Abstract
Peanut (Arachis hypogaea L.) is a geocarpic plant, which absorbs nutrients not only through roots but also through pods. However, the effect of nitrogen fertilizer application in pod area on pod gene expression and peanut yield is unknown. In this study, we determined the pod yield, dry matter accumulation, nitrogen accumulation in different organs under different nitrogen treatment in root zone and pod zone. The results showed that pod area application of nitrogen led to significantly higher pod yield to compare with the control. The application of 60 kg/hm2 nitrogen at the pod area led to the highest pod yield, full fruit rate and plant nitrogen content. 15N directly absorbed by pods was mainly accumulated in the shell at early stage of pods development, and then transferred to seeds at later period. However, the ratio of 15N absorbed directly by pods from soil was low, only about 10%. To understand the underlying molecular mechanisms, immature pods of different developmental stages were collected for gene expression analysis. The expression levels of several genes encoding ABC type transporter family, nitrate reductase, nitrite reductase and glutamine synthetase increased in different stages of pod development upon pod area nitrogen application. These results indicated that the efficiency of N assimilation and glutamate metabolic cycle in pods was increased under pod nitrogen application. The expression levels of several genes in gibberellin and brassinolide biosynthesis pathways were also up-regulated, suggesting that these two hormones were involved in the promoting effect of pod nitrogen application on pod growth.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
INTRODUCTION
Peanut is one of the important oil crops in the world. Nitrogen (N) fertilizer as base fertilizer is an important measure to obtain high yield in peanut production [1]. Nitrogen sources of peanut include soil, fertilizer and nodule nitrogen fixation. Recent studies have shown that peanut nodule fixed N can only provide 40 to 50% of the N required for peanut growth, which indicates that more than half of the N required for peanut is from soil and fertilizer [2, 3]. Nitrogen affects the growth and development of various organs of peanut, and further affects the yield and quality. In order to increase the yield, more and more N fertilizer is applied in peanut production. In a certain range, total N uptake and N use efficiency (NUE) of peanut increased with the increased level of N application, however, NUE decreased when the N level is too high [4].
NUE is related to N uptake efficiency (NUpE; above-ground N uptake per unit N available) and N‑utilization efficiency (NUtE; grain dry matter yield per unit above-ground N uptake). Nitrate nitrogen (\({\text{NO}}_{3}^{ - }\)) and ammonium nitrogen (\({\text{NH}}_{4}^{ + }\)) are the two main forms of N uptake by roots and utilization by plants. The most used N fertilizers in the modern agriculture was urea [CO(NH2)2] because of the low price and the high N content. Urea can be directly absorbed by plant, part of it is rapidly broken down into ammonium by urease, which is in turn converted into nitrate nitrogen in soils [5]. Nitrogen assimilation process first requires specific transporters including nitrate transporter family (NRT), the chloride channel family (CLC), ammonium transporters (AMTS)and tonoplast intrinsic proteins (TIPs) for N uptake from the surrounding environment [6–9]. In plant cells, nitrate is reduced by nitrate reductase (NR) and nitrite reductase (NiRA) to ammonia, and more than 95% of \({\text{NH}}_{4}^{ + }\) was then incorporated into glutamine synthetase/glutamate synthetase (GS/GOGAT) pathway to generate glutamine, which then transferred into other amino acids or biomolecules [10–12].
More and more N fertilizer is used in agriculture. The excess N could be converted to N oxide gases that contribute to global warming, and it could also cause water eutrophication, global acidification and stratospheric ozone loss [13]. Improving NUE in crops is critical for sustainable agriculture. Different from most of other crops, peanut has a unique feature of that flowering and fertilization are accomplished above ground, while the fruit formation is underground [14]. In the process of pod expansion, nutrients are absorbed not only through the root, but also through the pod, providing more nutrients for the peanut [15]. The 45Ca labeling experiments revealed that during pod development, 90% of calcium is absorbed directly by pods from the soil. Sufficient calcium in the pod area is very important for pod development, yield formation and nutritional quality. Calcium content of the pod increased sharply in the early stage of pod development. If calcium supply in pod area was insufficient 20–30 d after the gynophore buried in the soil, the calcium content of pods would decrease, and eventually the pod size and dry weight decreased significantly [16, 17]. However, whether pod plays critical roles in nitrogen absorption is unknown. In this project, 15N labeled nitrogen fertilizer was applied in pod area which was separated from the roots. The pod yield, dry matter accumulation, total nitrogen of different organs and pod gene expression were investigated upon pod area nitrogen application. The expression of genes related to nitrogen absorption, metabolism, hormone synthesis and signal transduction were affected, suggesting the global changes after pod area nitrogen application, and eventually improved pod yield. This study provided important information for efficient N fertilizer application during peanut growth.
MATERIALS AND METHODS
Plant growth conditions and sampling methods. The cultivated peanut ‘Fenghua 1’ (FH1) was selected as the experimental material. The experiment was conducted in pot culture in experimental station of Shandong Academy of Agricultural Sciences from May to September of 2019. The size of the pot was 40 cm in inner diameter and 35 cm in depth. Each pot was filled with 25 kg of air-dried soil, the basic nutrient contents of the soil were as follows: organic matter 12.36 g/kg, total nitrogen 1.08 g/kg, alkali-hydrolyzed nitrogen 89 mg/kg, available phosphorus 45.46 mg/kg, and available potassium 98.83 mg/kg. Three seeds were planted in the center of the pot, and two plants were final singling one week after the seedlings emerged. Before the peanut flowering, the designed plastic box is put into the pot, and filled with clean sand. The peanut gynophores sand pods could grow with in the box, so that the root and pod could grow under different nutritional environments. In the root zone, two nitrogen application levels were set, the low (60 kg/hm2 of urea) and the high (120 kg/hm2 of urea) levels, and the levels of 90 kg/hm2 and 150 kg/hm2 P2O5 and K2O were applied to the root zone soil. The nitrogen level in the pod area was 0, 30, 60 and 90 kg/hm2, respectively. Ten pots for each treatment. The nitrogen source was 15N labeled urea with 15N abundance of 10.12% and nitrogen content was 46.03%, purchased from Shanghai Research Institute of Chemical Industry. Nitrogen fertilizer was applied to the root zone before sowing and pod zone fertilizer was applied before the gynophores growing into soil. During the growth period, referring to the method of Inanaga et al. [15], nutrient solution was applied in the pod area, and the nutrient composition was: 0.6 g P as K2HPO4 and KH2PO4, 0.8 g Ca as CaCl2, 0.5 g Mg as MgSO4, 2 mg Bas H3BO3, 2 mg Mn as MnCl2·4H2O, 0.3 mg Cu as CuSO4·5H2O, 0.4 mg Zn as ZnSO4·7H2O, 0.3 mg Mo as (NH4)6Mo7O24·4H2O, 0.4 mg Fe as EDTA-Fe per pot. The peanut sowed on May 15 and harvested on September 20.
Recorded the date when the gynophores were buried into the soil. Samples were collected from the whole plant according to stem, leaf, gynophores, fruit shell and seeds at three periods: the pod expanding stage (PES, 20 d after soil penetration), the pod filling stage (PFS, 40 d after insertion) and the harvesting stage (HS, 60 d after soil penetration). Samples were fixing dried at 105°C for 0.5 h and then fully dried at 80°C before weighted. The dry weight, plant nitrogen content and 15N abundance were determined. The nitrogen content was measured by Kjeltec 2300 automatic nitrogen determination instrument, and 15N abundance was measured by Isoprime-100 stable isotope analyzer.
To study the effect of nitrogen application on changes of gene expression in pods development and nitrogen absorption and utilization, nitrogen fertilizer was applied to the pod zone when the gynophores were buried into the soil. The pure nitrogen concentrations were 0 (FHO) and 60 kg/hm2 (FHN) for the control and the treated group. Nutrient solution was applied at the same time. The pods were sampled at 10 d (S1), 20 d (S2) and 30 d (S3) after gynophores soil penetration.
Calculation method for nitrogen uptake in root and pod zones. Nitrogen accumulation amount per plant, (NAA, g/plant) = ∑ Nitrogen content of each organs (%) × dry matter accumulation (g), the Nitrogen content was converted into nitrogen elements from urea;
15N atom % excess = 15N abundance – 15N natural abundance;
The ratio of nitrogen accumulation from labeled nitrogen (N proportion derived from pod zone, NPDF, %) = 15N atom % excess in plant sample/15N atom % excess in fertilizer × 100;
Plant 15N accumulation (N amount derived from fruiting zone, NADF, g/plant) = total nitrogen accumulation (g/plant) × NPDF%.
Statistical analyses. The statistical analysis and independent t-test was performed using SPSS, version 17.0 software (SPSS, United States). When three or more groups were compared, a one-way analysis of variance (ANOVA) was performed, followed by Tukey’s multiple-comparison test, with the comparisons between different treatments considered significantly different at a P-value of 0.05. Then SigmaPlot 14.0 was used to make diagrams.
RNA extraction, cDNA synthesis and high-throughput sequencing. Total RNA was extracted from seeds using Trizol Reagent (TaKaRa, China) according to the manufacturer’ s instructions. RNA samples were treated with DNase I to remove genomic DNA contamination. RNA quality and quantity were analyzed using Agilent 2100 and NanoDrop. mRNA was enriched using magnetic beads with Oligo (dT) and cleaved into short fragments (~200 nt) in fragmentation buffer. The reverse transcription was conducted with random hexamer primer and then the second strand cDNA was synthesized. After end repair, the 5' tails were phosphorylated, the 3' tails were added with an adenine. Sequencing adaptors were ligated to the double-stranded DNA fragments. Then the fragments were amplified by PCR to construct cDNA library. The library was sequenced using BGISEQ-500 platform by Beijing Genomics Institute (BGI).
Bioinformatics analysis of RNA sequence data. Raw reads were generated from each cDNA library. To obtain clean reads, the adaptor sequences, low-quality reads and reads containing more than 5% unknown bases were removed using SOAPunke and trimmomatic software. Clean reads were mapped to the reference genome of Arachis hypogaea ‘Tifrunner’ (https://www.peanutbase.org/data/public/Arachis_hypogaea/) using HISAT2 program [18]. The clean reads were aligned to reference gene by Bowtie2 (RNA-Seq by Expectation Maximization) [19]. The statistical analyses of randomness, degree of coverage, and sequencing saturation were also accomplished. The gene expression level was calculated with RSEM method and normalized to FPKM (Fragments per Kb per million reads) [20]. The relative gene expression level between two samples was counted by log2 ratio. Pearson’s correlation coefficient between every two samples and principal component analysis (PCA) were performed. Differentially expressed genes (DEGs) were identified using DEGseq2 method and screened with the criteria of fold change ≥2 and Q-value ≤0.001 [21]. Gene Ontology (GO) annotation was carried out by Blast2GO program through comparing DEGs with GO terms in the GO database and GO functional classification was performed using WEGO software. KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway analysis was conducted by mapping DEGs to KEGG database. The GO and KEGG functional enrichment were performed using hypergeometric test. The P-value formula in hyper geometric test can be acquired from https://en.wikipedia.org/wiki/Hypergeometric distribution in detail. FDR (False Discovery Rate) correction of all p-values was conducted. The GO terms and KEGG pathways whose FDR ≤0.01 were defined as significant enriched. BLASTX (E <0.00001) against NCBI Nr database was carried out.
The RNA-seq data in this study were available at NCBIBioProject: PRJNA764875 (https://www.ncbi. nlm.nih.gov/bioproject/PRJNA764875).
Verification of RNA-seq data by qRT-PCR. qRT-PCR was used to verify the expressional levels of 17 selected genes. RNA samples were those used for high-throughput sequencing and the reverse transcription was performed using PrimeScript II 1st Strand cDNA Synthesis Kit (TaKaRa). The gene-specific primers were designed using PerlPrimer software and were listed in Additional file 8: Table S7. qRT-PCR reaction was performed on ABI7500 Real Time System (Applied Biosystems) using TB Green™ Premix Ex Taq™ II (TaKaRa). The parameters of thermal cycle were 94°C for 10 min, followed by 40 cycles of 94°C for 15 s and 60°C for 1 min in a 20 µL volume. Three biological replications were performed for each reaction with actin gene as internal reference. The relative expressional level of each gene between FHN and FHO was calculated by 2–ΔΔCt method.
RESULTS
Nitrogen Fertilizer Application in Pod Area Increased Peanut Yield and Dry Matter Accumulation of the Plants
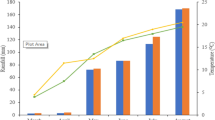
In the case of applying low nitrogen (LN) to the root zone, with the increase of N application in the pod area, the peanut pod yield was increased significantly. However, there was no significant difference among three different concentrations of pod N application. In the case of applying high nitrogen (HN) to the root zone, the pod yield was increased significantly after pod area N application. When 60 kg/hm2 nitrogen was applied in the pod zone, the maximum yield was reached (Fig. 1). There was no significant difference among three different concentration of pod N application. At root zone HN condition, the yield increase of pod N application was lower than that of root zone LN condition.
Pod yield per plant under different nitrogen application ate in root and pod zone. LN—Low nitrogen of 60 kg/hm2 in the root zone; HN—High nitrogen of 120 kg/hm2 in the root zone; N0, N30, N60, N90: Nitrogen application amount of 0, 30, 60, 90 kg/hm2 in pod zone. The same designations are below. (1) N0; (2) N30; (3) N60; (4) N90.
The whole peanut plants dry matter accumulation was also increase upon pod area N application (Fig. 2). At root zone LN condition, pod zone N application significantly promoted dry matter accumulation of the whole plant, with concentration dependent manner. At pod filling stage (PFS) and harvesting stage (HS) developmental stages, the dry matter accumulation was much higher than that of the pod expanding stage (PES) stage (Fig. 2). Similar results were observed at root zone HN condition. However, at root zone HN, high N application (90 kg/hm2) in pod zone resulted in a lower dry matter to compare the medium level of N application (60 kg/hm2), especially in HS stage. For example, in HS stage, at root zone LN condition, the dry matter of high N application (90 kg/hm2) was 0.46 g higher than medium N application (60 kg/hm2). However, at root zone HN condition, the dry matter of high N application (90 kg/hm2) was 0.73 g lower than medium N application (60 kg/hm2). The contribution of N application in pod area to dry matter accumulation of HN was smaller than that of LN. In conclusion, the effect of nitrogen supplementation in pod zone was more significant when there was insufficient N in root zone, while excessive N application was not conducive to dry matter accumulation when there was sufficient N in root zone.
Nitrogen Absorption and Distribution of Different Organs with Pod Zone N Application
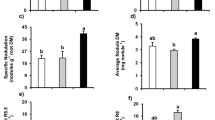
Nitrogen application in pod area increased the N content of total plant Table S1Fig 3). At PES, the main organ of nitrogen accumulation was the leaves and the N ratio in leaves was 49.1 ~ 50.6%. AtPFS, the N accumulation ratio in leaf and seeds were both high. AtHS, the nitrogen distribution ratio in stem or in leaves decreased rapidly, and a large amount of nitrogen flows to the seeds. At HS, under root LN and HN conditions, pod area N application led to 19.53–33.24% and 17.31 ~ 22.51% increasement of total plant nitrogen accumulation, respectively. It was obvious that the increased range under root LN condition was significantly greater than that of root HN condition. It was suggested that the N application in pod zone could in some extent compensate the deficiency of nitrogen in the root zone.
Nitrogen or 15N accumulation in different organs of peanut under different nitrogen application rate in root and pod zone (mg/plant). LNN—Nitrogen content under LN treatment; LN15N—15N content under LN treatment; HNN—Nitrogen content under HN treatment; HN15N—15N content under HN treatment. (1) N0; (2) N30; (3) N60; (4) N90.
Nitrogen uptake by pods was tracked by 15N in plants. The total absorbed amount of 15N per plant increased with the increase of N application rate in pod area (Table S2; Fig 3). AtPES and PFS, the total 15N uptake under LN60 and LN90 condition was higher than under HN condition. It suggested that the available N at LN condition was insufficient during these two developmental stages, and therefore the pod absorbed N was important for pod development. At HS, the 15N accumulation per plant under LN was lower than under HN, mainly due to the reduced biomass of plants under LN.
15N allocation proportion at different organs was researched. In PES, 15N directly absorbed by pods was mainly accumulated in the shell, and then transferred to seeds in PFS and HS periods (Table 1). The ratio of 15N in leaves under LN was higher than those under HN, indicated that when the root absorbed insufficient nitrogen, the pod absorbed nitrogen could distribute more to vegetative organs. However, during PFS and HS stages, when active biosynthesis of seed storage substances occurred, the proportion of nitrogen in shells decreased while the proportion of nitrogen in seeds increased gradually. At PES, the early filling stage of pods, the leaves shared more pod absorbed nitrogen, while significantly less was shared by leaves at HS stage, indicating that the pod absorbed nitrogen could be distributed according to the needs of different organs. The proportion of 15N distributed in seeds under HN was higher than that under LN, however, the proportion of 15N distributed in leaves and stems under HN was lower than that under LN, indicating that the plant could adjust the nitrogen supply when the nitrogen absorbed from root zone was insufficient, and the nitrogen absorbed by pod should first meet the needs of vegetative organs such as stem and leaf. At HS, 73.08~77.01% of pod absorbed 15N under LN condition was kept in pods, while 79.83~82.98% of pod absorbed 15N under HN condition remained in pods. In general, the pod absorbed N accounted a small portion (about 10% under LN90) of the total N absorbed by the plant at HS.
Differentially Expressed Genes between FHN and FHO
In order to elucidate the mechanisms underlie yield improvement by pod area N application, gene expression of the immature pods was analyzed. Nitrogen application in pod zone significantly increased fruit size, including the length, the width and the fresh weight of the pods in different stages (Fig. 4). Pods of FH1 after buried in soil for 10, 20 and 30 d were collected for total RNA extraction. Eighteen cDNA libraries were constructed and sequenced by BGISEQ-500 (Table 2). A total of 0.396 Gb raw reads were generated, and the average output data per sample was 22.02 Mb. After eliminating the low-quality sequences, the average clean reads of each library was 21.62 Mb, and the average clean reads ratio was 98.19%. About 20.93 Mb and 16.84 Mb clean reading in each library matched perfectly with reference genome and gene sets, with the average alignment rate of 96.82 and 77.90%, respectively.
Pod size at different development stages of nitrogen application in pod zone. N60—60 kg hm–2 nitrogen application; N0—0 kg hm–2 nitrogen application; S1—Pods of 10 days after the gynophores putted into the soil; S2—Pods of 20 days after the gynophores putted into the soil; S3—Pods of 30 days after the gynophores putted into the soil. Significant differences between treatments were indicated by one asterisk (P < 0.05) or two asterisks (P < 0.01). (1) N0; (2) N60
A total of 58443 genes were detected, most of the transcripts were completely covered, and the reads were evenly distributed in each region of the transcripts. Comparing gene expression during three developmental stages of FHN and FHO immature pods, 3918, 5943 and 2399 DEGs were identified, respectively (Tables S3–S5). At stage one, 1256 up-regulated genes and 2662 down-regulated genes in FHN were identified to compare with FHO. At stage two, 4724 up-regulated genes and 1219 down-regulated genes were identified in FHN to compare with FHO. Similarly, we found 1222 up-regulated genes and 1177 down-regulated genes in FHN to compare with FHO at stage three. Compared with FHO1, 1256 genes were up-regulated and 2662 genes were down regulated in FHN1; Compared with FHO2, the number of up-regulated and down regulated genes in FHN2 was 4724 and 1219, respectively; Compared with FHO3, the number of up-regulated and down regulated genes in FHN3 was 1222 and 1177, respectively (Fig. 5).
Differentially expressed genes of immature pods at three stages under nitrogen treatment in pod zone. (a) The number of differentially expressed genes in three developmental stages; (1) up-regulated; (2) down-regulated. (b) Venn diagram showed the common and specific differentially expressed genes in three developmental stages.
Functional Analysis of Differentially Expressed Genes
GO classification and KEGG pathway analysis of DEGs were carried out to further understand the molecular mechanisms of pod N application on yield. go classification showed that there were 1577, 2649 and 1086 DEGs in the three stages (S1, S2 and S3) of immature pods under nitrogen treatment. GO analysis divided these DEGs into three categories: biological process, cell composition and molecular function. In terms of molecular function, catalytic activity and binding were the most abundant classes; in terms of cell composition, membrane, cell, membrane part and organelle were the main classes; in terms of biological process, metabolic process and cellular process were the most abundant classes.
KEGG enrichment pathway analysis was carried out for the DEGs identified in this study (Fig. 6; Table S6). In S1, MAPK signal transduction pathway, plant hormone signal transduction and biosynthesis pathway were more abundant, indicating the active response to environmental stimuli. Several pathways related to the synthesis of secondary metabolites, sugars and lipids were enriched. For the secondary metabolism, we found that phenylpropane biosynthesis, flavonoid biosynthesis, isoflavonoid biosynthesis were enriched. For sugar metabolism, the results showed that pentose and glucuronide conversion, starch and sucrose metabolism, polysaccharides degradation, and galactose metabolism were enriched. For lipid metabolism, the enriched pathways were ether lipid metabolism, butanote metabolism, glyoxylate and dicarboxylate metabolism, ketone bodies synthesis and degradation, fatty acid degradation, linoleic acid metabolism, and C5-branched dibasic acid metabolism. At S2, the plant hormone signaling pathway, secondary metabolites, carbohydrate and lipid synthesis pathways were all enriched. In addition, photosynthesis, amino acid biosynthesis, nucleic acid biosynthesis pathways were enriched. In S3, in addition to the above-mentioned pathways, the stilbenoid biosynthesis, diarylheptanoid, gingerol and selenocompound metabolism pathway were enriched.
The first 20 KEGG enrichment pathways of DEGs in immature pods at three developmental stages under urea treatment. S1, S2 and S3 represent the three developmental stages of pods after 10, 20 and 30 days respectively. The x-axis represents the ratio of gene enrichment, that is, the ratio of the number of differential genes in this pathway to the number of specific genes annotated. The size of the bubble indicates the number of annotated genes in the KEGG pathway. The color represents the enriched Q-value. The darker the color, the smaller the Q-value.
DEGs Involved in Nitrogen Absorption and Metabolism
Plants absorbed urea partly by urea transporters, two types of plant membrane transporters, major intrinsic proteins (MIPs) and high-affinity urea transporter (DUR3). However, the expression levels of MIP and DUR3 genes were low and no obvious changed in peanut pods. ABC transporter is a kind of urea transporters which can consumes energy and transport urea against a concentration gradient in bacteria. We found that 48 ABC transporter genes were expressed in pods, and some of them were highly expressed in S2 (Table 3). The expressions of 25 ABC transporter family genes were significantly higher in FHN2 than that in FHO2. For example, the expression of ABC transporter A family member 7-like (HBIL5H) in FHN2 was 41.51 times higher than that in FHO2. In addition, the expressions of 10 ABC-type transporter genes were lower in FHN1 than that in FHO1, while their expressions were higher in FHN3 than that in FHO3. The differential expression of these genes suggests that pod zone N application active these special group of ABC transporter family genes, which may play roles in pod urea transport and utilization. Aquaporin may also play roles in pod urea absorption. Our results showed that upon pod N application, the expression of most aquaporin in FHN1 was lower than FHO1, however, their expression level increased significantly in FHN2 to compare with FHO2. For example, the expression of aquaporin TIP1-3-like in FHN2 was 4.81 times higher than that in FHO2.
Urea could be degraded to CO2 and NH3 by the function of urease in soil, which then absorbed by plants then in the form of ammonium nitrogen.GS/GOGAT cycleis the main pathway of ammonium assimilation in organisms. GS catalyzes the synthesis of glutamine from ammonium ion and glutamate, and then receives the carbon skeleton from α-ketoglutarate to produce glutamate under the action of glutamate synthase (GOGAT). The glutamate is converted into ammonia through the action of glutamate dehydrogenase (GDH), forming a complete glutamate cycle. GS is the rate limiting enzyme of this cycle. Under pod N application, among the 12 annotated GS genes, most of them did not change obviously, only two of them showed higher expression level in FHN3 to compare with FHO3. The expression of one GOGAT gene in FHN1 and four GDH in FHN2 was up-regulated to compare with FHO1 and FHO2, respectively. The results showed that the efficiency of \({\text{NH}}_{4}^{ + }\) assimilation and glutamate metabolic cycle increased upon pod N application. Nitrate nitrogen is another form of nitrogen absorption by plants. After pod zone urea application, the expression of nitrate transporter genes in pods changed slightly. The expression of three high affinity nitrate transporter 2.5 genes in FHN1 and FHN2 were two times higher than that in FHO1 and FHO2, while no significant difference was observed between FHN3 and FHO3. Nitrate reductase (NR), nitrite reductase (NiRA) and glutamine synthetase (GS) are important components of \({\text{NO}}_{3}^{ - }\) assimilation in plants, they are the rate limiting factor in nitrate reduction pathway. NR convert nitrate to nitrite and NiRA degraded nitrite to \({\text{NH}}_{4}^{ + }\) the expressions of one NR gene in FHN1, FHN2, FHN3 were 7.77, 2.37 and 15.19 times higher than that in three corresponding stages of FHO, respectively. One NIRA gene in FHN1, FHN2, FHN3 was 2.54, 4.40 and 28.57 times higher than that in three corresponding stages of FHO, respectively.
DEGs Related to Hormone Synthesis and Signal Transduction
Gibberellin and brassinolide play key roles in fruit enlargement. In the gibberellin synthesis pathway, the expression levels of two ent-kaurenoic acid oxidase (KAO1) genes in FHN2 were 6.14 and 12.28 times higher than that in FHO2 (Table 4). One CPS gene which coding ent-copalyl diphosphate synthase was strongly induced by pod N application, the expression level in FHN3 was 33.16-fold higher than that in FHO3. Most of gibberellin 20 oxidase (GA20ox) and gibberellin 3-beta-dioxygenase (GA3ox) encoding genes, which catalyze the formation bioactive GA, were up-regulated upon pod N application. For example, the expression of nine GA20ox genes in FHN2 and FHN3 were significantly up-regulated to compare with FHO2 and FHO3, respectively. The expression levels of the one Ga3ox genes were strongly up-regulated in FHN2 to compare with FHO2. Gibberellin 2-beta-dioxygenase (GA2ox) catalyzes the inactivation of bioactive GAs, some GA2ox genes were induced different degrees in S1 or S2 stage under N application. The expressions of three gibberellin-regulated protein were found increased in FHN2 than that in FHO2. The significant expression changes of these GA related genes suggested that pod N application promoted gibberellin metabolism, and eventually promoted pod growth and development.
Pod N application induced the changes in expression of several genes in the brassinolide synthesis pathway. One cytochrome p45090A1 like isoform X1 gene was significantly up-regulated in FHN3, which was 13.95-fold higher than that in FHO3. Two cytochrome p45085A like genes were highly up-regulated in FHN2, the expression levels in FHN2 were 99.01 and 1085.40-fold higher than in FHO2. The expression of brassinolide receptor (BRI1) gene was also induced by pod N application. The expression of one BES1/BZR1, the key positive regulator of brassinolide signaling pathway, increased significantly in FHN2 and FHN3 to compare with FHO2 and FHO3, respectively. The expression of two zeatin biosynthesis pathway genes (CYP734A1 and CYP735A1) in FHN2 was significantly increased, which were hardly detectable in FHO2. These results indicated that the synthesis of brassinolide was active under pod N application, which may play important roles in peanut pods enlargement.
Verification of DEGs Using qRT-PCR
In order to validate the RNA-Seq data, the expression levels of 14 DEGs were detected using Quantitative Realtime PCR (qRT-PCR). The relative expression levels of these genes estimated by qRT-PCR were generally consistent with those by RNA-seq. The correlation coefficient of a liner regression analysis was 0.9066 (Fig. 7; Table S7).
Verification of DEGs by qRT-PCR. (a) Transcript levels of 14 genes related to hormone signal transduction and cell wall organization. Data are means of three replicates, and error bars represent ± SE (n = 3). (b) Pearson’s correlation of gene expression ratios between RNA-seq and qRT-PCR results. The correlation of the fold change was analyzed by RNA-seq (x-axis) with qRT-PCR (y-axis) data. y = 7.037x + 0.1986; R2 = 0.9066.
DISCUSSION
Previous studies showed that with the increase of N application, the main stem height, lateral branch length, branch number, leaf number and dry matter accumulation of peanut were significantly affected [22]. During peanut pod development, 90% of calcium is absorbed directly by pods from the soil [16, 17]. However, our results showed that the ratio of N absorbed directly by pods from soil was low, only about 10%. In early development stage of the pods, the main organ of nitrogen accumulation was the leaves, but during pods maturation, lots of nitrogen flew to seeds. 15N labeled experiment showed that during early pod development, nitrogen absorbed by pods was mainly accumulated in the shell, followed by other organs, especially the leaves. At late development stages, the pod absorbed nitrogen was transferred to seeds mainly. This result is consistent with previous research results [22].
Application of N increased the content of soluble protein and free amino acid in various peanut organs, and also improve the activities of nitrogen assimilases such as nitrate reductase, glutamine synthase and glutamate dehydrogenase [23, 24]. Our results showed that the expression levels of NR, NIRA and GS were increased in the pods after pod zone N application. When applied urea in the soil, it can be directly absorbed by plants or decomposed into ammonium nitrogen (\({\text{NH}}_{4}^{ + }\)) and nitrate nitrogen (\({\text{NO}}_{3}^{ - }\)) in the soil, which are the two main forms of nitrogen absorption by plants. Plant uptake these substances requires a variety of transporters, such as ABC transporters, nitrate transporter family (NRT), ammonium transporters (AMTS) and tonoplast intrinsic proteins (TIPs), etc. [6–9]. ABC transporter family has been widely studied in mammals and microorganisms, but it is still a relatively new research field in plants. In this study, the expression of several ABC transporter genes and aquaporin genes were increased, especially in second stage. Nitrate transporter (NPT) were mainly distributed in the cortical cells of the root mature area of Arabidopsis [25]. In the absence of nitrogen, NRT2.1 and NRT2.2 could regulate the high affinity transport system of NO3–, thus affecting nitrate absorption and root development [26, 27]. Three high affinity nitrate transporter genes NRT2.5 are highly induced during nitrogen starvation and play a potential role in nitrate transport in phloem. They can improve the utilization efficiency of NO3–, make plants adapt to short-term nitrogen starvation and maintain normal life activities [28, 29]. In this study, twelve high affinity nitrate transporter genes were expressed in pods, and three of them were up-regulated upon pod N application. It showed that the expression of high affinity nitrate transporter was affected not only by nitrate deficiency, but also by the absorption and utilization of ammonium nitrogen y plants. Ureaseisa is a kind of enzymes that can convert urea in to \({\text{NH}}_{4}^{ + }\), which is a direct nitrogen source and participates in nitrogen reabsorption. The activation process of urease requires the participation of urease accessory protein UREFs. However, pod N application had no significant effect on the expression of urease and UREFs genes. The ABC transporter family, aquaporins and high affinity nitrate transporters (NRTs) may play key roles in urea or nitrate transport, and different transporters functions in different developmental stages. However, pod N application had no significant effect on AMT gene expression.
Phytohormones and nitrate transporters interact and coordinate with each other, and play important roles in plant growth and development [30]. Studies have shown that when plants grow from low nitrogen to high nitrogen condition, ethylene production was markedly promoted in roots. This is because the increase of \({\text{NO}}_{3}^{ - }\) supply can trigger ethylene biosynthesis by activating the transcription of ACS (ACC synthase) and ACO (ACC oxidase) genes. Ethylene synthesis can also inhibit the expression of AtNRT2.1 gene, thereby reducing the absorption of \({\text{NO}}_{3}^{ - }\) [30, 31]. Spraying ethephon on peanut plants could improve the enzymes activities of glutazamide synthase, nitrate reductase and the nitrogen use efficiency [32]. In the current study, the expression of ACO genes increased, it may be associated with the synthesis of ethylene to regulate the activity of nitrogen metabolism enzymes. The expression of EIN3 (Ethylene-insensitive3) and ERF1 in ethylene signal transduction increased. Exogenous application of GA can improve the activity of nitrogen metabolism enzymes in cucumber, enhance the ability of nitrogen metabolism, and eventually promote root nitrogen absorption [33]. GA can regulate the expression level of NPF in cucumber roots and NPF members have the ability to transport GA in Arabidopsis [34, 35]. In our study, after pod zone N application, NPF3.1 was activated. The expression of several genes of GAs and zeatin synthesis pathway and signal transduction pathway were increased, which may promote pod expansion.
Nitrogen fixation of peanut root nodules contributed greatly to plant nitrogen. However, nitrogen fixation related genes, such as Nod-factor receptor 1 (NRF1), Nod-factor receptor 5 (NRF5), Symbiotic Receptor Kinase (SYMRK) and \(Ca_{2}^{ + }\)/calmodulin-dependent protein kinase (CCaMK) genes, were expressed lowly in pods, and nitrogen application in pod areas affect little on their expression. Only some early nodulin protein genes were expressed higher under nitrogen deficiency than under nitrogen application in S1 stage of N application in pods areas. It suggested that the lack of nitrogen in early pod development promoted biological nitrogen fixation in some extent. It is consistent to the previous study that with the increase of nitrogen fertilizer application, nitrogen fixation of rhizobia was gradually inhibited [36].
REFERENCES
Wienhold, B.J., Trooien, T.P., and Reichman, G.A., Yield and nitrogen use efficiency of irrigated corn in the northern Great Plains, Agron. J., 2019, vol. 87, p. 842. https://doi.org/10.2134/agronj1995.00021962008700050010x
Ball, S.T., Wynne, J.C., Elkan, G.H., and Schneeweis, T.J., Effect of inoculation and applied nitrogen on yield, growth and nitrogen fixation of two peanut cultivars, Field Crop Res., 1983, vol. 6, p. 85. https://doi.org/10.1016/0378-4290(83)90050-3
Siri-Prieto, G., Reeves, D.W., and Raper, R.L., Tillage systems for a cotton–peanut rotation with winter-annual grazing: impacts on soil carbon, nitrogen and physical properties, Soil Tillage Res., 2007, vol. 96, p. 260. https://doi.org/10.1016/j.still.2007.06.010
Zhang, X., Zhang, X.Y., Zhang, Y.T., Mao, J.W., Li, G.P., and Zhao, L.J., Effects of nitrogen application rate on nodulation, nitrogen absorption and utilization of peanut, J. Peanut Sci., 2012, vol. 41, p. 12. https://doi.org/10.3969/j.issn.1002-4093.2012.04.003
Witte, C.P., Urea metabolism in plants, Plant Sci., 2011, vol. 180, p. 431. https://doi.org/10.1016/j.plantsci.2010.11.010
Hao, D.L., Yang, S.Y., Huang, Y.N., and Su, Y.H., Identification of structural elements involved in fine-tuning of the transport activity of the rice ammonium transporter OsAMT1;3, Plant Physiol. Biochem., 2016, vol. 108, p. 99. https://doi.org/10.1016/j.plaphy.2016.07.003
Kotur, Z., Mackenzie, N., Ramesh, S., Tyerman, S.D., Kaiser, B.N., and Glass, A.D.M., Nitrate transport capacity of the Arabidopsis thaliana NRT2 family members and their interactions with AtNAR2.1, New Phytol., 2012, vol. 194, p. 724. https://doi.org/10.1111/j.1469-8137.2012.04094.x
Loqué, D., Yuan, L.X., Kojima, S., Gojon, A., Wirth, J., Gazzarrini, S., Ishiyaina, K., Takahashi, H., and Wirén, N.V., Additive contribution of AMT1;1 and AMT1;3 to high-affinity ammonium uptake across the plasma membrane of nitrogen deficient Arabidopsis roots, Plant J., 2006, vol.48, p. 522. https://doi.org/10.1111/j.1365-313X.2006.02887.x
Liu, K.H. and Tsay, Y.F., Switching between the two action modes of the dual-affinity nitrate transporter CHL1 by phosphorylation, EMBO J., 2003, vol. 22, p. 1005. https://doi.org/10.1093/emboj/cdg118
Krapp, A., David, L.C., Chardin, C., Girin, T., Marmagne, A., Leprince, A.S., Chaillou, S., Ferrario-Méry, S., Meyer, C., and Daniel-Vedele, F., Nitrate transport and signaling in Arabidopsis, J. Exp. Bot., 2014, vol. 65, p. 789-798. https://doi.org/10.1093/jxb/eru001
Seabra, A.R., Silva, L.S., and Carvalho, H.G., Novel aspects of glutamine synthetase (GS) regulation revealed by a detailed expression analysis of the entire GS gene family of Medicago truncatula under different physiological conditions, BMC Plant Biol., 2013, vol. 13, p. 137. https://doi.org/10.1186/1471-2229-13-137
Nigro, D., Blanco, A., Anderson, O.D., and Gadaleta, A., Characterization of Ferredoxin-Dependent Glutamine-Oxoglutarate Amidotransferase (Fd-GOGAT) genes and their relationship with grain protein content QTL in wheat, PLoS One, 2014, vol. 9, p. 1. https://doi.org/10.1371/journal.pone.0103869
Gruber, N. and Galloway, J.N., An Earth-system perspective of the global nitrogen cycle, Nature, 2008, vol. 451, p. 293. https://doi.org/10.1038/nature06592
Xia, H., Zhao, C.Z., Hou, L., Li, A.Q., Zhao, S.Z., Bi, Y.P., An, J., Zhao, Y.X., Wan, S.B., and Wang, X.J., Transcriptome profiling of peanut gynophores revealed global reprogramming of gene expression during early pod development in darkness, BMC Genomics, 2013, vol. 14, p. 517. https://doi.org/10.1186/1471-2164-14-517
Inanaga, S., Utunomiya, M., Horiguchi, T., and Nishihara, T., Behavior of fertilizer-N absorbed through root and fruit in peanut, Plant Soil, 1990, vol. 122, p. 85. https://doi.org/10.1007/BF02851913
Yang, R., Howe, J.A., and Balkcom, K.B., Soil evaluation methods for calcium for peanut (Arachis hypogaea L.) production in the coastal plain, Peanut Sci., 2017, vol. 44, p. 1. https://doi.org/10.3146/PS16-5.1
Li, Y., Meng, J.J., Yang, S., Guo, F., Zhang, J.L., Geng, Y., Cui, L., Wan, S.B., and Li, X.G., Transcriptome analysis of calcium- and hormone-related gene expressions during different stages of peanut pod development, Front. Plant Sci., 2017, vol. 8, p. 1241. https://doi.org/10.3389/fpls.2017.01241
Kim, D., Langmead, B., and Salzberg, S.L., HISAT: a fast spliced aligner with low memory requirements, Nat. Methods, 2015, vol. 12, p. 357. https://doi.org/10.1038/nmeth.3317
Langmead, B. and Salzberg, S.L., Fast gapped-read alignment with bowtie 2, Nat. Methods, 2012, vol. 9, p. 357. https://doi.org/10.1038/nmeth.1923
Li, B. and Dewey, C.N., RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome, BMC Bioinf., 2011, vol. 12, p. 323. https://doi.org/10.1186/1471-2105-12-323
Wang, L., Feng, Z., Wang, X., Wang, X., and Zhang, X., DEGseq: an R package for identifying differentially expressed genes from RNA-seq data, Bioinformatics, 2010, vol. 26, p. 136. https://doi.org/10.1093/bioinformatics/btp612
Dai, L.X., Zhang, Z.M., Zhang, G.C., Zhang, Y., Ci, D.W., Qin, F.F., and Ding, H., Effects of nitrogen application on nitrogen uptake and distribution in peanut, J. Nucl. Agric. Sci., 2020, vol. 34, p. 0370.
Luo, H., Zhou, G.Y., Luo, Y.F., and Liang, X.Q., Traits analysis of high yield peanut varieties, J. Peanut Sci., 2009, vol. 38, p. 15. https://doi.org/10.3969/j.issn.1002-4093.2009.03.004
Zhang, Z.M., Wan, S.B., Ning, T.Y., and Dai, L.X., Effects of nitrogen level on nitrogen metabolism and correlating enzyme activity of peanut, J. Plant Ecol., 2008, vol. 32, p. 1407. https://doi.org/10.3773/j.issn.1005-264x.2008.06.022
Remans, T., Nacry, P., Pervent, M., Girin, T., Tillard, P., Lepetit, M., and Gojon, A., A central role for the nitrate transporter NRT2.1 in the integrated morphological and physiological responses of the root system to nitrogen limitation in Arabidopsis, Plant Physiol., 2006, vol. 140, p. 909. https://doi.org/10.1104/pp.105.075721
Cerezo, M., Tillard, P., Filleur, S., Muños, S., Daniel-Vedele, F., and Gojon, A., Major alterations of the regulation of root \({\text{NO}}_{3}^{ - }\) uptake are associated with the mutation of Nrt2.1 and Nrt2.2 genes in Arabidopsis, Plant Physiol., 2001, vol. 127, p. 262. https://doi.org/10.1104/pp.127.1.262
Filleur, S., Dorbe, M. F., Cerezo, M., Orsel, M., Granier, F., Gojon, A., and Daniel-Vedele, F., An Arabidopsis T-DNA mutant affected in Nrt2 genes is impaired in nitrate uptake, FEBS Lett., 2001, vol. 489, p. 220. https://doi.org/10.1016/s0014-5793(01)02096-8
Lezhneva, L., Kiba, T., Feria-Bourrelier, A.B., Lafouge, F., Boutet-Mercey, S., Zoufan, P., Sakakibara, H., Daniel-Vedele, F., and Krapp, A., The Arabidopsis nitrate transporter NRT2.5 plays a role in nitrate acquisition and remobilization in nitrogen-starved plants, Plant J., 2014, vol. 80, p. 230. https://doi.org/10.1111/tpj.12626
Kiba, T., Feria-Bourrellier, A.B., Lafouge, F., Lezhneva, L., Boutet-Mercey, S., Orsel, M., Bréhaut, V., Miller, A., Daniel-Vedele, F., Sakakibara, H., and Krapp, A., The Arabidopsis nitrate transporter NRT2.4 plays a double role in roots and shoots of nitrogen-starved plants, Plant Cell, 2012, vol. 24, p. 245. https://doi.org/10.1105/tpc.111.092221
Krouk, G., Hormones and nitrate: a two-way connection, Plant Mol. Biol., 2016, vol. 91, p. 599. https://doi.org/10.1007/s11103-016-0463-x
Tian, Q.Y., Sun, P., and Zhang, W.H., Ethylene is involved in nitrate-dependent root growth and branching in Arabidopsis thaliana, New Phytol., 2009, vol. 184, p. 918. https://doi.org/10.1111/j.1469-8137.2009.03004.x
Ge, J.J., Zhu, L., Zhang, G.L., Han, W.J., Yin, Y.H., and Li, Y., Effects of ethephon on nitrogen metabolism and photosynthesis characters of peanut, J. Peanut Sci., 2008, vol. 37, p. 22. https://doi.org/10.3969/j.issn.1002-4093.2008.02.006
Bai, L.Q., Deng, H.H., Zhang, X.C., Yu, X.C., and Li, Y.S., Gibberellin is involved in inhibition of cucumber growth and nitrogen uptake at suboptimal root-zone temperatures, PLoS One, 2016, vol. 23, p. 11, p. e0156188. https://doi.org/10.1371/journal.pone.0156188
Chiba, Y., Shimizu, T., Miyakawa, S., Kanno, Y., Koshiba, T., Kamiya, Y., and Seo, M., Identification of Arabidopsis thaliana NRT1/PTR FAMILY (NPF) proteins capable of transporting plant hormones, J. Plant Res., 2015, vol. 128, p. 679. https://doi.org/10.1007/s10265-015-0710-2
David, L.C., Berquin, P., Kanno, Y., Seo, M., Daniel-Vedele, F., and Ferrario-Méry, S., N availability modulates the role of NPF3.1, a gibberellin transporter, in GA-mediated phenotypes in Arabidopsis, Planta, 2016, vol. 244, p. 1315. https://doi.org/10.1007/s00425-016-2588-1
Wu, Z. F., Chen, D. X., Zheng, Y.M., Wang, C.B., Sun, X.W., Li, X.D., Wang, X.X., Shi, C.R., Feng, H., and Yu, T.Y., Supply characteristics of different nitrogen sources and nitrogen use efficiency of peanut, Chin. J. Oil Crop Sci., 2016, vol. 38, p. 207. https://doi.org/10.7505/j.issn.1007-9084.2016.02.011
Funding
This work was financially supported by grants from the National Natural Science Foundation of China (project nos. 31801301, 31861143009); Natural Science Foundation of Shandong Province (project nos. ZR2020MC104, ZR2020MC105); Young Talents Training Program; Agricultural scientific and technological innovation project of Shandong Academy of Agricultural Sciences (project no. CXGC2018E13); Taishan Scholar Foundation of Shandong Province (project no. ts20190964).
Author information
Authors and Affiliations
Contributions
L. Hou and R. Lin contributed equally to this work.
Corresponding author
Ethics declarations
Conflict of interests. The authors declare that they have no conflicts of interest.
Statement on the welfare of humans or animals. This article does not contain any studies involving humans or animals performed by any of the authors.
Additional information
Abbreviations: NUE—N use efficiency; NRT—nitrate transporter family; CLC—chloride channel family; AMTS—ammonium transporters; TIPs—tonoplast intrinsic proteins; NR— nitrate reductase; NiRA—nitrite reductase; GS/GOGAT—glutamine synthetase/glutamate synthetase; FH1—‘Fenghua 1’; PES—pod expanding stage; PFS—pod filling stage; HS—harvesting stage; LN—low nitrogen; HN—high nitrogen; DEGs— differentially expressed genes; FHN—FH1 treated with nitrogen in pod zone; FHO—FH1 control; MIPs—major intrinsic proteins; DUR3—high-affinity urea transporter; GDH—glutamate dehydrogenase; KAO1—ent-kaurenoic acid oxidase; GA20ox— gibberellin 20 oxidase 2; GA3ox—gibberellin 3-beta-dioxygenase; GA2ox—gibberellin 2-beta-dioxygenase; BRI1—brassinosteroid-insensitive1; BES1/BZR1—BRI1 EMS SUPPRESSOR 1/BRASSINAZOLE RESISTANT 1; EIN3—Ethylene-insensitive3; ACS—ACC synthase; ACO—ACC oxidase.
Supplementary Information
Rights and permissions
About this article
Cite this article
Hou, L., Lin, R.X., Wang, X.J. et al. The Mechanisms of Pod Zone Nitrogen Application on Peanut Pod Yield. Russ J Plant Physiol 69, 51 (2022). https://doi.org/10.1134/S1021443722030050
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1134/S1021443722030050