more than 750 000 cases per year. Although liver cancer is much more common in Southeast Asia, the incidence rate of liver cancer worldwide including Russia increases. The prognosis of the survival rate for liver cancer patients is unfavorable: approximately 15% of patients will live 5 years after the establishment of the diagnosis of liver cancer [72]. Such a poor prognosis is explained by the resistance of hepatocellular carcinoma (this type of tumor occurs in 90% of cases with liver cancer) to chemotherapy and the lack of possible biologically targeted methods of treatment. The only existing targeted therapy for liver cancer is the use of the drug sorafenib (a kinase inhibitor), which prolongs lives of such patients roughly up about 3 months [73]. One of the promising approaches of the anticancer therapy is the development of a targeted method based on AHR and its ligands. The results of in vitro and in vivo studies performed on hepatocellular carcinoma are summarized in Table 3.
In different strains of mice with AHR gene knockout, the liver size was much smaller, and there were defects in the development of the vascular network [74–76]. The genes required for normal growth and development often play an important role in oncogenesis. They function as oncogenes or tumor suppressors, and sometimes both as tumor suppressor and as oncogene. This depends on the genetic background or expression of other regulatory proteins. Mice with null AHR mutation do not develop spontaneous tumors in the liver. This suggests that AHR is not a classic tumor suppressor gene. Normally, a genetically programmed system of “checks” and “balances,” which is responsible for elimination of abnormal cells, resists oncogenesis. In the absence of any exogenous ligands endogenous AHR functions as a tumor modifier gene for liver cancer. The ability of AHR to act as a tumor modifier was examined in mice exposed to chemical carcinogens. Fan et al. used the genotoxic carcinogens, diethyl nitrosamine, to induce liver tumors in wild-type mice with normal AHR, and AHR knockout mice [77]. Diethyl nitrosamine is not an AHR ligand and its use has allowed the study of AHR functions that are independent of xenobiotics. In AHR knockout mice (AHR–/–) diethyl nitrosamine administration resulted in increased incidences of liver tumors as compared to wild-type mice AHR+/+. In addition, the number of tumors and their size were higher in AHR–/– mice than in AHR+/+ mice. Parameters of cell proliferation, cytokine expression, and DNA damage were significantly higher in wild-type mice. Based on these results the authors have concluded that AHR functions as a tumor suppressor or modifier in liver cancer [77].
Studies on hepatocellular carcinoma cell cultures have shown that the level of AHR expression is elevated in liver cancer cells, and its ligands inhibit cell proliferation and/or induce cancer cell death, and these effects depend on the AHR expression level [56]. Many efforts have been undertaken to find AHR ligands, which would exhibit the anticancer effect on hepatocellular carcinoma [19, 56, 78–80]. The specificity and selectivity of the identified small molecules for AHR have been confirmed in well characterized cellular systems. Moreover, these compounds were tested for AHR-dependent, growth inhibitory, effects in cancer cells. This resulted in identification of promising AHR ligands with potential anticancer effects, and raloxifene was one of such AHR ligands. Raloxifene is a selective estrogen receptor modulator used for osteoporosis prevention. Raloxifene binds to AHR, thus contributing to its nuclear translocation followed by activation of target genes [80, 81]. The AHR-dependent programmed death of breast and liver cancer cells that do not express the estrogen receptor promoted raloxifene-induced inhibition of cell growth. Despite the ability of TCDD to strongly activate AHR, it does not induce apoptosis. In this regard, other AHR ligands, for example raloxifene, are unique [80]. In contrast to TCDD, raloxifene is not a high-affinity ligand for AHR, so it is important to understand how this ligand-selective AHR pathway produces an anticancer effect. Raloxifene is well tolerated by patients, so for future clinical trials it is important to continue works on creating new drugs based on this substance with stronger affinity for AHR.
Analysis of chemicals within the ToxCast project, aimed at studies of the effects of chemicals that lead to adverse health effects, particularly, their ability to activate nuclear receptors, including AHR, showed no link between AHR activation and progression of the liver damage [82]. The growth of human hepatoma cells HCCLM3 was inhibited both in vitro and in vivo (xenograft) by the ligand AHR, ITE [83]. Flutamide, an antiandrogen, used for treatment of prostate cancer, is also an AHR ligand, and its ability to suppress growth of human hepatocellular carcinoma cells is determined by the AHR-dependent induction of TGFβ1 [79]. AHR-mediated activation of TGFβ1 signals led to activation of cell-cycle-inhibiting proteins p15 and p27, and AHR knockdown (by using siAHR) or TGFβ1 eliminated the antiproliferative effects of flutamide. This is an example of an AHR-dependent pharmacological agent that can be used in the treatment of not only prostate cancer, but also hepatocellular carcinoma.
2.6 Breast Cancer
Breast cancer is the most common cancer among women all over the world, and most of the fatal cases of this disease are caused by metastases. Breast cancer includes various subtypes with different molecular markers. Three main hormone-dependent breast cancer subtypes include: (1) hormone-sensitive cancer, in which the estrogen receptor (ER) and progesterone receptor (PR) are expressed, (2) HER2-positive cancer with overexpression of the HER2 protein (human epidermal growth factor receptor 2) and (3) triple-negative breast cancer (TNBC), in which neither the ER receptor nor the PR receptors are expressed, and HER2 may not be expressed at all or it may be normal [84, 85].
Approximately 20% of breast cancers are diagnosed as triply negative [86]. Breast cancer of this type has poor prognosis and is most difficult to treat. AHR is expressed in all three types [80]. Higher AHR expression positively correlated with a better prognosis, including an increase in the total life expectancy of patients and survival without distant metastasis in various forms of breast cancer [80].
The use of targeted AHR based therapy is a unique opportunity for breast cancer patients with limited variants for treatment. Table 3 lists examples of recent studies that consider the role of AHR in breast cancer. The results of many studies presented in this table confirm the use of AHR as an anticancer target for breast cancer. Pretreatment of CB6F1 mice with TCDD inhibited tumor formation caused by the chemical carcinogen 7,12-dimethylbenzo[a]anthracene (DMBA) in the mammary glands [87]. Diindoylmethane (DIM), a food ligand of AHR, also inhibited DMBA-induced mammary tumor formation in Sprague-Dawley rats [88].
In a syngeneic mouse model of breast cancer metastasis, TCDD has been shown to reduce metastasis of breast tumors to the lungs and other mammary glands [89]. Interestingly, TCDD treatment did not affect the primary tumor growth in these mice and did not affect cell proliferation studied in vitro experiments. The data from these studies show a positive trend in testing AHR targeting as an anticancer therapy both in vitro and in vivo. Since most breast cancer deaths are caused by complications of metastases to distant organs, systematic testing of various classes of AHR modulators can help to identify those modulators that effectively inhibit metastasis.
Omeprazole, a proton pump inhibitor, activated AHR and also reduced metastasis in triple negative breast cancer [78]. AHR activation by some agonists, including omeprazole, suppressed expression of CXCR4, the G-protein-coupled receptor, associated with formation of metastasis of breast tumors [78, 89–91]. miRNAs regulated by AHR also play a role in breast cancer metastasis. TCDD and MHDF induced expression of miR-335 in BT474 and MDA-MD-231 cells; this resulted in inhibition of the prometastatic SOX4 gene and inhibition of lung metastases in vivo [51]. A new flavonoid agonist flavipin also induced a microRNA cluster, miR-212/132, which inhibited migration and invasion of cancer cells [92].
The selective estrogen receptor modulator raloxifene induces apoptosis in triple negative breast cancer cells; this indicates that this compound or its analogs also have the potential of AHR-targeted therapy for breast cancer [80, 93].
In experiments with xenotransplantation of human breast adenocarcinoma cells (BP1, Hs578T, MDA-MB-231, and SUM149 cell lines) AHR knockdown in the fish Danio rerio (by siAHR) and application of AHR antagonists (CH223191, CB7993113) reduced the invasion and migration of human cancer cells, reduced tumor metastasis [94]. This was attributed to a decrease in expression of genes associated with invasion (for example, fibronectin, VCAM1, thrombospondin, MMP1) and an increase in expression of CDH1/E-cadherin previously associated with reduced tumor aggression. Paradoxically, the use of agonists (TCDD, DIM) in the same experiments also reduced invasion of cancer cells and blocked metastases in vivo, but accelerated cell migration [94]. These data show the difficulty of modulating AHR activity in cancer, suggesting that AHR inhibitors and, in some cases, AHR agonists, may be useful as cancer therapy.
2.7 Cancer Stem Cells
AHR plays a role in stem cell functioning, and results of recent studies show that AHR antagonists promoted expansion of hematopoietic stem cells [95–99]. Cancer stem cells are often drug resistant and important for maintaining and expanding certain types of tumors. There is evidence that AHR can be a target in some cancer stem cells. For example, AHR-activating pharmacological agent tranilast significantly inhibited growth of breast cancer stem cells and prevented lung metastasis in mice that were injected with cells of triple negative breast cancer line MDAMB-231, resistant to the antitumor drug mitoxantrone [100].
The other study demonstrated the AHR-dependent response of cancer stem cell derivatives of triple negative breast cancer Hs578T. It was shown that ligands induced AHR interaction with Sox2, a regulator of self-reproduction; this clearly demonstrates the role of AHR and its agonists as “amplifiers” of cancer stem cells [101]. These results differ from those obtained using tranlilast, thus suggesting different AHR functions dependent on the cellular context in breast cancer stem cells and possibly related to the differential expression of ARR, ARNT, HIF-1α, and other cofactors. Cheng et al. [102] described the effects of exposure to tryptophan derivatives, including ITE and demonstrating suppression of Oct4 transcription in stem cancer cells. The ITE ligand caused an AHR-dependent decrease in the expression of a stem cell marker, Oct4. After exposure to ITE, stem cell-like cancer cells were differentiated and their tumor potential decreased in subcutaneous and orthotopic xenograft tumor models. In contrast, AHR antagonists increased activity of leukemia stem cells [103]; this was consistent with the effects reported in the study of hematopoietic stem cells [95]. These and other studies [104–106] show that AHR and its responsive genes are important for cancer stem cells, and AHR ligands (agonists or antagonists) represent a unique set of agents for the treatment of cancer of stem cells.
3 CONCLUSIONS
The detected AHR function to act as a tumor modifier and the anticancer effects, stimulated by various classes of AHR ligands with diverse pharmacology, are strong arguments for studying the AHR signaling pathway in the context of its use in anticancer therapy. However, involvement of AHR in known negative effects of TCDD, realized via AHR activation, ruined hopes in this receptor and reduced support from large financial institutions and biotechnology companies for scientific work in the field of AHR based cancer therapy. The reason for the cautious approach to the use of AHR in anticancer therapy is understandable provided that there are other treatment options or other molecular routes are known for targeted therapy. However, for treatment of severe cancer and cancer with limited variants of treatment or with lack of appropriate treatment at all (pancreatic cancer, liver cancer, a hormone-independent form of breast and prostate cancer), it is time to use the AHR potential for development of a new class of anticancer drugs. It is important to determine the mode of AHR functioning, which contributes to its anticancer effect, and some general issues, including cell cycle gene regulation, interaction with various coregulatory molecules, and non-genomic pathways that contribute to the anticancer activity of AHR.
Design and selection of ligands based on the mechanism of AHR action will make it possible to identify new molecules that are valuable in the context of oncotherapy. Among the receptors and their ligands, there are many examples illustrating their successful use for clinical purposes such as the retinoid X receptor (drug bexarotin), the estrogen receptor (drugs tamoxifen and raloxifene), the androgen receptor (flutamide, enzalutamide) and the glucocorticoid receptor (fluticasone) [107–110]. It would be great to supplement this list with an aryl-hydrocarbon receptor.
4 ACKNOWLEDGMENTS
This work was carried out with the financial support of the Russian Foundation for Basic Research in the framework of the research project no. 18-34-00162 mol-a and the section of the State Task of the IDB RAS no. 0108-2018-0001.
5 COMPLIANCE WITH ETHICAL STANDARDS
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
REFERENCES
Fukunaga, B.N., Probst, M.R., Reisz-Porszasz, S., and Hankinson, O., J. Biol. Chem., 1995, vol. 270, pp. 29 270–29 278.
Feng, S., Cao, Z., and Wang, X., Biochim. Biophys. Acta, 2013, vol. 1836, pp. 197–210. https://doi.org/10.1016/J.BBCAN.2013.05.001
Tissue Expression of AHR—Summary—The Human Protein Atlas. https://www.proteinatlas.org/ ENSG00000106546-AHR/tissue
Ikuta, T., Kobayashi, Y., Kitazawa, M., Shiizaki, K., Itano, N., Noda, T., Pettersson, S., Poellinger, L., Fujii-Kuriyama, Y., Taniguchi, S., and Kawajiri, K., Carcinogenesis, 2013, vol. 34, pp. 1620–1627. https://doi.org/10.1093/carcin/bgt083
Beischlag, T.V., Morales, J.L., Hollingshead, B.D., and Perdew, G.H., Crit. Revs. Eukar. Gene Expr., 2008, vol. 18, pp. 207–250.
Cauchi, S., Stücker, I., Cénée, S., Kremers, P., Beaune, P., and Massaad-Massade, L., Pharmacogenetics, 2003, vol. 13, pp. 339–347. https://doi.org/10.1097/01.fpc.0000054093.48725.79
Wang, F., Samudio, I. and Safe, S., Mol. Cell. Endocrinol., 2001, vol. 172, pp. 91–103.
Tojo, M., Matsuzaki, K., Minami, T., Honda, Y., Yasuda, H., Chiba, T., Saya, H., Fujii-Kuriyama, Y., and Nakao, M., J. Biol. Chem., 2002, vol. 277, pp. 46 576–46 585. https://doi.org/10.1074/jbc.M205987200
Xu, C., Li, C.Y.-T., and Kong, A.-N.T., Arch. Pharmacal Res., 2005, vol. 28, pp. 249–268.
Gasiewicz, T.A., Singh, K.P., and Casado, F.L., Chemico-Biological Interactions, 2010, vol. 184, pp. 246–251. https://doi.org/10.1016/j.cbi.2009.10.019
Akahoshi, E., Yoshimura, S., and Ishihara-Suga-no, M., Environmental Health: A Global Access Science Source, 2006, vol. 5, 24. https://doi.org/10.1186/1476-069X-5-24
Akishina, A.A., Vorontsova, J.E., Cherezov, R.O., Mertsalov, I.B., Zatsepina, O.G., Slezinger, M.S., Panin, V.M., Petruk, S., Enikolopov, G.N., Mazo, A., Simonova, O.B., and Kuzin, B.A., Oncotarget, 2017, vol. 8. https://doi.org/10.18632/oncotarget.22173
Harper, P.A., Prokipcak, R.D., Bush, L.E., Golas, C.L., and Okey, A.B., Arch. Biochem. Biophys., 1991, vol. 290, pp. 27–36.
Roberts, E.A., Harper, P.A., Wong, J.M., Wang, Y., and Yang, S., Arch. Biochem. Biophys., 2000, vol. 384, pp. 190–198. https://doi.org/10.1006/abbi.2000.2059
Poland, A., Glover, E., Ebetino, F.H., and Kende, A.S., J. Biol. Chem., 1986, vol. 261, pp. 6352–6365.
Pohjanvirta, R., Viluksela, M., Tuomisto, J.T., Unki-la, M., Karasinska, J., Franc, M.A., Holowenko, M., Giannone, J.V., Harper, P.A., Tuomisto, J., and Okey, A.B., Toxicol. Appl. Pharmacol., 1999, vol. 155, pp. 82–95. https://doi.org/10.1006/taap.1998.8565
Bank, P.A., Yao, E.F., Swanson, H.I., Tullis, K., and Denison, M.S., Arch. Biochem. Biophys., 1995, vol. 317, pp. 439–448.
Ema, M., Ohe, N., Suzuki, M., Mimura, J., Sogawa, K., Ikawa, S., and Fujii-Kuriyama, Y., J. Biol. Chem., 1994, vol. 269, pp. 27 337–27 343.
Safe, S., Lee, S.-O., and Jin, U.H., Toxicol. Sci., 2013, vol. 135, pp. 1–16. https://doi.org/10.1093/toxsci/kft128
Tice, C.M. and Zheng, Y.J., Bioorg. Med. Chem. Letts., 2016, vol. 26, pp. 4157–4164. https://doi.org/10.1016/j.bmcl.2016.07.067
Haggiag, S., Ruggieri, S., and Gasperini, C., Ther. Adv. Neurol. Disord., 2013, vol. 6, pp. 343–352. https://doi.org/10.1177/1756285613499424
Loaiza-Pérez, A.I., Kenney, S., Boswell, J., Hollingshead, M., Alley, M.C., Hose, C., Ciolino, H.P., Yeh, G.C., Trepel, J.B., Vistica, D.T., and Saus-ville, E.A., Mol. Cancer Ther., 2004, vol. 3, pp. 715–725.
Bock, K.W. and Köhle, C., Biochem. Pharmacol., 2005, vol. 69, pp. 1403–1408. https://doi.org/10.1016/j.bcp.2005.02.004
Knerr, S. and Schrenk, D., Mol. Nutr. Food Res., 2006, vol. 50, pp. 897–907. https://doi.org/10.1002/mnfr.200600006
Kociba, R.J., Keyes, D.G., Beyer, J.E., Carreon, R.M., Wade, C.E., Dittenber, D.A., Kalnins, R.P., Frauson, L.E., Park, C.N., Barnard, S.D., Hummel, R.A., and Humiston, C.G., Toxicol. Appl. Pharmacol., 1978, vol. 46, pp. 279–303.
Fritz, W.A., Lin, T.-M., Safe, S., Moore, R.W., and Peterson, R.E., Biochem. Pharmacol., 2009, vol. 77, pp. 1151–1160. https://doi.org/10.1016/j.bcp.2008.12.015
Barnes-Ellerbe, S., Knudsen, K.E., and Puga, A., Mol. Pharmacol., 2004, vol. 66, pp. 502–511. https://doi.org/10.1124/mol.104.000356
Haque, M., Francis, J., and Sehgal, I., Cancer Letts., 2005, vol. 225, pp. 159–166. https://doi.org/10.1016/j.canlet.2004.11.043
Gluschnaider, U., Hidas, G., Cojocaru, G., Yutkin, V., Ben-Neriah, Y., and Pikarsky, E., PLoS One, 2010, vol. 5, e9060. https://doi.org/10.1371/journal.pone.0009060
Tran, C., Richmond, O., Aaron, L., and Powell, J.B., Biochem. Pharmacol., 2013, vol. 85, pp. 753–762. https://doi.org/10.1016/j.bcp.2012.12.010
Richmond, O., Ghotbaddini, M., Allen, C., Walker, A., Zahir, S., and Powell, J.B., PLoS One, 2014, vol. 9, e95058. https://doi.org/10.1371/journal.pone.0095058
Fritz, W.A., Lin, T.-M., Cardiff, R.D., and Peterson, R.E., Carcinogenesis, 2007, vol. 28, pp. 497–505. https://doi.org/10.1093/carcin/bgl179
Moore, R.W., Fritz, W.A., Schneider, A.J., Lin, T.-M., Branam, A.M., Safe, S., and Peterson, R.E., Toxicol. Appl. Pharmacol., 2016, vol. 305, pp. 242–249. https://doi.org/10.1016/j.taap.2016.04.018
Yu, J., Feng, Y., Wang, Y., and An, R., Arch. Biochem. Biophys., 2018, vol. 654, pp. 47–54. https://doi.org/10.1016/j.abb.2018.07.010
Ishida, M., Mikami, S., Kikuchi, E., Kosaka, T., Miyajima, A., Nakagawa, K., Mukai, M., Okada, Y., and Oya, M., Carcinogenesis, 2010, vol. 31, pp. 287–295. https://doi.org/10.1093/carcin/bgp222
Callero, M.A., Suárez, G.V., Luzzani, G., Itkin, B., Nguyen, B., and Loaiza-Perez, A.I., Int. J. Oncol., 2012, vol. 41, pp. 125–134. https://doi.org/10.3892/ijo.2012.1427
Ishida, M., Mikami, S., Shinojima, T., Kosaka, T., Mizuno, R., Kikuchi, E., Miyajima, A., Okada, Y., and Oya, M., Int. J. Cancer, 2015, vol. 137, pp. 299–310. https://doi.org/10.1002/ijc.29398
Gramatzki, D., Pantazis, G., Schittenhelm, J., Tabatabai, G., Köhle, C., Wick, W., Schwarz, M., Weller, M., and Tritschler, I., Oncogene, 2009, vol. 28, pp. 2593–2605. https://doi.org/10.1038/onc.2009.104
Opitz, C.A., Litzenburger, U.M., Sahm, F., Ott, M., Tritschler, I., Trump, S., Schumacher, T., Jestaedt, L., Schrenk, D., Weller, M., Jugold, M., Guillemin, G.J., Miller, C.L., Lutz, C., Radlwimmer, B., Lehmann, I., Deimling, A., von Wick, W., and Platten, M., Nature, 2011, vol. 478, pp. 197–203. https://doi.org/10.1038/nature10491
Silginer, M., Burghardt, I., Gramatzki, D., Bunse, L., Leske, H., Rushing, E.J., Hao, N., Platten, M., Weller, M., and Roth, P., Oncogene, 2016, vol. 35, pp. 3260–3271. https://doi.org/10.1038/onc.2015.387
Dever, D.P. and Opanashuk, L.A., Mol. Pharmacol., 2012, vol. 81, pp. 669–678. https://doi.org/10.1124/mol.111.077305
Jaffrain-Rea, M.-L., Angelini, M., Gargano, D., Tichomirowa, M.A., Daly, A.F., Vanbellinghen, J.-F., D’Innocenzo, E., Barlier, A., Giangaspero, F., Esposito, V., Ventura, L., Arcella, A., Theodoropoulou, M., Naves, L.A., Fajardo, C., Zacharieva, S., Rohmer, V., Brue, T., Gulino, A., Cantore, G., Alesse, E., and Beckers, A., Endocrine-Related Cancer, 2009, vol. 16, pp. 1029–1043. https://doi.org/10.1677/ERC-09-0094
Huang, T.-C., Chang, H.-Y., Chen, C.-Y., Wu, P.-Y., Lee, H., Liao, Y.-F., Hsu, W.-M., Huang, H.-C., and Juan, H.-F., FEBS Letts., 2011, vol. 585, pp. 3582–3586. https://doi.org/10.1016/j.febslet.2011.10.025
Sánchez-Martín, F.J., Fernández-Salguero, P.M., and Merino, J.M., NeuroToxicology, 2010, vol. 31, pp. 267–276. https://doi.org/10.1016/j.neuro.2010.03.005
Talari, N.K., Panigrahi, M.K., Madigubba, S., and Phanithi, P.B., J. Neuro-Oncology, 2018, vol. 137, pp. 241–248. https://doi.org/10.1007/s11060-017-2730-3
Wang, C.-K., Chang, H., Chen, P.-H., Chang, J.T., Kuo, Y.-C., Ko, J.-L., and Lin, P., Int. J. Cancer, 2009, vol. 125, pp. 807–815. https://doi.org/10.1002/ijc.24348
Shimba, S., Komiyama, K., Moro, I., and Tezuka, M., J. Biochem., 2002, vol. 132, pp. 795–802.
Chuang, C.-Y., Chang, H., Lin, P., Sun, S.J., Chen, P.H., Lin, Y.Y., Sheu, G.T., Ko, J.L., Hsu, S.L., and Chang, J.T., Gene, 2012, vol. 492, pp. 262–269. https://doi.org/10.1016/j.gene.2011.10.019
Portal-Nuñez, S., Shankavaram, U.T., Rao, M., Datrice, N., Atay, S., Aparicio, M., Camphausen, K.A., Fernández-Salguero, P.M., Chang, H., Lin, P., Schrump, D.S., Garantziotis, S., Cuttitta, F., and Zudaire, E., Cancer Res., 2012, vol. 72, pp. 5790–5800. https://doi.org/10.1158/0008-5472.CAN-12-0818
Hecht, E., Zago, M., Sarill, M., Rico de Souza, A., Gomez, A., Matthews, J., Hamid, Q., Eidelman, D.H., and Baglole, C.J., Toxicol. Appl. Pharmacol., 2014, vol. 280, pp. 511–525. https://doi.org/10.1016/j.taap.2014.08.023
Zhang, J., Zong, H., Li, S., Zhang, D., Zhang, L., and Xia, Q., Tumori, 2012, vol. 98, pp. 152–157. https://doi.org/10.1700/1053.11514
Vogel, C.F.A., Li, W., Sciullo, E., Newman, J., Hammock, B., Reader, J.R., Tuscano, J., and Matsumu-ra, F., Am. J. Pathol., 2007, vol. 171, pp. 1538–1548. https://doi.org/10.2353/ajpath.2007.070406
Mohammadi, S., Seyedhosseini, F.S., Behnampour, N., and Yazdani, Y., J. Receptor Signal Transduct. Res., 2017, vol. 37, pp. 506–514. https://doi.org/10.1080/10799893.2017.1360351
Scoville, S.D., Nalin, A.P., Chen, L., Chen, L., Zhang, M., McConnell, K., Beceiro Casas, S., Ernst, G., Traboulsi, A.A.-R., Hashi, N., Williams, M., Zhang, X., Hughes, T., Mishra, A., Benson, D.M., Saultz, J.N., Yu, J., Freud, A.G., Caligiuri, M.A., and Mundy-Bosse, B.L., Blood, 2018, https://doi.org/10.1182/blood-2018-03-838474
Contador-Troca, M., Alvarez-Barrientos, A., Barrasa, E., Rico-Leo, E.M., Catalina-Fernández, I., Menacho-Márquez, M., Bustelo, X.R., García-Borrón, J.C., Gómez-Durán, A., Sáenz-Santamaría, J., and Fernández-Salguero, P.M., Carcinogenesis, 2013, vol. 34, pp. 2683–2693. https://doi.org/10.1093/carcin/bgt248
O’Donnell, E.F., Kopparapu, P.R., Koch, D.C., Jang, H.S., Phillips, J.L., Tanguay, R.L., Kerkvliet, N.I., and Kolluri, S.K., PLoS One, 2012, vol. 7, e40926. https://doi.org/10.1371/journal.pone.0040926
Barretina, J., Caponigro, G., Stransky, N., Venkatesan, K., Margolin, A.A., Kim, S., Wilson, C.J., Lehár, J., Kryukov, G.V., Sonkin, D., Reddy, A., Liu, M., Murray, L., Berger, M.F., Monahan, J.E., Morais, P., Meltzer, J., Korejwa, A., Jané-Valbuena, J., Mapa, F.A., Thibault, J., Bric-Furlong, E., Raman, P., Shipway, A., Engels, I.H., Cheng, J., Yu, G.K., Yu, J., Aspesi, P., Silva, M., de Jagtap, K., Jones, M.D., Wang, L., Hatton, C., Palescandolo, E., Gupta, S., Mahan, S., Sougnez, C., Onofrio, R.C., Liefeld, T., MacConaill, L., Winckler, W., Reich, M., Li, N., Mesirov, J.P., Gabriel, S.B., Getz, G., Ardlie, K., Chan, V., Myer, V.E., Weber, B.L., Porter, J., Warmuth, M., Finan, P., Harris, J.L., Meyerson, M., Golub, T.R., Morrissey, M.P., Sellers, W.R., Schlegel, R., and Garraway, L.A., Nature, 2012, vol. 483, pp. 603–607. https://doi.org/10.1038/nature11003
Villano, C.M., Murphy, K.A., Akintobi, A., and White, L.A., Toxicol. Appl. Pharmacol., 2006, vol. 210, pp. 212–224. https://doi.org/10.1016/j.taap.2005.05.001
Tompkins, L.M., Li, H., Li, L., Lynch, C., Xie, Y., Nakanishi, T., Ross, D.D., and Wang, H., Biochem. Pharmacol., 2010, vol. 80, pp. 1754–1761. https://doi.org/10.1016/j.bcp.2010.08.016
Xie, G., Peng, Z., and Raufman, J.P., Am. J. Physiology-Gastrointestinal Liver Physiol., 2012, vol. 302, G1006–G1015. https://doi.org/10.1152/ajpgi.00427.2011
Ronnekleiv-Kelly, S.M., Nukaya, M., Díaz-Díaz, C.J., Megna, B.W., Carney, P.R., Geiger, P.G., and Kennedy, G.D., Cancer Letts., 2016, vol. 370, pp. 91–99. https://doi.org/10.1016/j.canlet.2015.10.014
Yin, J., Sheng, B., Han, B., Pu, A., Yang, K., Li, P., Wang, Q., Xiao, W., and Yang, H., Cell Biol. Int., 2016, vol. 40, pp. 560–568. https://doi.org/10.1002/cbin.10592
Díaz-Díaz, C.J., Ronnekleiv-Kelly, S.M., Nukaya, M., Geiger, P.G., Balbo, S., Dator, R., Megna, B.W., Carney, P.R., Bradfield, C.A., and Kennedy, G.D., Ann. Surg., 2016, vol. 264, pp. 429–436. https://doi.org/10.1097/SLA.0000000000001874
Kawajiri, K., Kobayashi, Y., Ohtake, F., Ikuta, T., Matsushima, Y., Mimura, J., Pettersson, S., Pollenz, R.S., Sakaki, T., Hirokawa, T., Akiyama, T., Kurosumi, M., Poellinger, L., Kato, S., and Fujii-Kuriyama, Y., Proc. Natl. Acad. Sci. USA, 2009, vol. 106, pp. 13 481–13 486. https://doi.org/10.1073/pnas.0902132106
Lai, D.-W., Liu, S.H., Karlsson, A.I., Lee, W.J., Wang, K.B., Chen, Y.C., Shen, C.C., Wu, S.M., Liu, C.Y., Tien, H.R., Peng, Y.C., Jan, Y.J., Chao, T.H., Lan, K.H., Arbiser, J.L., Sheu, M.L., Lai, D.W., Liu, S.H., Isabella Karlsson, A., Lee, W.J., Wang, K.B., Chen, Y.C., Shen, C.C., Wu, S.M., Liu, C.Y., Tien, H.R., Peng, Y.C., Jan, Y.J., Chao, T.H., Lan, K.H., Arbiser, J.L., and Sheu, M.-L., Oncotarget, 2014, vol. 5, pp. 7788–7804. https://doi.org/10.18632/oncotarget.2307
Yin, X.-F., Chen, J., Mao, W., Wang, Y.-H., and Chen, M.-H., Oncology Reports, 2013, vol. 30, pp. 364–370. https://doi.org/10.3892/or.2013.2410
Peng, T.L., Chen, J., Mao, W., Song, X., and Chen, M.H., BMC Cell Biology, 2009, vol. 10, 27. https://doi.org/10.1186/1471-2121-10-27
Yin, X.F., Chen, J., Mao, W., Wang, Y.H., and Chen, M.H., J. Exper. Clin. Cancer Res., 2012, vol. 31, 46. https://doi.org/10.1186/1756-9966-31-46
Su, M., Qian, C., Hu, Y., Lu, W., Huang, R., Chen, M., and Chen, J., Oncology Letts., 2017, vol. 14, pp. 8100–8105. https://doi.org/10.3892/ol.2017.7185
Andersson, P., McGuire, J., Rubio, C., Gradin, K., Whitelaw, M.L., Pettersson, S., Hanberg, A., and Poellinger, L., Proc. Natl. Acad. Sci. USA, 2002, vol. 99, pp. 9990−9995. https://doi.org/10.1073/pnas.152706299
Kuznetsov, N.V., Andersson, P., Gradin, K., Stein, P., von Dieckmann, A., Pettersson, S., Hanberg, A., and Poellinger, L., Oncogene, 2005, vol. 24, pp. 3216–3222. https://doi.org/10.1038/sj.onc.1208529
WHO, Informatsionnyy Byulleten’, Sotsial’nyye Aspekty Zdorov’ya Naseleniya, Elektronnyy Nauchnyy Zhurnal, January 2018. http://vestnik.mednet.ru/content/view/958/30/lang.ru/.
Bruera, G., Cannita, K., Giordano, A.V., Manetta, R., Vicentini, R., Carducci, S., Saltarelli, P., Iapadre, N., Coletti, G., Ficorella, C., and Ricevuto, E., BioMed Res. Int., 2014, 806 391. https://doi.org/10.1155/2014/806391
Fernandez-Salguero, P.M., Hilbert, D.M., Rudikoff, S., Ward, J.M., and Gonzalez, F.J., Toxicol. Appl. Pharmacol., 1996, vol. 140, pp. 173–179. https://doi.org/10.1006/taap.1996.0210
Lahvis, G.P. and Bradfield, C.A., Biochem. Pharmacol., 1998, vol. 56, pp. 781–787.
Mimura, J., Yamashita, K., Nakamura, K., Morita, M., Takagi, T.N., Nakao, K., Ema, M., Sogawa, K., Yasuda, M., Katsuki, M., and Fujii-Kuriyama, Y., Genes to Cells: Devoted to Molecular & Cellular Mechanisms, 1997, vol. 2, pp. 645–654.
Fan, Y., Boivin, G.P., Knudsen, E.S., Nebert, D.W., Xia, Y., and Puga, A., Cancer Res., 2010, vol. 70, pp. 212–220. https://doi.org/10.1158/0008-5472.CAN-09-3090
Jin, U.H., Lee, S.O., Pfent, C., and Safe, S., BMC Cancer, 2014, vol. 14, 498. https://doi.org/10.1186/1471-2407-14-498
Koch, D.C., Jang, H.S., O’Donnell, E.F., Punj, S., Kopparapu, P.R., Bisson, W.H., Kerkvliet, N.I., and Kolluri, S.K., Oncogene, 2015, vol. 34, pp. 6092–6104. https://doi.org/10.1038/onc.2015.55
O’Donnell, E.F., Koch, D.C., Bisson, W.H., Jang, H.S., and Kolluri, S.K., Cell Death Disease, 2014, vol. 5, e1038. https://doi.org/10.1038/cddis.2013.549
Bisson, W.H., Koch, D.C., O’Donnell, E.F., Khalil, S.M., Kerkvliet, N.I., Tanguay, R.L., Abagyan, R., and Kolluri, S.K., J. Med. Chem., 2009, vol. 52, pp. 5635–5641. https://doi.org/10.1021/jm900199u
Shah, I., Houck, K., Judson, R.S., Kavlock, R.J., Martin, M.T., Reif, D.M., Wambaugh, J., and Dix, D.J., PLoS One, 2011, vol. 6, e14584. https://doi.org/10.1371/journal.pone.0014584
Zhao, Q.W., Zhou, Y.W., Li, W.X., Kang, B., Zhang, X.Q., Yang, Y., Cheng, J., Yin, S.Y., Tong, Y., He, J.Q., Yao, H.P., Zheng, M., and Wang, Y.J., Oncology Reports, 2015, vol. 33, pp. 1621–1629. https://doi.org/10.3892/or.2015.3752
Santagata, S., Thakkar, A., Ergonul, A., Wang, B., Woo, T., Hu, R., Harrell, J.C., McNamara, G., Schwede, M., Culhane, A.C., Kindelberger, D., Rodig, S., Richardson, A., Schnitt, S.J., Tamimi, R.M., and Ince, T.A., J. Clin. Invest., 2014, vol. 124, pp. 859–870. https://doi.org/10.1172/JCI70941
Cancer Facts & Figures, American Cancer Society, 2016. https://www.cancer.org/research/cancer-facts-statistics/all-cancer-facts-figures/cancer-facts-figures-2016.html.
Lehmann, B.D., Bauer, J.A., Chen, X., Sanders, M.E., Chakravarthy, A.B., Shyr, Y., and Pietenpol, J.A., J. Clin. Invest., 2011, vol. 121, pp. 2750–2767. https://doi.org/10.1172/JCI45014
Wang, T., Gavin, H.M., Arlt, V.M., Lawrence, B.P., Fenton, S.E., Medina, D., and Vorderstrasse, B.A., Int. J. Cancer, 2011, vol. 128, pp. 1509–1523. https://doi.org/10.1002/ijc.25493
Chen, I., McDougal, A., Wang, F., and Safe, S., Carcinogenesis, 1998, vol. 19, pp. 1631–1639.
Wang, T., Wyrick, K.L., Meadows, G.G., Wills, T.B., and Vorderstrasse, B.A., Toxicol. Sci., 2011, vol. 124, pp. 291–298. https://doi.org/10.1093/toxsci/kfr247
Hall, J.M., Barhoover, M.A., Kazmin, D., McDonnell, D.P., Greenlee, W.F., and Thomas, R.S., Mol. Endocrinol., 2010, vol. 24, pp. 359–369. https://doi.org/10.1210/me.2009-0346
Hsu, E.L., Yoon, D., Choi, H.H., Wang, F., Taylor, R.T., Chen, N., Zhang, R., and Hankinson, O., Toxicol. Sci., 2007, vol. 98, pp. 436–444. https://doi.org/10.1093/toxsci/kfm125
Hanieh, H., Mohafez, O., Hairul-Islam, V.I., Alzahrani, A., Bani Ismail, M., and Thirugnanasambantham, K., PLoS One, 2016, vol. 11, e0167650. https://doi.org/10.1371/journal.pone.0167650
Jang, H.S., Pearce, M., O’Donnell, E.F., Nguyen, B.D., Truong, L., Mueller, M.J., Bisson, W.H., Kerkvliet, N.I., Tanguay, R.L., and Kolluri, S.K., Biology, 2017, vol. 6. https://doi.org/10.3390/biology6040041
Narasimhan, S., Stanford Zulick, E., Novikov, O., Parks, A.J., Schlezinger, J.J., Wang, Z., Laroche, F., Feng, H., Mulas, F., Monti, S., and Sherr, D.H., Int. J. Mol. Sci., 2018, vol. 19. https://doi.org/10.3390/ijms19051388
Boitano, A.E., Wang, J., Romeo, R., Bouchez, L.C., Parker, A.E., Sutton, S.E., Walker, J.R., Flaveny, C.A., Perdew, G.H., Denison, M.S., Schultz, P.G., and Cooke, M.P., Science, 2010, vol. 329, pp. 1345–1348. https://doi.org/10.1126/science.1191536
Casado, F.L., Singh, K.P., and Gasiewicz, T.A., Mol. Pharmacol., 2011, vol. 80, pp. 673–682. https://doi.org/10.1124/mol.111.071381
Hou, P., Li, Y., Zhang, X., Liu, C., Guan, J., Li, H., Zhao, T., Ye, J., Yang, W., Liu, K., Ge, J., Xu, J., Zhang, Q., Zhao, Y., and Deng, H., Science, 2013, vol. 341, pp. 651–654. https://doi.org/10.1126/science.1239278
Rentas, S., Holzapfel, N.T., Belew, M.S., Pratt, G.A., Voisin, V., Wilhelm, B.T., Bader, G.D., Yeo, G.W., and Hope, K.J., Nature, 2016, vol. 532, pp. 508–511. https://doi.org/10.1038/nature17665
Singh, K.P., Wyman, A., Casado, F.L., Garrett, R.W., and Gasiewicz, T.A., Carcinogenesis, 2009, vol. 30, pp. 11–19.https://doi.org/10.1093/carcin/bgn224
Prud’homme, G.J., Glinka, Y., Toulina, A., Ace, O., Subramaniam, V., and Jothy, S., PLoS One, 2010, vol. 5, e13831. https://doi.org/10.1371/journal.pone.0013831
Stanford, E.A., Wang, Z., Novikov, O., Mulas, F., Landesman-Bollag, E., Monti, S., Smith, B.W., Seldin, D.C., Murphy, G.J., and Sherr, D.H., BMC Biology, 2016, vol. 14, 20. https://doi.org/10.1186/s12915-016-0240-y
Cheng, J., Li W., Kang, B., Zhou, Y., Song, J., Dan, S., Yang, Y., Zhang, X., Li, J., Yin, S., Cao, H., Yao, H., Zhu, C., Yi, W., Zhao, Q., Xu, X., Zheng, M., Zheng, S., Li, L., Shen, B., and Wang, Y.J., Nature Commun., 2015, vol. 6, 7209. https://doi.org/10.1038/ncomms8209
Pabst, C., Krosl, J., Fares, I., Boucher, G., Ruel, R., Marinier, A., Lemieux, S., Hébert, J., and Sauvageau, G., Nature Methods, 2014, vol. 11, pp. 436–442. https://doi.org/10.1038/nmeth.2847
Kim, H.M., Kim, J.W., Choi, Y., Chun, H.S., Im, I., Han, Y.M., Song, C.W., Yoon, S., and Park, H.J., Scientific Reports, 2016, vol. 6, 216 84. https://doi.org/10.1038/srep21684
Tsai, C.-F., Hsieh, T.H., Lee, J.N., Hsu, C.Y., Wang, Y.C., Kuo, K.K., Wu, H.L., Chiu, C.C., Tsai, E.M., and Kuo, P.L., J. Agricult. Food Chem., 2015, vol. 63, pp. 10 388–10 398. https://doi.org/10.1021/acs.jafc.5b04415
Yan, B., Liu, S., Shi, Y., Liu, N., Chen, L., Wang, X., Xiao, D., Liu, X., Mao, C., Jiang, Y., Lai, W., Xin, X., Tang, C.-E., Luo, D., Tan, T., Jia, J., Liu, Y., Yang, R., Huang, J., Zhou, H., Cheng, Y., Cao, Y., Yu, W., Muegge, K., and Tao, Y., Cell Death Disease, 2018, vol. 9, 490. https://doi.org/10.1038/s41419-018-0542-9
Bambury, R.M. and Scher, H.I., Urologic Oncology: Seminars and Original Investigations, 2015, vol. 33, pp. 280−288. https://doi.org/10.1016/j.urolonc.2014.12.017
Helsen, C., Broeck, T.V., den Voet, A., Prekovic, S., Poppel, H.V., Joniau, S., and Claessens, F., Endocrine-Related Cancer, 2014, vol. 21, pp. T105−T118. https://doi.org/10.1530/ERC-13-0545
le Maire, A., Alvarez, S., Shankaranarayanan, P., Lera, A.R., de Bourguet, W., and Gronemeyer, H., Curr. Topics Med. Chem., 2012, vol. 12, pp. 505–527.
McDonnell, D.P. and Wardell, S.E., Curr. Opinion Pharmacol., 2010, vol. 10, pp. 620–628. https://doi.org/10.1016/j.coph.2010.09.007
Morrow, D., Qin, C., Smith, R., and Safe, S., J. Steroid Biochem. Mol. Biol., 2004, vol. 88, pp. 27–36. https://doi.org/10.1016/j.jsbmb.2003.10.005
Hrubá, E., Vondráček, J., Líbalová, H., Topinka, J., Bryja, V., Souček, K., and Machala, M., Toxicol. Letts., 2011, vol. 206, pp. 178–188. https://doi.org/10.1016/j.toxlet.2011.07.011
Yu, J.-S., Leng, P.-F., Li, Y.-F., Wang, Y.-Q., Wang, Y., An, R.-H., and Qi, J.-P., DNA Cell Biology, 2017, vol. 36, pp. 1010–1017. https://doi.org/10.1089/dna.2017.3783
Ide, H., Lu, Y., Yu, J., Noguchi, T., Kanayama, M., Muto, S., Yamaguchi, R., Kawato, S., and Horie, S., Human Cell, 2017, vol. 30, pp. 133–139. https://doi.org/10.1007/s13577-016-0158-2
Sun, F., Indran, I.R., Zhang, Z.W., Tan, M.H.E., Li, Y., Lim, Z.L.R., Hua, R., Yang, C., Soon, F.F., Li, J., Xu, H.E., Cheung, E., and Yong, E.L., Carcinogenesis, 2015, vol. 36, pp. 757–768. https://doi.org/10.1093/carcin/bgv040
Zhu, M., Wu, J., Ma, X., Huang, C., Wu, R., Zhu, W., Li, X., Liang, Z., Deng, F., Zhu, J., Xie, W., Yang, X., Jiang, Y., Wang, S., Geng, S., Xie, C., and Zhong, C., Toxicol In Vitro, 2019, vol. 54, pp. 82–88. https://doi.org/10.1016/j.tiv.2018.09.007
Iida, K., Mimura, J., Itoh, K., Ohyama, C., Fujii-Kuriyama, Y., Shimazui, T., Akaza, H., and Yamamoto, M., J. Biochem., 2010, vol. 147, pp. 353–360. https://doi.org/10.1093/jb/mvp169
Luzzani, G.A., Callero, M.A., Kuruppu, A.I., Trapani, V., Flumian, C., Todaro, L., Bradshaw, T.D., and Loaiza Perez, A.I., J. Cell. Biochem., 2017, vol. 118, pp. 4526−4535. https://doi.org/10.1002/jcb.26114
Gu, A., Ji, G., Jiang, T., Lu, A., You, Y., Liu, N., Luo, C., Yan, W., and Zhao, P., Toxicol. Sci., 2012, vol. 128, pp. 357–364. https://doi.org/10.1093/toxsci/kfs158
Guastella, A.R., Michelhaugh, S.K., Klinger, N.V., Fadel, H.A., Kiousis, S., Ali-Fehmi, R., Kupsky, W.J., Juhász, C., and Mittal, S., J. Neuro-Oncology, 2018, vol. 139, pp. 239–249. https://doi.org/10.1007/s11060-018-2869-6
Procházková, J., Strapáčová, S., Svržková, L., Andrysík, Z., Hýžďalová, M., Hrubá, E., Pěnčíková, K., Líbalová, H., Topinka, J., Kléma, J., Espinosa, J.M., Vondráček, J., and Machala, M., Toxicol. Letts., 2018, vol. 292, pp. 162–174. https://doi.org/10.1016/j.toxlet.2018.04.024
Cheng, Y.H., Huang, S.C., Lin, C.J., Cheng, L.C., and Li, L.A., Toxicol. Appl. Pharmacol., 2012, vol. 259, pp. 293–301. https://doi.org/10.1016/j.taap.2012.01.005
Chang, J.T., Chang, H., Chen, P.H., Lin, S.L., and Lin, P., Clin. Cancer Res., 2007, vol. 13, pp. 38–45. https://doi.org/10.1158/1078-0432.CCR-06-1166
Duan, Z., Li Y., and Li, L., Mol. Cell. Biochem., 2018, https://doi.org/10.1007/s11010-018-3323-y
Ye, M., Zhang, Y., Gao, H., Xu, Y., Jing, P., Wu, J., Zhang, X., Xiong, J., Dong, C., Yao, L., Zhang, J., and Zhang, J., Clinical Cancer Research, 2018, vol. 24, pp. 1227–1239. https://doi.org/10.1158/1078-0432.CCR-17-0396
Bunaciu, R.P. and Yen, A., Cancer Res., 2011, vol. 71, pp. 2371–2380. https://doi.org/10.1158/0008-5472.CAN-10-2299
Hayashibara, T., Yamada, Y., Mori, N., Harasawa, H., Sugahara, K., Miyanishi, T., Kamihira, S., and Tomonaga, M., Biochem. Biophys. Res. Commun., 2003, vol. 300, pp. 128–134.
Mulero-Navarro, S., Carvajal-Gonzalez, J.M., Herranz, M., Ballestar, E., Fraga, M.F., Ropero, S., Esteller, M., and Fernandez-Salguero, P.M., Carcinogenesis, 2006, vol. 27, pp. 1099–1104. https://doi.org/10.1093/carcin/bgi344
To, K.K.W., Yu, L., Liu, S., Fu, J., and Cho, C.H., Molecular Carcinogenesis, 2012, vol. 51, pp. 449–464. https://doi.org/10.1002/mc.20810
Villard, P.H., Caverni, S., Baanannou, A., Khalil, A., Martin, P.G., Penel, C., Pineau, T., Seree, E., and Barra, Y., Biochem. Biophys. Res. Commun., 2007, vol. 364, pp. 896–901. https://doi.org/10.1016/j.bbrc.2007.10.084
Jin, U.-H., Park, H., Li, X., Davidson, L.A., Allred, C., Patil, B., Jayaprakasha, G., Orr, A.A., Mao, L., Chapkin, R.S., Jayaraman, A., Tamamis, P., and Safe, S., Toxicol. Sci., 2018, vol. 164, pp. 205–217. https://doi.org/10.1093/toxsci/kfy075
Alzahrani, A.M., Hanieh, H., Ibrahim, H.-I.M., Mohafez, O., Shehata, T., Bani Ismail, M., and Alfwuaires, M., Int. Immunopharmacol., 2017, vol. 52, pp. 342–351. https://doi.org/10.1016/j.intimp.2017.09.015
Becker, R.A., Patlewicz, G., Simon, T.W., Rowlands, J.C., and Budinsky, R.A., Regulatory Toxicol. Pharmacol., 2015, vol. 73, pp. 172–190. https://doi.org/10.1016/j.yrtph.2015.06.015
de Tomaso Portaz, A.C., Caimi, G.R., Sánchez, M., Chiappini, F., Randi, A.S., Kleiman de Pisarev, D.L., and Alvarez, L., Toxicology, 2015, vol. 336, pp. 36–47. https://doi.org/10.1016/j.tox.2015.07.013
Yamaguchi, M. and Hankinson, O., Int. J. Oncol., 2018, vol. 53, pp. 1657–1666. https://doi.org/10.3892/ijo.2018.4507
Tian, W., Fu, H., Xu, T., Xu, S.L., Guo, Z., Tian, J., Tao, W., Xie, H.Q., and Zhao, B., Environmental Pollution (Barking, Essex: 1987), 2018, vol. 237, pp. 508–514. https://doi.org/10.1016/j.envpol.2018.02.079
O’Donnell, E.F., Jang, H.S., Pearce, M., Kerkvliet, N.I., and Kolluri, S.K., Oncotarget, 2017, vol. 8, pp. 25 211–25 225. https://doi.org/10.18632/oncotarget.16056
Schreck, I., Deigendesch, U., Burkhardt, B., Marko, D., and Weiss, C., Arch. Toxicol., 2012, vol. 86, pp. 625–632. https://doi.org/10.1007/s00204-011-0781-3
Weiss, C., Faust, D., Dürk, H., Kolluri, S.K., Pelzer, A., Schneider, S., Dietrich, C., Oesch, F., and Göttlicher, M., Oncogene, 2005, vol. 24, pp. 4975–4983. https://doi.org/10.1038/sj.onc.1208679
Harrill, J.A., Parks, B.B., Wauthier, E., Rowlands, J.C., Reid, L.M., and Thomas, R.S., Hepatology, 2015, vol. 61, pp. 548–560. https://doi.org/10.1002/hep.27547
Kennedy, G.D., Nukaya, M., Moran, S.M., Glover, E., Weinberg, S., Balbo, S., Hecht, S.S., Pitot, H.C., Drinkwater, N.R., and Bradfield, C.A., Toxicol. Sci., 2014, vol. 140, pp. 135–143. https://doi.org/10.1093/toxsci/kfu065
Volkov, M.S., Bolotina, N.A., Evteev, V.A., and Koblyakov, V.A., Biochemistry (Moscow), 2012, vol. 77, pp. 248–255. https://doi.org/10.1134/S0006297912020125
Kolluri, S.K., Balduf, C., Hofmann, M., and Göttlicher, M., Cancer Res., 2001, vol. 61, pp. 8534–8539.
Pesatori, A.C., Consonni, D., Rubagotti, M., Grillo, P., and Bertazzi, P.A., Environmental Health, 2009, vol. 8, 39. https://doi.org/10.1186/1476-069X-8-39
Fukasawa, K., Kagaya, S., Maruyama, S., Kuroiwa, S., Masuda, K., Kameyama, Y., Satoh, Y., Akatsu, Y., Tomura, A., Nishikawa, K., Horie, S., and Ichikawa, Y., Mol. Cancer Ther., 2015, vol. 14, pp. 343–354. https://doi.org/10.1158/1535-7163.MCT-14-0158
D’Amato, N.C., Rogers, T.J., Gordon, M.A., Greene, L.I., Cochrane, D.R., Spoelstra, N.S., Nemkov, T.G., D’Alessandro, A., Hansen, K.C., and Richer, J.K., Cancer Res., 2015, vol. 75, pp. 4651–4664. https://doi.org/10.1158/0008-5472.CAN-15-2011
Li, Z.D., Wang, K., Yang, X.W., Zhuang, Z.G., Wang, J.J., and Tong, X.W., Int. J. Clin. Exper. Pathol., 2014, vol. 7, pp. 7931–7937.
Brinkman, A.M., Wu, J., Ersland, K., and Xu, W., BMC Cancer, 2014, vol. 14, 344. https://doi.org/10.1186/1471-2407-14-344
Tomblin, J.K. and Salisbury, T.B., Biochem. Biophys. Res. Commun., 2014, vol. 443, pp. 1092–1096. https://doi.org/10.1016/j.bbrc.2013.12.112
Zhang, S., Kim, K., Jin, U.H., Pfent, C., Cao, H., Amendt, B., Liu, X., Wilson-Robles, H., and Safe, S., Mol. Cancer Ther., 2012, vol. 11, pp. 108–118. https://doi.org/10.1158/1535-7163.MCT-11-0548
Hýžd’alová, M., Pivnicka, J., Zapletal, O., Vázquez-Gómez, G., Matthews, J., Neca, J., Pencíková, K., Machala, M., and Vondrácek, J., Toxicol. Sci., 2018, https://doi.org/10.1093/toxsci/kfy153
Vacher, S., Castagnet, P., Chemlali, W., Lallemand, F., Meseure, D., Pocard, M., Bieche, I., and Perrot-Applanat, M., PLoS One, 2018, vol. 13, e0190619. https://doi.org/10.1371/journal.pone.0190619
Al-Dhfyan, A., Alhoshani, A., and Korashy, H.M., Molecular Cancer, 2017, vol. 16, 14. https://doi.org/10.1186/s12943-016-0570-y
Saito, R., Miki, Y., Hata, S., Ishida, T., Suzuki, T., Ohuchi, N., and Sasano, H., Breast Cancer Research Treatment, 2017, vol. 161, pp. 399–407. https://doi.org/10.1007/s10549-016-4063-x
Author information
Authors and Affiliations
Corresponding author
Additional information
Translated by A. Medvedev
Rights and permissions
About this article
Cite this article
Vorontsova, J.E., Cherezov, R.O., Kuzin, B.A. et al. Aryl-Hydrocarbon Receptor as a Potential Target for Anticancer Therapy. Biochem. Moscow Suppl. Ser. B 13, 36–54 (2019). https://doi.org/10.1134/S1990750819010116
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1990750819010116