Abstract
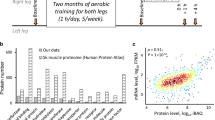
We aimed to investigate the effect of 8 weeks of moderate endurance training without considerable mechanical stress on the activation of extracellular matrix (ECM) gene expression in human skeletal muscles. Mechanical stress activates ECM biogenesis in the skeletal muscles, therefore aerobic exercise on a cycling ergometer with concentric muscle contractions only was used in the study. Skeletal muscle samples from m. vastus lateralis were taken from seven young untrained men before and after 8 weeks of aerobic training. Changes in the transcriptome (RNA sequencing) and proteome (shotgun quantitative proteomics analysis) were assessed in the samples; ECM-associated proteins (or matrisome) were determined using the Matrisome DB database. After the training period, a change (mainly an increase) in the content of 14 ECM proteins and 134 mRNAs of ECM proteins was found. The largest increase in protein content was found for collagens type 1 and 3 (1.7 and 2.2 times, respectively), the main proteins of the human skeletal muscle’s ECM, which was consistent with an increase in the corresponding mRNA by 10–20 times. In addition, an increase in the expression of more than a hundred mRNAs of collagens, glycoproteins, proteoglycans, and enzymatic regulators of ECM was found, which occurs simultaneously with an increase in the expression of genes of growth factors (IGF1, PDGFs, TGFB1, MDK, etc.), which play a main role in ECM biogenesis regulation. In conclusion, 8-week aerobic exercise training without considerable mechanical stress is a powerful stimulus for the activation of ECM biogenesis in skeletal muscle.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
One of the most common categories of injury to the musculoskeletal system is damage to skeletal muscles, tendons and ligaments of varying severity. The risk of such injuries is high in middle-aged and elderly people during daily physical activity, as well as in people with reduced functional capabilities of the musculoskeletal system (reduced strength and performance capabilities, as well as strength and elasticity of ligaments and tendons). A decrease in the functionality and tolerance of skeletal muscles to daily physical activity, as well as an increase in their damage, is observed after a few days of immobilization of the limb, bedrest, or exposure to microgravity [1–3]. This is due to a number of changes, including a decrease in the rate of synthesis of muscle proteins and muscle mass, mitochondrial density and oxidative capacity of muscles, an increase in inflammation, edema, the appearance of pain, as well as a violation of the biogenesis of intramuscular connective tissue structures, including the extracellular matrix (ECM) [1, 4–8]. The ECM plays a key role in transferring force from contracting muscle fibers to tendons, preventing damage to muscle membranes during exercise, delivering and retaining various biomolecules (including enzymes and growth factors), and during recovery from muscle injury. The ECM is responsible for orienting muscle fibers [9–13]. Therefore, the development of approaches to the activation of ECM biogenesis is relevant not only for preventing muscle injuries with reduced functionality, but also for rehabilitation after injuries of the musculoskeletal system.
Regular strength training (short, high-intensity exercise) is effective in increasing muscle mass and strength and in activating ECM biogenesis [14–17]. However, these exercises in most cases are of little use in rehabilitation after injuries and/or after prolonged physical inactivity due to high trauma. Regular aerobic exercise (low-intensity and prolonged physical activity) increases aerobic performance (endurance) but has little effect on muscle mass and strength. At the same time, such training reduces the damage to muscle membranes in response to a single load/exercise [18–20], which is presumably associated with the activation of ECM biogenesis. However, the molecular mechanisms responsible for the activation of ECM biogenesis during aerobic training have been studied fragmentarily [21, 22].
Objective—To investigate the effect of 8 weeks of moderate-intensity, non-impact aerobic training on the activation of ECM gene expression in exercised skeletal muscle (m. vastus lateralis). Eccentric muscle contractions (such as running) may be a trigger for activation of ECM biogenesis [23]. To exclude such effects in our study, we used aerobic exercise on a bicycle ergometer, including only concentric muscle contractions. ECM includes about 300 different proteins divided into functional groups: collagens (fibrillar glycoproteins, which are most represented in the ECM of all human tissues and play a key role in the formation of the connective tissue structure), structural glycoproteins and proteoglycans (form the main substance of the ECM), as well as hundre of ECM-associated proteins: enzymatic regulators (enzymes directly involved in ECM remodeling) and secreted factors (proteins secreted by various muscle tissue cells during ECM remodeling, including growth factors). All of the above functional groups of proteins are united by a common term, the matrisome. In view of the large number of matrisome proteins, the use of omics approaches (RNA sequencing and mass spectrometric proteomics) seems to be a logical approach to assess changes in the expression of almost all mRNAs of ECM-related proteins and changes in the content of highly abundant proteins (such as collagens). The physiological effects of this training program and the results of transcriptomic and proteomic analyses have been presented and discussed previously [24, 25]. In the present study, we conducted an in-depth analysis of the effect of training on the gene expression of all ECM-related proteins. The list of these genes was taken from the database MatrisomeDB, containing comprehensive information about the various functional groups of proteins associated with ECM [26–29].
METHODOLOGY
Seven untrained young men (age, 21–24 years, body weight, 72–79 kg, body mass index, 22–25 kg/m2) took part in the experiment. Volunteers performed aerobic exercises on an electromagnetic bicycle ergometer (Ergoselect 200, Ergoline, Germany) for 8 weeks: 5 times a week, 1 h a day, as described by us earlier [25]. Briefly, before the training period and every 2 weeks, the volunteers performed an incremental test on a bicycle ergometer (15 W/min). During the test, capillary blood samples were taken every 2 min to assess lactate concentration; the anaerobic threshold (a marker of the organism’s aerobic capacity) was assessed as power at a lactate level at 4 mM (LT4) [30]. During the training period, the volunteers alternately performed exercises with a constant (60 min, 70% LT4) and variable (3 min 50% LT4 + 2 min 85% LT4) × 12) power. Samples from m. vastus lateralis were taken before and after the training period in the baseline state (48 h after the last exercise) using a needle biopsy under local anesthesia (2 mL of 2% lidocaine), as described by us previously [25].
Shotgun proteomics. Sample preparation and proteomic analysis have been described by us previously [24]. Briefly, a fragment of frozen tissue (~15 mg) was homogenized in buffer (4% sodium dodecyl sulfate in 0.1 M Tris-HCl, pH 7.6, 0.1 M dithiothreitol), incubated for 5 min at 95°C, sonicated (2 times for 10 s at 100 W) and centrifuged (5 min, 16 000 g). Alkylation and trypsinolysis of proteins (12 h, trypsin 1 : 100 (Tripsin Gold, Promega, United States) in 40 μL of 0.1 M triethylammonium bicarbonate) was carried out on a YM-30 filter (Millipore, Ireland) using the filter aided sample preparation (FASP) method. The peptides were washed off the filter by centrifugation (10 min, 14 000 g) and labeled with an isobaric iTRAQ 8-plex label (Sciex, United States). The mixture of labeled peptides was concentrated and fractionated using XBridge C18 columns (250 × 4.6 mm, particle size 5 µm, Waters, Ireland) on an Agilent 1200 Series chromatograph (Agilent, United States). The resulting fractions (30 pcs) were concentrated and combined into 10 mixed fractions. Each fraction was separated three times on an Ultimate 3000 RSLCnano chromatograph (pre-column Acclaim (0.5 × 3 mm, particle size 5 µm) and Acclaim Pepmap C18 column (75 µm × 150 mm, particle size 2 µm); everybody was then put in a gradient elution mode (90 min) and analyzed on a Q Exactive HF mass spectrometer (Thermo Scientific, United States).
The search and identification of reporter ions was carried out using the MaxQuant platform (1.5.7.4) with default settings for false discovery rate (FDR) of 1%. The data was processed on the Perseus platform (1.6.1.2): after filtration, for each protein, the ratio of reporter ion intensities was calculated (intensity “after” training to intensity “before” training); then, changes in the intensities of reporter ions (protein content) were evaluated using the Wilcoxon signed rank test at padj < 0.05 (Benjamini–Hochberg multiple comparison correction).
RNA sequencing. Sample preparation and analysis was described by us previously [25]. Briefly, frozen tissue samples (~20 mg) were homogenized, RNA was extracted using RNeasy Mini Kit columns (Qiagen, Germany). RNA concentration was measured on a Qubit 3.0 fluorimeter (Thermo Scientific, United States), RNA integrity was assessed by capillary electrophoresis (Bioanalyzer 2100, Agilent, United States). Libraries were prepared from 300 ng of RNA using the NEB Next Ultra II RNA kit (New England Biolabs, United States) according to the manufacturer’s protocol. The concentration of libraries was measured on a Qubit 3.0 fluorimeter, the length distribution of library fragments was estimated using a Bioanalyzer 2100. The effective concentration of the libraries was assessed by real-time PCR. Libraries sequenced on NextSeq 500 instrument (Illumina, United States) in the single-ended read mode. The average number of reads per sample was 47 million.
Data quality was assessed using FASTQC (v0.11.4); adapter sequences and low-quality reads were removed with Timmomatic (v0.36). The scans were mapped to the human GRCh38 genome. Protein-coding genes were isolated and the change in their expression was analyzed using the DESeq2 R package (paired sample analysis) with cut-off criteria padj < 0.05 (Benjamini–Hochberg correction) and |log2(Fold Change)| > log2 (1.25).
Analysis of functional enrichment. To search for genes/proteins belonging to various functional groups of ECM, all genes/proteins detected by us were compared with the database MatrisomeDB containing information about ECM proteins: 44 collagens, 195 glycoproteins, and 35 proteoglycans, and about ECM-associated proteins: 238 enzymatic regulators, 171 ECM-affiliated proteins, and 344 secreted factors. To identify functional groups enriched in genes that changed their expression after 8 weeks of aerobic training, the χ2 (chi-square) with Bonferroni correction at p < 0.05 was used.
RESEARCH RESULTS
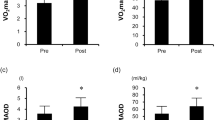
As previously described by us [31], 8 weeks of training led to a marked increase in aerobic capacity of the organism: power at the anaerobic threshold increased by 35% (p < 0.01).
In muscle tissue samples taken before and after the training period, 13 279 mRNAs of protein-coding genes and 795 proteins were detected. Among them, we identified 501 mRNAs and 32 matrisome-related proteins. After 8 weeks of training, a change (mainly an increase) in the content of 1650 mRNA and 250 proteins was found (Table 1). Of these, 9 detected proteins and 34 mRNA belonged to the collagen group, 7 and 22, respectively, to proteoglycans, 5 and 139, to glycoproteins, 8 and 137, to enzymatic regulators, and 3 and 169, to secreted factors (Table 1).
A significant increase (padj < 0.05) in protein content was found mainly for the collagen group (8 proteins), as well as for 3 proteoglycan proteins and 1 protein in each of the groups of glycoproteins, secreted factors and, ECM regulators (Table 1). Among collagens, the largest increase in protein content was found for COL1A1/2 (1.7 times), and COL3A1 (2.2 times) (Fig. 1), the main ECM proteins of human skeletal muscle [32], which was consistent with an increase in the corresponding mRNAs of 19.7 for COL1A1, 2.9 times for COL1A2, and 9.8 times for COL3A1. In addition, the increase (padj < 0.01) in mRNA content was shown for 14 other collagen genes, the protein products of which were not detected (Fig. 1).
Unlike shotgun proteomics, RNA sequencing makes it possible to evaluate the expression of almost all expressed genes, including those related to the matrisome. This made it possible to isolate the enriched functional groups of the matrisome, i.e., groups in which the proportion of genes that increased expression after a period of training, in relation to all genes that increased expression, is significantly greater than the proportion of genes belonging to the same functional group in relation to all detected protein-coding genes. Significant enrichment (p < 0.05) was found for the groups of collagens, enzymatic regulators, and glycoproteins (Fig. 2).
When considering each functional group, some of them (enzymatic regulators, secreted factors) could be divided into subgroups of genes with different functions. Thus, among the enzymatic regulators (46 genes) that have changed mRNA expression, there are subgroups of metalloproteinases, regulators of the collagen network, and regulators of the IGF1 signaling cascade and regeneration of muscle fibers (Fig. 3).
Among the secreted factors (24 genes) that increased mRNA expression, there are subgroups of growth factors and bone metabolism regulators; the remaining secreted factors include several cytokines (CCL2, CCL18, CXCL9 and IL34) (Fig. 4).
mRNAs that increased expression (padj < 0.01) in the groups of glycoproteins and proteoglycans are presented in Fig. 5. It should be noted that in the group of glycoproteins, the gene ELN, encodes elastin, the fourth most abundant protein in the ECM of skeletal muscle [32].
RESULTS AND DISCUSSION
Collagens make up two-thirds of all ECM proteins, with type 1 and 3 collagens accounting for more than 50% of the mass of ECM proteins in human skeletal muscle [32]. Previously, in experiments on animals and with the volunteers, it was shown that regular high-intensity short-term physical activity, leading to an increase in muscle mass (strength exercise), causes an increase in the expression of collagen-coding genes and activates ECM biogenesis. Thus, 12-week strength training in rats (climbing stairs with weights) led to an increase in the expression of mRNA of collagen types 1 and 3 in skeletal muscle [14], and expression and activity of metalloproteinases (MMP 2 and 9) in skeletal muscle and blood [14, 15]. Strength training of the lower extremities (3 times a week, 11 weeks) of young (27 years old) untrained men led to a significant increase in gene expression in m. vastus lateralis, in particular, the content of mRNA increased by 5.2 times for COL1A1 [16]. In another study, 10 weeks of strength training (leg press) in young (26 year old) untrained men resulted in a significant increase in the expression of ECM-related genes [17]. These data are consistent with the fact that in human skeletal muscle, genes that increase expression after regular strength training associated with ECM-related terms (meta-analysis of transcriptome data) [33].
In our study, it was shown that training without the use of high-intensity and eccentric contraction, namely, moderate-intensity aerobic training on a bicycle ergometer, is a sufficient stimulus for a pronounced activation of ECM biogenesis, an increase in the content of the main ECM proteins of collagen types 1 and 3 (2–3 times), as well as other collagens that perform mainly structural and regulatory functions (collagen types 4 and 6 and types 11, 14, and 15, respectively; 1.2–1.7 times). These changes occurred against the background of an increase in the expression of the corresponding mRNAs, as well as mRNAs of other collagens, the protein products of which were not detected by us, and about a hundred mRNAs encoding glycoproteins, proteoglycans, and enzymatic regulators of ECM biogenesis. In addition, similar changes in the transcriptome were observed in various studies with volunteers. Thus, 12-week (5 times a week, 60 min a day) exercise on a bicycle ergometer by young (19–32 years old) untrained men led to an increase in the expression of ECM genes, in particular, type 1 collagen [34].
It is interesting to note that similar effects were observed for older people (64 years): 6-week aerobic training on a bicycle ergometer (5 times a week, 60 min per day, moderate aerobic exercise) led to a pronounced increase in mRNA expression, including collagen types 3 and 4 [35]. Twelve weeks of aerobic exercise on a bicycle ergometer (3 times a week, 45 min per day) of elderly volunteers (68 years old) led to changes in the expression of 397 genes, among which the expression of collagen mRNA was also increased COL3A1 and COL6A3 and secreted factors TNFSF10 and CRLF3, a decreased expression of glycoprotein mRNA IGFBP6 and DPT, enzymatic regulators F10 and ADAMTS5, and secreted factors S100A6 and CXCL14 [36].
We have shown (Figs. 3 and 4) that, in addition to activating collagen expression, regular aerobic physical activity causes a large-scale and pronounced increase in the expression of genes for metalloproteinases and other enzymes involved in ECM remodeling, mainly due to the degradation of old collagen bonds and molecules (especially types 1, 3, and 4) [37–41]. In addition, we found an increase in the expression of a large number of glycoproteins (including elastin, the third most abundant protein in human skeletal muscles) that perform various structural and regulatory functions. This is in complete agreement with the results of a meta-analysis that studied transcriptomic responses to regular aerobic training and showed that in human skeletal muscles, the set of genes that increased expression after regular aerobic training is associated with ECM-related terms [42], however these data are not consistent with other meta-analysis [33].
Despite the absence of significant enrichment in the genes of secreted factors in our study (Fig. 3), we found an increase (~2 times) in the expression of individual genes of growth factors (Fig. 5). Model studies with altered gene expression have shown that these factors play an important role in the regulation of ECM biogenesis both in skeletal muscles: IGF1, MDK [43, 44], PDGFB, PDGFD [45], and TGFB1 [46, 47], and in other tissues: HGF [48, 49], INHBB [50], and PGF [51]. These data are consistent with the results of studies on rodents with tenotomy of synergistic muscles (a model of chronic increase in muscle load), which showed an increase in the concentration of ECM proteins and expression Igf1 and Tgf in plantar muscle [52–54].
CONCLUSIONS
Our results show that, despite low intensity, regular non-impact aerobic exercise is a powerful stimulus for the activation of ECM biogenesis. In particular, a pronounced increase in the content of the main ECM proteins, collagen types 1 and 3, was found, as well as an increase in the expression of more than a hundred mRNAs of collagens, glycoproteins, proteoglycans, and enzymatic regulators of the ECM, occurring against the background of an increase in the expression of the genes of the main growth factors that regulate ECM biogenesis (IGF1, PDGFs, TGFB1, MDK, etc.). Thanks to the use of omics technics, it was possible for the first time to evaluate the effect of long-term aerobic training on the expression of all ECM genes, as well as on the content of key ECM proteins. Non-impact aerobic exercise, as opposed to strength training and exercise with an impact and eccentric contraction (such as running), is applicable to a larger contingent of people who need to restore skeletal muscle (and possibly tendons and ligaments) function, people with reduced functional opportunities, patients after injuries and/or a long period of physical inactivity, cosmonauts in the recovery period after a flight, etc. Therefore, the use of aerobic exercises can be promising both for optimizing existing approaches to recovery after injuries of the musculoskeletal system, and for preventing injuries with reduced functionality.
REFERENCES
Narici, M.V. and de Boer, M.D., Disuse of the musculo-skeletal system in space and on earth, Eur. J. Appl. Physiol., 2011, vol. 111, no. 3, p. 403.
Hackney, K.J. and Ploutz-Snyder, L.L., Unilateral lower limb suspension: integrative physiological knowledge from the past 20 years (1991–2011), Eur. J. Appl. Physiol., 2012, vol. 112, no. 1, p. 9.
Hyatt, H., Deminice, R., Yoshihara, T., Powers, S.K., et al., Mitochondrial dysfunction induces muscle atrophy during prolonged inactivity: a review of the causes and effects, Arch. Biochem. Biophys., 2019, vol. 662, p. 49.
Bamman, M.M., Clarke, M.S.F., Feeback, D.L., et al., Impact of resistance exercise during bed rest on skeletal muscle sarcopenia and myosin isoform distribution, J. Appl. Physiol., 1998, vol. 84, no. 1, p. 157.
Crossland, H., Skirrow, S., Puthucheary, Z.A., et al., The impact of immobilisation and inflammation on the regulation of muscle mass and insulin resistance: different routes to similar end-points, J. Physiol., 2019, vol. 597, no. 5, p. 1259.
Hortobágyi, T., Dempsey, L., Fraser, D., et al., Changes in muscle strength, muscle fibre size and myofibrillar gene expression after immobilization and retraining in humans, J. Physiol., 2000, vol. 524, part 1, p. 293.
Rudrappa, S.S., Wilkinson, D.J., Greenhaff, P.L., et al., Human skeletal muscle disuse atrophy: effects on muscle protein synthesis, breakdown, and insulin resistance—a qualitative review, Front. Physiol., 2016, vol. 7, p. 361.
Yasuda, N., Glover, E.I., Phillips, S.M., et al., Sex-based differences in skeletal muscle function and morphology with short-term limb immobilization, J. Appl. Physiol., 2005, vol. 99, no. 3, p. 1085.
Webster, M.T., Manor, U., Lippincott-Schwartz, J., and Fan, C.M., Intravital imaging reveals ghost fibers as architectural units guiding myogenic progenitors during regeneration, Cell Stem Cell, 2016, vol. 18, no. 2, p. 243.
Gillies, A.R. and Lieber, R.L., Structure and function of the skeletal muscle extracellular matrix, Muscle Nerve, 2011, vol. 44, no. 3, p. 318.
Heredia, J.E., Mukundan, L., Chen, F.M., et al., Type 2 innate signals stimulate fibro/adipogenic progenitors to facilitate muscle regeneration, Cell, 2013, vol. 153, no. 2, p. 376.
Joe, A.W.B., Yi, L., Natarajan, A., et al., Muscle injury activates resident fibro/adipogenic progenitors that facilitate myogenesis, Nat. Cell Biol., 2010, vol. 12, no. 2, p. 153.
Trotter, J.A. and Purslow, P.P., Functional morphology of the endomysium in series fibered muscles, J. Morphol., 1992, vol. 212, no. 2, p. 109.
Guzzoni, V., Ribeiro, M.B.T., Lopes, G.N., et al., Effect of resistance training on extracellular matrix adaptations in skeletal muscle of older rats, Front. Physiol., 2018, vol. 9, p. 374.
Sousa Neto, I.V. de Durigan, J.L.Q., Guzzoni, V., et al., Effects of resistance training on matrix metalloproteinase activity in skeletal muscles and blood circulation during aging, Front. Physiol., 2018, vol. 9, p. 190.
Norheim, F., Raastad, T., Thiede, B., et al., Proteomic identification of secreted proteins from human skeletal muscle cells and expression in response to strength training, Am. J. Physiol. Endocrinol. Metab., 2011, vol. 301, no. 5. E1013
Damas, F., Ugrinowitsch, C., Libardi, C.A., et al., Resistance training in young men induces muscle transcriptome-wide changes associated with muscle structure and metabolism refining the response to exercise-induced stress, Eur. J. Appl. Physiol., 2018, vol. 118, no. 12, p. 2607.
Vincent, H.K. and Vincent, K.R., The effect of training status on the serum creatine kinase response, soreness and muscle function following resistance exercise, Int. J. Sports Med., 1997, vol. 18, no. 6, p. 431.
Fehrenbach, E., Niess, A.M., Schlotz, E., et al., Transcriptional and translational regulation of heat shock proteins in leukocytes of endurance runners, J. Appl. Physiol., 2000, vol. 89, no. 2, p. 704.
Brancaccio, P., Lippi, G., and Maffulli, N., Biochemical markers of muscular damage, Clin. Chem. Lab. Med., 2010, vol. 48, no. 6, p. 757.
Kritikaki, E., Asterling, R., Ward, L., et al., Exercise training-induced extracellular matrix protein adaptation in locomotor muscles: a systematic review, Cells, 2021, vol. 10, no. 5, p. 1022.
Csapo, R., Gumpenberger, M., and Wessner, B., Skeletal muscle extracellular matrix—what do we know about its composition, regulation, and physiological roles? A narrative review, Front. Physiol., 2020, vol. 11, p. 253.
Willis, C.R.G., Deane, C.S., Ames, R.M., et al., Transcriptomic adaptation during skeletal muscle habituation to eccentric or concentric exercise training, Sci. Rep., 2021, vol. 11, no. 1, p. 23930.
Makhnovskii, P.A., Zgoda, V.G., Bokov, R.O., et al., Regulation of proteins in human skeletal muscle: the role of transcription, Sci. Rep., 2020, vol. 10, no. 1, p. 3514.
Popov, D.V., Makhnovskii, P.A., Shagimardanova, E.I., et al., Contractile activity-specific transcriptome response to acute endurance exercise and training in human skeletal muscle, Am. J. Physiol. Endocrinol. Metab., 2019, vol. 316, no. 4. E605
Naba, A., Pearce, O.M.T., and Del Rosario, A., et al., Characterization of the extracellular matrix of normal and diseased tissues using proteomics, J. Proteome Res., 2017, vol. 16, no. 8, p. 3083.
Naba, A., Clauser, K.R., Ding, H., et al., The extracellular matrix: tools and insights for the “omics” era, Matrix Biol., 2016, vol. 49, p. 10.
Shao, X., Taha, I.N., Clauser, K.R., et al., MatrisomeDB: the ECM-protein knowledge database, Nucleic Acids Res., 2020, vol. 48, no. D1, p. D1136.
Naba, A., Clauser, K.R., Hoersch, S., et al., The matrisome: in silico definition and in vivo characterization by proteomics of normal and tumor extracellular matrices, Mol. Cell. Proteomics, 2012, vol. 11, no. 4, p. M111.014647.
Stegmann, H. and Kindermann, W., Comparison of prolonged exercise tests at the individual anaerobic threshold and the fixed anaerobic threshold of 4 mmol.l(-1) lactate, Int. J. Sports Med., 1982, vol. 3, no. 2, p. 105.
Popov, D.V., Lysenko, E.A., Bokov, R.O., et al., Effect of aerobic training on baseline expression of signaling and respiratory proteins in human skeletal muscle, Physiol. Rep., 2018, vol. 6, no. 17. e13868
McKee, T.J., Perlman, G., Morris, M., and Komarova, S.V., Extracellular matrix composition of connective tissues: a systematic review and meta-analysis, Sci. Rep., 2019, vol. 9, no. 1, p. 10542.
Pillon, N.J., Gabriel, B.M., Dollet, L., et al., Transcriptomic profiling of skeletal muscle adaptations to exercise and inactivity, Nat. Commun., 2020, vol. 11, no. 1, p. 470.
Nishida, Y., Tanaka, H., Tobina, T., et al., Regulation of muscle genes by moderate exercise, Int. J. Sports Med., 2010, vol. 31, no. 9, p. 656.
Riedl, I., Yoshioka, M., Nishida, Y., et al., Regulation of skeletal muscle transcriptome in elderly men after 6 weeks of endurance training at lactate threshold intensity, Exp. Gerontol., 2010, vol. 45, no. 11, p. 896.
Radom-Aizik, S., Hayek, S., Shahar, I., et al., Effects of aerobic training on gene expression in skeletal muscle of elderly men, Med. Sci. Sports Exerc., 2005, vol. 37, no. 10, p. 1680.
Cui, N., Hu, M., and Khalil, R.A., Biochemical and biological attributes of matrix metalloproteinases, Prog. Mol. Biol. Transl. Sci., 2017, vol. 147, p. 1.
Visse, R. and Nagase, H., Matrix metalloproteinases and tissue inhibitors of metalloproteinases: structure, function, and biochemistry, Circ. Res., 2003, vol. 92, no. 8, p. 827.
Serra, R., Matrix metalloproteinases in health and disease, Biomolecules, 2020, vol. 10, no. 8, p. 1138.
Alameddine, H.S., Matrix metalloproteinases in skeletal muscles: friends or foes? Neurobiol. Dis., 2012, vol. 48, no. 3, p. 508.
Corcoran, M.L., Hewitt, R.E., Kleiner, D.E., and Steuer-Stevenson, W.G., MMP-2: expression, activation and inhibition, Enzyme Protein, 1996, vol. 49, nos. 1—3, p. 7.
Makhnovskii, P.A., Bokov, R.O., Kolpakov, F.A., and Popov, D.V., Transcriptomic signatures and upstream regulation in human skeletal muscle adapted to disuse and aerobic exercise, Int. J. Mol. Sci., 2021, vol. 22, no. 3, p. 1208.
Jones, J.C., Kroscher, K.A., and Dilger, A.C., Reductions in expression of growth regulating genes in skeletal muscle with age in wild type and myostatin null mice, BMC Physiol., 2014, vol. 14, p. 3.
Ikutomo, M., Sakakima, H., Matsuda, F., et al., Midkine-deficient mice delayed degeneration and regeneration after skeletal muscle injury, Acta Histochem., 2014, vol. 116, no. 2, p. 319.
Duffy, F.J., Seiler, J.G., Gelberman, R.H., and Hergrueter, C.A., Growth factors and canine flexor tendon healing: Initial studies in uninjured and repair models, J. Hand Surg. Am., 1995, vol. 20, no. 4, p. 645.
Majewski, M., Porter, R.M., Betz, O.B., et al., Improvement of tendon repair using miscle grafts transduced with TGF-β1 cDNA, Eur. Cells Mater., 2015, vol. 23, p. 94.
Klein, M.B., Yalamanchi, N., Pham, H., et al., Flexor tendon healing in vitro: effects of TGF-β on tendon cell collagen production, J. Hand Surg. Am., 2002, vol. 27, no. 4, p. 615.
González, M.N., Mello, W. de Butler-Browne, G.S., et al., HGF potentiates extracellular matrix-driven migration of human myoblasts: involvement of matrix metalloproteinases and MAPK/ERK pathway, Skeletal Muscle, 2017, vol. 7, no. 1, p. 20.
Karalaki, M., Fili, S., Philippou, A., and Koutsilieris, M., Muscle regeneration: cellular and molecular events, In Vivo (Brooklyn), 2009, vol. 23, no. 5, p. 779.
Arai, K.Y. and Nishiyama, T., Developmental changes in extracellular matrix messenger RNAs in the mouse placenta during the second half of pregnancy: possible factors involved in the regulation of placental extracellular matrix expression, Biol. Reprod., 2007, vol. 77, no. 6, p. 923.
Chen, C.P., Yang, Y.C., Su, T.H., et al., Hypoxia and transforming growth factor-β1 act independently to increase extracellular matrix production by placental fibroblasts, J. Clin. Endocrinol. Metab., 2005, vol. 90, no. 2, p. 1083.
Williams, P.E. and Goldspink, G., Connective tissue changes in surgically overloaded muscle, Cell Tissue Res., 1981, vol. 221, no. 2, p. 465.
Zamora, A.J. and Marini, J.F., Tendon and myo-tendinous junction in an overloaded skeletal muscle of the rat, Anat. Embryol. (Berlin), 1988, vol. 179, no. 1, p. 89.
White, J.P., Reecy, J.M., Washington, T.A., et al., Overload-induced skeletal muscle extracellular matrix remodelling and myofibre growth in mice lacking IL-6, Acta Physiol., 2009, vol. 197, no. 4, p. 321.
Funding
The study was carried out under of the Russian Science Foundation (grant no. 14-15-00768) and the budget topic of the research of the Faculty of Fundamental Medicine of Moscow State University “Systemic, Cellular and Molecular Mechanisms of Functioning of the Organism in Extreme Conditions.” Mass spectrometric measurements were performed on the equipment of the Center for Collective Use, Human Proteom, Institute of Biomedical Chemistry (Moscow).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
All studies were carried out in accordance with the principles of biomedical ethics formulated in the Declaration of Helsinki of 1964 and its subsequent updates and approved by the Commission on Biomedical Ethics of the Institute of Biomedical Problems of the Russian Academy of Sciences (Moscow) (Protocol No. 404).
CONFLICT OF INTEREST
The authors declare that they do not have a clear and potential conflict of interest associated with the publication of this article.
Contribution of authors to the publication. E.M. Lednev, E.A. Lysenko, O.L. Vinogradova, V.E. Dubrov, and D.V. Popov took part in organizing and conducting a physiological experiment with volunteers. E.M. Lednev, D.V. Popov, and V.E. Dubrov organized and performed biopsies of skeletal muscle tissue. V.G. Zgoda performed panoramic mass spectrometric proteomic analysis. G.R. Gazizova and E.I. Shagimardanov performed RNA sequencing. E.M. Lednev, E.A. Lysenko, P.A. Makhnovsky, O.L. Vinogradova, V.E. Dubrov, and D.V. Popov performed bioinformatics and statistical analysis of the obtained data and prepared the text of the manuscript.
INFORMED CONSENT
Each participant in the study provided a voluntary written informed consent signed by him after explaining to him the potential risks and benefits, as well as the nature of the upcoming study.
Rights and permissions
About this article
Cite this article
Lednev, E.M., Lysenko, E.A., Zgoda, V.G. et al. Eight-Week Aerobic Training Activates Extracellular Matrix Biogenesis in Human Skeletal Muscle. Hum Physiol 49, 129–137 (2023). https://doi.org/10.1134/S0362119722600436
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0362119722600436