Abstract
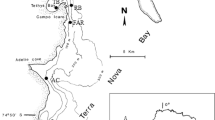
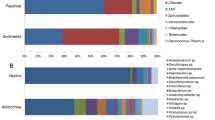
Bacterial diversity of two Lake Baikal endemic sponges characterized by different life forms, branching Lubomirskia baicalensis and encrusting Baikalospongia sp., was studied using 454 pyrosequencing of the 16S rRNA gene fragments. In the communities associated with L. baicalensis and Baikalospongia sp., 426 and 428 OTUs, respectively, were identified. In microbial associations of these sponges, 24 bacterial phyla with predominance of Bacteroidetes, Proteobacteria, and Actinobacteria were identified. Analysis of the taxonomic composition of bacterial communities of the sponges was carried out by searching the dominant phylotypes within the clusters of phylum level. Comparison of bacterial associations of the sponges with Lake Baikal bacterioplankton revealed both the shared OTUs and the unique ones characteristic of the studied species.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Taylor, M.W., Radax, R., Steger, D., and Wagner, M., Sponge-associated microorganisms: evolution, ecology, and biotechnological potential, Microbiol. Mol. Biol. Rev., 2007, vol. 71, pp. 295–347.
Mohamed, N.M., Rao, V., Hamann, M.T., Kelly, M., and Hill, R.T., Monitoring bacterial diversity of the marine sponge Ircinia strobilina upon transfer into aquaculture, Appl. Environ. Microbiol., 2008, vol. 74, pp. 4133–4143.
Thomas, T.R., Kavlekar, D.P., and LokaBharathi, P.A., Marine drugs from sponge-microbe association—a review, Mar. Drugs, 2010, vol. 8, pp. 1417–1468.
Henstchel, U., Piel, J., Degnan, S.M., and Taylor, M.W., Genomic insights into the marine sponge microbiome, Nat. Rev. Microbiol., 2012, vol. 10, pp. 641–654.
Manconi, R. and Pronzato, R., Suborder Spongillina subord. nov.: freshwater sponges, in Systema Porifera: A Guide to the Classification of Sponges, Hooper, N.J. and van Soest, R.W., Eds., NewYork: Kluwer Academic/Plenum, 2002, pp. 921–1019.
Lee, O.O., Wang, Y., Yang, J., Lafi, F.F., Al-Suwailem, A., and Qian, P.-Y., Pyrosequencing reveals highly diverse and species-specific microbial communities in sponges from the Red Sea, ISME J., 2011, vol. 5, pp. 650–664.
Parfenova, V.V., Terkina, I.L., Kostornova, T.Ya., Nikulina, I.G., Chernykh, V.I., and Maksimova, E.A., Microbial community of freshwater sponges in Lake Baikal, Biol. Bull., 2008, vol. 35, no. 4, pp. 374–379.
Kalyuzhnaya, O.V., Krivich, A.A., and Itskovich, V.B., Diversity of 16S rRNA genes in metagenomic community of the freshwater sponge Lubomirskia baicalensis, Russ. J. Genet., 2012, vol. 48, no. 8, pp. 855–858.
Gernert, C., Glockner, F.O., Krohne, G., and Hentschel, U., Microbial diversity of the freshwater sponge Spongilla lacustris, Microb. Ecol., 2005, vol. 50, pp. 206–212.
Costa, R., Keller-Costa, T., Newton, C.M., Gomes, N.T., da Rocha, U.N., van Overbeek, L., and van Elsas, J.D., Evidence for selective bacterial community structuring in the freshwater sponge Ephydatia fluviatilis, Microb. Ecol., 2013, vol. 65, no. 1, pp. 232–244.
Parfenova, V.V., Gladkikh, A.S., and Belykh, O.I., Comparative analysis of biodiversity in the planktonic and biofilm bacterial communities in Lake Baikal, Microbiology (Moscow), 2013, vol. 82, no. 1, pp. 91–101.
Chun, J., Kim, K.Y., Lee, J.H., and Choi, Y., The analysis of oral microbial communities of wild-type and toll-like receptor 2-deficient mice using a 454 GS FLX Titanium pyrosequencer, BMC Microbiol., 2010, 10:101. doi: 10.1186/1471-2180-10-101
Kim, O.S., Cho, Y.J., Lee, K., Yoon, S.H., Kim, M., Na, H., Park, S.C., Jeon, Y.S., Lee, J.H., Yi, H., Won, S., and Chun, J., Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species, Int. J. Syst. Evol. Microbiol., 2012, vol. 62, pp. 716–721.
Jackson, S.A., Kennedy, J., Morrissey, J.P., O’Gara, F., and Dobson, A.D., Pyrosequencing reveals diverse and distinct sponge-specific microbial communities in sponges from a single geographical location in Irish waters, Microb. Ecol., 2012, vol. 64, no. 1, pp. 105–116.
White, J.R., Patel, J., Ottesen, A., Arce, G., Blackwelder, P., and Lopez, J.V., Pyrosequencing of bacterial symbionts within Axinella corrugate sponges: diversity and seasonal variability, PLoS One, 7(6): e38204. doi: 10.1371/journal.pone.0038204
Ehrlich, H., Maldonado, M., Spindler, K.-D., Eckert, C., Hanke, T., Born, R., Goebel, C., Simon, P., Heinemann, S., and Worch, H., First evidence of chitin as a component of the skeletal fibers of marine sponges. Part I. Verongidae (Demospongia: Porifera), J. Exp. Zool. (Mol. Dev. Evol.), 2007, vol. 308B, pp. 347–356.
Newton, R.J., Jones, S.E., Eiler, A., McMahon, K.D., and Bertilsson, S., A guide to the natural history of freshwater lake bacteria, Microbiol. Mol. Biol., 2011, vol. 75, no. 1, pp. 14–49.
Nikitin, D.I., Strömp, C., Oranskaya, M.S., and Abraham, W.-R., Phylogeny of the ring-forming bacterium Arcicella aquatica gen. nov., sp. nov. (ex Nikitin et al. 1994), from a freshwater neuston biofilm, Int. J. Syst. Evol. Microbiol., 2004, vol. 54, pp. 681–684.
Bel’kova, N.L., Parfenova, V.V., Kostornova, T.Ya., Denisova, L.Ya., and Zaichikov, E.F., Microbial biodiversity in the water of Lake Baikal, Microbiology (Moscow), 2003, vol. 72, no. 2, pp. 203–212.
Parfenova, V.V., Mal’nik, V.V., Boiko, S.M., Sheveleva, N.G., Logacheva, N.F., Evstigneeva, T.D., Suturin, A.N., and Timoshkin, O.A., Communities of hydrobionts developing at the water-rock interface in Lake Baikal, Russ. J. Ecol., 2008, vol. 39, no. 3, pp. 198–204.
Zwart, G., Crump, B., Agterveld, M., Hagen, F., and Han, S., Typical freshwater bacteria: an analysis of available 16S rRNA gene sequences from plankton of lakes and rivers, Aquat. Microb. Ecol., 2002, vol. 28, pp. 141–155.
Bauer, M., Kube, M., Teeling, H., Richter, M., Lombardot, T., Allers, E., Würdemann, C.A., Quast, C., Kuhl, H., Knaust, F., Woebken, D., Bischof, K., Mussmann, M., Choudhuri, J.V., Meyer, F., Reinhardt, R., Amann, R.I., and Glöckner, F.O., Whole genome analysis of the marine Bacteroidetes ‘Gramella forsetii’ reveals adaptations to degradation of polymeric organic matter, Environ. Microbiol., 2006, vol. 8, pp. 2201–2213.
Berg, K.A., Lyra, C., Sivonen, K., Paulin, L., Suomalainen, S., Tuomi, P., and Rapala, J., High diversity of cultivable heterotrophic bacteria in association with cyanobacterial water blooms, ISME J., 2009, vol. 3, pp. 314–325.
Gleische, C., Fesefeldt, A., and Hirsch, P., Genus I. Hyphomicroblum Stutzer and Hartleb, 1898, in Bergey’s Manual of Systematic Bacteriology, Garrity, G.M., Ed., 2005, vol. 2, pp. 476–494.
Jezbera, J., Sharma, K.A., Brandt, U., Doolittle, W.F., and Hahn, M.W., Candidatus ‘Planktophila limnetica,’ an actinobacterium representing one of the most numerically important taxa in freshwater bacterioplankton, Int. J. Syst. Evol. Microbiol., 2009, vol. 59, pp. 2864–2869.
Fuerst, J.A. and Sagulenko, E., Beyond the bacterium: planctomycetes challenge our concepts of microbial structure and function, Nat. Rev. Microbiol., 2011, vol. 9, no. 6, pp. 403–413.
Zhang, H., Xu, Y., Ding, Y., Yang, Q., Ren, S., Li, X., Chen, X., Xu, Y., and Hao, H., Molecular diversity analysis of planctomycete-like bacteria in inosine fermentation and municipal wastewater treatment systems, Afr. J. Microbiol. Res., 2012, vol. 6, no. 37, pp. 6635–6641.
Lage, O.M., Bondoso, J., and Viana, F., Isolation and characterization of Planctomycetes from the sediments of a fish farm wastewater treatment tank, Arch. Microbiol., 2012, vol. 194, pp. 879–885.
Woebken, D., Teeling, H., Wecker, P., Dumitriu, A., Kostadinov, I., Delong, E.F., Amann, R., and Glöckner, F.O., Fosmids of novel marine Planctomycetes from the Namibian and Oregon coast upwelling systems and their cross-comparison with planctomycete genomes, ISME J., 2007, vol. 1, pp. 419–435.
Kulichevskaya, I.S., Ivanova, A.O., Belova, S.E., Baulina, O.I., Bodelier, P.L., Rijpstra, W.I., Sinninghe Damsté, J.S., Zavarzin, G.A., and Dedysh, S.N., Schlesneria paludicola gen. nov., sp. nov., the first acidophilic member of the order Planctomycetales, from Sphagnum-dominated boreal wetlands, Int. J. Syst. Evol. Microbiol., 2007, vol. 57, pp. 2680–2687.
Schmid, M., Schmitz-Esser, S., Jetten, M., and Wagner, M., 16S-23S rDNA intergenic spacer and 23S rDNA of anaerobic ammonium-oxidizing bacteria: implications for phylogeny and in situ detection, Environ. Microbiol., 2001, vol. 3, no. 7, pp. 450–459.
Mohamed, N.M., Saito, K., Tal, Y., and Hill, R.T., Diversity of aerobic and anaerobic ammonia-oxidizing bacteria in marine sponges, ISME J., 2010, vol. 4, no. 1, pp. 38–48.
Elshahed, M.S., Najar, F.Z., Aycock, M., Qu, C., Roe, B.A., and Krumholz, L.R., Metagenomic analysis of the microbial community at Zodletone spring (Oklahoma): insights into the genome of a member of the novel candidate division OD1, Appl. Environ. Microbiol., 2005, vol. 71, no. 11, pp. 7598–7602.
Wrighton, K.C., Thomas, B.C., Sharon, I., Miller, C.S., Castelle, C.J., VerBerkmoes, N.C., Wilkins, M.J., Hettich, R.L., Lipton, M.S., Williams, K.H., Long, P.E., and Banfield, J.F., Fermentation, hydrogen, and sulfur metabolism in multiple uncultivated bacterial phyla, Science, 2012, vol. 337, no. 6102, pp. 1661–1665.
Peura, S., Eiler, A., Bertilsson, S., Nykänen, H., Tiirola, M., and Jones, R.I., Distinct and diverse anaerobic bacterial communities in boreal lakes dominated by candidate division OD1, ISME J, 2012, vol. 6, pp. 1640–1652.
Schmitt, S., Deines, P., Behnam, F., Wagner, M., and Taylor, M.W., Chloroflexi bacteria are more diverse, abundant, and similar in high than in low microbial abundance sponges, FEMS Microbiol. Ecol., 2011, vol. 78, pp. 497–510.
Philips, S., Laanbroek, H.J., and Verstraete, W., Origin, causes, and effects of increased nitrite concentrations in aquatic environments, Rev. Environ. Sci. Biotechnol., 2002, vol. 1, pp. 115–141.
van der Gucht, K., Vandekerckhove, T., Vloemans, N., Cousin, S., Muylaert, K., Sabbe, K., Gillis, M., Declerk, S., De Meester, L., and Vyverman, W., Characterization of bacterial communities in four freshwater lakes differing in nutrient load and food web structure, FEMS Microbiol. Ecol., 2005, vol. 53, no. 2, pp. 205–220.
Liu, Y., Yao, T., Jiao, N., Kang, S., Zeng, Y., and Huang, S., Microbial community structure in moraine lakes and glacial meltwaters, Mount Everest, FEMS Microbiol. Lett., 2006, vol. 265, no. 1, pp. 98–105.
Bultel-Poncé, V., Berge, J.P., Debitus, C., Nicolas, J.L., and Guyot, M., Metabolites from the sponge-associated bacterium Pseudomonas species, Mar. Biotechnol., 1999, vol. 1, pp. 384–390.
Lee, Y.K., Lee, J.-H., and Lee, H.K., Microbial symbiosis in marine sponges, J. Microbiol., 2001, vol. 39, no. 4, pp. 254–264.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © A.S. Gladkikh, Ok.V. Kalyuzhnaya, O.I. Belykh, T.S. Ahn, V.V. Parfenova, 2014, published in Mikrobiologiya, 2014, Vol. 83, No. 6, pp. 682–693.
Rights and permissions
About this article
Cite this article
Gladkikh, A.S., Kalyuzhnaya, O.V., Belykh, O.I. et al. Analysis of bacterial communities of two Lake Baikal endemic sponge species. Microbiology 83, 787–797 (2014). https://doi.org/10.1134/S002626171406006X
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S002626171406006X