Abstract
A manual labeling of 20 layers of the known open dataset EPFL for six classes is prepared. These classes are: (1) mitochondria, including their boundaries; (2) boundaries of mitochondria; (3) cell membranes; (4) postsynaptic densities (PSD); (5) axon sheaths; and (6) vesicles. Software for generating synthetic labeled datasets and the dataset itself balancing the representativeness of classes are created. Results of multiclass segmentation of brain electron microscopy (EM) data for each class for the case of binary segmentation and segmentation into five and six classes using a modified U-Net model are investigated. The model was trained on 256 × 256 fragments of the original EM resolution. In the case of six-class segmentation, mitochondria were segmented with the Dice–Sørensen coefficient of 0.908, which is somewhat lower than in the case of binary (0,911) and five-class segmentation (0.91). An extension of the dataset by synthesized images improved the classification results in an experiment. The extension of the manually labeled dataset (860 images of size 256 × 256) by the synthesized dataset (100 images of size 256 × 256 containing the poorly represented classes—axons and PSD) gave a significant increase of accuracy in the six-class U-Net model from 0.228 to 0.790 and from 0.553 to 0.745, respectively.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 INTRODUCTION
Considering the application of artificial intelligence methods and especially deep neural networks (DNN) for reconstructing brain electron microscopy (EM) data during the last ten years, we begin with the publication [1] of 2010. This paper actually announced the beginning of the use of serial block scanning electron microscopy as a source of high-resolution three-dimensional nanohistology for cells and tissues. A subsequent series of works was aimed at creating datasets for training deep learning networks and DNN methods and models for EM data segmentation designed for binary segmentation of brain cell organelles—neural membranes [2] and supervoxel segmentation of mitochondria [3]. Simultaneously, the problem of 3D reconstruction of the brain neural network and the problem of brain connectomics on the basis of neuron organelles and connections between neurons (synapses) is stated [4]. In this problem, of particular importance is the segmentation of such oganelles as postsynaptic densities (PSD), vesicles, and axons.
In [5], a team of 24 authors involved in the organization of the first international competition in 2D-segmentation of brain EM images claims that already at the conference on connectomics in 2014 organized by the Howard Hughes Medical Institute and Max Planck Society it became clear that convolutional networks became a dominating approach to detecting cell boundaries in serial EM images. The authors also suggest focusing on 3D processing of EM images and joined efforts in connectomics; however, they note that even the best modern algorithms for 3D reconstruction still require significant manual correction effort, which is available only for crowd sourcing. This opinion is supported by the earlier paper [6] of 21 authors from leading USA universities, which reports about the creation by joined effort of a saturated 3D reconstruction of a small (0.13 mm3) portion of an EM mouse neorcortex and a database of 1700 synapses in this portion.
The invention of U-Net in 2015 [7] opened a series of novel models and adaptations for segmenting brain EM data. The source of U-Net success is in involving the contextual information of the input image at all levels of processing. Almost immediately, the publication [8] experimentally confirmed that the skip connection of the U-Net architecture is effective for solving segmentation problems in biomedicine. U-Net also provided a basis for creating models with parallel inputs that make it possible to use correlation between inputs and, in particular, between EM layers in the 3D space [9, 10]. Next, attempts were made to use the capabilities of 3D convolutions for a multiple increase of the context amount in U-Net and U-Net-like networks—3DU-Net [11] (2016), V-Net [12] (2016), DeepMedic [13] (2017), and HighRes3DNet [14] (2017). This also gave a considerable effect, since the amount of context data for the 3D neighborhood of radius one of a voxel increases by a factor of three, and for the neighborhood of radius two of a voxel it increases by a factor of five.
An interesting direction of development of semantic segmentation implemented using fully convolutional networks is described in [15, 16]. The latter paper is most interesting and promising. For reconstructing the 3D interconnections of a system of neurons, a novel deep contextual network with a threefold reduction in resolution is proposed, which analyzes multiscale contextual information in a hierarchical structure of resolutions. The network architecture includes auxiliary classifiers that analyze the semantic meaning of image hierarchy and restrict themselves to low-level contextual features. As a dataset for the segmentation problem, ISBI 2012 is used. This method is aimed at minimizing human involvement and demonstrates a drift to explainable artificial intelligence (XAI).
The advantages of using 3D data analysis are undeniable; however, the use of 3D convolutional neural networks (CNN) with a 3D convolution kernel significantly increases the number of training parameters, computational cost, and memory consumption, which is especially sensitive for GPU applications. For this reason, the architectures using 3D convolutions are gradually replaced by the architectures that decrease the number of training parameters, the amount of memory, and increase the training speed, while preserving the quality of training and regulating balance between networks with 3D and 2D convolutions. In this process, various preprocessing methods are usually used, which often give an effect of 5% or more [17–19]. For example, in [20] contrast is enhanced using the adaptive gamma correction with wait distribution (AGCWD) [21]. Another trend is the factorization of low-rank convolutional kernels [22–25].
The paper [26] of 2019 reports the creation of a UNI-EM system with an interface convenient for subject matter experts. After labeling a small number of training samples, the system uses 2D and 3D deep learning networks and produces a segmentation of brain EM images for correcting the labeling and training parameters. UNI-EM comes with a set of 2D DNNs—U-Net, ResNet, HighwayNet, and DenseNet.
The paper [27] of 2019 determines the best version of U-Net using as an example the detection of vesicles in the data EM Transmission Electron Microscopy (TEM) with the resolution of 1.56 nm (two–three times better than the usual one) by comparing U-Net and Fully Residual U-Net (FRU-Net) architectures. It is found that the latter one improves accuracy by 4–5%. In the case of binary classification on three different datasets TEM, the error for FRU-Net did not exceed 10%. For the U-Net, the errors were 17, 27, and 17%.
The paper [28] of 2021 investigates the capabilities of Fully Residual U-Net (FRU-Net) with four levels resolution reduction (the original resolution of 640 × 640 is reduced four times by a factor of two to 40 × 40) using binary 2D segmentation of cell membranes as an example. Augmentation that increased the dataset by eight times due to rotations and reflections is created. On Drosophila EM dataset (ISBI 2012 EM segmentation challenge leaderboard, June 2020), the accuracy of about 98–99% of segmenting membranes was achieved. The publication [29] of 2021proposes a more complex network structure called hierarchical view-ensemble convolutional (HVEC) network as an alternative to a simple 3D structure. This structure inherits the abovementioned idea of [16] with three levels of resolution reduction and additional outputs for each level; next, the resolution reduction architecture is completed with a branch of resolution increase, which is typical for U-Net.
The application of artificial intelligence method for EM data processing is largely hampered by a small amount of labeled data for training and testing DNNs. Open EM data as a whole are represented by only a few labeled dataset, both due to the laboriousness of preparing samples for an electron microscope, and due to the lack of specialists for manual labeling. We found four open EM datasets the earliest and most popular of which are labeled only for one class (mitochondria or membranes). In the two other datasets, several classes are distinguished. As a result, the majority of neural networks used in EM processing are trained only to perform binary segmentation.
In connection with the above, the main aim of this work is to (1) create a dataset with manual multiclass labeling for a list of classes that provides a solution to the main modern tasks of EM data segmentation; (2) to develop algorithms for automatic generation of a dataset of synthetic objects of the specified main classes and create a dataset of synthetic objects, primarily those objects that are scarcely represented in the traditional datasets; (3) to study the capabilities of multiclass segmentation of U-Net-like architectures, starting with U-Net (in this work), using datasets with manual labeling and additional synthetic labeling.
2 DATA AND METHODS
In this section, we describe publicly available datasets. The most popular datasets for assessing the segmentation of mitochondria were collected by Lucchi et al. in [3].
It is seen that in three of the four labeled open datasets, only one class is labeled. Only one dataset contains more than one labeled class. For this reason, the vast majority of neural networks in EM are trained to classify only two classes (object and background).
We used the dataset EPFLВ or the data set of mitochondria segmentation Lucchi available at https://www.epfl.ch/labs/cvlab/data/data-em/. Initially, these data contain masks only for mitochondria. For this reason, to assess multiclass segmentation algorithms, we manually labeled 20 layers in the training sample (1024 × 768) and three layers for the following classes: (1) mitochondria, including their boundaries; (2) boundaries of mitochondria; (3) cell membranes; (4) postsynaptic densities (PSD); (5) axon sheaths; and (6) vesicles.
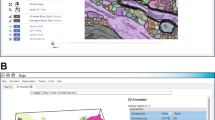
Accurate manual labeling of one layer takes 5–8 hours. Our labeling of the dataset EPFL is available at https://github.com/GraphLabEMproj/unet. We plan to continue the work on labeling and do this for both datasets. An example of labeling a layer fragment is shown in Fig. 1.
It just so happens that the axon sheath in the training dataset is present only in the first 36 layers and looks completely different from the axon sheath in the test dataset (Fig. 2). In the test dataset, the axon is represented in the first 70 layers, changes its shape for elongated to more rounded, and also has a darker interior and inner ring.
For the synthesized dataset, we generated 100 images of size 256 × 256 pixels containing the least represented classes—postsynaptic densities and axon sheaths. An example of data is shown in Fig. 3. The program for data generation is written in C#. The shape, size, and gray levels of compartments are chosen to be similar to the shape, size, and gray levels of the test dataset EPFL. To make the generated images more similar to real-life images, these images was blurred with a Gaussian filter with a kernel of radius seven, and Gaussian noise with a level of 20 was added. The advantage of a synthetic set is that you can get any number of images you need along with their labeling automatically.
2.1 Network Architecture
U-Net is considered to be a standard convolutional network architecture for image segmentation tasks. This architecture consists of a contracting path for capturing the global context and a symmetric expanding path that enables accurate localization. The basis of this network is the project U-Net https://github. com/zhixuhao/unet. In the original project, U-Net was used for binary classification of membranes. In this work, we use U-Net for multiclass segmentation. We copied the original repository, and made modifications in it, which are available at https://github.com/GraphLabEMproj/unet together with our labeling of the Lucci data.
Following the author of the code at https://github.com/zhixuhao/unet, the implementation of U-Net has some differences from the classical U-Net network [7]:
• The network input is an image of size 256 × 256 × 1.
• The network output is 256 × 256 × \(num\_classes\), where \(num\_classes\) is the number of classes.
• The sigmoid activation function guarantees that the mask is in the range [0, 1].
In addition, we added batch normalization after each ReLU convolution and activation layers.
3 EXPERIMENTAL RESULTS
3.1 Assessment Criteria
We use the Dice–Sørensen coefficient (DSC) and Jaccard’s coefficient (JAC), which are usually used for segmenting biomedical images. Define the number of correctly classified pixels as belonging to the target class (true positive) TP, the number of correctly classified background pixels (true negative) TN, the number of erroneously classified pixels as belonging to the target class (false positive) FP, and the number of erroneously classified background pixels (false negative) FN. Then, define the metrics as follows:
The values of the DSC and JAC vary from zero to one. By contrast with Jaccard’s coefficient, the corresponding difference function is not a correct distance metric since it does not satisfy the triangle inequality. JAC and DSC are equivalent in the sense they may be represented in terms of each other:
Since we consider multiclass segmentation in this work, we are interested in multiclass metrics. Since the Jaccard (or Dice) metrics compare two sets, in the case of multiclass classification the result will be a vector of Jaccard (or Dice) metrics for each class. For training a neural network, a scalar error function is used. Therefore, for multiclass segmentation, we should convolve the metric vector. To convolve a vector into a scalar, we use the linear convolution
where \({{\lambda }_{i}}\) is a weighting coefficient and \({{W}_{i}}\) is the value of the distance coefficient for the ith class. \({{W}_{{scalar}}}\) is a scalar value or convolution of a distance vector, and N is the number of classes.
In this work, we use the linear convolution of DSC with the weighting coefficients \({{\lambda }_{i}}\) equal to 1/N.
3.2 Experiments
To obtain a new training sample, twenty high-resolution images of the original training sample were cut into 256 × 256 fragments with an overlap of a quarter of the fragment size. In total, 860 fragments were obtained. To additionally increase the training sample, we made random rotations of images, random shifts, and random scale changes in a small range (5%).
To obtain a mixed training sample, we added to the original 860 fragments 100 synthesized fragments; thus, in total we have 960 fragments.
We selected 20% of images from the training sample into a validation sample with the batch size equal to seven. The model was tested on three layers (129 fragments). We used Adam’s optimizer with the training rate of 2 × 10–5. The training curves for different experiments are presented in Figs. 4 and 5.
Experiment 1. Five segmentation classes—mitochondrion with its boundary, membranes, PSD, axon sheaths, and vesicles. The number of epochs is 1000.
Experiment 2. Six segmentation classes—mitochondrion with its boundary, boundary of the mitochondrion, membranes, PSD, axon, and vesicles. The number of epochs is 1000.
One more class of mitochondria boundaries is added.
Experiment 3. One segmentation class—mitochondrion with its boundary. The number of epochs is 200.
It is seen from Table 2 that the quality of multiclass segmentation is only slightly inferior to binary segmentation.
The class mitochondria boundaries is a sublass of the class mitochondria with their boundaries, and the additional edge enhancement improves the segmentation results of the unifying class. The network was trained on unbalanced classes, since the sizes of compartments and their occurrence differ by dozens of times.
4 DISCUSSION
In this section, we discuss Table 3 “Comparison of mitochondria segmentation results,” in which we placed the most representative results on membrane segmentation using binary and multiclass models.
We tested our models on the entire dataset EPFL and used these values instead of the results presented in Table 2. We cannot directly compare the results in Table 3, since our models were trained on a significantly reduced version of EPFL. However, we can put forward several hypotheses that need to be tested. The worst results were obtained in layers containing an axon, fuzzy membranes, incomplete mitochondria, mitochondria with darker borders and darker inclusions than on labeled layers.We assume that labeling more layers or generating synthetic data with proper characteristics will improve the results.
5 CONCLUSIONS
We manually carried out the multiclass labeling of 20 layers of the training set and three layers of the test set for the well-known dataset EPFL, which includes the following classes: (1) mitochondria, including their boundaries; (2) boundaries of mitochondria; (3) membranes; (4) postsynaptic densities (PSD); (5) axon sheaths; and (6) vesicles. Software for generating synthetic labeled datasets with the same classes was developed. A synthetic labeled dataset that includes axons, PSD, and membranes was created.
Results of segmentation of multiclass brain electron microscopy data obtained using a modified U-Net with decomposition of data layers into 256 × 256 fragments while preserving the original resolution are presented.
The study showed that the results of binary, five-class and six-class segmentation are similar in quality: 0.911, 0.910 and 0.908, respectively. The quality of segmentation is affected by the presence of a sufficient number of specific features that distinguish the selected classes, and the representation of these features in the training sample.
The expansion of datasets by synthesized images improves the classification results. The expansion of a manually labeled dataset (860 256 × 256 images) by a synthesized dataset consisting of (100 256 × 256 images containing less represented classes (axons, PSD, and membranes) significantly improved the accuracy of the six-class model (see Table 2)—from 0.228 to 0.790, from 0.553 to 0.745, and from 0.743 to 0.750, respectively, in proportion to the liquidated deficit.
REFERENCES
Deerinck, T. et al., Enhancing serial block-face scanning electron microscopy to enable high resolution 3D nanohistology of cells and tissues, Microscopy Microanal., 2010, vol. 16, no. 2, pp. 1138–1139. https://doi.org/10.1017/S1431927610055170
Ciresan, D.C. et al., Deep neural networks segment neuronal membranes in electron microscopy images, IN NIPS, 2012, pp. 2852–2860.
Lucchi, A. et al., Supervoxel-based segmentation of mitochondria in EM image stacks with learned shape features, IEEE Trans. Medical Imaging, 2012, vol. 31, no. 2, pp. 474–486. https://doi.org/10.1109/TMI.2011.2171705
Helmstaedter, M. and Mitra, P.P., Computational methods and challenges for large-scale circuit mapping, Current Opinion Neurobiol., 2012, vol. 22, no 1, pp. 162–169. http://www.sciencedirect.com/science/article/pii/ S0959438811002133.https://doi.org/10.1016/j.conb.2011.11.010
Arganda-Carreras I. et al., Crowdsourcing the creation of image segmentation algorithms for connectomics, Frontiers Neuroanatomy, 2015, vol. 9, pp. 1–13. https://www.frontiersin.org/article/10.3389/fnana. 2015.00142.https://doi.org/10.3389/fnana.2015.00142
Kasthuri, N. et al., Saturated reconstruction of a volume of neocortex, Cell, 2015, vol. 162, pp. 648–661.
Ronneberger, O., Fischer, P., and Brox, T., U-Net: Convolutional Networks for Biomedical Image Segmentation, 2015. arXiv: 1505.04597 [cs.CV].
Drozdzal, M. et al. The importance of skip connections in biomedical image segmentation, 2016. arXiv: 1608.04117 [cs.CV].
Fakhry, A.E., Zeng, T., and Ji, S., Residual deconvolutional networks for brain electron microscopy image segmentation, IEEE Trans. Medical Imaging, 2017, vol. 36, pp. 447–456.
Xiao, C. et al., Deep contextual residual network for electron microscopy image segmentation in connectomics, 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), 2018, pp. 378–381. https://doi.org/10.1109/ISBI.2018.8363597.
Çiçek, Ö. et al., 3D U-Net: Learning Dense Volumetric Segmentation from Sparse Annotation. 2016. arXiv: 1606.06650 [cs.CV].
Milletari, F., Navab, N., and Ahmadi, S.-A., V-Net: Fully Convolutional Neural Networks for Volumetric Medical Image Segmentation, 2016 Fourth International Conference on 3D Vision (3DV), 2016, pp. 565–571.
Kamnitsas, K. et al., Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation, Med. Image Anal., 2017, vol. 36, pp. 61–78.
Li, W. et al., On the compactness, efficiency, and representation of 3D convolutional networks: Brain parcellation as a pretext task, Inf. Process. Med. Imaging, Ed. by Niethammer, M. Cham Springer, 2017, pp. 348–360.
Long, J., Shelhamer, E., and Darrell, T., Fully convolutional networks for semantic segmentation, 2015. arXiv: 1411.4038 [cs.CV].
Chen, H. et al., Deep contextual networks for neuronal structure segmentation, Proc. of the Thirtieth AAAI Conference on Artificial Intelligence (AAAI-16), 2016, pp. 1167–1173. https://ojs.aaai.org/index.php/AAAI/article/view/ 10141/10000.
Liu, T. et al., A modular hierarchical approach to 3D electron microscopy image segmentation, J. Neurosci. Meth., 2014, vol. 226, pp. 88–102.
Liu, J. et al., Automatic detection and segmentation of mitochondria from SEM images using deep neural network, 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), 2018, pp. 628–631.
Oztel, I. et al., Mitochondria segmentation in electron microscopy volumes using deep convolutional neural network, 2017 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 2017, pp. 1195–1200. https://doi.org/10.1109/BIBM.2017.8217827.
Žerovnik, Mekuč M. et al., Automatic segmentation of mitochondria and endolysosomes in volumetric electron microscopy data, Comput. Biol. Med., 2020, vol. 119, p. 103693. https://doi.org/10.1016/j.compbiomed.2020.103693. https://www.sciencedirect.com/science/article/pii/S0010482520300792.
Huang, S.-C., Cheng, F., and Chiu, Y., Efficient contrast enhancement using adaptive gamma correction with weighting distribution, IEEE Trans. Image Process., 2013, vol. 22, pp. 1032–1041.
Szegedy, C. et al., Rethinking the Inception Architecture for Computer Vision, 06/2016. https://doi.org/10.1109/CVPR.2016.308
Chollet, F., Xception: Deep Learning with depthwise separable convolutions, IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2017, pp. 1800–1807.
Xie, S. et al., Aggregated residual transformations for deep neural networks, 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2017, pp. 5987–5995.
Cheng, H.-C. and Varshney, A., Volume segmentation using convolutional neural networks with limited training data, 2017 IEEE International Conference on Image Processing (ICIP), 2017, pp. 590–594. https://doi.org/10.1109/ICIP.2017.8296349.
Urakubo, H. et al., UNI-EM: An environment for deep neural network-based automated segmentation of neuronal electron microscopic images, Sci. Rep., 2019, vol. 9, p. 19413. https://www.biorxiv.org/content/biorxiv/early/2019/04/12/607366.full.pdf.https://doi.org/10.1038/s41598-019-55431-0
Gómez-de-Mariscal, E. et al., Deep-learning-based segmentation of small extracellular vesicles in transmission electron microscopy images, Sci. Rep., 2019, vol. 9. https://doi.org/10.1038/s41598-019-49431-3
Quan, T.M., Hildebrand, D.G.C., and Jeong, W.-K., FusionNet: A deep fully residual convolutional neural network for image segmentation in connectomics, Frontiers Comput. Sci., 2021, vol. 3, pp. 34. https://www.frontiersin.org/article/10.3389/fcomp.2021.613981.https://doi.org/10.3389/fcomp.2021.613981
Yuan, Z. et al., HIVE-Net: Centerline-aware HIerarchical view-ensemble convolutional network for mitochondria segmentation in EM images, Comput. Meth. Programs Biomed., 2021, vol. 200, p. 105925.
ACKNOWLEDGMENTS
The study was supported by a grant from the strategic academic leadership program “Priority 2030” (project N‑483-99_2021-2022).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare that they have no conflicts of interest.
Additional information
Translated by A. Klimontovich
Rights and permissions
About this article
Cite this article
Getmanskaya, A.A., Sokolov, N.A. & Turlapov, V.E. Multiclass U-Net Segmentation of Brain Electron Microscopy Data Using Original and Semi-Synthetic Training Datasets. Program Comput Soft 48, 164–171 (2022). https://doi.org/10.1134/S0361768822030057
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0361768822030057