Abstract
The increasing ecological significance of Planctomycetes and the still limited knowledge of this group prompted us to obtain cultured isolates from the sediment of a treatment water recycling tank of a marine fish farm. Presence of strains from this group was assessed in the sediments and water column of the tank. Eleven isolates were obtained from the sediment sample by exploiting Planctomycetes natural resistance to several antibiotics and their capacity to degrade organic matter. Based on morphological characteristics and resistance to antibiotics, Planctomycetes were identified. Their phylogenetic affiliation was confirmed by the sequence analysis of the 16S rRNA gene that revealed the presence of a group of 6 isolates closely related to Rhodopirellula baltica and a cluster of 5 isolates with 97.7–97.9 % of similarity to this species, which probably are a different species of Rhodopirellula. ERIC-PCR profiles showed a higher discrimination within the two groups and allowed the identification of nine different genotypes within the isolated strains. This work corroborates the association of Rhodopirellula spp. with fish farm environments.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The Planctomycetes comprise one phylum of the PVC superphylum that also contains the phyla Verrucomicrobia, Chlamydiae, and Lentisphaerae (Wagner and Horn 2006). These remarkable bacteria are generally characterized by budding reproduction, membrane-bounded compartments within the cells, and absence of peptidoglycan in their proteinaceous cell walls (Ward et al. 2006). Other cellular eukaryote-like features, presence of endocytosis-like protein uptake, and membrane coat-like proteins (Lonhienne et al. 2010; Santarella-Mellwig et al. 2010) were observed in one genus evidencing the evolutionary relevance of this group.
Molecular ecology and culture studies have revealed the widespread distribution of Planctomycetes in a broad range of terrestrial and aquatic environments and in association with several organisms namely sponges, the giant tiger prawn, corals, macroalgae, and the human gut (Nelson et al. 2010; and see Fuerst and Sagulenko 2011 for a review). Higher representation of Planctomycetes was observed in marine sediments (Rusch et al. 2003; Musat et al. 2006; Chipman et al. 2010; Hu et al. 2010) and on the biofilm of the kelp Laminaria hyperborea (Bengtsson and Ovreas 2010; Bengtsson et al. 2010) and Fucus vesiculosus (Lachnit et al. 2011) when compared with coastal waters or the marine pelagic environment (Rusch et al. 2007). But the highest density of Planctomycetes in natural environments was found in the hindgut of soil-feeding termites Cubitermes spp., where they reach concentrations of 2.6 × 109 cells mL−1 (Kohler et al. 2008).
Planctomycetes isolates have been obtained from oligotrophic to highly polluted environments suggesting a broad metabolic adaptation. They are believed to play an important role in the global carbon and nitrogen cycles (Strous et al. 1999; Glockner et al. 2003) through, for example, the mineralization of marine snow particles (DeLong et al. 1993) and the anaerobic ammonia oxidation—anammox (Strous et al. 1999). Furthermore, the unexpected presence of key genes for 110 sulfatases in Rhodopirellula baltica (Glockner et al. 2003) suggests their involvement in the transformation of inorganic sulfur. Up-to-now, the majority of the known Planctomycetes strains is chemoheterotrophic or chemoautotrophic (anammox). Despite the information gathered in the last years, the physiology and metabolism of Planctomycetes are still not well understood (Ward et al. 2006), and an effort to improve the number of new cultivable strains is important. Improvements in the isolation methodology of Planctomycetes have been achieved (Schlesner 1994; Zengler et al. 2002; Winkelmann and Harder 2009; Lage and Bondoso 2011).
In the last decade, the great diversity of the microbial communities associated with aquaculture systems have been revealed using molecular tools (Tal et al. 2003; Cytryn et al. 2005; Sugita et al. 2005; Michaud et al. 2009; Gao et al. 2012). Although not the most frequent, Planctomycetes seem to be usual inhabitants of aquaculture environments (Sugita et al. 2005; Bissett et al. 2006; Kawahara et al. 2009; Michaud et al. 2009; Gao et al. 2012). In this study, we describe the isolation of Planctomycetes from the sediment of a treatment water recycling tank of a marine fish farm.
Materials and methods
Isolation, cultivation, and maintenance
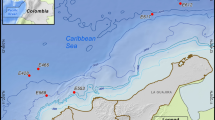
Samples were collected from a fish farm located in the north region of Portugal (41°27′10″N and 8°46′27″W) nearby the sea. The fish tanks used for the growth of turbot, Scophthalmus maximus, European sea bass, Dicentrarchus labrax, and sole, Solea senegalensis, were feed with a mixture of seawater captured directly from the sea and the water coming from the treatment water recycling tanks. The water coming from the fish tanks goes through sedimentation and nitrification in the treatment water recycling tank where samples were collected. The dimensions of the tank are 10 × 25 × 1.2 m. The temperature of the water varied between 7 and 15 °C (mean 11 °C), and the pH was 8. The samples (water and sediments, n = 1) were collected directly using sterile 1-L glass bottles or 50-mL sterile collection vials and brought immediately to the laboratory. The sediments, 0.1 g, were washed in sterile seawater to remove the remaining water from the tank and resuspended in 1 mL of PBS with vortexing to suspend the attached bacteria. About 500 mL of the water was concentrated by centrifugation at 5,000 rpm during 30 min, and the concentrated cells were resuspended in 1 mL PBS. Aliquots of the prepared sediment and water samples (100 μl) were spread into five solid media, modified M13, M14, Zobell, M590 (Table 1), and R2A (BD-Difco™). For the selective isolation of Planctomycetes, the media were supplemented with 200 μg mL−1 ampicillin, 500 μg mL−1 streptomycin, and 20 μg mL−1 cycloheximide. The cultures were incubated in the dark at 20 °C, for one month. Screening for Planctomycetes was done by light microscopy. Pure cultures were obtained by subcultivation in the same isolation media and routinely maintained in M13. For long-term purpose storage, pure cultures were stored in M13 liquid medium with 20 % (w/v) glycerol and kept at −80 °C.
Electron microscopy
For transmission electron microscopy, cells were harvested from 3-day-old cultures in M13, fixed in 2.5 % (w/v) glutaraldehyde in marine buffer (pH 7.2) for 2 h, and post-fixed in 1 % (v/v) osmium tetroxide in the water for 4 h and 1 % uranyl acetate for 1 h. Cells were dehydrated through a graded ethanol series, then propylene oxide, and embedded in Epon resin. Ultrathin sections were stained for 10 min in 1 % (v/v) uranyl acetate and for 10 min in Reynolds lead citrate. The sections were examined in a JEOL 100CXII transmission electron microscope.
Molecular identification of the isolates
Identification of the isolates was done by 16S rRNA gene analysis after amplification with the primers 27f and 1492r (Lane 1991) by direct colony PCR as described by Lage and Bondoso (2011). Briefly, the PCR mixture consisted of 1× PCR buffer, 1.5 mM MgCl2, 1 unit of GoTaq Flexi DNA Polymerase, 200 μM of each deoxynucleoside triphosphate, 2 μM of each primer, and material from a single colony to a final volume of 50 μl. Thermal PCR profile was done in MyCycler™ Thermo Cycler (Bio-Rad) and consisted in an initial denaturing step of 5 min at 95 °C, 30 cycles of 1 min at 94 °C, 1 min at 52 °C, and 90 s at 72 °C, and a final extension of 5 min at 72 °C. The obtained PCR products were purified using the GFX™ PCR DNA and Gel Band purification Kit (Amersham Biosciences), cloned into pGEM-T Easy vector (Promega) following standard protocols, and sequenced with universal primers M13F and M13R. Sequencing was performed at STAB Vida (Oeiras, Portugal). The 16S rDNA sequences obtained were assembled with Vector NTI Advance™ 10.3, compared with known sequences in GenBank and aligned using ClustalW (Thompson et al. 1994). The sequences were corrected manually using the alignment generated by ClustalW. Phylogenetic and molecular evolutionary analyses were conducted using MEGA version 5.03 (Tamura et al. 2007). Phylogenetic trees were generated with different calculation methods including neighbor joining (NJ), maximum parsimony (MP), and maximum likelihood (ML) to test for stability of the tree. Final ML tree applying Jukes–Cantor model was calculated with 33 sequences. Validation of reproducibility of the branching patterns was made by bootstrap based on 1,000 resamplings.
ERIC-PCR fingerprinting
DNA of the isolates was extracted using E.Z.N.A Bacterial DNA Isolation Kit (Promega) according to the manufacturer’s instructions from 7-day cultures on agar plates. The PCR was performed as described before (Rademaker and De Bruijn 1997) with the pair of primers ERIC1R and ERIC2 (Versalovic et al. 1991), and 100 ng of template DNA was used in each reaction. The annealing temperature was 48 °C. The PCR was carried out in a MyCycler™ Thermo Cycler (Bio-Rad). Each ERIC-PCR product (ten microliters) was separated by electrophoresis in a 2 % agarose gel at 60 V for 90 min in Tris–acetate–EDTA buffer. The gel was stained with GelRed 1:100,000 (v/v) during 45 min, and images were captured using GenoPlex (VWR). Computer-assisted analysis of the fingerprintings was performed using the software BioNumerics 6.5 (Applied Maths, Belgium).
Nucleotide sequence accession numbers
The 16S rRNA gene sequences of the isolated strains determined in this study were submitted to the GenBank and have been given accession numbers DQ851134, EF012748 to EF012750, and EF421447 to EF421453.
Results and discussion
Isolation and morphological identification
After about 1-month incubation, eleven pink colonies appeared on the solid medium M13 inoculated with sediment sample from the water recycling treatment tank. These isolates were designated as strains OJF1, 4, 5, 6, 9, 10, 11, 12, 14, and 15. No isolation was achieved on the other solid media or from the water column of the tank. The selectivity of the medium was provided by the ampicillin, a β-lactamic antibiotic that is ineffective against Planctomycetes, as they do not possess peptidoglycan in their cell walls, and by streptomycin as described by Schlesner (1994). These antibiotics prevented the overgrowth of other bacteria. The grown colonies were small round and convex with a granular appearance and a pink pigmentation characteristic of some Planctomycetes.
The cells of all the isolates were generally pear-shaped having the rosette-like cell aggregation (Fig. 1) with a variable number of cells characteristic of Rhodopirellula-like strains (Schlesner et al. 2004; Ward et al. 2006; Winkelmann and Harder 2009). Cells were about 0.7 × 1.5 μm. They reproduced by budding. In young cultures, many unicellular forms were mobile as observed by light microscopy. This is the first phase of some Planctomycetes life cycles, which occur prior to the loss of cell mobility when the cells become sessile (Staley 1973; Gade et al. 2005b). Under the electron microscope, cells presented polarity due to the asymmetric location of the DNA and fimbriae (Fig. 2). On this pole, crateriform structures are also evident. These morphologic characteristics are representative of this phylum and were indicative of the presence of Planctomycetes.
Isolation was only obtained in M13 medium, the media that has been used for the successful isolation and cultivation of strains of Rhodopirellula (Schlesner 1986; Winkelmann and Harder 2009). Medium M14 is appropriate for Blastopirellula (Schlesner 1986) and did not provide the isolation of Rhodopirellula spp. The other three tested media commonly used in isolation from marine environments (R2A, M590, and Zobell) were unable to provide growth probably due to the lack of vitamin B12 needed by Rhodopirellula spp. (Schlesner et al. 2004).
Molecular identification by 16S rDNA and ERIC fingerprinting of the strains
The analysis of the 16S rRNA gene confirmed the affiliation of all the obtained isolates to the Planctomycetes (Fig. 3). The strains were affiliated to Rhodopirellula and were divided in two clusters. Cluster A, which has a 99.5–99.9 % similarity to R. baltica, represents strains of this species. Cluster B, although also closely related to R. baltica, only has 97.7–97.9 % similarity to this species. This degree of similarity may be indicative of a new species of Rhodopirellula. Stackebrandt and Ebers (2006) considered a threshold of 98.7 % similarity in the 16S rRNA gene sequence as the separation for a new species. A similarity of 99.8–100 % exists among the members of cluster B. This putative new species has also been found in the epiphytic community of macroalgae from the north coast of Portugal (Lage and Bondoso 2011). Despite the differences in the geographic and environmental conditions, strain OJF1 is very similar to strain Rhodopirellula sp. LF2 as confirmed by the BOX-PCR fingerprinting (data not shown). Rhodopirellula sp. LF2 was isolated from the surface of Laminaria sp. collected in Foz, Porto (Lage and Bondoso 2011). R. baltica is a cosmopolitan species of Planctomycetes as confirmed by 16S rRNA gene sequence analysis, DNA–DNA hybridization, and MLSA analysis (Schlesner et al. 2004; Winkelmann et al. 2010). It was found for the first time as a free-living organism in the water column in the Baltic Sea (Schlesner 1994) and, posteriorly, in the tissue of the Mediterranean sponge Aplysina aerophoba (Gade et al. 2004), in marine snow (Pacific Ocean) (DeLong et al. 1993), in the biofilm of macroalgae (Bengtsson and Ovreas 2010; Lage and Bondoso 2011), and in several European Seas (Winkelmann and Harder 2009).
Phylogenetic 16S rDNA tree generated by maximum-likelihood analysis based on Jukes–Cantor model indicating the relationship between the strains isolated to members of the Planctomycetes. Isolates obtained in this study are shown in bold. GenBank accession numbers are shown in parenthesis. The anammox “Candidatus” genera were used as outgroup. The tree with the highest log likelihood is shown. The percentage of trees in which the associated taxa clustered together is shown above the branches. Bar 0.05 substitutions per 100 nucleotides
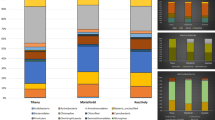
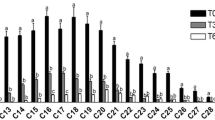
Intraspecies variability was accessed with the ERIC-PCR fingerprinting (Fig. 4). With an 80 % similarity cutoff in the ERIC-PCR fingerprinting, it is possible to differentiate 9 genotypes. In cluster A, we can discriminate 5 genotypes, with OJF6 and OJF14 being part of the same genotype; in cluster B, we can see 4 genotypes in which strains OJF9 and OJF10 are the same. This fingerprinting analysis indicated that strains OJF1 and OJF12 are more similar to R. baltica strains than to cluster B strains. Simpson index for the whole community is 0.018, with cluster B presenting a higher diversity (0.05) than cluster A (0.033). Genetic diversity in Rhodopirellula was also reported by Winkelmann et al. (2010), confirming the presence of different genotypes within this genus.
Dendrogram of ERIC-PCR patterns of the isolated Rhodopirellula sp. strains, using curve-based Pearson-UPGMA method. The clusters A and B based on the 16S rDNA phylogenetic tree are evidenced. Gel images were analyzed with Bionumerics 6.5. Numbers on the tree refer to the similarity between the isolated strains
Planctomycetes in aquacultures environments
Several Planctomycetes were reported in association with aquacultures environments namely Planctomyces, Pirellula, Rhodopirellula, and the “Candidatus” anammox (Sugita et al. 2005; Bissett et al. 2006; Kawahara et al. 2009; Michaud et al. 2009; Gao et al. 2012) reinforcing our findings. The ecological role of Planctomycetes, namely R. baltica, in fish farm environments may be due to their ability to perform the dissimilatory nitrate reduction to ammonium (DNRA) as reveled by in silico analysis of R. baltica genome (Mohan et al. 2004; Simon et al. 2011). This hypothesis is supported by the fact that in carbon-rich habitats, like fish farm sediments, DNRA pathway is greater than denitrification (Christensen et al. 2000; Nizzoli et al. 2006). Furthermore, they might also be important players in mineralization of organic matter as heterotrophic bacteria are known to play a key role in the biodegradation of organic material in marine environments (Deming and Baross 1993). R. baltica is a known specialized aerobic carbohydrate utilizer (Schlesner et al. 2004; Gade et al. 2005a), and strains belonging to Cluster B also utilized a wide variety of carbohydrates (unpublished data).
Information on the occurrence of antibiotic-resistant bacteria in marine sediments near fish farms and their relationship with population composition in the microbial community is still scarce (Schmidt et al. 2000; Tendencia and de la Peña 2001). According to Chelossi et al. (2003), in Italy, ampicillin is one of the most employed antibiotics against fish pathologies due to Vibrio spp., Pasteurella piscicida, and Myxobacterium spp. This and other classes of antibiotics, namely streptomycin, are used in aquaculture practice for the control of diseases (Kummerer 2009). Our work evidences the presence of Rhodopirellula spp. on the treatment of effluents or recirculation water in marine hatcheries. Their ability to degrade a variety of carbon sources (Gade et al. 2005a) and their natural resistance to high levels of streptomycin and β-lactamic antibiotics may be indicative of an ecological role in these kinds of habitats.
Conclusions
The isolated strains from the sediments of a water recycling treatment tank of a marine fish farm were identified as Planctomycetes by the combination of their morphological characteristics and antibiotic resistance. Further investigation using molecular techniques revealed the presence of two phylogenetic groups belonging to Rhodopirellula. A potential new species of Rhodopirellula was isolated. The ERIC fingerprinting showed a higher diversity within the isolates than the one revealed by the 16S rDNA.
References
Bengtsson MM, Ovreas L (2010) Planctomycetes dominate biofilms on surfaces of the kelp Laminaria hyperborea. BMC Microbiol 10:261
Bengtsson MM, Sjøtun K, Øvreås L (2010) Seasonal dynamics of bacterial biofilms on the kelp Laminaria hyperborea. Aquat Microb Ecol 60:71–83
Bissett A, Bowman J, Burke C (2006) Bacterial diversity in organically-enriched fish farm sediments. FEMS Microbiol Ecol 55:48–56
Chelossi E, Vezzulli L, Milano A, Branzoni M, Fabiano M, Riccardi G, Banat IM (2003) Antibiotic resistance of benthic bacteria in fish-farm and control sediments of the Western Mediterranean. Aquaculture 219:83–97
Chipman L, Podgorski D, Green S, Kostka J, Cooper W, Huettel M (2010) Decomposition of plankton-derived dissolved organic matter in permeable coastal sediments. Limnol Oceanogr 55:857–871
Christensen PB, Rysgaard S, Sloth NP, Dalsgaard T, Schwaerter S (2000) Sediment mineralization, nutrient fluxes, denitrification and dissimilatory nitrate reduction to ammonium in an estuarine fjord with sea cage trout farms. Aquat Microb Ecol 21:73–84
Cohen-Bazire G, Sistrom WR, Stanier RY (1957) Kinetic studies of pigment synthesis by non-sulphur purple bacteria. J Cell Comp Physiol 49:25–68
Cytryn E, van Rijn J, Schramm A, Gieseke A, de Beer D, Minz D (2005) Identification of bacteria potentially responsible for oxic and anoxic sulfide oxidation in biofilters of a recirculating mariculture system. Appl Environ Microbiol 71:6134–6141
DeLong EF, Franks DG, Alldredge L (1993) Phylogenetic diversity of aggregate-attached vs. free-living marine bacterial assemblages. Limnol Oceanogr 38:924–934
Deming JW, Baross JA (1993) The early diagenesis of organic matter: bacterial activity. In: Engel MH, Macko SA (eds) Organic geochemistry: principles and applications. Plenum Press, New York, pp 119–144
Fuerst JA, Sagulenko E (2011) Beyond the bacterium: planctomycetes challenge our concepts of microbial structure and function. Nat Rev Microbiol 9:403–413. doi:10.1038/nrmicro2578
Gade D, Gobom J, Rabus R (2005a) Proteomic analysis of carbohydrate catabolism and regulation in the marine bacterium Rhodopirellula baltica. Proteomics 5:3672–3683
Gade D, Stührmann T, Reinhardt R, Rabus R (2005b) Growth phase dependent regulation of protein composition in Rhodopirellula baltica. Environ Microbiol 7:1074–1084
Gao X-Y, Xu Y, Liu Y, Liu Z-P (2012) Bacterial diversity, community structure and function associated with biofilm development in a biological aerated filter in a recirculating marine aquaculture system. Mar Biodivers 1–11
Glockner FO, Kube M, Bauer M, Teeling H, Lombardot T, Ludwig W, Gade D, Beck A, Borzym K, Heitmann K, Rabus R, Schlesner H, Amann R, Reinhardt R (2003) Complete genome sequence of the marine planctomycete Pirellula sp. strain 1. Proc Natl Acad Sci USA 100:8298–8303
Hu YF, Fu CZ, Yin YS, Cheng G, Lei F, Yang X, Li J, Ashforth EJ, Zhang LX, Zhu BL (2010) Construction and preliminary analysis of a deep-sea sediment metagenomic fosmid library from Qiongdongnan Basin, South China Sea. Mar Biotechnol 12:719–727
Kawahara N, Shigematsu K, Miyadai T, Kondo R (2009) Comparison of bacterial communities in fish farm sediments along an organic enrichment gradient. Aquaculture 287:107–113
Kohler T, Stingl U, Meuser K, Brune A (2008) Novel lineages of Planctomycetes densely colonize the alkaline gut of soil-feeding termites (Cubitermes spp.). Environ Microbiol 10:1260–1270
Kummerer K (2009) Antibiotics in the aquatic environment—a review—part I. Chemosphere 75:417–434
Lachnit T, Meske D, Wahl M, Harder T, Schmitz R (2011) Epibacterial community patterns on marine macroalgae are host-specific but temporally variable. Environ Microbiol 13:655–665
Lage OM, Bondoso J (2011) Planctomycetes diversity associated with macroalgae. FEMS Microbiol Ecol 78:366–375
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, New York, pp 115–175
Lonhienne TGA, Sagulenko E, Webb RI, Lee KC, Franke J, Devos DP, Nouwens A, Carroll BJ, Fuerst JA (2010) Endocytosis-like protein uptake in the bacterium Gemmata obscuriglobus. Proc Natl Acad Sci USA 107:12883–12888
Michaud L, Lo Giudice A, Troussellier M, Smedile F, Bruni V, Blancheton JP (2009) Phylogenetic characterization of the heterotrophic bacterial communities inhabiting a marine recirculating aquaculture system. J Appl Microbiol 107:1935–1946
Mohan SB, Schmid M, Jetten M, Cole J (2004) Detection and widespread distribution of the nrfA gene encoding nitrite reduction to ammonia, a short circuit in the biological nitrogen cycle that competes with denitrification. FEMS Microbiol Ecol 49:433–443
Musat N, Werner U, Knittel K, Kolb S, Dodenhof T, van Beusekom JEE, de Beer D, Dubilier N, Amann R (2006) Microbial community structure of sandy intertidal sediments in the North Sea, Sylt-Rømø Basin, Wadden Sea. Syst Appl Microbiol 29:333–348
Nelson KE et al (2010) A catalog of reference genomes from the human microbiome. Science 328:994–999
Nizzoli D, Welsh DT, Fano EA, Viaroli P (2006) Impact of clam and mussel farming on benthic metabolism and nitrogen cycling, with emphasis on nitrate reduction pathways. Mar Ecol Prog Ser 315:151–165
Rademaker JLW, De Bruijn FJ (1997) Characterization and classification of microbes by REP-PCR genomic fingerprinting and computer-assisted pattern analysis. In: Caetano-Anollés G, Gresshoff PM (eds) DNA markers: protocols, applications and overviews. Wiley, New York, pp 1–26
Rusch A, Huettel M, Reimers CE, Taghon GL, Fuller CM (2003) Activity and distribution of bacterial populations in Middle Atlantic Bight shelf sands. FEMS Microbiol Ecol 44:89–100
Rusch DB et al (2007) The Sorcerer II global ocean sampling expedition: northwest Atlantic through eastern tropical Pacific. PLoS Biol 5:e77
Santarella-Mellwig R, Franke J, Jaedicke A, Gorjanacz M, Bauer U, Budd A, Mattaj IW, Devos DP (2010) The compartmentalized bacteria of the planctomycetes-verrucomicrobia-chlamydiae superphylum have membrane coat-like proteins. PLoS Biol 8:e1000281
Schlesner H (1986) Pirella marina sp. nov., a budding, peptidoglycan-less bacterium from brackish water. Syst Appl Microbiol 8:177–180
Schlesner H (1994) The development of media suitable for the microorganisms morphologically resembling Planctomyces spp., Pirellula spp., and other Planctomycetales from various aquatic habitats using dilute media. Syst Appl Microbiol 17:135–145
Schlesner H, Rensmann C, Tindall BJ, Gade D, Rabus R, Pfeiffer S, Hirsch P (2004) Taxonomic heterogeneity within the Planctomycetales as derived by DNA–DNA hybridization, description of Rhodopirellula baltica gen. nov., sp. nov., transfer of Pirellula marina to the genus Blastopirellula gen. nov. as Blastopirellula marina comb. nov. and emended description of the genus Pirellula. Int J Syst Evol Microbiol 54:1567–1580
Schmidt AS, Bruun MS, Dalsgaard I, Pedersen K, Larsen JL (2000) Occurrence of antimicrobial resistance in fish-pathogenic and environmental bacteria associated with four Danish rainbow trout farms. Appl Environ Microbiol 66:4908–4915
Simon J, Kern M, Hermann B, Einsle O, Butt JN (2011) Physiological function and catalytic versatility of bacterial multihaem cytochromes c involved in nitrogen and sulfur cycling. Biochem Soc Trans 39:1864–1870
Stackebrandt E, Ebers J (2006) Taxonomic parameters revisited: tarnished gold standards. Microbiol Today 33:152–155
Staley JT (1973) Budding bacteria of the Pasteuria-Blastobacter group. Can J Microbiol 19:609–614
Strous M, Fuerst JA, Kramer EHM, Logemann S, Muyzer G, Pas-Schoonen KTVD, Webb R, Kuenen JG, Jetten MSM (1999) Missing lithotroph identified as new Planctomycete. Nature 400:446–449
Sugita H, Nakamura H, Shimada T (2005) Microbial communities associated with filter materials in recirculating aquaculture systems of freshwater fish. Aquaculture 243:403–409
Tal Y, Watts JEM, Schreier SB, Sowers KR, Schreier HJ (2003) Characterization of the microbial community and nitrogen transformation processes associated with moving bed bioreactors in a closed recirculated mariculture system. Aquaculture 215:187–202
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tendencia EA, de la Peña LD (2001) Antibiotic resistance of bacteria from shrimp ponds. Aquaculture 195:193–204
Thompson JD, Higgins DG, Gibson TJ (1994) Clustal-W—improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Versalovic J, Koeuth T, Lupski JR (1991) Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res 19:6823–6831
Wagner M, Horn M (2006) The Planctomycetes, Verrucomicrobia, Chlamydiae and sister phyla comprise a superphylum with biotechnological and medical relevance. Curr Opin Biotechnol 17:241–249
Ward N, Staley JT, Fuerst JA, Giovannoni S, Schlesner H, Stackebrandt E (2006) The order Planctomycetales, including the genera Planctomyces, Pirellula, Gemmata and Isosphaera and the Candidatus genera Brocadia, Kuenenia and Scalindua. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The prokaryotes: a handbook on the biology of bacteria. Springer, New York, pp 757–793
Winkelmann N, Harder J (2009) An improved isolation method for attached-living Planctomycetes of the genus Rhodopirellula. J Microbiol Methods 77:276–284
Winkelmann N, Jaekel U, Meyer C, Serrano W, Rachel R, Rossello-Mora R, Harder J (2010) Determination of the diversity of Rhodopirellula isolates from European seas by multilocus sequence analysis. Appl Environ Microbiol 76:776–785
Zengler K, Toledo G, Rappe M, Elkins J, Mathur EJ, Short JM, Keller M (2002) Cultivating the uncultured. Proc Natl Acad Sci USA 99:15681–15686
Zobell CE (1941) Studies on marine bacteria. I. The cultural requirements of heterotrophic aerobes. J Mar Res 4:42–75
Acknowledgments
We would like to thank Fernando Tavares for phylogenetic support, Helena Abreu and Rui Pereira for sample collection and Rogério Tenreiro (BioFig) for the utilization of the BioNumerics and interpretation of data. This study was financially supported by the pluriannual program of Fundação para a Ciência e Tecnologia (FCT).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
Rights and permissions
About this article
Cite this article
Lage, O.M., Bondoso, J. & Viana, F. Isolation and characterization of Planctomycetes from the sediments of a fish farm wastewater treatment tank. Arch Microbiol 194, 879–885 (2012). https://doi.org/10.1007/s00203-012-0821-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-012-0821-2